Abstract
Background:
DNAJC6 gene is one of the Parkinson’s disease (PD) related genes, but relationship between DNAJC6 polymorphisms and PD remains unclear.
Objective:
We aims to examine the association between genetic variations in DNAJC6 gene and idiopathic early-onset PD (EOPD) in the Chinese population.
Methods:
Exons and intron/exon boundaries of DNAJC6 gene was amplified and sequenced in 135 EOPD patients and 212 healthy controls. Single nucleotide polymorphisms (SNP)-based and haplotype-based association study between EOPD and DNAJC6 was conducted. SNP-SNP interactions were investigated using the generalized multifactor dimensionality reduction (GMDR) method. We further evaluated the effect of variants on gene function and expression using online in silico algorithms and databases.
Results:
We found fourteen previously reported SNPs in the DNAJC6 gene. The frequencies of variant alleles in rs11208644, rs4582839 and rs4915691 were observed significantly higher in EOPD group compared to healthy controls, while in rs6588144 was significantly lower. Additionally, haplotype analysis indicated that the CTCACTCGGC, CTTACTCGGC and TTTGTTCGAC haplotypes were associated with higher EOPD risk in EOPD patients. SNP-SNP interaction analysis showed that rs12077111-rs4592284 SNP combination was the best model with higher EOPD risk. Based on the in silico analysis results, these SNPs were predicted to be no harm to the protein function, but might lead to possible changes in splice site and alter the expression level of DNAJC6.
Conclusion:
Our study indicated that EOPD was associated with several SNPs and haplotypes of DNAJC6 gene.
INTRODUCTION
Parkinson’s disease (PD) is a chronic, progressive, multisystem neurodegenerative movement disorder characterized by complex symptoms including motor symptoms (resting tremor, bradykinesia, rigidity, gait abnormalities) [1], and non-motor symptoms (hyposmia, sleep disorders, affective disorders) [2]. At present, there are over two million PD patients in China, about half of the total number of patients in the world, and the prevalence of PD is almost 1.7% among people older than 65 in China, among which, the patients with early onset age under the age 50 accounted for 5–10% [3]. Onset age of PD less than 50 years was defined as early-onset PD (EOPD) in most literatures with clinical features such as slower disease progression, less cognitive decline, earlier motor complications and higher genetic tendency [4]. Three genes had been identified to be responsible for EOPD with typical symptoms, including Parkin [5], PINK1 [6] and DJ1 [7] genes, while atypical parkinsonism caused by mutations in ATP13A2 [8], PLA2G6 [9], FBXO7 [10] and SYNJ1 [11] genes displaying more complex phenotypes.
The DnaJ heat shock protein family (Hsp40) member C6 (DNAJC6) gene encodes auxilin, which regulates molecular chaperone activity by stimulating ATPase activity in clathrin uncoating [12]. DNAJC6 has been certified as a juvenile parkinsonism gene. Previous studies have indicated that DNAJC6 mutation is associated with the risk for juvenile parkinsonism [12–14] and PD [15, 16]. However, no clear pathogenic mutation in DNAJC6 gene has been found in Chinese classic idiopathic EOPD patients [17, 18], and the relationship between DNAJC6 polymorphisms and EOPD remains unclear. Therefore, we performed this SNP- and haplotype-based case-control study to illustrate the association between DNAJC6 and EOPD risk in Chinese population.
MATERIALS AND METHODS
Subjects
Subjects included in this study consisted of 135 EOPD patients with mean age of 52.82±1.63 years (age at onset 43.84±7.69, 73 male and 62 female) and 212 age- and sex-matched healthy controls (mean age 53.17±0.51 years, 118 male and 94 female), all recruited from the Second Affiliated Hospital, School of Medicine, Zhejiang University of China. The patients were diagnosed with The International Parkinson and Movement Disorder Society (MDS) clinical diagnostic criteria for PD [19]. This study has been approved by ethics committee of Second Affiliated Hospital, School of Medicine, Zhejiang University.
Genotyping
Blood samples were collected from all subjects, and genomic DNA was extracted from blood leukocytes by phenol chloroform extracting method. We amplified the exons and intron/exon boundaries of DNAJC6 gene by polymerase chain reaction (PCR) and then directly sequenced using an ABI 3730 XL genetic analyzer (Applied Biosystems, Foster City, USA). Alignment and analysis of the sequencing results was carried out with DNAStar (DNAStar, In Madison, WI).
Statistical analysis
Statistical analysis was performed using the GraphPad prism software (version 6.0). Hardy-Weinberg equilibrium (HWE) test was used to test for the population stratification and other forms of non-random mating. Allele frequencies and genotype distributions of detected SNPs in DNAJC6 gene were compared between the EOPD and healthy control groups using chi-squared test or Fisher’s exact tests as appropriate, and the results were considered as statistically significant if p-value < 0.05. The strength of the association with EOPD was estimated by odds ratio (OR) with its 95% confidence interval (CI). We used the Bonferroni correction to account for multiple testing and a two-tailed p-value < 0.0036 (= 0.05/14 SNPs) was considered statistically significant. We also extracted data from two online databases, 1000 Genomes Project (http://www.1000genomes.org/) and ExAC Browser (Exome Aggregation Consortium; http://exac.broadinstitute.org/) for comparison between our EOPD patients and healthy controls in East Asian population. Three logistic regression models (dominant, recessive and additive) were also used to analyze the SNPs, and a two-tailed p-value < 0.0012 (= 0.05/42 tests) was used for the Bonferroni correction.
Linkage disequilibrium (LD) and haplotype-based case–control analysis were performed using Haploview 4.2 software [20], results were considered as statistically significant if p-value < 0.05. Bonferroni corrected p-value < 0.0042 (= 0.05/12 haplotypes) was considered statistically significant. For SNP-SNP interactions, we used a generalized multifactor dimensionality reduction (GMDR) model to investigate the multiplicative interaction effect and find the best interaction combination among SNPs [21]. The GMDR software provides some parameters including cross-validation consistency score, testing balance accuracy, and p-value for sign test as described by Kim et al. [22]. The cross-validation consistency score is a measure of the degree of consistency defined as the number of times in a particular validated run that a given attribute combinations was selected as the best model. The testing balance accuracy measures the degree to which the interaction accurately predicts case or control status with a score between 0.5 (indicating the model predicts no better than chance) and 1.0 (indicating perfect prediction). The sign test for prediction accuracy can be used to measure the significance of an identified model.
In silico analysis
To predict the effect of variants on gene function, we used the in silico algorithms: Mutation Taster (http://www.mutationtaster.org) [23]. Possible changes of the splice sites were predicted using NNSplice program encoded in Mutation Taster website. Genetic regulation of both wild-type and variant-type genes were determined using the Brain eQTL Almanac (Braineac) dataset (http://caprica.genetics.kcl.ac.uk/BRAINEAC/), which contains brain tissues from ten regions including thalamus (THAL), frontal cortex (FCTX), substantia nigra (SNIG), temporal cortex (TCTX), medulla inferior olivary nucleus (MEDU), hippocampus (HIPP), occipital cortex (OCTX), putamen (PUTM), intralobular white matter (WHMT) and cerebellum (CRBL).
RESULTS
SNP-based analysis results
We have investigated the exons and exon-intron boundaries of DNAJC6 in this study. After sequencing of the PCR-amplified fragments, we found fourteen known SNPs in this gene: one missense SNP (rs4915691), four synonymous SNPs (rs4325172, rs11208644, rs12077111, rs4582839), and nine intron SNPs in intron/exon boundaries (rs2296480, rs2296481, rs10889546, rs6588142, rs6588143, rs6588144, rs4592284, rs2274437) (Fig. 1). Before genetic analysis, it is preferred to check whether our data are free from sample level substructure or genotyping error using the HWE test. We proved that the genotype frequencies in healthy control group were all accorded with HWE (p > 0.05) (Table 1). We found five SNPs, rs4325172 (p = 0.00001), rs11208644 (p = 0.018), rs12077111 (p = 0.00001), rs4582839 (p = 0.0063) and rs4592284 (p = 0.037) did not fit with HWE in EOPD group, indicating these polymorphisms might be associated with EOPD.
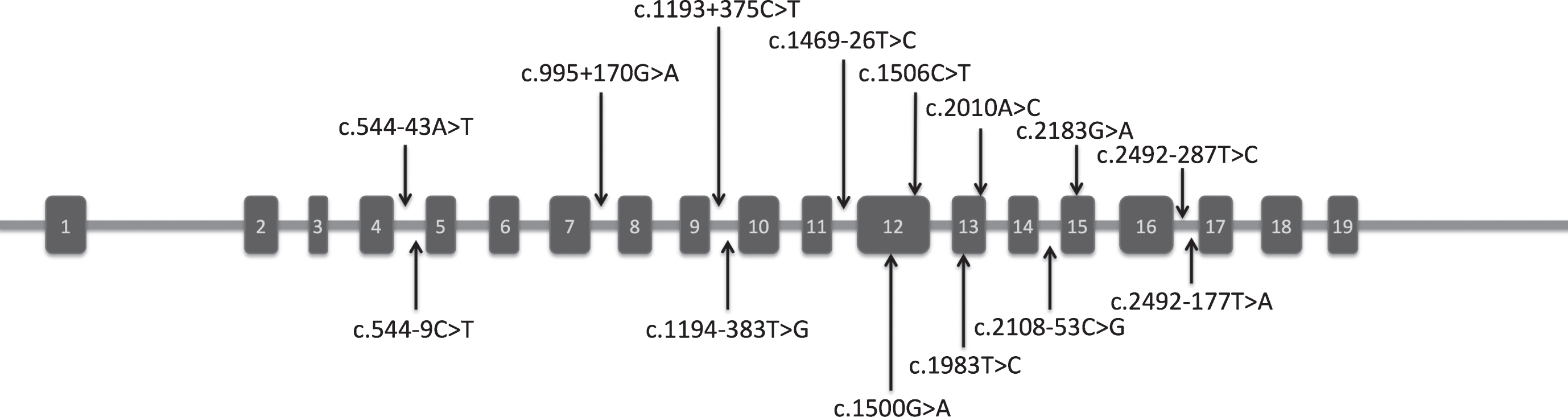
Schematic representation of the DNAJC6 gene. Black arrows indicate the locations of SNPs detected in this gene.
Allele and genotype distribution of SNPs in EOPD patients and controls
P-value≤0.05 considered as statistically significant (in bold). *means the p-values still maintained their significant after Bonferroni correction (0.05/14 = 0.0036).
Genotype distribution and allele frequencies of these fourteen SNPs are shown in Table 1. The frequency of C allele in rs6588144 (p = 0.0001) was significantly lower in EOPD group compared to our healthy controls. The frequencies of T allele in rs11208644 (p = 0.010) and A allele in rs4915691 (p = 0.003) were observed significantly higher in EOPD group compared to healthy controls, but the former SNP didn’t survive the Bonferroni correction. The frequencies of genotypes of rs6588144 (p = 0.0004), rs4325172 (p = 0.009), rs11208644 (p = 0.0007), rs12077111 (p = 0.00002) and rs4915691 (p = 0.003) were significantly different between EOPD patients and healthy controls. Distributions of genotypes and alleles of other SNPs showed no significant differences between two groups. At the same time, we estimated the differences of allele and genotype frequencies between our EOPD patients and control group from online database, similar results were obtained, but didn’t survive the Bonferroni correction (Table 5).
Furthermore, the genotype frequencies of these fourteen SPNs were further analyzed based on three genetic models: dominant, recessive and additive, model (Table 2). We found significant association between SNP rs4915691 and higher EOPD risk in both recessive model (OR = 2.196, 95% CI = 1.397–3.451, p = 0.0006, survived after Bonferroni correction) and addictive model (OR = 1.666, 95% CI = 1.187–2.339, p = 0.003). In the case of SNP rs11208644, significant positive correlation between this SNP and EOPD risk were also found in recessive model (OR = 2.268, 95% CI = 1.439–3.575, p = 0.0004, survived after Bonferroni correction) and addictive model (OR = 1.547, 95% CI = 1.107–2.162, p = 0.010). In the case of SNP rs6588144, we found significant association between this SNP and lower EOPD risk in dominant model (OR = 0.334, 95% CI = 0.192–0.582, p = 0.0001) and addictive model (OR = 0.387, 95% CI = 0.237–0.631, p = 0.0001), both survived after Bonferroni correction. A lower risk of EOPD was found in dominant model (OR = 0.439, 95% CI = 0.249–0.776, p = 0.004) of SNP rs12077111.
Association between DNAJC6 genotypes and EOPD based on three genetic models
Abbreviations: OR, odds ratio; p-value≤0.05 considered as statistically significant (in bold). *means the p-values still maintained their significant after Bonferroni correction (0.05/42 = 0.0012).
Haplotype-based analysis results
LD was assessed between the fourteen DNAJC6 SNPs genotyped within our data using the Haploview software. As shown in Fig. 2A, D’ values = 1 indicates complete LD and D’>0.8 is strong LD as represented by a red-colored square. Complete LD or nearly complete was detected between the following SNPs: rs2296480 and rs2296481; rs10889546, rs6588142, rs6588143 and rs6588144; rs6588144, rs4325172, rs3818513 and rs2274437. All these 14 SNPs were located in one haplotype block because most D’ were beyond 0.5. The r2 values are shown in Fig. 2B, r2 values = 1 indicates complete LD and r2 > 0.8 is strong LD as represented by a black-colored square. Following SNPs were in complete or nearly complete LD: rs2296480 and rs2296481; rs10889546, rs6588142 and rs6588143; rs3818513 and rs2274437. We selected a representive SNP from each of the above three groups separately, then performed the association analysis based on ten SNPs (rs2296481, rs6588142, rs6588144, rs4325172, rs11208644, rs12077111, rs4582839, rs4592284, rs4915691 and rs3818513).
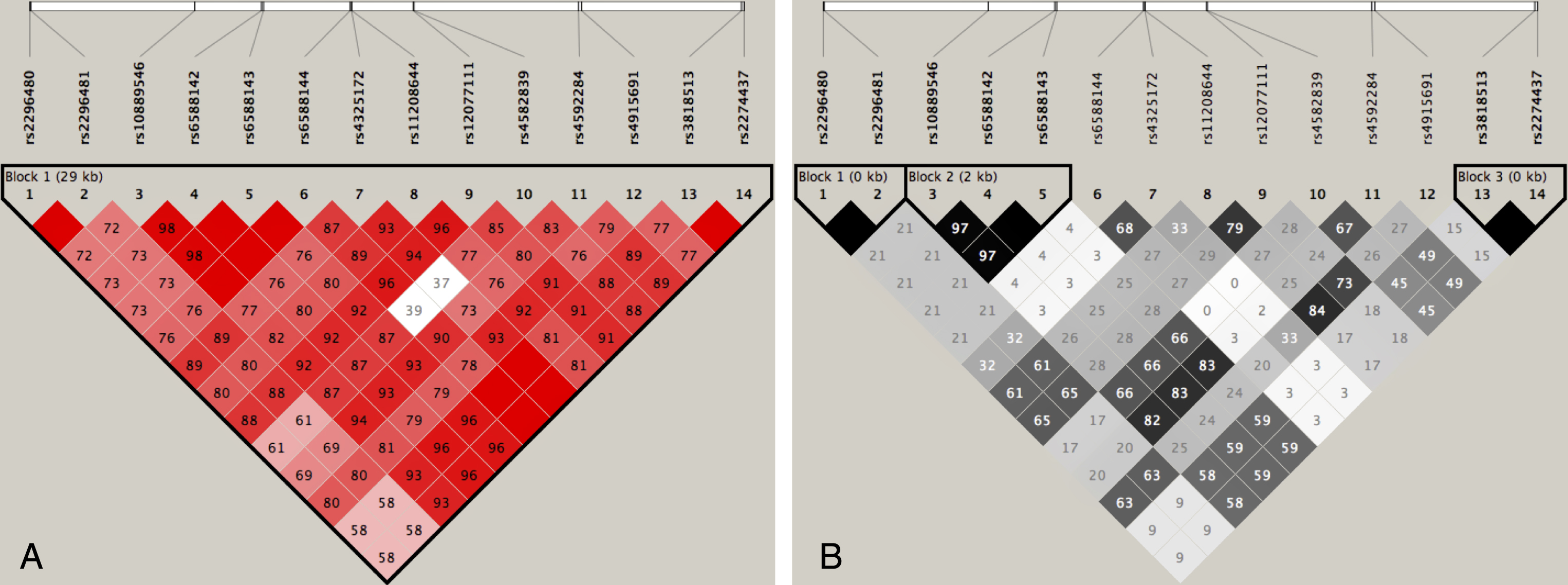
The patterns of linkage disequilibrium in the DNAJC6 gene, with their |D’| (A) and r2 values (B).
Haplotypes of DNAJC6 were generated using the Haploview software based on our genotyping data of the ten SNPs and haplotype-based association analysis was performed between the EOPD and healthy control groups. In the haplotype-based case–control analysis, CTCACTCGGC haplotype (p = 0.000013), CTTACTCGGC haplotype (0.00000014) and TTTGTTCGAC haplotype (p = 0.0005) were associated with significantly higher risk for EOPD, while TTCGCTCGAC haplotype (p = 0.0463) was found lower risk for EOPD. The other haplotypes showed no significant differences between two groups (Table 3). We further conducted a Bonferroni correction to control for the overall type I error rates, since our sample size was small. After the Bonferroni correction, significant associations still maintained for these three haplotypes with higher EOPD risk.
Distribution of Haplotypes in EOPD patients and controls
SNP-IDs of haplotypes: rs2296481, rs6588142, rs6588144, rs4325172, rs11208644, rs12077111, rs4582839, rs4592284, rs4915691 and rs3818513. Abbreviations: freq, frequency; p-value≤0.05 considered as statistically significant (in bold). *means the p-values still maintained their significant after Bonferroni correction (0.05/12 = 0.0042).
SNP-SNP interaction analysis results
The GMDR model was used to estimate the multiplicative interaction effect and find the best interaction combination among fourteen detected SNPs of DNAJC6 gene. Table 4 summarizes the results obtained from GMDR analysis using 2- to 5-locus models for SNP-SNP interaction. A significant 2-locus model (p = 0.0010) involving rs12077111 and rs4592284 SNPs was selected as the best combination with cross-validation consistency of 7/10, indicating a potential SNP–SNP interaction between rs12077111 and rs4592284. There were also a significant 3-locus model (p = 0.0010) involving rs6588144, rs4582839 and rs4915691 SNPs, as well as a significant 5-locus model (p = 0.0010) involving rs10889546, rs6588144, rs432517, rs12077111 and rs4592284 SNPs were identified with cross-validation consistency of 5/10, indicating potential SNP–SNP interactions among these SNPs of DNAJC6 gene. The 2- to 5-locus models all had the testing accuracy over 60.0%.
GMDR analysis on the best SNP-SNP interaction models
In silico analysis results
Pathogenicity of these variants was predicted using Mutation Taster program (Table 5), all were predicted to be polymorphism. NNSplice program predicted that most SNPs would lead to possible changes of the splice sites, except SNP rs6588142, rs6588144 and rs4592284. We obtained the brain expression pattern of DNAJC6 gene from the Braineac database, which showed significant higher expression level (1.8-fold change, p = 1.8*10–22) in SNIG than CRBL (Fig. 3). Variant-type alleles of most SNPs were associated with significant higher expression level of DNAJC6 in several brain regions (Fig. 3).
Details of the SNPs detected during the present study
aNNsplice analysis possible changes in splice site, i.e. Donor gained (DG), Donor increased (DI), Donor marginally increased (DMI), Accept gained (AG), Accept increased (AI), Accept marginally increased (AMI), No abrogation of potential splice sites (NO); bMutationTaster predictions: Polymorphism (P). P-value≤0.05 considered as statistically significant (in bold). *means the p-values still maintained their significant after Bonferroni correction (0.05/14 = 0.0036).
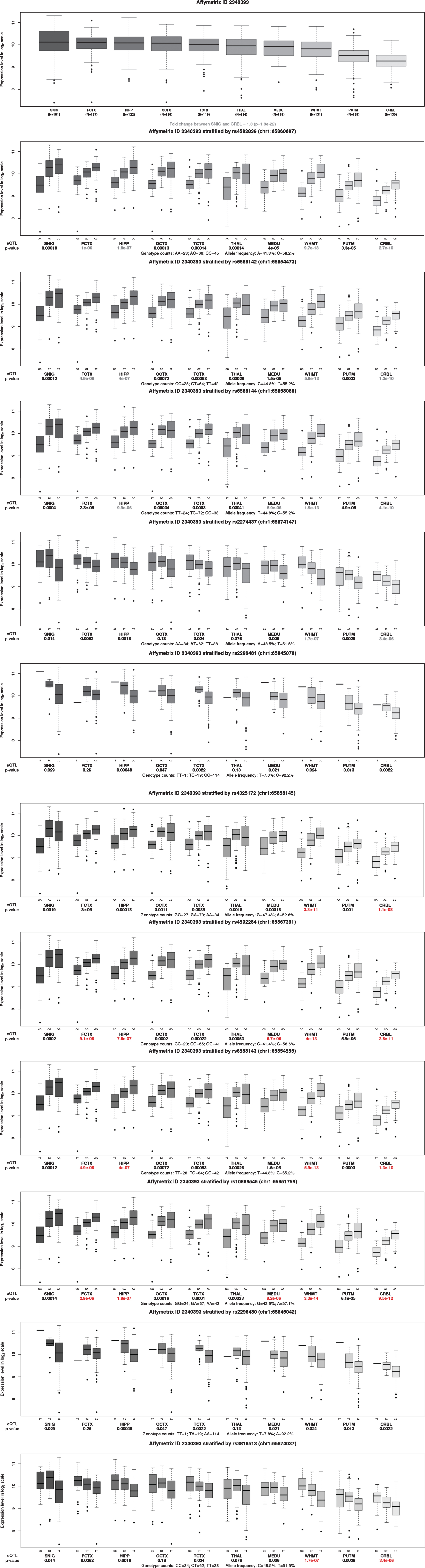
Effect of different variants on the expression levels of DNAJC6 in Braineac database. Box plot of mRNA expression levels in ten brain regions. THAL, thalamus; FCTX, frontal cortex; SNIG, substantia nigra; TCTX, temporal cortex; MEDU, medulla inferior olivary nucleus; HIPP, hippocampus; OCTX, occipital cortex; PUTM, putamen; WHMT, intralobular white matter; CRBL, cerebellum.
DISCUSSION
In this study, the exons and intron/exon boundaries of DNAJC6 gene was sequenced among a sample of Chinese EOPD and healthy controls. Single SNP based, haplotype-based case-control association study of DNAJC6 gene, and the SNP-SNP interaction analysis were performed for the first time in our study. We found SNPs and haplotypes of DNAJC6 gene were associated with EOPD risk in Chinese population. Findings of this study may provide further evidence for the contribution of DNAJC6 gene in PD pathogenesis.
DNAJC6 gene encodes auxilin, which is expressed in neurons and enriched in presynaptic termini. It is a synaptic chaperone protein, plays a role in recycling synaptic vesicles and contributes to the maintenance of the nerve terminal [24]. Previous studies have indicated that DNAJC6 mutation is associated with the risk for juvenile parkinsonism [12–14] and PD [15, 16]. A homozygous splicing mutation c.8010–2A>G in the DNAJC6 gene was previously reported in two brothers with autosomal-recessive juvenile parkinsonism [12]. A homozygous truncating mutation c.2200C>T (p.Q734X) in DNAJC6 gene was identified in a Turkish family with juvenile parkinsonism, these four affected family members also presented clinical features including mental retardation, pyramidal signs and epilepsy [13]. A nonsense mutation in DNAJC6 c.2365C>T (p.Gln789X) were then reported in juvenile PD patients [14]. Olgiati et al. [16] identified three mutations, c.2779A>G (p.Arg927Gly), c.2223A>T (p.Thr741X) and c.2038 + 3A>G/c.1468 + 83del in EOPD patients, and those DNAJC6-mediated PD patients presented atypical symptoms including rapid disease course with poor response to L-DOPA, pyramidal signs, seizures, mental retardation, and dystonia. Homozygous mutations in the DNAJC6 gene significantly reduced the expression levels of auxilin, which may disturb synaptic vesicle endocytosis and trafficking in the pathogenesis of PD [16]. Previous studies haven’t reported the presence of pathogenic DNAJC6 mutation in Chinese idiopathic EOPD patients [17, 18], and the relationship between DNAJC6 polymorphisms and EOPD remains unclear.
In the present study, we detected fourteen known SNPs in DNAJC6 gene, while no clear pathogenic mutation was found in our study. We found that rs11208644 and rs4915691 of the DNAJC6 gene were significantly associated with the risk of EOPD. The variant genotypes of four SNPs (rs4325172, rs11208644, rs12077111 and rs4915691) significantly differed between EOPD patients and healthy controls, indicating an increased risk for PD with these variants. Haplotype analysis can provide more information on complex relationship between variations and disease. We established haplotypes for DNAJC6 gene from different combinations of selected ten SNPs. CTCACTCGGC, CTTACTCGGC and TTTGTTCGAC haplotypes significantly increased risk of EOPD. SNP-SNP interaction analysis showed that rs12077111-rs4592284 SNP combination is the best model with higher EOPD risk.
The DNAJC6 gene is relatively higher expressed in substantia nigra, the neurodegeneration of which is the main pathological hallmark in PD. Based on the results of the in silico analysis, we found that these SNPs were predicted to be no harm to the protein function, but most of them might lead to possible changes in splice site and alter the expression level of DNAJC6 in several region (Table 5).
In conclusion, we aimed to illustrate the roles of DNAJC6 polymorphisms in modifying the risk of EOPD in Chinese population. This study indicated that EOPD was associated with several SNPs and haplotypes of DNAJC6 gene. Our data may contribute to a better understanding of the genetics of EOPD, while further genetic analysis and molecular biological studies still needs to be carried out with a larger population of subjects.
CONFLICT OF INTEREST
The authors declare no conflict of interest.
Footnotes
ACKNOWLEDGMENTS
We would like to thank all individuals participating in the study. This work was supported by grant 81520108010 and 81600982 from the National Natural Science Foundation of China.
