Abstract
Somatic instability of the huntingtin (
Keywords
Variation in inherited length of an expanded (>35) CAG repeat in the Huntington’s disease (HD) gene
Current data support a two-step theory of pathogenesis in HD where the rate of somatic CAG repeat expansion controls the timing/rate of disease onset [3]. This theory proposes that the inherited expanded
We quantified somatic variation in CAG repeat length in the OVT73 transgenic sheep model of HD in striatum and liver. These two tissues exhibit high instability both in HD knock-in and transgenic mouse models [25–27] and in patients [6, 9–11]. OVT73 are a premanifest model expressing an 11,625 bp transgene consisting of full length human
Transgene repeat instability was first assessed in six post-mortem OVT73 sheep aged 5-years (3 ewes, 3 rams; G1 and G2) (Supplementary Table 1). High resolution small pool-PCR (SP-PCR) was used to analyze CAG repeat length in single DNA molecules containing the OVT73 transgene, based on limiting dilution and Poisson analysis. All tissue samples reported in this manuscript were obtained from the South Australian Research and Development Institute (SARDI) and were sampled in accordance with approval of the Department of Primary Industries and Regions (PIRSA) Animal Ethics Committee (Approval number 19/02).The SP-PCR protocol was based on that previously described [8], with the use of sheep- (first round amplification) and transgene- (second round amplification) specific primers. The transgene-specific forward primer was 6-FAM labelled to enable resolution of the PCR product and quantification of repeat length using an automated ABI3130XL DNA sequencer with GeneScantrademark 600 LIZ® Size Standard (Applied Biosystems). Pure CAG repeat length was calculated from the tallest peak (modal repeat) of each trace compared against an OVT73 sheep standard with structure (CAG)69(CAACAG)2 (Supplementary Material). At least 50 single molecules were genotyped per sample.
For all samples, the modal repeat was determined to be 69 CAG units (43.1–71.1% of molecules per sample; Supplementary Dataset 1). This aligns with genotyping of bulk genomic DNA (1000 genome equivalents) from these animals and previous reports for the OVT73 line [28, 29]. A smaller peak was also detected at a lower frequency (6.6–20.8% of molecules per sample). Sanger sequencing confirmed this is a transgene copy with structure (CAG)26(CAACAG)2. This short transgene copy is also detected in genomic DNA from the OVT73 founder animal HD260 (Supplementary Figure 1). SP-PCR of fibroblast cells derived from founder HD260 (Supplementary Material) shows that both (CAG)26(CAACAG)2 and (CAG)69(CAACAG)2 transgenes are present in all cells (1:4 ratio). Previously reported capture sequencing estimates there are 10 full-length transgene copies at the single OVT73 locus [30]. It therefore appears that a fragment of a single copy of the transgene integrated, or that the construct DNA used in microinjection to create the founder OVT73 animal HD260 [28] inadvertently contained a short repeat clone in addition to the predominant cDNA which integrated along with the other copies at the OVT73 locus. The latter is most likely as other potential founder animals generated at the same time as HD260 also carry the short (CAG)26(CAACAG)2 transgene (Supplementary Figure 1). The absence of an mRNA corresponding to the (CAG)26(CAACAG)2 transgene in OVT73 indicates that it is not expressed.
As the (CAG)26(CAACAG)2 repeat is not expanded it was removed from the SP-PCR dataset for instability analysis. For the remaining products, a size distribution all alleles with >56 repeats, the modal repeat for each sample remained as 69 CAG units with modest variation (57 –80 CAG) observed (Fig. 1, Supplementary Dataset 1).
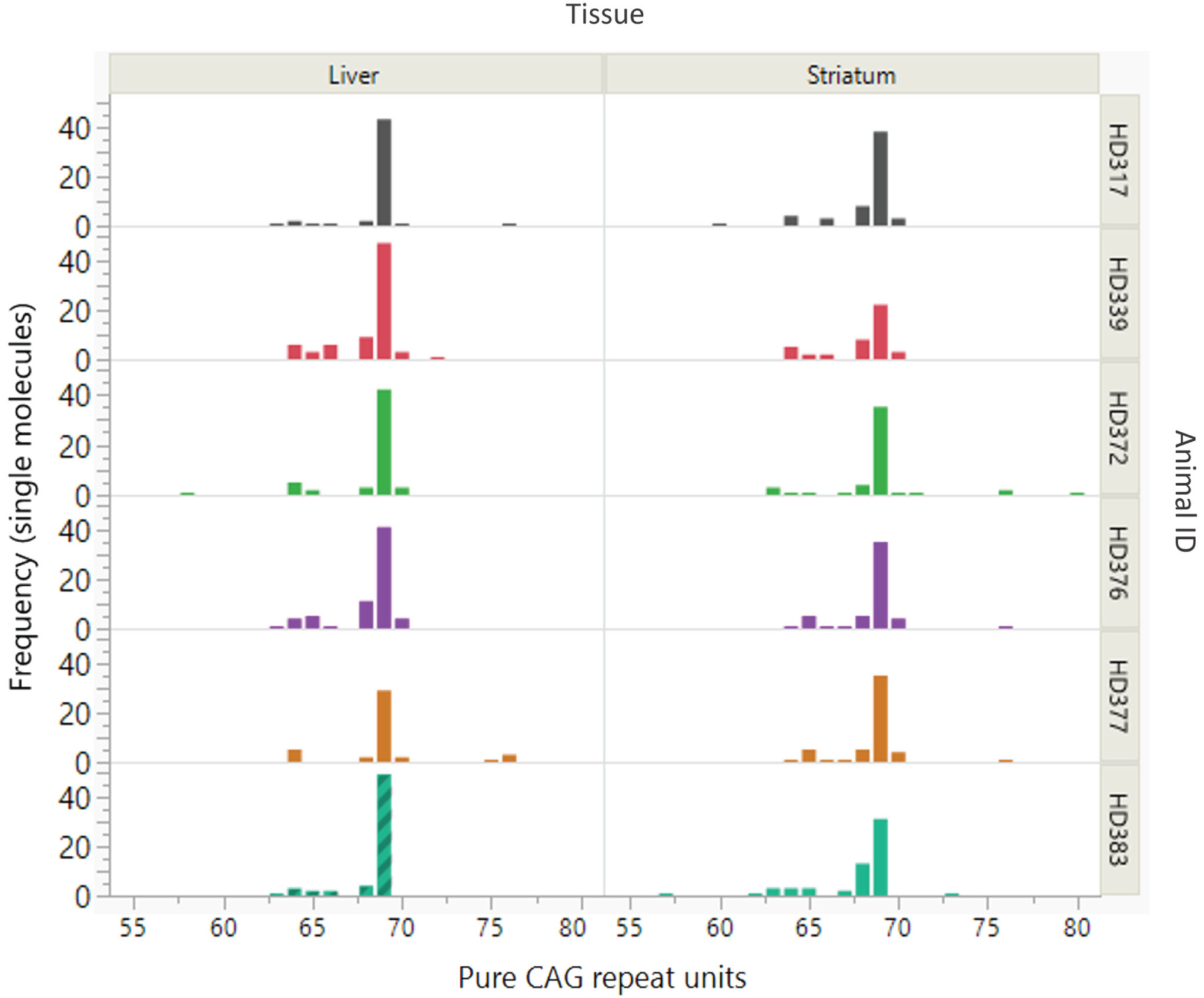
Somatic variation of the expanded CAG repeat of the OVT73 sheep transgene. Distribution and frequency of somatic variation in pure CAG repeat length of the OVT73 transgene is shown for liver and striatum tissues from six OVT73 sheep (5 years old) with constitutive polyglutamine-coding repeat structure (CAG)69(CAACAG)2. Repeat lengths were assessed in single molecules by small pool PCR. Modal repeat length for all animals was 69 CAG.
Interestingly, the SP-PCR data also revealed a cluster of alleles ∼65 CAGs as well as a few alleles between 75 and 80 CAGs. Further analyses, presented below, revealed that additional copies of the transgene at the multi-copy locus had repeat lengths in these ranges. Given this, it is likely that CAG-containing alleles around these sizes are products of other transgene copies, rather than somatic derivatives of the 69 CAG repeat-containing transgene. The presence of multiple transgene copies precludes accurate quantification of repeat instability, e.g., using Instability Index methods [8, 25]. This is because, for repeats that are closely spaced in length it is difficult to distinguish expansion peaks originating from shorter alleles from contraction peaks originating from longer alleles. Further, for transgenes with the same repeat length, it is impossible to discern the specific transgene(s) from which expansion or contraction peaks originate. The small sample number in this study also precluded meaningful assessment for effects of generation, age, or inherited CAG repeat length on instability, as has been observed in mouse models [6, 42]. Regardless, the data clearly show that the polyglutamine-coding repeat in the dominant OVT73 sheep transgene (CAG)69(CAACAG)2 is remarkably stable. Notably, a repeat of this length is expected to exhibit significant levels of somatic expansion after 5 years, given the instability present in a knock-in HD mouse model with 72 CAG repeats [6].
To examine whether older sheep might show evidence for somatic expansion, we performed bulk PCR and MiSeq analysis in three OVT73 sheep of advanced age (10-year-old, G3 ewes). These animals were recently utilized in a non-invasive cohort study [39] where OVT73 sheep at 9 and 10 years of age were found to have elevated mHTT levels in CSF and changes in brain white matter structure as assessed by MRI. The microstructural white matter changes correlated with declining gait and mHTT levels of CSF, indicating measurable disease progression over the one-year period. To assess potential brain somatic expansion more broadly bulk PCR was performed here on seven brain regions (brainstem, caudate, cerebellum, motor cortex, piriform cortex, putamen, temporal lobe) as well as liver (Supplementary Material). Interestingly, at least four discrete peaks were observed in tissues from all three animals, corresponding to pure CAG repeat length estimates of 27, 65–66, 69–70 and 82–83 units (respectively), and a peak with 76-CAG evident only in animal HD909 (Fig. 2, Supplementary Figures 2 and 3).
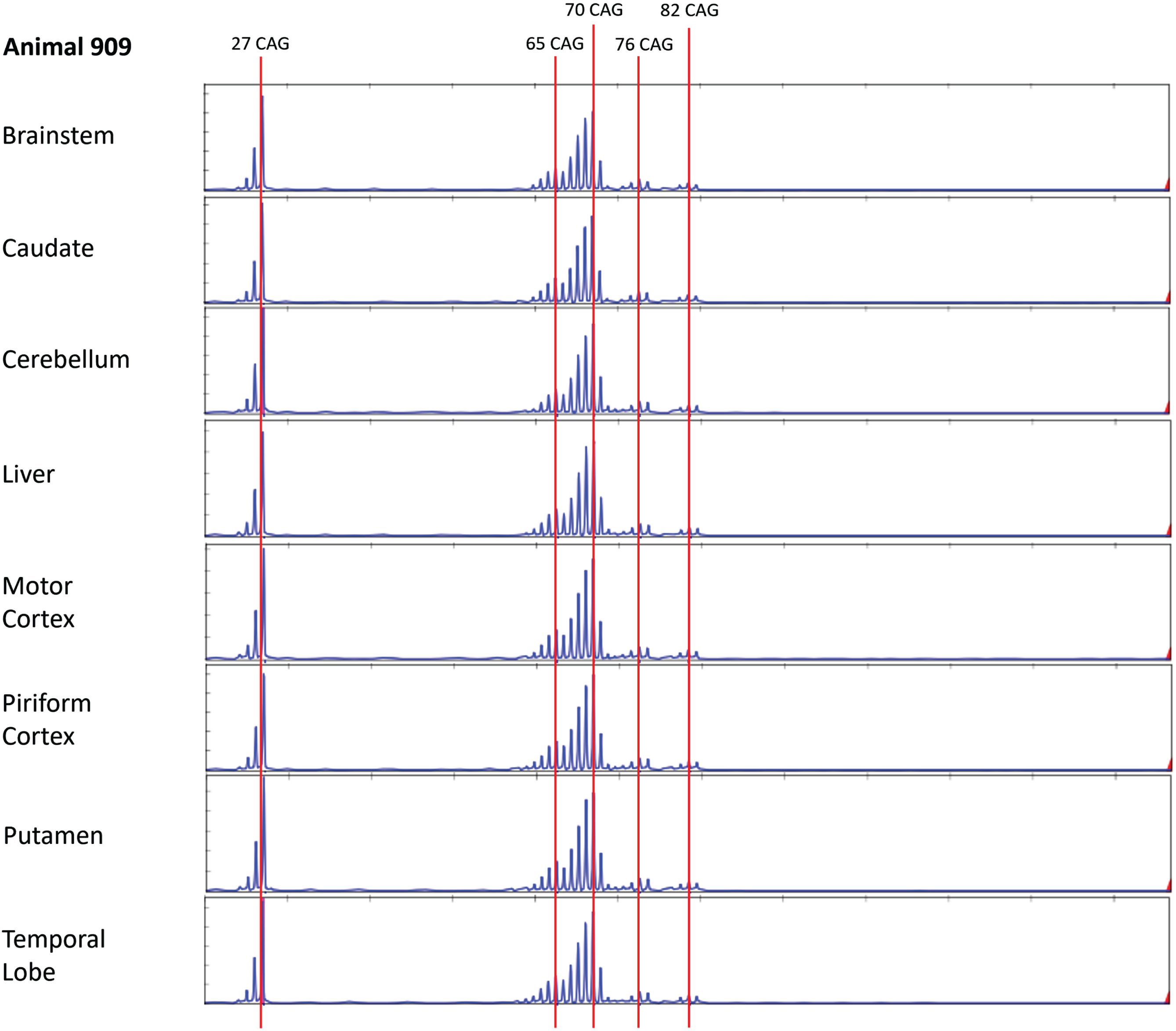
Alternative copies of the OVT73 transgene have different polyglutamine coding repeat lengths (Animal 909). Bulk PCR traces from striatal genomic DNA of three 10-year-old (G3) OVT73 sheep indicate presence of multiple copies of the OVT73 transgene with differing polyglutamine-coding repeat lengths, inserted at the ovine chromosome 10 locus. Bulk PCR traces from brain and liver tissues are shown for a single animal (animal 909). The dominant pure CAG repeat length was 70. Repeat lengths of 27 (short allele), 65, 76 and 82 are also observed. Traces were consistent across the eight tissues examined. GeneScan 500 LIZ internal size standard was used to determine product size. CAG repeat size was estimated against a knock-in mouse model standard, with adjustment for known differences in the polyglutamine-polyproline repeat sequence structure between the mouse and sheep models (refer to Supplementary Material).
The bulk PCR electrophoretogram peaks correspond with the 5-year-old SP-PCR data and MiSeq data, although there is a discrepancy in the short transgene copy repeat length which was called as 26 CAGs in the SP-PCR data and 27 CAGs by MiSeq. CAG repeat length calls for both fragment sizing datasets were calculated based on the OVT73 transgene polyglutamine-polyproline coding repeat structure (CAG)n(CAACAG)2(CCG)9(CCT)3, previously determined by sanger sequencing of all OVT73 generations (data not shown) and validated here by MiSeq sequencing of bulk PCR products from the 10-year-old animals HD909 and HD913 (Supplementary Figure 4). Review of the shape of SP-PCR electrophoretograms (Supplementary Figure 1) supports that the predominant short transgene copy in the 5-year-old animals likely also contains 27 CAGs, as the height of the 27 CAG peak is almost equal to that of the 26 CAG peak. The modal call of 26 CAGs for the short transgene in the SP-PCR dataset may therefore reflect PCR slippage caused by overamplification of the short allele during SP-PCR. Other repeat lengths detected in the 10-year-old bulk PCR fragment analysis (65–66, 76 and 82–83 units) were present with lower abundance as evident by the electrophoretogram peak heights (Fig. 2, Supplementary Figures 2 and 3) consistent with constitutive CAG repeat lengths of lower abundance transgene copies, rather than instability of the predominant (CAG)69(CAACAG)2 transgene copies. The presence of transgene copies with these CAG repeat lengths was confirmed in the MiSeq data (Supplementary Figure 4). The electrophoretogram peak indicating a 76 CAG repeat length in animal HD909 (Fig. 2) may reflect germline expansion of a copy of the transgene. As explained above, CAG-containing alleles detected by SP-PCR in the 5-year sheep around the repeat sizes of these major repeat length clusters are most likely attributable to individual 65–66-, 76-, and 82–83-unit polyglutamine coding transgene copies. Analysis of many more single molecules by SP-PCR would confirm the presence of these alternative repeat-length transgene copies in the 5-year-old animals, although the narrow repeat range (65–87) prevents the accurate assignment of variation to specific transgene alleles.
It was notable that two of the three 10-year-old animals (all G3) had a 70-unit CAG repeat allele, compared to 69 units in all 5-year-old sheep (G1 and G2). This agrees with previously reported one-unit generational creep between G2 and G3 animals [29]. Identifying generational and/or individual differences in CAG repeat length is relevant for breeding of the OVT73 model. We also note that the shapes of the 69–70 CAG repeat distributions differed between animals, e.g., in animal 909, the peak to the right of the modal peak drops sharply in height relative to the modal 70-repeat peak, while in animal 912, the peak to the right of the modal peak is relatively close in height to the modal 69-repeat peak. This may reflect subtly different degrees of somatic expansion between these sheep; however, the shapes of these CAG length distributions might also suggest the presence of transgene copies with both 69 and 70 CAGs and overlapping PCR products generated from each (Fig. 2, Supplementary Figures 2 and 3). Regardless, these data reveal that at 10 years the OVT73 transgene CAG repeat is still remarkably stable in all tissues tested.
Taken together, these data provide new insight into OVT73 transgene locus composite sequence, where at least 4–5 different repeat lengths are apparent in different full-length copies of the transgene, with a majority of copies having (CAG)69-70(CAACAG)2 polyglutamine-coding repeats. As explained above, the presence of multiple transgene copies is a complicating factor for quantifying somatic variation in this model. However, minimal repeat length variation observed overall indicates that all transgene copies are quite stable, including between tissues, individuals, generations, and ages. As the OVT73 sheep are a prodromal model with no overt neurological symptoms or cell death, these findings provide further evidence that somatic instability of the
Other
Sheep may also lack the appropriate complement of
With limited availability of human tissue, characterizing the handful of animal models capturing prodromal and early HD is important to resolve the intracellular threshold CAG repeat length that triggers cellular dysfunction and the transition from pre-symptomatic to symptomatic [23]. Understanding the mechanism of instability may further reveal possible targets to suppress expansion. Mouse models of HD and other repeat disorders have demonstrated that DNA repair genes influence repeat instability [13–16, 55]. The reason for the stability of the OVT73 sheep transgene expanded CAG repeat is not currently clear but may involve access by the DNA repair gene complement. Attempting to introduce instability via modulation/knock-out of DNA repair genes would test this idea and potentially enable study of the CAG length pathogenic threshold, with the complex brain and longer lifespan of sheep providing a realistic window to assess the mechanism of the prodromal-symptomatic transition. A humanized knock-in sheep would complement this by resolving complications of transgene composition and transgenesis.
Footnotes
ACKNOWLEDGMENTS
We thank the staff at the South Australian Research and Development Institute for all animal management and preparation of tissues. Marian DiFiglia and Ellen Sapp (Massachusetts General Hospital) kindly provided tissue samples for the analysis of 10-year-old OVT73 animals, originally donated from a longitudinal study cohort by Heather Grey-Edwards (University of Massachusetts). Kristine Boxen of the University of Auckland Centre for Genomics, Proteomics and Metabolomics (University of Auckland) performed Sanger sequencing and genescan services. We thank Emanuela Elezi of the MGH Mission Driven Service Core for DNA Fragment and MiSeq analyses.
FUNDING
This work was supported by the CHDI Foundation (A-2476, A-2690) and NIH grants R01 NS049206 and R01 NS091161.
CONFLICT OF INTEREST
V.C.W. was a founding scientific advisory board member with financial interest in Triplet Therapeutics Inc. Her financial interests were reviewed and are managed by Massachusetts General Hospital and Mass General Brigham in accordance with their conflict-of-interest policies. She is a scientific advisory board member of LoQus23 Therapeutics Ltd. and has provided paid consulting services to Acadia Pharmaceuticals Inc., Alnylam Inc., Biogen Inc., Passage Bio and Rgenta Therapeutics. She has received research support from Pfizer Inc.
DATA AVAILABILITY
The data supporting the findings of this study are available within the article and/or its supplementary material.
