Abstract
Neuromuscular disorders (NMDs) are a large group of diseases associated with either alterations of skeletal muscle fibers, motor neurons or neuromuscular junctions. Most of these diseases is characterized with muscle weakness or wasting and greatly alter the life of patients. Animal models do not always recapitulate the phenotype of patients. The development of innovative and representative human preclinical models is thus strongly needed for modeling the wide diversity of NMDs, characterization of disease-associated variants, investigation of novel genes function, or the development of therapies. Over the last decade, the use of patient’s derived induced pluripotent stem cells (hiPSC) has resulted in tremendous progress in biomedical research, including for NMDs. Skeletal muscle is a complex tissue with multinucleated muscle fibers supported by a dense extracellular matrix and multiple cell types including motor neurons required for the contractile activity. Major challenges need now to be tackled by the scientific community to increase maturation of muscle fibers in vitro, in particular for modeling adult-onset diseases affecting this tissue (neuromuscular disorders, cachexia, sarcopenia) and the evaluation of therapeutic strategies. In the near future, rapidly evolving bioengineering approaches applied to hiPSC will undoubtedly become highly instrumental for investigating muscle pathophysiology and the development of therapeutic strategies.
Keywords
INTRODUCTION
The 2012 Nobel Prize in Medicine awarded to Prs Gurdon and Yamanaka highlighted 60 years of research on pluripotency and pluripotency maintenance together with method for the derivation of induced pluripotent stem cells (iPSC) from any human being [1, 2]. Over the last 15 years, this major through has transformed biomedical research and opened the gate to the elaboration of novel in vitro models of human diseases. The very first consequence of the development of the iPSC technology was to bring pluripotent stem cells to many laboratories worldwide together with the development of increasingly innovative differentiation procedures covering the wide repertoire of human cell types. Most importantly, because they can be derived directly from patients, human induced pluripotent stem cells (hiPSC) offer promising opportunities for modeling and investigating human genetic diseases, which many hoped could prove to be more relevant than animal models that don’t necessarily recapitulate human physiology or diseases. The development of hiPSC-based model also opens a broad range of novel opportunities for enforcement of the 3 R rules to replace or refine the use of animal models. By easing access to relevant human tissues not otherwise accessible, hiPSC slowly became a powerful source of biological material for enhancing our understanding of physiological and pathological processes, disease modeling, drug discovery and optimization of cell-based therapies (Fig. 1A).
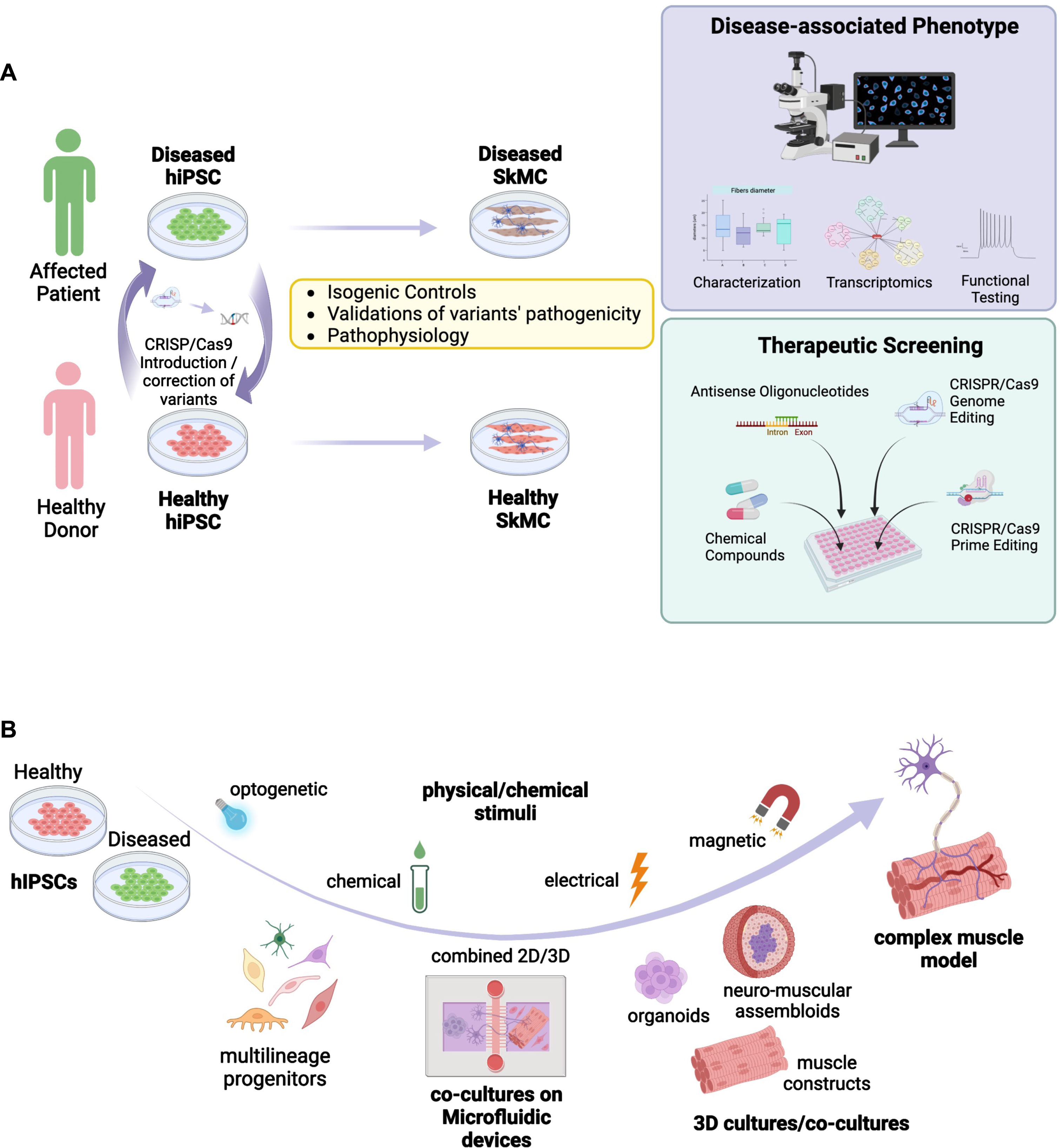
Skeletal muscle is a complex tissue composed of different cell types that all contribute to its function. Mouse models and murine cell lines such as C2C12 cells do not accurately reflect all aspects of human muscle development and biology. Human myoblasts can be obtained by isolation of muscle stem cells from human biopsies [3, 4]. These cells can be readily expanded but may lose their capacity over time with a reduced proliferation potential and cell senescence. Cell immortalization using SV40 T antigen, CDK4 and hTERT overexpression can circumvent these limitations [5, 6]. Alternatively, myoblasts can be obtained from primary fibroblasts after overexpression of PAX7 or MYOD1 [7, 8]. However, these 2-dimensional (2D) primary cell culture systems also have major limitations such as a limited proliferation rate and differentiation capacity, a lack of functionality, the absence of the other cell types present in the muscle such as stromal cells, motor neurons, vascular cells or of the extracellular matrix (ECM) components. They also lack mechanical constraints and 3D organization that characterizes the muscle tissue in vivo. Two-dimensions muscle cells cultures are also poorly amenable for functional assessment such as electro-mechanical coupling, measurement of strength and contraction or calcium handling. Altogether, this implies the need for the development of more elaborate tissue culture models with settings able to mimic more efficiently the in vivo complexity (Fig. 1B). A number of reports describe settings or assemblies of cells of different origin or even species to evaluate myo-innervation, myo-inflammation or myo-vascularization by co-culture between muscle cells together with other cells types. The challenge resides now in building more complex “bio-constructs” with appropriate cell types ratio and optimal ECM for improving cell survival, differentiation and maturation, a challenge that remains to date only partially addressed by available hiPSC-based models. Tackling these various challenges opens up a wide range of opportunities for the development of the next-generation models able to recapitulate physiological and pathological muscle development but also to respond to unmet medical needs with the increasing identification of new disease-causing genes or variants and the emergence of novel non-pharmacological therapies.
FROM DEVELOPMENTAL CUES TO THE MODELING OF MYOGENESIS IN VITRO
Compared to other lineages, hiPSC-based skeletal muscle differentiation is still lagging behind. As a consequence, modeling the large repertoire of NMDs [9] has been hampered by the absence of efficient protocols able to generate fully mature skeletal Muscle Cells (SkMC) in vitro. As for other cell lineages, differentiation of human embryonic stem cells (hESC) or hiPSC toward the skeletal muscle lineage relies on our knowledge of the differentiation of the mesodermal lineage that gives rise to the somites from the anterior to the caudal part of the embryo. During development, the dermomyotome that expresses the PAX3 and PAX7 transcription factors in the dorsal part of the embryo will form skeletal muscles, the dermis, endothelial cells and vascular smooth muscle cells [10, 11]. Under the influence of the neural tube and notochord, the dorsomedial lip of the dermomyotome progressively acquires features specific to the skeletal muscle lineage with expression of the MYOD and MYF5 transcription factors in myoblasts, able to migrate from the myotome and fuse to form embryonic muscle fibers [12, 13]. Based on this sequential activation of myogenic factors, initial protocols for differentiation of hiPSC toward the skeletal muscle lineage made use of induced expression of master myogenic transcription factors such as PAX3 [14] or MYF5 [15] but more efficiently, PAX7 [16–19] or MYOD [20–28] following viral transduction [16–18, 24–28], mRNA transfection [29] with eventually co-expression of epigenetic modifiers, BAF60 C [30] or JMJD3 [31]. MYOD induction was more largely employed for modeling muscle diseases, in particular Duchenne Muscular Dystrophy (DMD) and the evaluation of therapeutic approaches (Table 1). However, most of these protocols had the disadvantage of using viral induced expression of myogenic factors with the risk of random transgene insertion that can interfere with the host cells genome activity when considering regenerative medicine approaches.
Example of hiPSC- or hESC-based models of Neuromuscular Disorders
DMD: Duchenne Muscular Dystrophy, DM1: Myotonic Dystrophy; FSHD: FacioScapuloHumeral Dystrophy; LGMD: Limb Girdle Muscular Dystrophy; SMA: Spinal Muscular Atrophy.
A second generation of approaches consisted in the stepwise addition of small molecules to induce mesodermal differentiation, with activation of the Wnt, Sonic hedgehog (Shh) or Notch pathways for the conversion of PAX3 + /PAX7 + progenitors into MYOD+/MYF5 + committed myoblasts [32–34] together with inhibition Bone Morphogenic Proteins (BMPs) that maintains PAX3/PAX7 expression in myogenic progenitors [35–37]. Wnt activation mediated by addition of Wnt3a or more commonly CHIR99021, a GSK3β inhibitors, was efficient in driving muscle differentiation but required Pax3 + /Pax7 + cell sorting, limiting large scale explorations [38, 39]. Alternatively, activation of the Wnt pathway using CHIR99021 together with inhibition of the lateral mesoderm using the LDN193189 BMP inhibitor [40] or followed by inhibition of the Notch signaling using the DAPTγ secretase inhibitor [41] efficiently induced formation of multinucleated and contractile muscle fibers with sarcomeric organization in 50 days and 30 days, respectively. Another GSK3β inhibitor, Bio, was also used in combination with Forskolin, an Adenylyl Cyclase activator that activates cAMP and CREB signaling, to stimulate myoblast proliferation in vitro and led to the production of multinucleated myotubes with Myosin Heavy Chain proteins in 36 days [42]. Treatment of hiPS cell aggregates with a high concentration of bFGF and EGF converted stem cells into 40–50% of myogenic cells expressing MYOD, PAX7 and Myogenin in 6 weeks [43]. Pre-induction of hiPSC in the presence of DMSO followed by the sequential addition of small molecules including CHIR99021, BMP4 and Lys294002 (a PI3K inhibitor) yielded the production of myocytes in 12 days, followed by muscle fusion after 34 days and spontaneous contraction upon addition of 15% of fetal bovine serum [44]. TGFβ inhibitors (SB431542 or A83-01) were also shown as capable of enhancing fusion and maturation of ERBB3 + /NGFR+hiPSC-derived muscle progenitors expressing PAX7 and Myf5 [41, 45]. Alternatively, hiPSC plated on Collagen I in the presence of 5% horse serum and treated with CHIR99021, Ascorbic acid, Alk5 inhibitor, dexamethasone, EGF and insulin, yielded up to 80% of PAX3-positive cells in 10 days and 50–60% of MyoD-positive cells after addition of SB431542 (to inhibit Alk4, 5 and 7), PDGF, EGF, HGF, bFGF, Oncostatin and IGF1 [46] with fusion and terminal differentiation later improved by addition of myokines and anabolic factors [47] (Fig. 2).
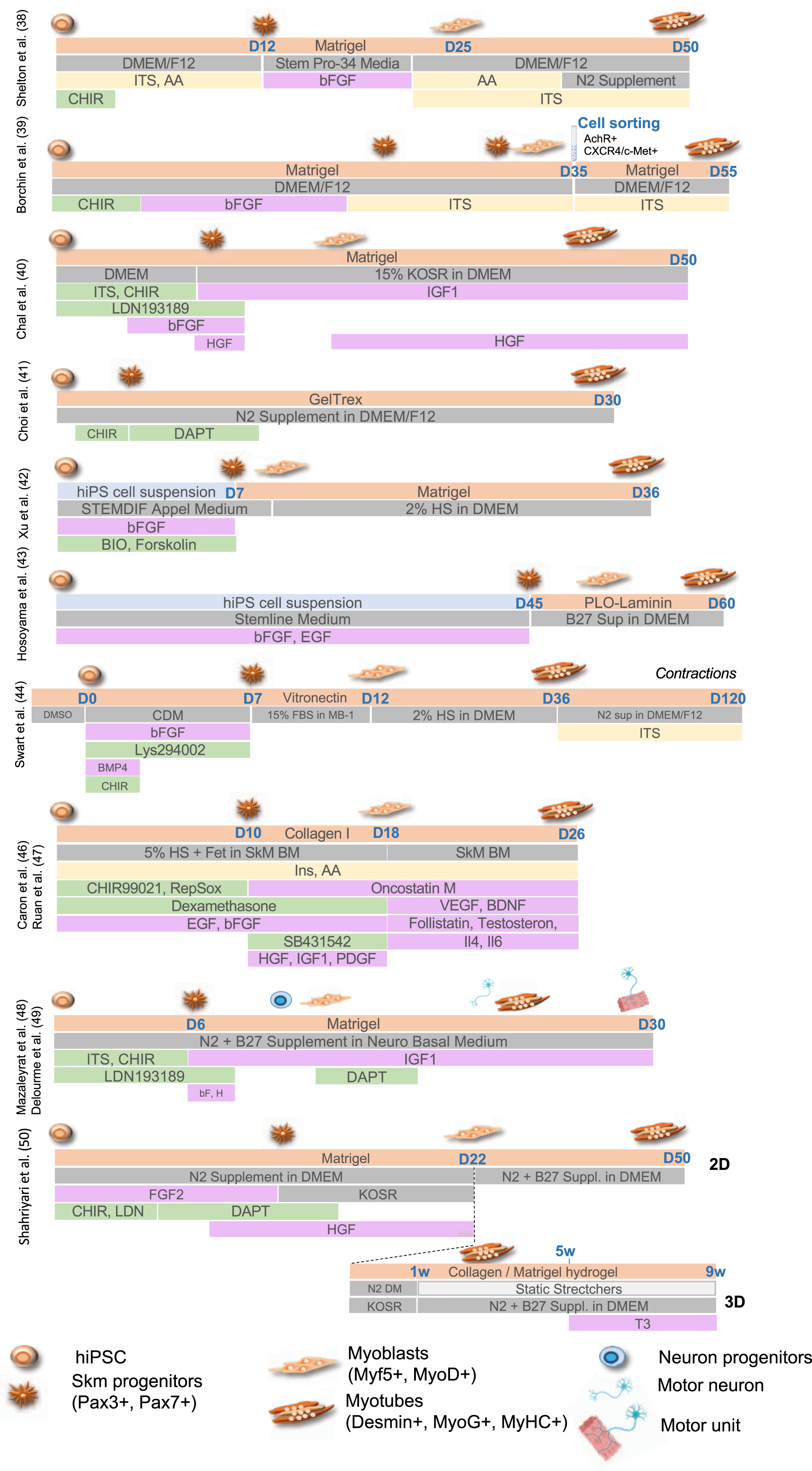
Chal et al. [40] and Hosoyama et al. [43] reported the presence of neural cells differentiating along with muscle cells suggesting an intermediate population of neuro-mesodermal progenitors during the differentiation process. Taking advantage of the transient presence of these two populations of cells, Mazaleyrat et al. [48, 49] proposed a protocol combining activation of the Wnt Pathway (addition of the CHIR99021 GSK3β inhibitor) to induce mesoderm commitment followed by BMP inhibition (LDN193189) for production of muscle and motor neurons progenitors in 8 days. The subsequent addition of DAPT induces the production of motor neurons and formation of contractile millimeter-long multinucleated muscle fibers with sarcolemmal organization in 20 days [48, 49]. Another transgene-free and serum-free protocol and a combination of small molecules to achieve 2D and 3D differentiation was more recently reported [50]. As described by others [40, 48], gene expression profiling at different stages showed that muscle cells differentiation follows developmental trajectories with paraxial mesoderm induction (days 1–4), myogenic specification (days 4–13) and myogenic expansion (days 13–22). At day 22, cells can be plated on Matrigel coated plates for 2D differentiation or into a hydrogel containing Collagen I and Matrigel to achieve 3D differentiation with secondary myogenesis occurring between days 22 to 60. In this setting, only expression of embryonic Myosins (MyH3) or at a lower level, fetal Myosin (MyH8) are detectable but addition of Tricodo-L-Thyronine (T3) at the final maturation step (weeks 5–9 post-differentiation), increase contraction strength together with expression of fast mature MyHC isoforms, sarcomere organization, an important mitochondrial network and a dense ECM [50].
For several of these protocols, cultures can be maintained in the long term thanks to the production of a dense ECM and the presence of PAX7 + satellite cells [23, 51], an advantage in the perspective of regenerative therapy. Furthermore, treatment of myogenic cells [46] with recombinant DLL4 (Delta Like 4, Notch activation) and PDGF-BB (a PDGF receptor β ligand) limits terminal differentiation but favors myogenic progenitors motility and migration, including trans-endothelial migration, opening new perspectives for the systemic delivery of hiPSC-derived muscle progenitors for regenerative therapy [52].
The capacity of self-assembly between muscle and motor neurons was nicely illustrated in human 3D cortico-motor assembloids formed by combining hiPSC-derived cortical and motor neurons together with muscle cells embedded in an extracellular matrix to form 3D spheroids placed on top of a transwell insert. This configuration allows the formation of cortico-spinal connections and neuromuscular junctions (NMJ) between spinal neurons and muscle fibers that triggers muscle contraction up to 10 weeks post-assembly, mimicking the cortico-spinal-muscle circuit that controls muscle activity [53]. In addition, hiPSC are more and more largely used for the production of organoids or mini organs, defined as three-dimensional in vitro cell-derived structures able to self-organize to mimic a given organ. To date, only one report describes the production of neuromuscular organoids composed of spinal cord neurons and skeletal muscle cells that self-organize into 3D trunk neuromuscular organoids that can be maintained for months [54]. These organoids are characterized by the presence of neural and mesodermal cells and formation of NMJ containing pre-synaptic vesicles supported by the presence of S100β-positive Schwann cells, required for NMJ maturation [54].
In agreement with all previous reports, showing that co-culture of muscle cells with fibroblasts, fibro-adipogenic progenitors (FAPs) [55–57], macrophages [58–60] but more importantly motor neurons forming NMJ [61, 62], co-differention and self-organization of different hiPSC-derived cell types increase the differentiation rate and fiber maturation but also permits functional muscle assessments [48, 54] indicating that the presence of non-muscle cells is a key element in a successful in vitro myogenesis.
PLURIPOTENT STEM CELLS FOR MODELING OF NEUROMUSCULAR DISEASES
Neuromuscular diseases (NMDs) form a large and heterogeneous group of genetic diseases that cause progressive degeneration of striated muscles. Most NMDs result in chronic long-term disability imposing a significant burden for patients, families and public healthcare. These pathologies are present in all populations, affecting children as well as adults with a prevalence of 1 in a 1000 people [63, 64]. For many of these diseases, the causative gene is known but the impact of a given mutation on the molecular mechanisms of the disease often remains puzzling and requires further experimental validations for development of targeted and personalized therapies. For a number of hereditary NMDs, well-established animal models contributed to our understanding of the disease process or to the evaluation of therapies. However, many animal models imperfectly recapitulate pathophysiological features of a given disease with more than 95% of all animal-tested therapeutics, inefficiently transferred for the cure of patients.
Relative to the large number of genes associated to NMDs [9], only a few of them have been modelled using hiPSC, with the vast majority of reports focusing on Duchenne Muscular Dystrophy (DMD) for the development or evaluation of therapeutic approaches, including gene therapy (Table 1). In the context of genetic NMDs, hiPS-based models represent unprecedented opportunities to investigate the different aspects of a given neuromuscular disorders, identify new genes, validate the pathogenicity of variants or define early stages of muscle wasting. HiPSC-based models also supports the development of innovative treatments for muscle conditions and may open the door for preclinical screening of a personalized panel of drug candidates to improve these debilitating disorders, as exemplified for instance with the use of neuromuscular organoids to evaluate Myastenia gravis (MG)-associated autoantibodies against Acetylcholine receptor at the NMJ [54] (Fig. 1A).
Regarding the validation of gene variants, the latest advances in next-generation sequencing have uncovered a plethora of genomic variants of unknown significance (VUS) defined as rare or novel genetic variants for which the association with a disease has not been conclusively demonstrated [65]. One of the greatest advances of the twenty-first century is undoubtedly the clustered regularly interspaced short palindromic repeats (CRISPR)/Cas 9 genome editing technology that allows for a precise endogenous genomic correction [66, 67] but also the introduction of single point mutations, insertions, or deletions. The combination of stem cell-based approaches with CRISPR-mediated genomic editing turned hiPSC into powerful tools to study the impact of disease-associated mutations on specific cell types as well as to determine the correlation genotype-phenotype (Fig. 1A). Most importantly, using genome-editing strategies, the pathogenicity of a VUS can be demonstrated by comparing edited hiPSC with their unedited control lines. This approach will likely contribute to the validation of genetic mechanisms underlying NMDs, as previously done for instance in hiPSC-cardiomyocytes (CM) for cardiac diseases [68], whilst maintaining relevance to the human condition and making research findings readily translatable to the clinic.
PLURIPOTENT STEM CELLS FOR THERAPEUTIC DEVELOPMENT
High-throughput drug screening and in vitro preclinical testing
Developing a novel drug to treat a disease is a lengthy and costly process. This is particularly true for muscle disorders, often due to a lack of adequate cellular and animal models that accurately reflect the human muscle pathology. Directly identifying compounds that act on human muscle would facilitate and accelerate drug discovery. HiPSC that provide a plentiful source of patient-derived specialized cells, effective for drug identification and preclinical testing of future therapeutics and captured a growing interest from the pharmaceutical industry and slowly transformed the landscape of drug discovery [69]. As an example, hiPSC-derived neural stem cells from patients with familial dysautonomia, a rare genetic but fatal neurodegenerative disorder, were used a decade ago to screen nearly 7,000 small molecules and led to the identification of a drug hit [70].
However, the optimization of protocols for cell differentiation, maturation and maintenance still represents a significant challenge, in particular for hiPSC-SkMC due to the difficulties in generating and expanding hiPSC-myoblast cultures or obtaining mature differentiated myotubes. As a result, muscle drug discovery approaches remained to date limited to target-based screening in non-muscle cells or rodent myoblasts [71] with testing often focused on known drugs such as androgen receptor modulators (e.g. testosterone), myostatin inhibitors (e.g. Follistatin) or anti-inflammatory agents. The improvement of drug discovery for the broad range of muscle conditions and the development of new therapies for the millions of patients that are currently untreated strongly requires large amount of human skeletal muscle cells that demonstrate a fully mature phenotype.
HiPSC-derived Myoblasts/Myotubes and cardiac myocytes derived from DMD patients have been extensively used to model disease-specific phenotypes that can be reversed by pharmacological compounds (Table 1). DMD hiPSC-myotubes respond to hypertrophic proteins such as IGF-1 and Wnt7a [24], both tested as potential treatments for DMD, and can be pharmacological rescued by “dual-SMAD” inhibition [41] or Prednisolone [72], a corticosteroid and the current standard of care for DMD patients to delay the disease progression by reducing inflammation-induced muscle damage and muscle strength loss. HiPSC-based model demonstrated that Prednisolone has a direct beneficial impact on myofibers and not only on inflammation [72]. Similarly, stress-induced cell death and increased apoptosis can be pharmacologically modulated in DMD hiPSC-CM and improve DMD-associated dilated cardiomyopathy [73, 74]. With the goal of identifying new potential drugs for DMD treatment, a large screen of over 1500 compounds was later carried out on hiPSC-myoblasts carrying a nonsense mutation (c.457 C>T) which abolishes production of all Dystrophin isoforms [75]. Two compounds that restored fusion deficiency of these nonsense DMD hiPSC-myoblasts were identified and further validated in DMD hiPSC–myoblasts carrying different mutations (c.5533 G>C and Ex45del). Fenofibrate and ginsenoside Rd related to TGFβ signaling and FLT3 signaling, also showed efficacy in mdx 5cv mice by reducing fibrosis and improving muscle function. Patient-derived hiPSC-myotubes were also used as a drug screening platform for Dysferlinopathy with Nocodazole as a potential drug candidate for clinical applications aimed at increasing Dysferlin protein level [76].
Overall, by efficiently responding to compound testing, hiPSC-derived muscle cells open broad perspectives for drug screening or to evaluates potential therapies on a large-scale in human models of NMDs, maximizing their translational potential, whilst reducing the cost and time of novel therapeutic development.
Development of novel therapeutics
HiPSC-based technologies are also expected to provide platforms for preclinical testing of innovative therapeutics, including gene therapy approaches. Exon skipping by antisense oligonucleotides (ASOs) has gained a lot of interest and proven to be a powerful tool for mRNA splicing modulation to restore open reading frames. ASOs have now extensively been tested as therapeutic for various NMDs. Dystrophin expression was restored in approximately 30% of DMD hiPSC-CM treated with an ASO targeting exon 51 [77], or in DMD hiPSC-myotubes treated with ASO skipping exon 45 [26]. Similarly, an hiPSC-model of DM1 was used to screen ASOs and successfully identified a candidate that effectively abolished RNA foci and rescued mis-splicing in DM1 hiPSC-myotubes [18].
As mentioned earlier, (CRISPR)/Cas 9-genetically corrected patients hiPSC lines provide a powerful source of isogenic control lines for in vitro disease modeling studies. Most importantly, CRISPR/Cas9 genome editing can also be used as therapeutics to restore in vivo deletions or single point mutations. A large number of proof-of-concept studies have already demonstrated the therapeutic use of CRISPR genome editing for NMDs [78, 79], in particular in DMD (Table 1 and reviewed in [80–82]). We do not discuss them all here but will just highlight a few strategies. Myo-editing of the DMD gene mutations was for example elegantly demonstrated in hiPSC-CM corrected for exon 44 deletion, which disrupts the open reading frame of Dystrophin by causing splicing of exons 43 to 45 and introducing a premature termination codon. The reading frame can be restored by using Cas9 with guide RNAs that permits deletion of the splice acceptor or exons 43 and 45 donor sites, allowing splicing between exons 42 and 45 or 43 and 46 respectively [83]. CRISPR/Cas9-mediated excision of exon 51 in DMDΔ52-patient’s hiPSC-derived myoblasts and cardiomyocytes also restores Dystrophin expression and ameliorated skeletal myotube formation as well as cardiac function [84]. CRISPR/cas9 gene correction is not restricted to DMD. As another example, Telethonin (TCAP) expression could be efficiently restored in LGMDR7 (LGMD2 G) hiPSC-derived myoblasts by homology-mediated repair using Cas9 with a guide RNA targeting the microduplication [85]. HiPSC-myoblasts derived from FKRP dystroglycanopathies and Limb-Girdle Muscular Dystrophy (LGMD) R7 patients recapitulated their disease molecular phenotypes responsive to small molecules and gene editing therapeutics [86, 87].
However, CRISPR/Cas9 genome editing is known to create off targets that can be deleterious to the genome. An alternative CRISPR-based technology can now precisely edit individual nucleotides. With the prime editing approach, the therapeutic gene is knocked-in into a safe harbor locus allowing the correction of diseased genes independently of the underlying mutations. This new method was able to successfully restore Dystrophin expression in a DMD-hiPSC model [88] and adenine base editing (ABE) strategy, to restore Dystrophin expression in hiPSC-CM harboring a deletion of exon 51 (ΔEx51) of the Dystrophin gene in DMD patients [89].
Taken together, these different studies show that hiPSC-derived muscle cells represent efficient tools for evaluating the efficacy of drug compounds or CRISPR/Cas9- or exon-skipping-based approaches tailored to individual genetic variants or MD patients. However, except for some specific configurations [90], each of these approaches are mutations specific and therefore have to be re-engineered and adapted for each patient or each variant, a lengthy and costly process. Nonetheless, in the recent years, hiPSC-derived sensory neurons generated from patients with inherited [91] or non-genetic [92] pain disorders highly refractory to all available treatments showed that specific sodium-blocking compounds reduced the hyperexcitability of hiPSC-neurons in vitro while decreasing pain in the same patients. These studies represent proofs of principle of successful clinical translation from patient’s hiPSC in vitro models to individualized treatments. The development of similar assays for NMDs could initiate a new drug discovery paradigm moving us closer toward the era of personalized medicine as valuable tools to predict whether a given patients would respond to a particular drug.
TOWARD THE DEVELOPMENT OF TISSUE ENGINEERING FOR DISEASE MODELING AND REGENERATIVE MEDICINE
Three-dimension (3D) cultures reproducing an extracellular environment which better promotes physiological-like processes allows for the prolonged maintenance together with better level of maturation of the resulting myotubes [93, 94]. The use of hiPSC brings several advantages for the production of 3D models of human skeletal muscle tissue (Fig. 1B). In the last decade, some research groups started to work in this direction, although this field of research remains somewhat unexplored.
One of the first example of 3D engineered muscle tissue from hiPSC dates from 2013, when Tchao and colleagues made a comparative study between muscle derived stem cells and hiPSC-derived cardiac cells. In a collagen-based scaffold, 3D muscle constructs revealed several similarities compared to the development process of skeletal muscle and cardiac progenitors, providing a hybrid model for the study of cardiovascular diseases [95].
In 2018, Rao and colleagues developed a 3D culture model of hiPSC-derived skeletal muscle, starting from induced muscle progenitors embedded in a fibrin-based matrix [96]. With this approach, properly differentiated 3D skeletal muscle bundles were able to respond to electrical or acetylcholine stimulation [96]. The same year, Osaki et al. produced an organ-on-chip 3D model of amyotrophic lateral sclerosis (ALS) using hiPSC-derived motor neurons (MN) and muscle cells. Tedesco and his team were also pioneers in the development of complex skeletal muscle models using fibrin hydrogels loaded with different cellular components (myogenic, endothelial, pericytes and neural hiPSC-derived human progenitors) to generate 3D artificial muscles in normal or pathological contexts [97–99].
The role of the NMJ in muscle disorders was further investigated by exploiting 3D co-cultures of myoblasts-motor neurons and optogenetic technique [100–102]. A 3D culture of healthy muscle cells embedded in a Collagen/Matrigel matrix seeded in one chamber, with light sensitive Channel Rhodopsin-2 (ChR2)-induced ALS-derived MN spheroids in the adjacent and interconnected chamber was used to evaluate the impact of affected MN on muscle functionality and drug screening [101]. Electrophysiological recording and calcium release analyses of 2D vs 3D co-cultures of primary human myoblasts (in a fibrin/Geltrex hydrogel) and hiPSC-derived MN demonstrated a better maturation of NMJs in 3D condition, with a shift of expression from the embryonic to the adult form of nicotinic acetylcholine receptors [102]. A similar approach used a bioreactor for studying the response of 3D muscle constructs to the stimuli transmitted via the NMJ by ChR2-modified MNs in response to a blue light exposure to evaluate the effect of MG auto-antibodies on NMJ functionality [100].
The effect of magnetic fields on 3D muscle construct development and functionality was also investigated by magnetic force-based tissue engineering (Mag-TE). Magnetically labeled cells embedded in a collagen/Matrigel matrix are capable of self-assembly when exposed to the effect of a magnet. The contractile activity of hiPSC-derived “miniaturized skeletal muscle tissue” was increased compared to non-magnetically stimulated constructs in response to stimulating drugs [103].
As demonstrated by the multiple strategies used to obtain a reliable “platform” for studying NMDs, these different developments highlight the growing interest of the scientific community for 3D muscle modeling using hiPSC as the starting material but also the benefit of interdisciplinarity in tissue bioengineering (Fig. 1B). The different techniques cited here such as microfluidic devices, organoids, optogenetic, and magnetic stimulation, represent excellent examples of applying the newest bio-technological throughs to a specific biological application. However, 3D bioprinting, one of the more promising techniques for building a complex 3D artificial muscle tissue [104–106], still remains under exploited. Among the currently available techniques, the extrusion-based bioprinting [107] turned out to be the most suitable for skeletal muscle tissue engineering approaches, thanks to the possibility to produce continuous fibers of cell laden scaffold printed in parallel lines, layer by layer, perfectly reproducing skeletal muscle fibers architecture [108–110]. In the future, 3D bioprinting of hiPSC-derived muscle progenitors to obtain highly organized muscle tissue, combining known proportions of muscle and non-muscle cells will likely open new avenues in modeling NMDs and provide invaluable tools for various medical applications, including regenerative medicine.
CONCLUSION
Together with the undeniable advances provided by cell reprogramming, for the definition of cell lineages trajectories at early developmental stages, in muscle [111, 112], the development of patient-derived hiPSC has resulted in tremendous progress in disease modeling and the design of new therapies. Furthermore, over the recent years, in vivo reprogramming also emerged as a promising approach for tissue regeneration, including muscle [113, 114]. Importantly and interestingly, thanks to their capacity of differentiation toward different cell lineages, hiPSC also offer the advantage of allowing the investigation of the multisystemic aspects of diseases and to cover disease complexity as a whole, including in the course of finding a therapeutic strategy. However, as it is the case for all models, hiPSC-based disease modeling also has several limitations. Among them, one can cite the variability between clones, their respective genomic stability or intrinsic differentiation capacity, but also the need to obtain proper control in a number of cases. To this aim, the most widely used strategy consists at generating isogenic clones either to induce or correct a gene variant using a gene editing technique. However, obtention of truly isogenic clones faces multiple pitfalls as off target effects, copy number variations or point mutations induced by clone selection but also the difficulty in obtaining heterozygous mutations or to induce/correct splice variants. The often-limited efficiency of differentiation protocols in term of yield or maturation also relate to our capacity in recapitulating the in vivo environment largely absent in 2D cultures involving a single cell type. Nevertheless, in a continuously growing field, many of these limitations will be slowly compensated by the fast development of more and more sophisticated cell culture models together with high resolution approach for decoding differentiation steps in physiological and pathological contexts and correcting related defects. The establishment of standardized protocols will also help alleviate the variabilities observed between the different methods and the different batches within the same method.
FUNDING
L.C. received funding from the European Union’s Horizon 2020 research and innovation program under the Marie Sklodowska-Curie grant, agreement number 101025510. S.T. is the recipient of a post-doctoral fellowship awarded through the ICELARE program (ANR-20-ASTR-004). The work in FM’s laboratory is funded by AFM-Telethon through the MoThARD strategic plan.
CONFLICT OF INTEREST
The authors declare no conflict of interest.
