Abstract
Keywords
INTRODUCTION
Multiple system atrophy (MSA) is a rare neurodegenerative disease that usually develops in the sixth-seventh decade of life with rapidly progressive symptoms typically leading to death between six-to-nine years after onset [1–6]. Consensus criteria for MSA diagnosis distinguish MSA with predominant Parkinsonism (MSA-P) and predominant cerebellar ataxia (MSA-C) [3]. Some of these subjects eventually develop features of both MSA-P and MSA-C [3, 6–8]. Unfortunately, the underlying mechanism of the neuroinflammation and neurodegeneration in MSA is not known, and symptoms occur late in the course of the disease when significant neuronal loss has already occurred.
Our group [9–11] and others [12] have data suggesting that gut-derived inflammation may sustain peripheral and neuroinflammation leading to neurodegeneration in Parkinson’s disease (PD). We hypothesized that like PD, subjects with MSA also have abnormal intestinal barrier function and dysbiosis leading to gut-derived inflammation. This gut-derived, endotoxin-mediated, inflammatory cascade could lead to local/systemic inflammation, elevated pro-inflammatory cytokines, and oxidative stress contributing to MSA pathology [13–17]. This hypothesis is based on: (1) the importance of the gut-brain axis and intestinal microbiota in neurological disorders [11, 18–20]; (2) similarities of pathological characteristics between MSA and PD subjects [21]; and (3) like PD, subjects with MSA-P and MSA-C suffer from multiple gastrointestinal (GI) symptoms including constipation [22–26], fecal incontinence [27–29], delayed gastric emptying [28], slow colonic transit, and abnormal electromyography of the external anal sphincter [27, 29–32].
To test our hypothesis, we assessed intestinal tight junction (TJ) barrier integrity, endotoxin-related inflammation markers, and sigmoid mucosal-associated and fecal microbiota compositions comparing MSA and healthy control (HC) subjects. We found preliminary evidence that MSA subjects had intestinal disruption of the TJ barrier, increased marker of endotoxin-related intestinal inflammation, and an abnormal microbiota composition with increased abundance of putative, “pro-inflammatory” bacteria.
MATERIALS AND METHODS
Subject enrollment
Healthy controls (N = 11) were recruited via flyers and research advertisements at Rush University Medical Center (RUMC, Chicago, IL) and MSA (N = 6) were recruited from the Movement Disorders clinic at RUMC. All subjects gave informed consent for the use of their samples and data becoming part of an RUMC Institutional Review Board (IRB)-approved GI repository and to the research studies performed for this investigation. All work for this study was carried out in accordance with the Code of Ethics of the World Medical Association (Declaration of Helsinki) for experiments involving humans and Uniform Requirements for manuscripts submitted to biomedical journals published by the International Committee of Medical Journal Editors.
MSA was diagnosed according to consensus guideline criteria [3, 33]. MSA subjects were diagnosed by a movement disorders expert (KMS) and had Parkinsonism with early autonomic dysfunction and a poor levodopa response. Exclusion criteria for MSA subjects were: (1) the use of probiotics or antibiotics within three months prior to sample collection, (2) primary GI pathology, (3) unstable medical, neurological, or psychiatric illness, (4) low platelet count (<80k), uncorrectable prolonged prothrombin time (>15 sec), or history of bleeding that precludesbiopsies.
Inclusion criteria for healthy control (HC) subjects (N = 11) was as follows: (1) normal physical exam, complete blood count (CBC), and comprehensive metabolic profile (2) no gastrointestinal (GI) complaints, symptoms, or documented GI chronic disease, (3) no neurodegenerative disease, (4) consumption of no more than a moderate amount of alcohol (NIAAA definition) [34], (5) no probiotic or antibiotic use for at least three months; nor NSAID or high dose aspirin use for at least four weeks prior to sample collection. Low dose (81 mg/day) aspirin was allowed. Five HC subjects were on medication for hyperlipidemia (Oretic, Lipotor, Zextia), hypertension (Diovan, Diltiazem, Metoprolol), or anxiety/depression (Lexapro) at the time of sample collection. Removal of the HC subjects on medication did not alter the between group differences (data not shown); thus, these samples were included for all data analysis.
Exclusion criteria for HC subjects: (1) primary gastrointestinal pathology, (2) unstable medical, neurological, or psychiatric illness, (3) daily alcoholic beverage consumption; (4) at risk alcohol drinkers based on NIAAA criteria (i.e., females more than three drinks per day on a regular basis; men more than four drinks per day) [35]; (5) unreliable drinking history; (6) the use of probiotics or antibiotics within three months prior to sample collection, and (7) low platelet count (<80k), uncorrectable prolonged prothrombin time (>15 seconds), or history of bleeding that precludes biopsies.
Mucosal and fecal specimen collection
Sigmoid mucosal biopsies were collected via endoscopy at the Rush University Medical Center (RUMC; Chicago, IL) Endoscopy Lab. A limited, un-prepped sigmoidoscopy was performed using a standard adult upper endoscope (Olympus America Inc., Center Valley, PA) to 20–25 cm from the anal verge. Suction was not used during advancement of the scope and the biopsy forceps was not taken out of the channel of the scope until sample collection. Biopsies were taken from pink mucosa without visible feces at the sigmoid colon about 20 cm from the anal verge and were either snap frozen in liquid nitrogen or processed with 10% formalin fixation in the endoscopy room.
Fecal samples were collected by subjects, four weeks after the sigmoidoscopy, using an anaerobic home collection kit (BD Gaspak, Becton Dickinson and Company, Sparks, MD). Each subject had one mucosa and one fecal sample collected, if they consented to both types of sample collections. Sigmoid mucosal biopsies and fecal samples were stored at – 80°C until use.
Immunohistochemistry and microscopic analyses
Histological analysis was performed on sigmoid biopsies collected from all six MSA and ten out of eleven HC subjects. Immunofluorescence staining of Zonula Occludens-1 (ZO-1) and immunoperoxidase staining for toll-like-receptor-4 (TLR4) in formalin-fixed-paraffin-embedded sigmoid mucosa samples were performed.
Immunohistochemistry
After formalin fixation, colonic sigmoid mucosal biopsies were processed for paraffin embedding. The paraffinized tissue blocks were then stored at room temperature, until use. Six micrometer thick sections were made using the microtome to perform immunohistochemistry. Immunoperoxidase/immunofluorescence staining was performed on these colonic sigmoid mucosal sections, according to previously established protocols [36, 37]. Briefly, the paraffin was melted at 60°C using an oven. Xylene and different grades of alcohol were used to remove melted paraffin from the tissue and samples were brought into water. Antigen retrieval was performed for 20 minutes using citrate buffer (Dako, cat#S1699) in a pressure cooker at high pressure. Samples were then allowed to cool at room temperature. Sodium periodate incubation was performed to quench endogenous peroxidase and serum block was performed for an hour to prevent non-specific antibody binding. Primary antibody incubation was performed for overnight at room temperature (TLR4: cat#ab22048, Abcam, 1 : 500; ZO-1 cat#617300, Abcam, 1 : 200). Next day, the secondary antibody was applied for an hour at room temperature. Immunofluorescence staining [37] was performed for Zonula Occludens-1 (ZO-1) tight junction protein marker using fluorophore tagged secondary antibody (Donkey anti-Rabbit 555: A31572, Life Technologies, 1 : 400). Fluorescence staining was visualized under confocal microscope. Immunoperoxidase staining was performed for Toll-like-receptor-4 (TLR4) marker using biotinylated secondary antibody (Horse anti-Mouse, BA2000, 1 : 200), followed by actin biotin complex incubation (ABC vectastain, PA1000) and color development in the presence of DAB chromogen as well as hydrogen peroxide. Immunofluorescence sections were counter stained using 4’,6-diamidino-2-Phenylindole (DAPI) (1 : 10000, D1306, Thermo Fischer Scientific, Carlsbad, CA) solution for five minutes. Immunoperoxidase sections were counter stained using hematoxylin (SH26-500D, Fisher Scientific, Atlanta, GA) according to manufacturer’s protocol. Immunofluorescence slides were mounted using aqueous mounting media (F4680, Sigma-Aldrich,St. Louis, MO) and stored in 4°C. Immunoperoxidase slides were dehydrated using different grades of alcohol and three treatments of xylenes. These slides were then cover slipped using xylene based mounting media (8310-16, Thermofisher Scientific, Waltham, MA) and stored at room temperature. Fluorescent images were taken using Zeiss confocal microscope (LSM700, Zeiss, Germany) and images were overlaid using Adobe Photoshop (v.CS3, Adobe, San Jose, CA) software. Light microscope (BX61, Olympus, Waltham, MA) was used to evaluate immunoperoxidase performed sections.
Quantification of Zonula Occludens-1 integrity
Confocal microscopic analyses were performed to evaluate colonic sigmoid mucosa’s ZO-1 tight junction barrier integrity. To keep consistencies among all samples, all fluorescence images were taken at constant levels: laser intensity, confocal aperture, photomultiplier voltage, offset, scan speed, image size, filter and magnification. These settings were maintained throughout the entire microscopic analysis procedure. Using a 20×1.2 or 40×1.2 magnification objective and a 555 nm excitation source, in combination with 405 nm (DAPI) excitation source, images were acquired at each sampling site from lamina propria (colonic epithelial lining and crypts) of MSA and HC subjects.
Quantification of the colonic sigmoid mucosa’s tight junction barrier integrity was performed using scale (0 – 3) for ZO-1 immunofluorescence sections. Scale (0 – 3) scores represent: [0 = no ZO-1 immunofluorescence; 1 = very light and discontinuous ZO-1 immunofluorescence; 2 = intense and discontinuous ZO-1 immunofluorescence; 3 = smooth continuous and well organized ZO-1 immunofluorescence]. One slide containing colonic sigmoid mucosa was used per case subject for imaging analyses. The slides were randomized and coded to collect fluorescence images to avoid any biases. Minimum of twenty crypts were analyzed per case subject and theaverage values were collected for comparisons between groups. ZO-1 integrity scoring data were represented average score+SEM. Mann-Whitney U (MWU) test was performed to compare MSA and HC subjects using GraphPad Prism (v5.03) software. Significance was accepted at P < 0.05.
Stereological Toll-like-receptor-4 cell evaluation
Light microscope attached to stereoinvestigator cell count probe (S.I.-V.1.5, MBF Bioscience, Williston, VT) was used to evaluate colonic sigmoid mucosa’s TLR4 positive cells into the lamina propria of MSA compared to HC subjects. One slide containing colonic sigmoid mucosa was used to evaluate TLR4 positive cells in the lamina propria. Stereology was performed according to an unbiased counting principle [38, 39]. Briefly, the lamina propria (area between epithelial crypts and the muscularis layer) was selected and outlined under low magnification (10x) in each colonic sigmoid mucosa sample. Using high magnification (60x), all counting sites (with grid size 100μmX100μm and counting frame size 90μmX90μm) were visited to assess the lamina propria, with an average of 250 counting frames per case. Using probe run analyses, data for estimated number of cells and total counting area (mm2) were collected to compare the data between groups. The data were presented as the estimated number of cells/counting area. Two-tailed unpaired t-test was performed to compare between MSA and HC subjects in GraphPad Prism (v5.03) software. Significance was accepted at P < 0.05.
Microbiota profiling and bioinformatics analyses
Total DNA was extracted from colonic mucosa and feces (FastDNA bead-beating Spin Kit for Soil, MP Biomedicals, Solon, OH), amplified the V4 variable region of the microbial 16S ribosomal RNA (rRNA) gene (Earth Microbiome Project primer set, adapted for the Illumina platform) [40], and sequenced on an Illumina MiSeq (2×150 reads) at Argonne National Laboratory. Negative controls were used with each set of amplifications, which indicated no contamination. Raw sequence data (FASTQ files) were deposited in the NCBI Sequence Read Archive under project PRJNA268515. Mapping data for demultiplexing are provided in (Supplementary Table 1).
Forward and reverse reads were merged and were quality trimmed and sequences longer than 200 bases were exported (CLC Genomics Workbench, v7.0, CLC Bio, Qiagen, Boston, MA). Sequences were screened for chimeras (usearch61 algorithm) [41], and putative chimeric sequences were removed from the dataset (QIIME v1.8.0) [42]. Number of reads and chimeric reads per sample are shown in (Supplementary Tables 2 and Tables 3). Each sample sequence set was rarefied to 8,700 sequences [43] and data were pooled, renamed, and clustered into operational taxonomic units (OTU) at 97% similarity (usearch61 algorithm). Representative sequences from each OTU were extracted and classified using the uclust consensus taxonomy assigner (Greengenes 13_8 reference database). A biological observation matrix (BIOM) [44] was generated at each taxonomic level (“make OTU table” algorithm) and analyzed and visualized using the software package Primer6 [45]. Differences in the relative abundance (RA) of individual taxa between a priori defined groups were detected using Kruskal-Wallis non-parametric one-way analysis of variance, implemented within the software package QIIME [42]. Taxa with an average abundance of <1% across the sample set were removed from the analysis. The RA of individual taxa reported was accepted at a significance of (P < 0.05), but failed when corrected for false discovery rate (FDR-P < 0.05). In addition, statistical testing of the differences in the RA of individual taxa between groups was performed using the parametric DESeq2 negative binomial Wald test, implemented within the software package R, generating an adjust P-value. In SPSS (v.22), two-tailed unpaired Independentt-test was used to analyze differences for parametric data satisfying test assumptions; Fisher’s Exact or Mann-Whitney U (MWU) tests were used to analyze nonparametric data. Graphs were created using GraphPad Prism (v5.03) software.
In silico community functional predictions were performed using PICRUSt (Phylogenetic Investigation of Communities by Reconstruction of Unobserved States) [46] and differences in Kyoto Encyclopedia of Genes and Genomes (KEGG) ortholog (KO) abundances between groups were identified after filtering to remove low abundance KOs (Kruskal-Wallis test, 1% filter threshold) [47]. KEGG orthologs were analyzed using the KEGG Mapper pathway search function [48]. PICRUSt analysis significance was accepted at P < 0.05.
Classification of putative “pro-inflammatory” and “anti-inflammatory” bacteria taxa were based on preceding reports [49–53]. Colonic mucosal and fecal samples were examined because the bacterial RA of individual taxa is shown to differentiate between sample sites [10, 54]. Thus, it is important to collect and analyze both fecal and mucosa samples to examine the entire microbial composition within the GItract.
RESULTS
Subject comparisons
The demographic characteristics of the study subjects are depicted in Supplementary Tables 2 and Tables 3. These variables were similar between the six MSA and 11 HC subjects (age P = 0.09, sex P = 0.33, Body Mass Index (BMI) P = 0.11, race P = 0.51). There were no significant correlations between bacteria with all demographic characteristics (feces/mucosa vs. age/sex/BMI/race; phylum/family/genus) (data not shown). Parkinsonism in MSA subjects was assessed using the Unified Parkinson’s disease rating scale (UPDRS) and the modified Hoehn and Yahr scale (H&Y Stage) [55, 56]. MSA subjects had average (mean±SEM) disease duration of 5.50±1.36 years, UPDRS of 55.33±7.77, H&Y Stage of 3.42±0.37. Constipation in MSA subjects was assessed using a standard visual analog scale from 0–10 (0 = no symptoms, 10 = severe constipation). Heathy control subjects reported no constipation; while MSA subjects complained of constipation with constipation duration of 3.38±1.03 years, and constipation severity score of3.83±1.33.
Tight junction protein Zonula Occludens-1 integrity in the sigmoid mucosa
MSA subjects have evidence of disrupted ZO-1 integrity compared to HC subjects. Immunofluorescence staining for ZO-1 showed distribution of this TJ protein restricted to the epithelial lining of the colonic lamina propria and epithelial apical cell junctions of the crypts. Further analyses using confocal microscope showed a typical pattern of smooth and organized integrity for ZO-1 structure at the apical surface of the crypts (Fig. 1A: a’) and the epithelial lining of the lamina propria (Fig. 1A: a”) in the HC. In comparison to HC, MSA subjects showed complete disruption or very minimal immunofluorescence, if any, of ZO-1 protein structure at the apical surface of the crypts (Fig. 1A: b’) and the epithelial lining of the lamina propria (Fig. 1A: b”). Scoring for the ZO-1 integrity was performed in a blind manner using 0–3 scale: (0: no ZO-1 immunofluorescence to 3: normal ZO-1 integrity). Analyses revealed significantly lower MWU test: (median values) HC: 2.833; MSA: 0.1667; U = 0.000; P = 0.0031) combined mean score for ZO-1 integrity at the apical surface of the crypts and the epithelial lining of the lamina propria (Fig. 1B) in MSA compared to HC. These data show disruptions of one of the major TJ proteins in MSA subjects, which support the concept that MSA subjects have abnormal intestinal permeability.
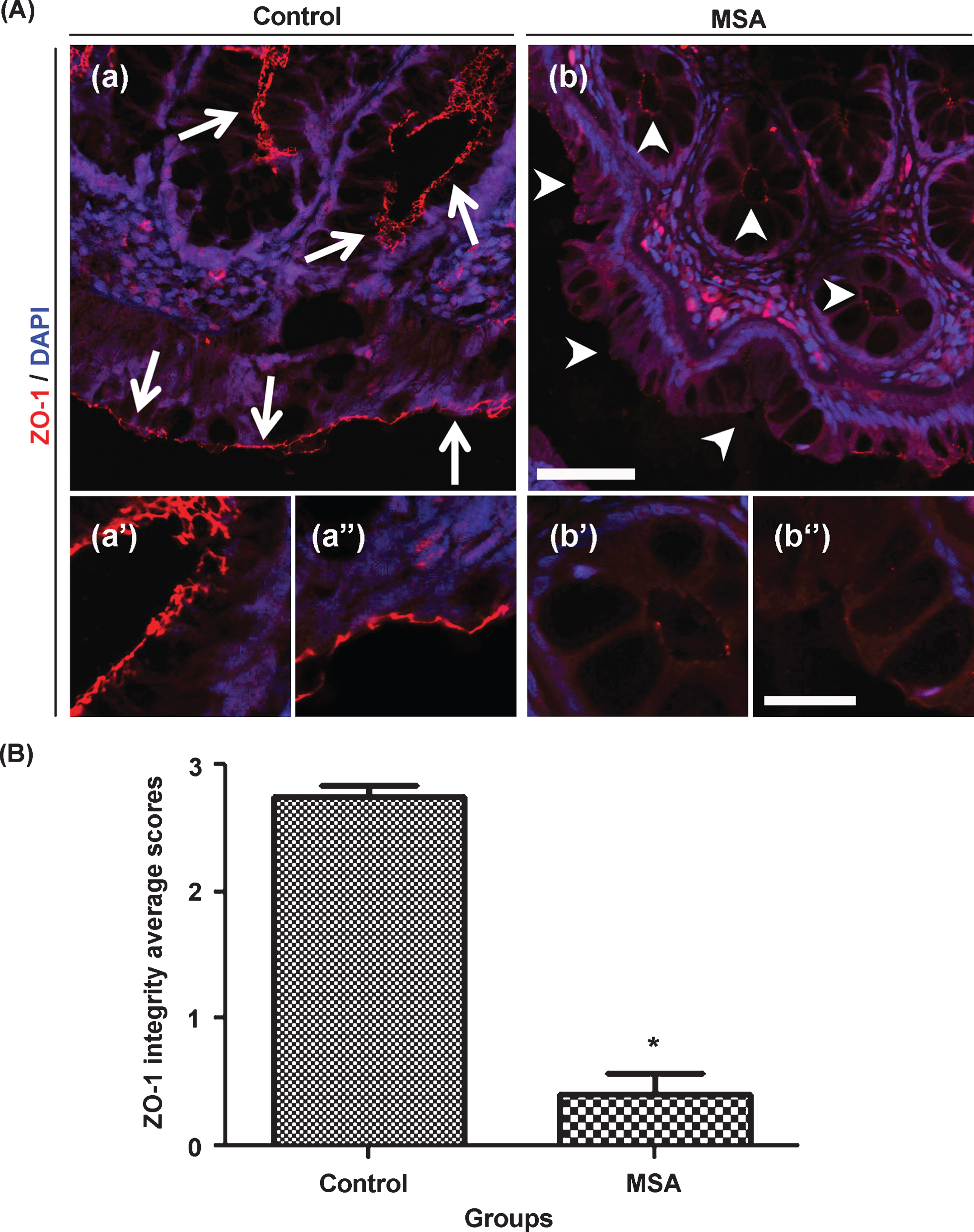
Tight junction protein zonula occludens-1 integrity is significantly disrupted in MSA sigmoidal biopsies compared to healthy control subjects. Photomicrographs (A) represent TJ protein ZO-1 (red) and a nuclear counterstain using DAPI (blue) in HC (a) and MSA (b) sigmoid biopsies. Panels (a’) and (b’) represent apical surface of the crypt while (a”) and (b”) represent epithelial lining of the lamina propria in HC and MSA subjects respectively. White arrows indicate smooth and organized distribution of ZO-1 in HC. In contrast, white arrowheads indicate minimal or no expression of ZO-1 in MSA sigmoid biopsies. Graph (B) represents mean score for ZO-1 integrity in MSA compared to HC. ZO-1 scoring scale represent: [0 = no ZO-1 immunofluorescence; 1 = very light and discontinuous ZO-1 immunofluorescence; 2 = intense and discontinuous ZO-1 immunofluorescence; 3 = smooth continuous and well organized ZO-1 immunofluorescence]. Scale bar in panel (b) and (b”) represent 50 and 20μm and applies to (a, b) and (a’, a”, b’, b”) respectively. Data are presented as mean score values±SEM. *(MWU; P = 0.0031).
Toll-like-receptor-4 expression in sigmoid mucosa
Immunoperoxidase staining for TLR4 showed significantly increased expression of this endotoxin-related inflammation marker in MSA subjects. Microscopic analyses showed very few, if any, TLR4 positive cells in the lamina propria of the HC subjects (Fig. 2A: a, b). Compared to the HC, MSA subjects showed an increased expression of TLR4 positive cells in the lamina propria (Fig. 2A: c, d). Stereo-investigator analyses using MicroBrightField Bioscience showed significantly higher number of TLR4 cells (Unpaired two-tailed t test: t(13) = 3.808, P = 0.0025) in the MSA (Mean±SEM: 66.64±6.64 cells/mm2) lamina propria compared to HC (Mean±SEM: 23.73±7.47 cells/mm2)(Fig 2B).
To further characterize the cell populations expressing TLR4, we investigated enteric glia cells using Glial Fibrillary Acidic Protein (GFAP) marker staining and a double label immunofluorescence protocol. Very few TLR4+ cells were positive for GFAP (data not shown) suggesting that other cells, such as lymphocytes, could be involved with TLR4 expression/activation.
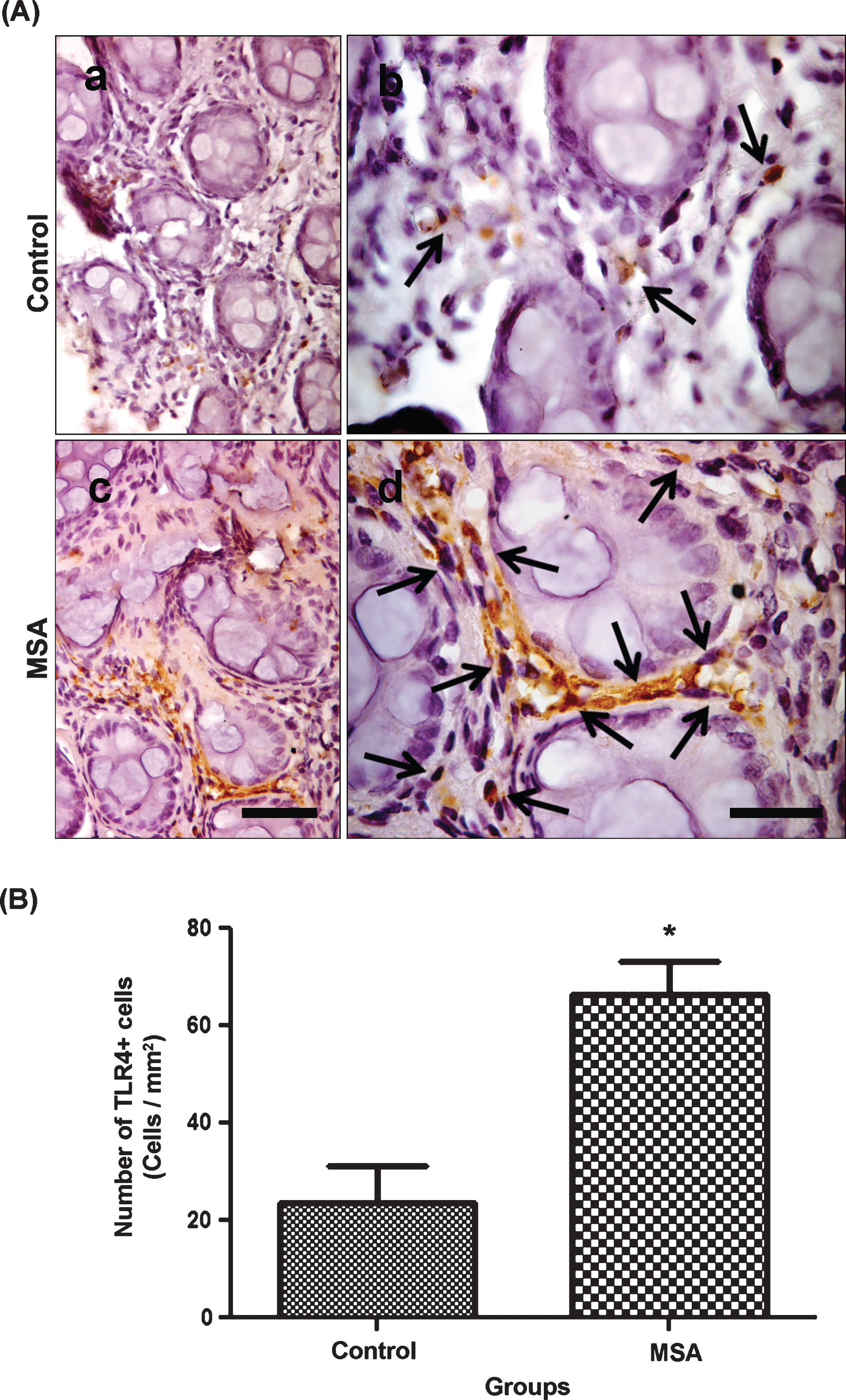
Mucosal Toll-like-receptor-4 expression is significantly higher in MSA sigmoidal biopsies compared to healthy control subjects. Photomicrographs (A) represent TLR4 (brown) immunoperoxidase staining in the sigmoid mucosa of HC (a, b) and MSA (c, d) with hematoxylin counterstain (purple). Microscopic analyses showed higher expression of TLR4 in MSA (c, d) compared to HC (a, b). Black arrows in panels (b) and (d) indicate TLR4 positive cell structures into the lamina propria of the HC and MSA colonic samples respectively. Stereology counting data (B) showed significantly higher number of TLR4 cells in the MSA sigmoid mucosa compared to HC. Scale bars in panels (c) and (d) represent 50μm and 25μm and applies to (a, c) and (b, d) respectively. Data are presented as (average number of cells/area)±(SEM). *(two-tailed unpaired t-test: P = 0.0025).
Fecal microbiota
The fecal microbiota analysis indicated that MSA subjects had higher alpha diversity [57]. Alpha diversity revealed a significant difference in Simpson’s index (MWU; P = 0.04), between MSA and HC subjects at the taxonomic level of phylum (Supplementary Table 4). Additionally, the RA of two phyla and three families were different between MSA and HC (Fig. 3A, Fig. 4A, and Supplementary Table 5) [58]. At the taxonomic level of phylum, the RA of Bacteroidetes was higher in MSA, while the RA of Firmicutes was lower in MSA (Fig. 3A, Supplementary Table 5). Additionally, the ratio of Firmicutes-to-Bacteroidetes was significantly different between MSA and HC (MWU; P = 0.03) (Fig. 3B). The RA of putative pro-inflammatory bacteria from the families Clostridiaceae (p_Firmicutes;c_Clostridia) and Rikenellaceae (p_Bacteroidetes;c_Bacteroidetes) were higher in MSA than in HC (Fig. 4A,Supplementary Table 5).
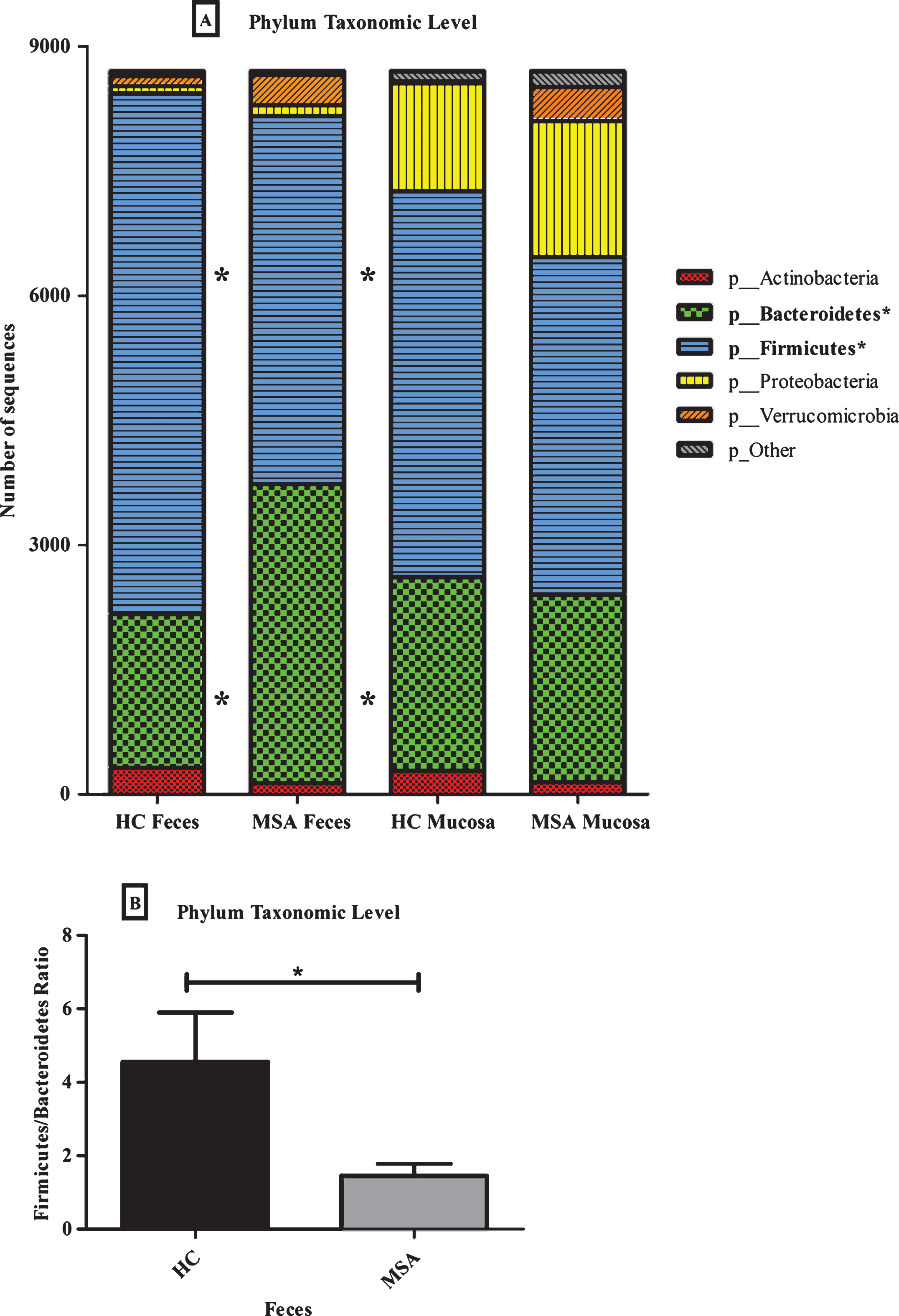
Different relative abundant (RA) phylum microbial taxa in the fecal and mucosal microbiomes of healthy control and MSA subjects. (A) The RA of Bacteroidetes-to-Firmicutes are different between MSA and HC feces. Bacteroidetes are increased, while Firmicutes are decreased in MSA feces. The average number of sequences was rarefied to 8,700 sequences per sample. (*) denotes differences between HC and MSA fecal samples (Kruskal-Wallis, P < 0.05). (B) Firmicutes-to-Bacteroidetes ratio is significantly different (MWU; P = 0.03) between HC and MSA fecal samples.
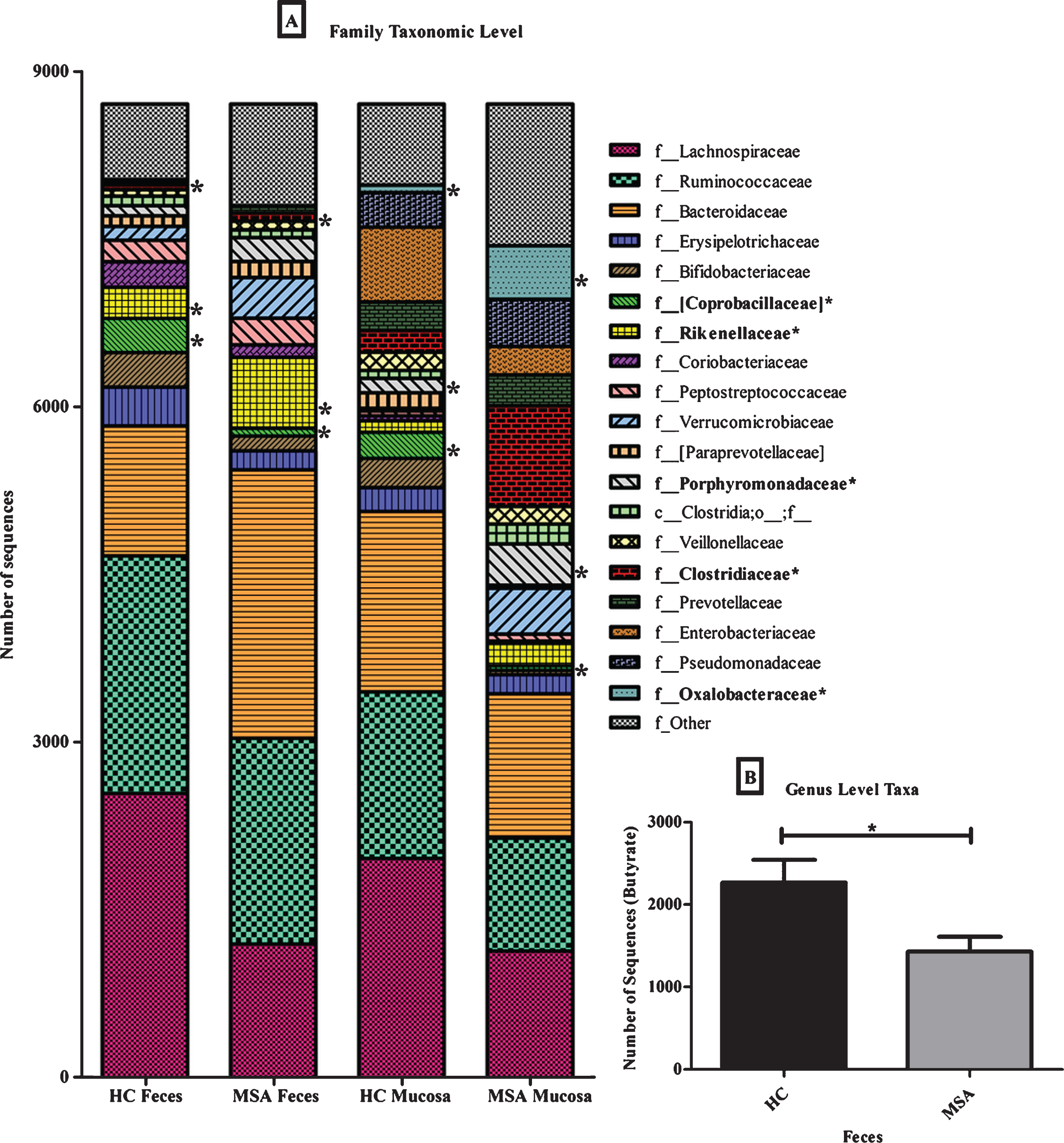
Different relative abundant (RA) family microbial taxa in the fecal and mucosal microbiomes of healthy control and MSA subjects. (A) The RA of different family level taxa are indicated between MSA and HC feces and mucosa. The average number of sequences was rarefied to 8,700 sequences per sample. (*) denotes differences between HC and MSA fecal samples (Kruskal-Wallis, P < 0.05), and differences between HC and MSA mucosa samples. (B) The RA of genus level SCFA butyrate-producing taxa is significantly different (MWU; P = 0.04) between HC and MSA fecal samples.
The RA of putative anti-inflammatory bacteria from the families Lachnospiraceae and Ruminococcaceae (p_Firmicutes;c_Clostridia) were lower in MSA than in HC (Fig. 4A, Supplementary Table 5). The RA of bacteria from the acetogenic and human intestinal microbiota phylogenetic core genera Blautia, Dorea, (p_Firmicutes;c_Clostridia;f_Lachnospiraceae) was lower in MSAthan HC. In addition to the non-parametric Kruskal-Wallis test, a parametric statistical test (negative binomial Wald test) was used to analyze the bacterial RA data. Using this test, three taxa were identified with significant adjusted P-values (adjusted P < 0.014; f_Coprobacillaceae) or nearly significant adjusted P-values (adjusted P < 0.054; genera Blautia and Dorea) (Supplementary Table 5). Using the non-parametric Kruskal-Wallis test, the RA of bacteria from the genera Blautia and Dorea was not shown to be significantly different between MSA and HC subjects (FDR P-value = 0.54–0.56; Supplementary Table 5). Additionally, the RA of sequences from putative short-chain-fatty-acid (SCFA) butyrate-producing anti-inflammatory genera [Ruminococcus], Roseburia, Coprococcus (p_Firmicutes;c_Clostridia;f_Lachnospiraceae), andFaecalibacterium (p_Firmicutes;c_Clostridia;f_Ruminococcaceae) were significantly lower (MWU; P = 0.04) in MSA compared to HC (Fig. 4B). Finally, all fecal taxon-specific differences were observed within the context of overall microbial community analyses that revealed no significant differences between MSA and HC individuals, at the taxonomic level of family, using analysis of similarity (ANOSIM) [45] (Global R = 0.04, P = 0.35) (Supplementary Table 6).
Overall, these findings indicate that the fecal microbiota of MSA subjects had higher alpha diversity, a different F/B ratio, higher RA of putative pro-inflammatory bacteria, and lower RA of putative SCFA-producing bacteria, especially putative SCFA butyrate-producing bacteria, when compared to HC subjects.
Sigmoid mucosa microbiota
Analysis of mucosal-associated microbiota communities revealed that MSA subjects had higher alpha diversity and a higher RA of putative pro-inflammatory bacteria. Alpha-diversity [57] indices indicated significant differences in Shannon index (MWU; P = 0.01), evenness (MWU; P = 0.01), and Simpson’s index (MWU; P = 0.01) between MSA and HC at the taxonomic level of family (Supplementary Table 4). The ratio of Firmicutes-to-Bacteroidetes was not significantly different between MSA and HC (MWU; P = 0.69). The RA of three microbial taxa trended different between MSA and HC (Supplementary Table 5) [58]. Specifically, the RA of putative pro-inflammatory bacteria from the family Oxalobacteraceae (p_Proteobacteria;c_Betaproteobacteria), Porphyromonadaceae (p_Bacteroidetes;c_Bacteroidia) and genus Ralstonia (p_Proteobacteria;c_Betaproteobacteria;f_Oxalobacteraceae) was higher in MSA than in HC (Fig. 4A, Supplementary Table 5). Using the negative binomial Wald test, a single taxon was identified with a nearly significant adjusted P-value (adjusted P < 0.051; f_Porphyromonadaceae) (Supplementary Table 5). Finally, all mucosal taxon-specific differences were observed within the context of overall microbial community analyses that revealed no significant differences between MSA and HC, at the taxonomic level of family, using ANOSIM (Global R = 0.11, P = 0.14) (Supplementary Table 6) [45].
These results show that the colonic sigmoid mucosa of MSA subjects had higher alpha diversity, no difference in the F/B ratio, and a higher RA of putative pro-inflammatory bacteria, when compared to HC subjects.
Predictive functional (PICRUSt) analysis of fecal and mucosal microbiota
In addition to the microbiota composition analysis, we attempted to predict metagenome functional gene content from 16S rRNA gene amplicon sequences, between MSA and HC subjects using the software package PICRUSt. Although the sample size is small, and the results should be taken in this context, the RA of a majority of reference gene pathways were different in fecal samples collected from MSA when compared to HC, especially metabolic and biosynthetic pathways (Supplementary Table 7). In contrast, the RA of LPS-biosynthesis, LPS-protein, and ubiquinone biosynthesis pathways were higher in the fecal microbiota of MSA compared to those of HC (Supplementary Table 7). Thirty-three genes were annotated in the LPS-biosynthesis pathway; the RA of 12 KOs trended different between MSA and HC (Supplementary Figure 1 and Table 8).
Additionally, the RA of LPS-biosynthesis pathway trended different in the mucosal associated microbiota of MSA compared to those of HC(Supplementary Table 7). Thirty-three genes were annotated in the LPS-biosynthesis pathway; the RA of nine KOs trended different between MSA and HC (Supplementary Figure 1 and Table 8). Based on a predictive analysis of microbiome functional genes (PICRUSt), the RA of LPS-biosynthesis pathways were enriched in both MSA feces and mucosa. Further studies using shotgun metagenomics analysis will be necessary to confirm these preliminaryfindings.
DISCUSSION
We hypothesized that similar to PD subjects, MSA subjects suffer from gut dysfunction such as alterations in the intestinal microbiota composition and/or disruption of intestinal barrier function. In our proof-of-concept study, we found that MSA subjects have evidence of TJ barrier dysfunction, increased number of TLR4 expressing cells in the lamina propria in sigmoid mucosa, and trends suggesting a putative “pro-inflammatory” microbiota. Our findings support our hypothesis suggesting that subjects with MSA have endotoxin mediated gut-derived inflammation. Our data warrants future longitudinal and/or interventional studies for an exact role of gut-derived inflammation in MSApathogenesis.
We are the first to investigate a TJ marker in the colonic sigmoid mucosa of MSA subjects. We found significantly disrupted levels of the TJ protein ZO-1 in MSA sigmoid mucosa compared to HC subjects. ZO-1 is a TJ protein that binds to other TJ proteins occludin and claudin-1 [59, 60]. Defects in ZO-1 expression could lead to abnormal permeability as shown in various GI disorders by increasing paracellular permeability [61–64] or by modulating functions of occluding [63] and claudins [64]. Intestinal TJ proteins play a vital role in maintaining the gut barrier and the loss of intestinal barrier integrity that could increase the invasion of pro-inflammatory products (i.e., LPS-producing-bacteria). Due to abnormal TJ barrier disruption in MSA subjects, gram-negative LPS-producing bacteria (or LPS) could translocate from the GI tract and infiltrate into peripheral tissue as well as organs to induce local and systemic inflammatory responses [65–68]. This chronic inflammation, by LPS exposure, could initiate MSA pathology in the enteric nervous system (ENS) and central nervous system (CNS) simultaneously or consecutively.
Previous reports show that LPS induce cascades of inflammatory events via TLR4 activation leading to inflammatory diseases [69–75], loss of dopaminergic neurons in vitro [76], and TLR4-stimulated neurodegeneration within in vivo models [13, 75]. Toll-like-receptor-4 mRNA is elevated in the brain of MSA subjects and is believed to act as a mediator for the induction of α-SYN induced microglial phagocytosis, pro-inflammatory cytokine release, pro-inflammatory astroglial responses [13, 77], and activation of multiple TLRs which may lead to neurodegeneration [13, 79]. We found a significant increase in TLR4 expressing cells in the lamina propria within the colonic sigmoid mucosa of MSA subjects. Increased mucosal TLR4 suggests increased exposure of LPS to intestinal epithelial and mucosal immune cells. Toll-like-receptor-4 is expressed by microglia, macrophages, astrocytes, oligodendrocytes, and neurons in response to innate inflammation [16, 81]. In our study, we found few enteric glia cells positive for TLR4 in the colonic sigmoid mucosa, and future studies focusing on colonic TLR4 are warranted. Considering gram-negative LPS-producing bacteria/bacterial products are major ligands for TLR4 activation [69–75, 82], TLR4 activation and the subsequent release of pro-inflammatory cytokines could cause neurotoxicity related to MSA pathology [83].
To date, no study has compared the colonic microbial profiles between MSA and HC subjects. We found that the RA of putative SCFA-producing bacteria, such as those from the Clostridial cluster XIV genera Blautia and Dorea, were lower in MSA feces compared to HC feces. Additionally, the significant overall lower RA of identified SCFA-butyrate-producing-bacteria could suggest loss of beneficial effects of SCFA on the intestinal barrier and immune function in the MSA subject colon, as shown in other diseases/models [49]. Predominately, SCFAs are generated in the colon and impact various functions of the colon, but can also affect other organs including the brain via the circulatory system [84]. This occurs via multiple mechanisms including epigenetic modifications [85] or activation of SCFA receptors that are anti-inflammatory, anti-microbial, and reduce intestinal leakiness [86, 87]. Further studies using metabolomic, metagenomic, and metatranscriptomic techniques will be required to better understand the role of SCFAs in intestinal inflammation and barrier dysfunction in MSA.
Both PD and MSA are classified as synucleinopathies, and overall, the microbiota profiles of these two syncleinopathies showed increased putative “pro-inflammatory” and decreased putative “anti-inflammatory” bacteria compared to HCsubjects [10]. There was an increase in the RA of the LPS-biosynthesis pathway in both MSA feces and mucosa samples, whereas the RA was only increased in PD subject feces compared to HC. The common bacteria (family and genus levels) that were significantly affected in both PD and MSA were: increased Oxalobacteraceae and Ralstonia in mucosa, increased Clostridiaceae in feces, and decreased Coprobacillaceae in feces and mucosa. In contrast, the RA of Rikenellaceae and Porphyromonadaceae, both Gram-negative LPS-producing bacteria, were higher in MSA feces or mucosa, while not significantly different in PD feces or mucosa. These individual bacterial taxa produce different strains of LPS [88] possibly related to specific immune activation pathways that might promote disease specific neuroinflammatory processes. Additionally, the feces of MSA subjects did not show a difference in RA of type III bacterial secretion systems, via PICRUSt analysis, whereas PD subject feces showed a significant increase in the abundance of type III bacterial secretion systems compared to HC feces. Differences in the RA of LPS-producing bacterial taxa between these two syncleinopathies need further investigation. The difference in LPS strains could possibly explain the specific pathogenesis relating to their well characterized neuronal (PD) vs glial (MSA) [21, 89–91] α-SYN inclusions associated inflammatory reactions. Lipopolysaccharides from the intestine might generate α-SYN aggregation [92] in the enteric nervous system (ENS) alone, which then could transfer to the brain in a prion fashion [93] or affect ENS and CNSsimultaneously [94].
There are several limitations with our study. One limitation is small sample size. MSA is a rare neurodegenerative disease with rapidly progressive course leading to significant frailty and eventually death six-to-nine years from first symptoms [4–6]. These factors restrict recruitment of large numbers of subjects in most MSA-related clinical studies that require sample collection. Another limitation is the lack of consistent dietary intake data. Two different food frequency questionnaires were administered to our HC and MSA subjects, preventing us from detailed comparisons between the eleven HC subjects to the six MSA subjects. Nonetheless, there was no significant difference in total fat or carbohydrate intake between the two groups. Our MSA subjects showed signs of constipation while none of the HC reported constipation. Therefore, the microbial profiles in MSA subjects could be impacted by their constipation condition. However, published microbiome studies in constipated subjects [95–97] reported different patterns of microbial profiles, than the significant difference observed in our reported data with MSA and HC subjects. This evidence suggests constipation might not be a driving factor in the reported microbial changes. Lastly, our MSA microbiota data are focused on colonic mucosa and fecal microbiota. MSA is associated with numerous GI abnormalities including intestinal dysmotility and impaired circadian rhythm of gastric myoelectrical activity [27, 98–100], yet no studies have analyzed the microbiota in the small intestine in MSA. Future studies characterizing microbiota in other parts of the GI tract are needed to determine the influence of the upper GI tract on microbiota in MSA. We also acknowledge the bi-directional communication between gut-brain and brain-gut. It is possible that earlier CNS dysfunction could impact ENS function and reported changes could be associated with CNS dysfunction occurring in MSA subjects. Future pre-clinical studies are warranted to further explore the MSA pathogenesis focusing on gut-brain and brain-gut axis.
In summary, our proof-of-concept study provides preliminary data that like PD, MSA subjects display evidence of disrupted intestinal barrier integrity, endotoxin mediated gut-derived inflammation, and putative pro-inflammatory microbiota. Further studies are needed to determine a potential causal link between the observed intestinal abnormalities and neuroinflammation in MSA. Future gut-brain-axis studies are warranted to elucidate the mechanism which could lead to the identification of novel therapeutic targets for this progressive and untreated disease.
FUNDING AGENCIES
This study was supported by research grants from Mr. and Mrs. Larry Field, Mr. Silas Keehn,Mr. Richard M. Burridge, Carole and Gordon Segal, Philip and Ellen Glass, Parkinson Disease Foundation, and Michael J. Fox Foundation.
CONFLICTS OF INTEREST/FINANCIAL DISCLOSURE
The authors have no conflict to report.
AUTHOR ROLES
PAE co-wrote the first draft of the manuscript, extracted DNA, assisted in data analysis, data interpretation, prepared figures for final publication, drafted manuscript for final publication, and reviewed and critiqued the manuscript and data analysis; HBD co-wrote the first draft of the manuscript, performed immunohistochemistry and microscopic analyses, assisted in data analysis, data interpretation, drafted manuscript, prepared final figures for publication, and reviewed and critiqued the manuscript and data analysis; AN co-wrote the first draft of manuscript, directed and performed the microbiota bioinformatics and analysis, reviewed and critiqued the manuscript; CBF supervised DNA extraction and all laboratory based measures and assisted in data interpretation, reviewed and critiqued the manuscript; SJG assisted in performing the microbiota bioinformatics and analysis, assisted in data interpretation, and reviewed and critiqued the manuscript; RMV assisted in data analysis, data interpretation, and reviewed and critiqued the manuscript; JHK supervised histology staining and all laboratory based measures and assisted in data interpretation, reviewed and critiqued the manuscript; EM helped design the microbiota study, collected the clinical samples for HC subjects, and assisted in data interpretation; KMS designed the clinical study, recruited and assessed MSA subjects, wrote the IRB and reviewed and critiqued the manuscript; AK developed the clinical and microbiota related hypothesis, designed the study, collected samples from MSA and HC subjects, assisted in data interpretation, co-wrote the first draft of the manuscript and reviewed and critiqued the manuscript. All authors were responsible for the study design, data analysis, data interpretation, writing of the report, and the decision to submit for publication. All authors had complete access to the database and vouch for the completeness and accuracy of thedata.
FULL FINANCIAL DISCLOSURE
Nothing to report.
Footnotes
ACKNOWLEDGMENTS
We are grateful to Ms. Rawan Abbasi, for her assistance with subject enrollment and sample collection.
