Abstract
BACKGROUND:
Large epidemiological studies have shown that diets high in fruits reduce the risk of esophageal squamous cell carcinoma (ESCC).
OBJECTIVE:
The current study investigated the effects of black raspberries (BRBs) on gene expression during the development of preneoplastic esophagi in rats.
METHODS:
Using a post-initiation protocol, F344 rats were injected with N-nitrosomethylbenzylamine (NMBA) and then fed either a control diet or 5% BRBs. At weeks 9, 15, and 35, we euthanized subgroups of the rats and collected preneoplastic esophagi to isolate RNA samples for DNA microarray.
RESULTS:
Along the development of NMBA-induced preneoplastic esophagi, NMBA injections led to differential expression of 1181 genes comparing to control rats, and dietary BRBs modulated 428 genes in esophagi from NMBA-treated rats. There are 137 common genes between 1181 and 428 gene sets, and BRBs significantly reversed the expression of 133 genes. These genes are associated with multiple gene oncology functions. BRBs induced an 8.8-fold gene enrichment on the pathway of inflammatory response and regulated 10 genes involved in this pathway. Among them, BRBs significantly reversed the expression of pro-inflammatory cytokines, such as CCL2, S100A8, and IL19.
CONCLUSIONS:
BRBs exhibit strong anti-inflammatory effects against NMBA-induced rat esophageal tumorigenesis.
Abbreviations
black raspberries
C-C motif chemokine ligand 2
CC chemokine receptor 2
confidence interval
dietary inflammatory index
dimethyl sulfoxide
esophageal adenocarcinoma
esophageal cancer
esophageal squamous cell carcinoma
false discovery rate
The International Agency for Research on Cancer
interleukin 19
keratin 16
monocyte chemoattractant protein-1
melanoma differentiation-associated antigen 7
myeloid-related protein 8
N-nitrosomethylbenzylamine
natriuretic peptide precursor type B
prostaglandin E2
risk ratio
S100 calcium-binding protein A8.
Introduction
Esophageal cancer (EC) is the 6th leading cause of cancer-related death worldwide [1]. The two main types of EC are esophageal squamous cell carcinoma (ESCC) and esophageal adenocarcinoma (EAC). The International Agency for Research on Cancer (IARC) estimates that, in 2012, 88% of the EC cases were ESCC and 12% were EAC [2]. The geographic distribution of ESCC varies greatly, with more than 10-fold differences between countries [3]. Regions of high incidence include Eastern to Central Asia, along the Rift Valley in East Africa, and into South Africa [3]. Many risk factors have been identified to explain the high incidence of ESCC in these regions, such as smoking, heavy alcohol consumption, pro-inflammatory diet, high-temperature foods, and poor oral health [3].
A healthy dietary pattern is an important contributing factor to reduce the risk of many types of cancer, including colorectal cancer [4] and EC [5]. For example, one meta-analysis involving a total of 57 cohort studies concluded that the risk of ESCC was inversely associated with fruit intake [risk ratio (RR) for 100 g/day increment: 0.84; 95% confidence interval (CI): 0.75–0.94], and positively associated with intakes of processed meats (RR for 50 g/day increment: 1.59; 95% CI: 1.11–2.28), processed and red meats (RR for 100 g/day increment: 1.37; 95% CI: 1.04–1.82), and alcohol (RR for 10 g/day increment:1.25; 95% CI: 1.12–1.41) [6]. Another meta-analysis involving 11 prospective cohort studies and 29 case-control studies evaluated the risk of developing ESCC using the dietary inflammatory index (DII) score, which represents stronger pro-inflammatory potential of diet with higher scores [7]. This study observed that 24% increased risk of developing ESCC for each unit higher of the DII scores (pooled RR: 1.24; 95% CI: 1.10–1.38) [7]. Similar results were found in Japan [8], China [9], Iran [10], Italy [11], and Sweden [12].
Berries including strawberries, blueberries, blackberries and raspberries, are widely consumed fruits worldwide and have many benefits to our health [13–15]. Our laboratory has long been evaluating the chemopreventive effects of berries against cancer including ESCC [16–38]. The widely used animal model for ESCC studies is the induction of esophageal papillomas in Fisher 344 (F344) rats with N-nitrosomethylbenzylamine (NMBA). NMBA is present in many foods, including grilled and smoked meat, pickled and preserved vegetables [39], and moldy corn in some high-incidence regions of China [40]. Our previous studies demonstrated that subcutaneous (s.c.) injections of NMBA (0.3 mg/kg body weight, three times per week for 5 weeks) induced esophageal preneoplasia by 15–17 weeks [41], and 100% esophageal tumor incidence by 25–35 weeks in F344 rats [42]. Most tumors were squamous papillomas because the occlusive effects of the developing papillomas led to significant body weight loss before carcinomas could develop in these rats [30, 42]. Black raspberries (BRBs) contain high levels of anthocyanins, ellagitannins, and dietary fiber, and have been shown to suppress the development of NMBA-induced esophageal papillomas in rats [36]. In addition, BRBs modulated the expression of proteins associated with proliferation, apoptosis, inflammation, and angiogenesis in NMBA-induced rat esophageal papillomas [42]. In the current study, we reported the results of companion animals from our previous studies [41, 42] and investigated the effects of BRBs on gene expressions during the development of NMBA-induced preneoplastic esophagi in rats. Our results showed that BRBs reversed NMBA-induced dysregulation of genes regulating inflammation.
Materials and methods
Animals, reagents, and diets
Four- to five-week-old male F344 rats were obtained from Harlan Sprague–Dawley (Indianapolis, IN). All rats were housed two per cage under standard conditions (20±2°C, 50±10% relative humidity, 12-h light/dark cycles). Food and water were available ad libitum. Hygienic conditions were monitored and maintained by twice weekly cage changes. All animal procedures followed the recommendations of the American Association of Laboratory Animal Care (AALAC).
NMBA was purchased from Ash Stevens (Detroit, MI), and dimethyl sulfoxide (DMSO) was purchased from Sigma (St. Louis, MO). NMBA was dissolved in a solution of DMSO/water (20:80). All reagents were prepared immediately prior to usage.
The American Institute of Nutrition-76A (AIN-76A) synthetic diet served as the control diet and was purchased from Dyets, Inc. (Bethlehem, PA). BRB powder was purchased from Berri Products LLC (Corvallis, OR) and stored at 4°C. The sugar and starch content of the BRB diet was adjusted to create an isocaloric diet [18–21, 42].
Animal experiments
The animal use protocol was approved by Institutional Animal Care and Use Committee at The Ohio State University in 2008 under the protocol number “2008A0076” and title “Efficacy of berries on NMBA-induced esophageal tumorigenesis in the F-344 rat.”
A post-initiation protocol was used to investigate BRBs’ chemopreventive effects on rat esophageal tumorigenesis (Fig. 1) [41, 42]. Two weeks after arrival in the animal facility, rats were randomly assigned into three groups (Fig. 1). Rats in Group 1 (G1) were given the control AIN-76A diet and were injected subcutaneously (s.c.) with DMSO/water (20:80). Rats in Groups 2 (G2) and 3 (G3) were given the control diet and were injected s.c. with NMBA (0.3 mg/kg body weight) three times per week for 5 weeks. After the injections, the rats in G2 continued on the control diet, whereas the rats in G3 were fed 5% BRB diet. Nine rats from each group were euthanized at weeks 9, 15, and the end of the study (week 35). The esophagus of each rat was collected, opened longitudinally, and the whole epithelium (except large papillomas) was snap-frozen in liquid nitrogen for RNA isolation and DNA microarray analysis.
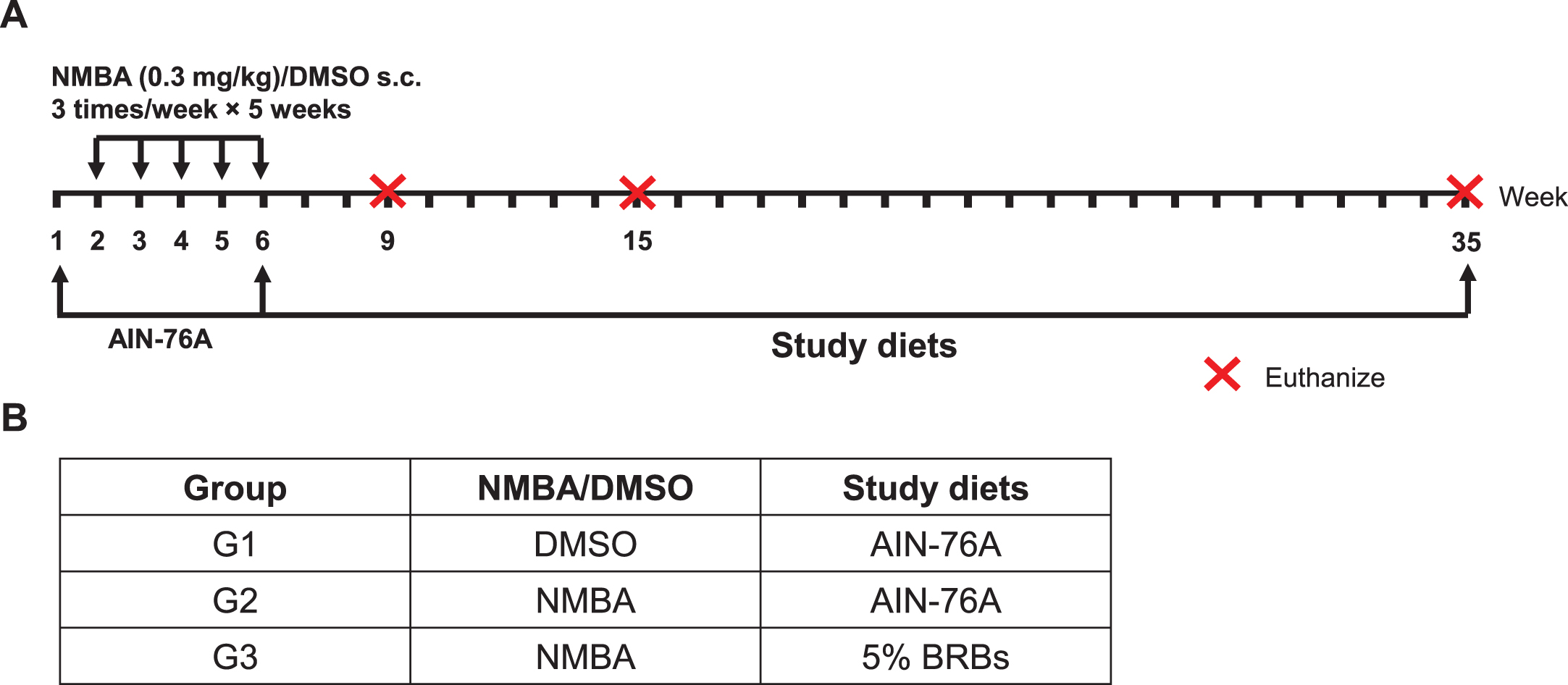
A post-initiation protocol was used to investigate the effects of BRBs on gene expression during the development of NMBA-induced preneoplastic esophagi in rats. (A) Study protocol of the post-initiation model. Rats were injected s.c. with NMBA (0.3 mg/kg body weight) or DMSO three times per week for 5 weeks. Different study diets were given after NMBA injections till the end of study (week 35). Subgroups of rats were euthanized at weeks 9, 15, and 35. (B) Group assignment of the different study diets.
RNA isolation and extraction, and DNA microarray analysis have been described [41, 42]. Total RNA was isolated according to manufacturer’s instructions (AllPrep DNA/RNA Mini kit, Qiagen, Valencia, CA) and stored at –80°C for further analysis of transcription changes. Agilent Rat Whole-Genome oligonucleotide arrays (Agilent Technologies, Santa Clara, CA) were used and 41,000 transcripts were detected in each microarray. For each time point, each group contained 9 rats. To reduce biological variation, samples from every 3 rats were combined, then triple microarrays were performed on each combined sample. This approach ensures appropriate biological replications within a limited number of microarrays [41, 42]. Normalized data for each microarray were imported into Rosetta Resolver for analysis (version 5.1.0.1.23; Rossetta Biosoftware). ANOVA was performed on log ratios using the Rosetta Resolver error model and weighting [43]. One-way ANOVA was used to identify significantly changed genes. A p-value less than 0.001 was considered statistically significant in each comparison. Genes with significant changes were filtered to identify those with more than 1.5-fold change. Then the selected genes were further analyzed for functional annotations using DAVID (Database for Annotation, Visualization and Integrated Discovery, https://david.ncifcrf.gov/).
Results and discussion
BRBs significantly regulated expression levels of 137 genes that were changed by NMBA
The current study used a post-initiation protocol of NMBA-induced esophageal tumors in rats to investigate BRBs’ effects on gene expression during the development of preneoplastic esophagi (Fig. 1). We have previously published the results of esophageal tumors from this study in which 5% BRBs were found to significantly decrease the number and size of papillomas in the esophagi of NMBA-treated rats [42]. Using esophageal samples collected at weeks 9, 15 and 35 during this study, we isolated RNA and performed DNA microarray. Three groups were DMSO + AIN-76A diet (G1), NMBA + AIN-76A diet (G2) and NMBA + 5% BRB diet (G3). We selected significantly (p < 0.001) changed genes that were present in all three time-points for all three groups. Comparing G1 and G2, we identified 2014 genes with a minimum 1.5-fold change. After restricting the false discovery rate (FDR) to 0.1, we obtained 1181 significantly changed genes. We applied the same analysis to the comparison of G2 and G3 and identified 428 significantly changed genes. There are 137 common genes between 1181 and 428 gene sets (Fig. 2, Supplementary Table 1). Among them, 133 genes were reversed towards the control values by BRBs, suggesting that these genes contribute to the development of NMBA-induced preneoplastic esophagi and reversed by BRBs.
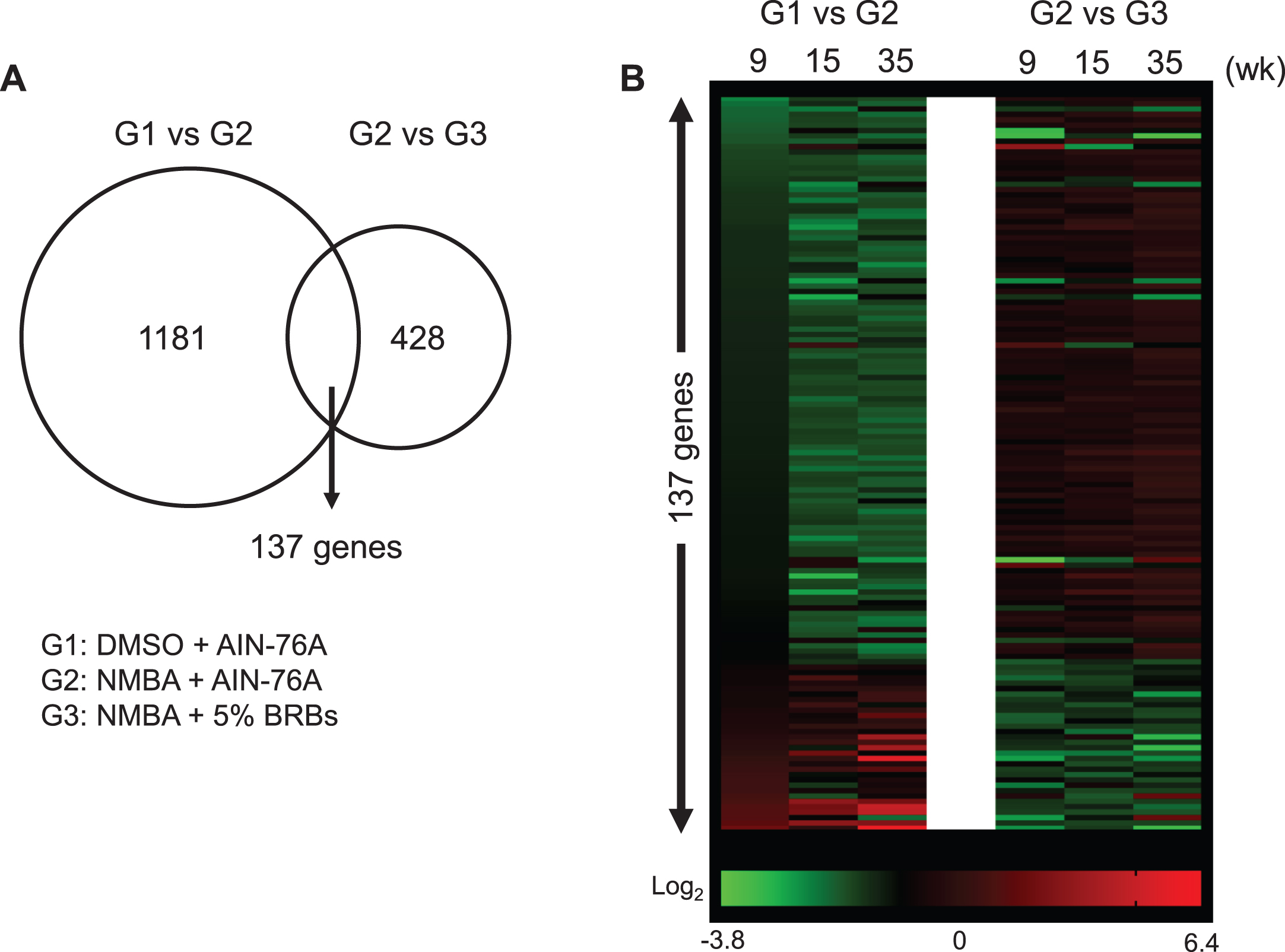
BRBs significantly regulated expression levels of 137 genes that were changed by NMBA. (A) 1181 and 428 genes were significantly changed in the comparison between G1 and G2, and between G2 and G3, respectively. There are 137 common genes between 1181 and 428 gene sets. (B) Heatmap showed relative expression levels of 137 genes.
We further analyzed the 137 genes for functional annotations using DAVID. Multiple gene oncology functions were identified (Table 1). Particularly, we detected an 8.8-fold gene enrichment in genes involved in the inflammatory response. Ten genes were associated with the inflammatory response pathway, including C-C motif chemokine ligand 2 (CCL2), S100 calcium-binding protein A8 (S100A8), tumor necrosis factor receptor superfamily member 19L (coded by RELT gene), keratin 16 (KRT16), apolipoprotein C-III (APOC3), interleukin 19 (IL19), natriuretic peptide precursor type B (NPPB), IL24, galanin (GAL), and selectin (SELE) (Fig. 3).
Functional annotations of 137 genes that were significantly changed in NMBA-induced preneoplastic esophagi in rats and regulated by BRBs
Functional annotations of 137 genes that were significantly changed in NMBA-induced preneoplastic esophagi in rats and regulated by BRBs
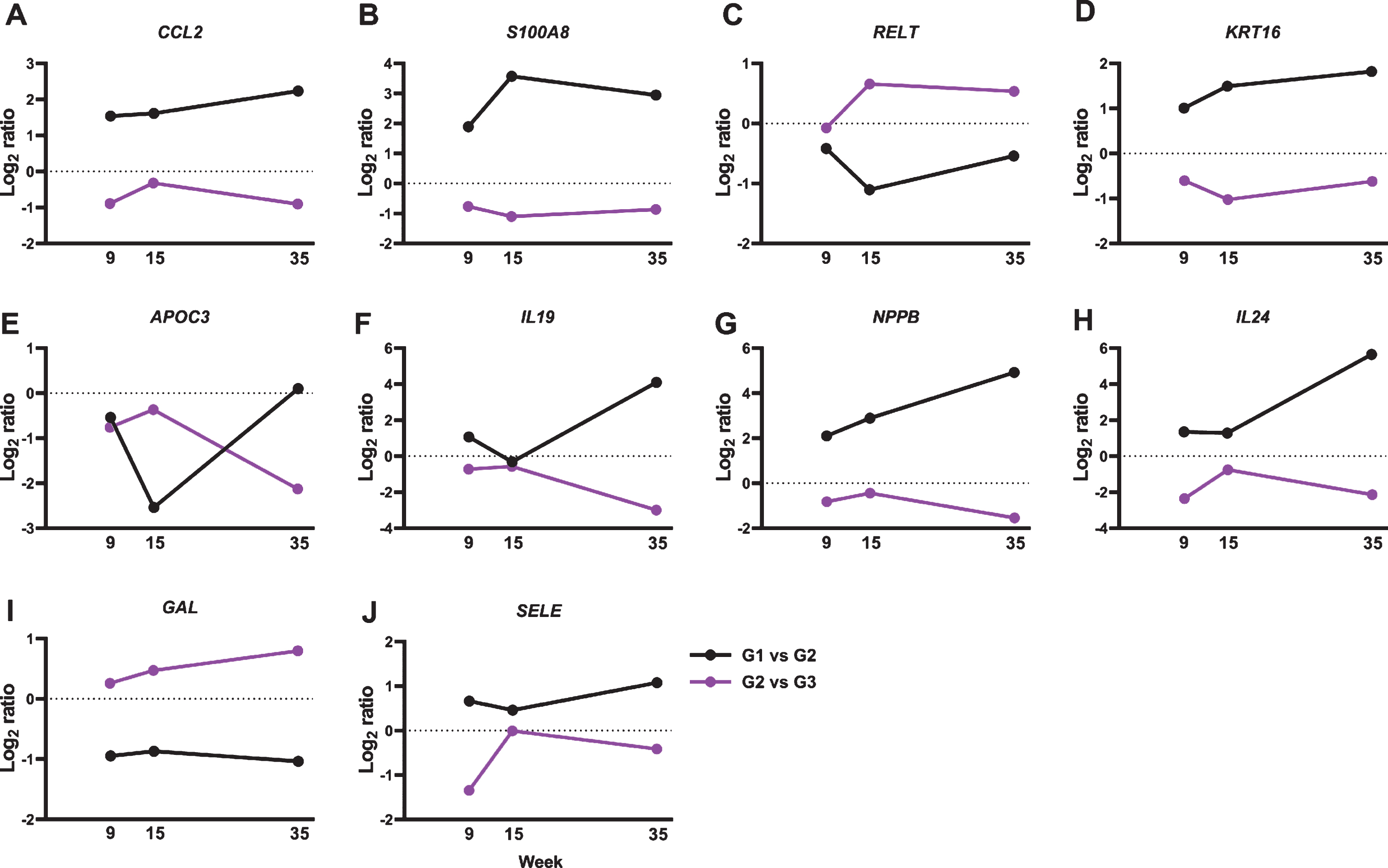
BRBs significantly regulated expression levels of 10 genes that associate with inflammatory response. Normalized Log2 ratio values were shown for (A) CCL2; (B) S100A8; (C) RELT; (D) KRT16; (E) APOC3; (F) IL19; (G) NPPB; (H) IL24; (I) GAL; and (J) SELE.
CCL2, also known as monocyte chemoattractant protein-1 (MCP-1), is a chemokine that binds to the CC chemokine receptor 2 (CCR2) and regulates monocyte chemotaxis and T-lymphocyte differentiation [44]. It plays a pivotal role in the pathogenesis of cancer and inflammatory diseases, such as asthma, systemic lupus erythematosus, atherosclerosis, and colitis [44]. ESCC patients have high levels of IL17A, which stimulates ESCC cells to release CCL2 in vitro [45, 46]. Moreover, upregulated mRNA expression levels of CCL2 have been reported in ESCC tumors than in tumor-free tissues [47]. CCL2-CCR2 signaling could promote angiogenesis thereby contributing to esophageal carcinogenesis [48]. Indeed, ESCC patients who expressed high levels of CCL2 had a worse 5-year survival rate compared to those with negative CCL2 expression [49]. In the current study, we observed a significant increase in CCL2 gene expression in NBMA-treated rat esophagus and BRBs reversed that (Fig. 3A), suggesting that BRBs might inhibit angiogenesis through suppressing CCL2-CCR2 signaling pathway in the NMBA-treated rats [50].
S100A8, also known as myeloid-related protein 8 (MRP8), is a Ca2 +-binding protein that belongs to the S100 family. S100A8 and S100A9 (MRP14) usually co-exist in the form of heterodimer, known as calprotectin, that is constitutively expressed in neutrophils and monocytes. The heterodimer functions to stimulate leukocyte recruitment and cytokine secretion thus playing an important role in modulating the inflammatory response [51]. Interestingly, studies of ESCC patients observed decreased levels of S100A8 and S100A9 in ESCC tumors compared to that in normal esophageal mucosa [52–54], whereas their expression levels increased in NMBA-induced esophageal tumors in rats [55, 56]. Similar with those rat studies, our current study observed a significant increase in the expression of S100A8 in the NMBA-treated rat esophagi, which was reversed by BRBs (Fig. 3B). Therefore, the inconsistency between human and rat studies needs further investigation to clarify the role of S100A8/S100A9 in the development of ESCC.
IL19 and IL24 [also known as melanoma differentiation-associated antigen 7 (MDA-7)] are members of the interleukin 10 (IL10) family of cytokines, which are involved in infectious and inflammatory diseases [57–61]. ESCC patients with advanced tumor stage showed increased levels of IL19 [62], and we observed higher IL19 gene expression in the NMBA-treated rat esophagus in the current study. In addition, BRBs significantly decreased levels of IL19 gene expression (Fig. 3F), suggesting that BRBs might suppress IL19-promoted inflammation and carcinogenesis. In contrast, IL24 serves as an anti-tumor cytokine, which inhibits tumor growth, invasion, metastasis and angiogenesis [63]. A Phase I/II clinical trial of patients with multiple advanced cancers demonstrated that intra-tumoral delivery of INGN 241 (a nonreplicating adenoviral construct expressing the mda-7 transgene) induced tumor apoptosis and exerted immune-activating effects [64]. However, ESCC patients were not included in this clinical trial. Our current study detected higher levels of IL24 gene expression in the NMBA-treated rat esophagus, and BRBs significantly decreased its expression (Fig. 3H). Therefore, the function of IL24 in esophageal tumorigenesis needs further investigation.
Our previous studies also demonstrated BRBs’ anti-inflammatory effects on NMBA-induced rat esophageal tumors. For example, 5% BRBs significantly decreased the levels of pro-inflammatory cytokines, such as prostaglandin E2 (PGE2) [42] and IL5 [36]. In addition, using a prevention model, we administered BRBs to rats before, during, and after NMBA injections, and observed a lower level of pentraxin-3 by BRBs [37]. These results suggest that anti-inflammation is a significant signaling pathway underlying BRBs’ anti-tumor effects.
Limited studies have investigated the role of other genes in the tumorigenesis. For example, increased serum levels of APOC3 were detected in EC patients among Kazakh people in Xinjiang region of China [65]. GAL-9 was shown to inhibit cell proliferation and promote apoptosis in EAC cell lines [66]. In addition, RELT was reported as one of the tumor-associated proteins that can be used for early diagnosis of breast cancer [67]. KRT16 has been included in a list of 24 important hub genes as potential diagnostic biomarkers of metastatic melanoma [68]. NPPB was decreased by tolfenamic acid, which suppressed cell proliferation and increased apoptosis in colon tumors [69]. E-selectin (SELE) promoted the matrix adhesion of pancreatic cancer cells [70]. These results suggest that RELT, KRT16, NPPB, and SELE are involved in the tumorigenesis. More research is warranted to examine their roles in the development of EC.
In summary, results from the present study demonstrated that BRBs reversed multiple pathways involved in the development of NMBA-induced preneoplastic esophagi in rats. In particular, BRBs significantly regulated expression levels of 10 genes involved in the pathway of inflammatory response such as CCL2, S100A8, and IL19. These results suggest that inflammatory pathway plays an important role in the development of NMBA-induced rat esophageal preneoplasia, and BRBs induce strong anti-inflammation effects. Further studies are needed to investigate the role of genes in other pathways in NMBA-induced rat esophageal tumors and to confirm their roles in human ESCC. Collectively, the nutrigenetic approach of our study is important because all the genes together can serve as the surrogate markers for early diagnosis of ESCC and determination of chemopreventive efficacy using dietary approaches.
Competing interests
No potential competing interest was disclosed.
Funding
We are grateful for financial support from NIH grants R01 CA103180 (to G. Stoner) and R01 CA148818 (to L-S. Wang).
Authors’ contribution
Study design and acquisition, analysis, and interpretation of data: Pan Pan, Alan A. Dombkowski, Gary D. Stoner, Li-Shu Wang.
Manuscript writing and revising: Pan Pan and Li-Shu Wang.
Final approval: Pan Pan, Li-Shu Wang, and Gary Stoner.
