Abstract
BACKGROUND:
The COVID-19 pandemic caused by SARS-CoV-2 is influenced by genetic and epigenetic factors, including miR-155, which affects immune cell and virus functions and laboratory biomarkers.
OBJECTIVE:
To evaluates miR-155’s role as a biomarker for SARS-CoV-2 detection and monitoring, examining its significance in identifying infection in both vaccinated and unvaccinated individuals using ROC curve analysis.
METHODS:
Blood samples were collected from 70 patients who attended Medical City Hospital in Baghdad from June 2022 to April 2023 and were determined to be associated with SARS-CoV-2 (35 patients were hospitalized at the Intensive Care Units due to the severity of their symptoms while the other 35 were left in the hospital upon treatment.). Additionally, 35 samples were collected as a healthy control group.
RESULTS:
The expression level of miR-155 in the serum of samples showed a high level (fold change: 9.81
CONCLUSIONS:
The miR-155 gene, overexpressed in SARS-CoV-2 patients, correlates with disease activity and severity, potentially serving as a biomarker for diagnosis and a potential therapeutic target.
Introduction
Acute respiratory syndrome coronavirus 2 (SARS-CoV-2) caused a pneumonia outbreak to spread throughout the world [1, 2] and became a widespread health issue [3, 4]. SARS-CoV-2 ranging symptoms from common cold to severe illnesses, Acute lung disease symptoms are common in SARS-CoV-2 patients and some of them may even be serious [5, 6].
MicroRNAs (miRNAs) are small, non-coding RNA molecules that play an essential role in post-transcript-ional gene regulation in organisms. These molecules are endogenously generated and are approximately 22 nucleotides in length. They are primarily involved in regulating gene expression by either facilitating the cleavage of target messenger RNAs (mRNAs) or repressing their translation [7, 8]. Once the virus infects cells, these miRNAs regulate viral infection and control viral replication [9]. MicroRNA involved in the cytokine storm induced by SARS-CoV-2 infection, have been identified as potential biomarkers for SARS-CoV-2 indicating their role in a wide range of clinical manifestations [10]. The viral load can be significantly decreased by the cellular miRNAs’ destruction of the viral RNA and the decreased expression of ACE2 receptors.
It has been discovered that certain miRNAs have antiviral properties that either force the virus into its latency stage or help the immune system fight off the infection in addition to reducing the harmful effects of the increased cytokine output [11], Viruses employ the cellular machinery of their host to copy their genetic material. Interactions between the viral genome and host miRNAs may take place throughout this process [12, 13].
MicroRNA-155 is involved in inflammation and post-transcriptional gene regulation [14]. concerning viral infections, miR-155 may be a viable therapeutic target [15], MiRNAs like miR-155 can target the host’s innate and adaptive immunity, serve as diagnostic tools enhancing patients screening, and potentially develop into miR-inhibitor or miR-mimics for therapeutic strategies [16], Cellular miRNAs can interact with RISC proteins to silence viral RNA, and non-coding RNAs (ncRNAs) can be used to treat SARS-CoV-2 infected host cells by inducing small interfering RNA (siRNA) through synthetic complementary siRNAs [11].
The use of laboratory biomarkers in predicting disease severity, hospitalization, intensive care unit (ICU) admission, and mortality has been well-documented [17]. By assessing and monitoring these biomarkers at an early stage of the disease, it may be possible to intervene and potentially halt disease progression and reduce mortality [18].
The study aims were to determine the analysis of the expression of miR-155 and its laboratory biomarkers using the ROC curve, the specificity and the sensitivity of these biomarkers. These finding can provide valuable insights into their potential as predictive markers of disease severity, hospitalization, ICU admission, and mortality. Moreover, it could have helped in developing the early intervention and monitoring strategies to effectively combat disease progression and reduce the risk of death.
Materials and methods
Study ethical considerations
The study was conducted per the ethical principles that have their origin in the Declaration of Helsinki, this research was approved by the Institute of Genetic Engineering and Biotechnology for the Postgraduate Studies Committee, as well as the Iraqi Ministry of Health Research Committee (The document number 1316 at 15/5/2022). Participant consent was obtained before enrolment in this study.
Subjects’ selection and sample collection
The study consisted of 105 individuals from Iraq, categorized into three distinct groups: patients: severe (
Primer Sequence for miRNA gene expression (Designed by NCBI)
Primer Sequence for miRNA gene expression (Designed by NCBI)
Every participant in this study had a venipuncture performed to obtain blood specimens. Five ml of blood were collected from both patients and the control. 3 ml of the blood was mixed with a gel tube, then transferred into a new sterile test tube, and 2
To analyze the Complete Blood Count, the blood specimen in the EDTA tube was shaken up and then examined as soon as possible in CELL-DYN Ruby automated hematology analyzers to count white blood cells, lymphocytes, HCT, HGB, and platelets. To estimate the C-reactive protein (CRP) and D. dimer, the gel tube containing serum was entered into the equipment and analyzed automatically The Roche Cobas C111 Chemistry Analyzer from Roche Diagnostics is used to detect C-reactive protein and D-dimer. CRP Normal Range: Less than 1 mg/l; High: greater than 13 mg/l and D-dimer Normal Range: Less than 255 ng/L; High: greater than 255 ng/L.
Molecular diagnosis of SARS-CoV-2
The study involved testing samples from a patient’s nasopharyngeal swab and extracting RNA. SARS-CoV-2 RNA was detected using a one-step RT-PCR method using a SARS-CoV-2 nucleic acid detection kit (Maccura Biotechnology Co. Ltd. 16#. China). The PCR amplification parameters were prepared according to the manufacturer’s recommendations. Results were considered positive when the CT of three targets (FAM, ROX, and CY5) was
MiR-155 quantitation
The miRNAs were quantified using the Qubit 4.0 fluorimeter platform, which exhibits outstanding selectivity for miRNA in comparison to other RNA types. For assessing initial sample concentrations ranging from 10 pg/L to 100 ng/L.
Quantitative PCR (qPCR)
High-capacity cDNA Kit (ProtoScript® II First Strand cDNA Synthesis Kit, NEW ENGLAND BIOLABS) was used to reverse transcribe the RNA.
RNA reverse transcription
The reverse transcription technique was used with the ProtoScript® II First Strand cDNA Synthesis Kit to evaluate the expression of PCR target genes with oligo-dT primers.
Measurement of miR-155 gene expression
The amplification of miRNA genes was carried out using specialized primers, as listed in Table 1. MacroGen® provided these primers in a lyophilized form. a working solution of the primers at 10 pmol/
The amplification conditions were established as follows: The second step of the methodology involved choosing and processing cDNA samples. Every sample was run through two different PCR tubes, one for miR-155 and the other for U6 snRNA, the study’s housekeeping gene. Specific ingredients were carefully combined to create the reaction mix components and their respective quantities, qPCR master mixer, Universal qPCR Master Mix: 10
Statistical analysis
A statistical study was conducted using the Statistical Package for Social Sciences (SPSS version 26, Inc., Chicago, IL, USA) and the Microsoft Excel Worksheet. The results and examples of the current study were analyzed.
Results and discussion
Subject analysis
In this study, the age range of the patients varied from 18 to 80 years, with an average age of 45.66
Comparison between SARS-CoV-2 patients and healthy control groups as regards CRP, D. dimer and CBC
Comparison between SARS-CoV-2 patients and healthy control groups as regards CRP, D. dimer and CBC
Values are expressed as mean
MiR-155 was measured using RT-PCR
This means having the different letters in the same column differed significantly, ** (
In terms of biomarkers, the patients had elevated levels of C-reactive protein (CRP), with severe cases exhibiting a mean value of 116.16
The miR-155 and the reference gene U6 were amplified in the molecular experiment to determine each threshold cycle (Ct) value. The melting temperature curve is calculated using the Ct values, which are also used to quantify real-time RT-PCR data that are inversely related to the amount of the beginning template.
The expression fold of miR-155 is displayed in Table 3. The results of a gene expression analysis using RT-PCR for miRNA-155 in vitro showed interesting findings. Specifically, the folding value of miR-155 in SARS-CoV-2 severe infection (9.81
Overall, the findings suggest that miR-155 may play a significant role in the progression and severity of SARS-CoV-2 infection.
Correlation between miR-155 and C-reactive protein.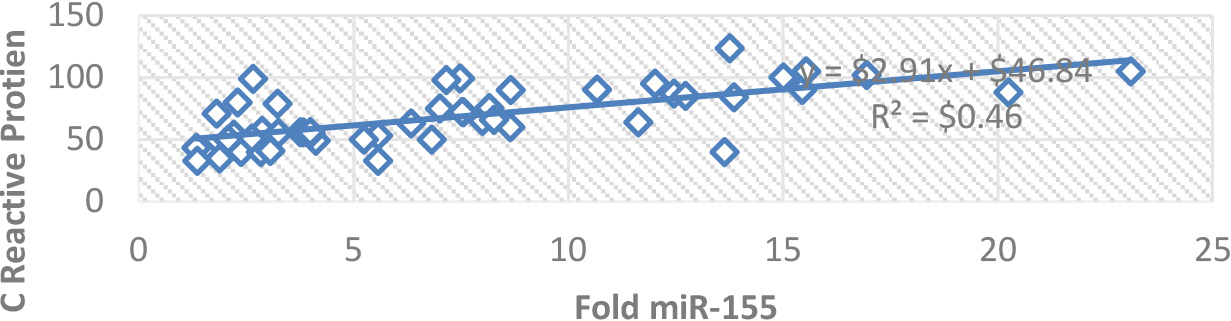
Vaccination according to the number of doses.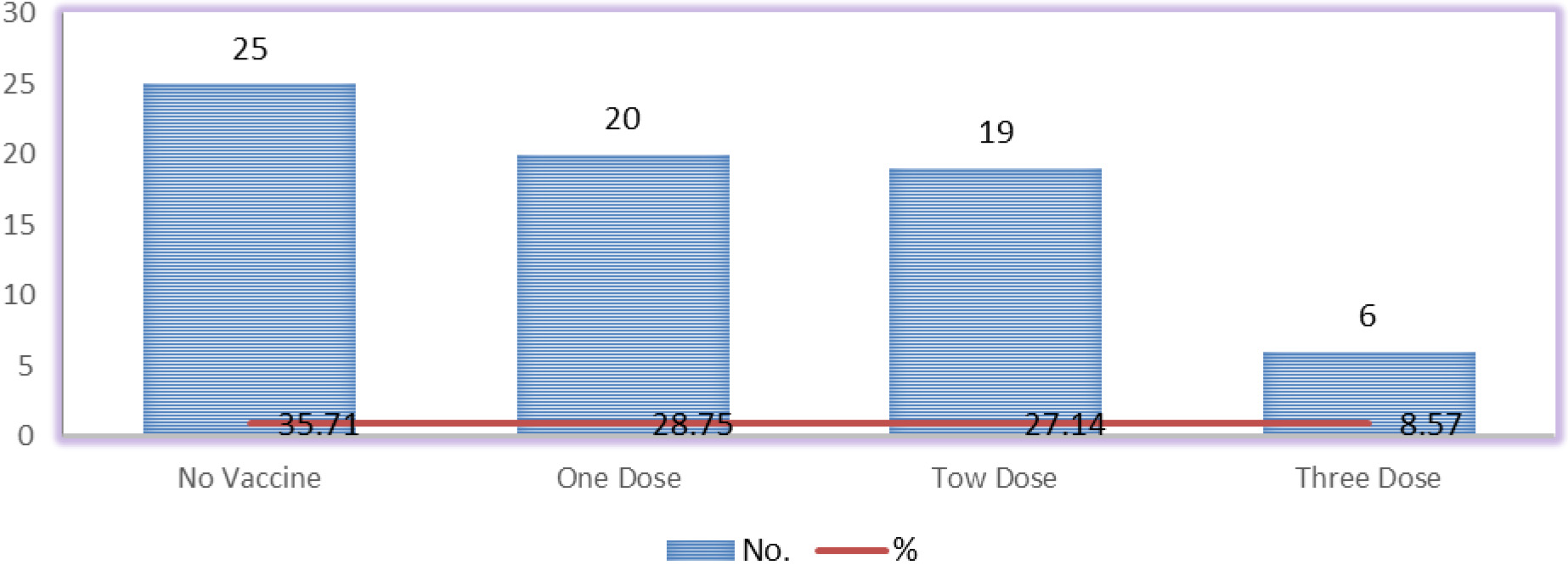
Compared with controls a significant upregulation of miR-155 was observed; miR-155 had a positive correlation with CRP depicted in Fig. 1. The positive correlation between miR-155 and C-reactive protein suggests that miR-155 may be involved in the regulation of C-reactive protein expression or its downstream signaling pathways. This finding provides further evidence for the potential involvement of miR-155 in the inflammatory response. Approximately 62.5 percent of individuals received the vaccinations, whereas 34.7% did not, Among SARS-CoV-2 patients, three different vaccines have been administered: Pfizer, AstraZeneca, and Sinopharm. These vaccines have been given in varying dosages, as indicated in Fig. 2.
The onset of the symptom and SARS-CoV-2 duration of survival
As for the onset of SARS-CoV-2 symptoms and the duration of survival for the SARS-CoV-2 virus among the patients. According to the data, the highest value for the onset of symptoms was observed for 40 out of 70 patients (57.1%) within a span of 3 days. Additionally, 27 patients (38.6%) experienced symptom onset within 2 days. In terms of the duration of survival for the SARS-CoV-2 virus, the maximum value was recorded at 10 days for 46 patients (65.7%). For 24 patients (34.3%), the virus reached its maximum duration of survival in 12 days. The complete details of these findings can be found in Table 4.
The onset of the symptom and SARS-CoV-2 duration of survival
* The mean difference is significant at the 0.05 level, NS: Not significant.
Sensitivity and specificity miR-155 and some parameters in SARS-CoV-2 patients
AUC is the area under the curve. A Highly significant at
The selection was taken according to gender, and age groups, so the
Receiver operating characteristic (ROC) curve of circulating miR-155 and other parameters for detection of (A) SARS-CoV-2 disease and (B) Severity (C) Lymphocyte SARS-CoV-2 disease (D) Lymphocyte severity (E) Vaccinated patients (F) Unvaccinated patients, for SARS-CoV-2 disease.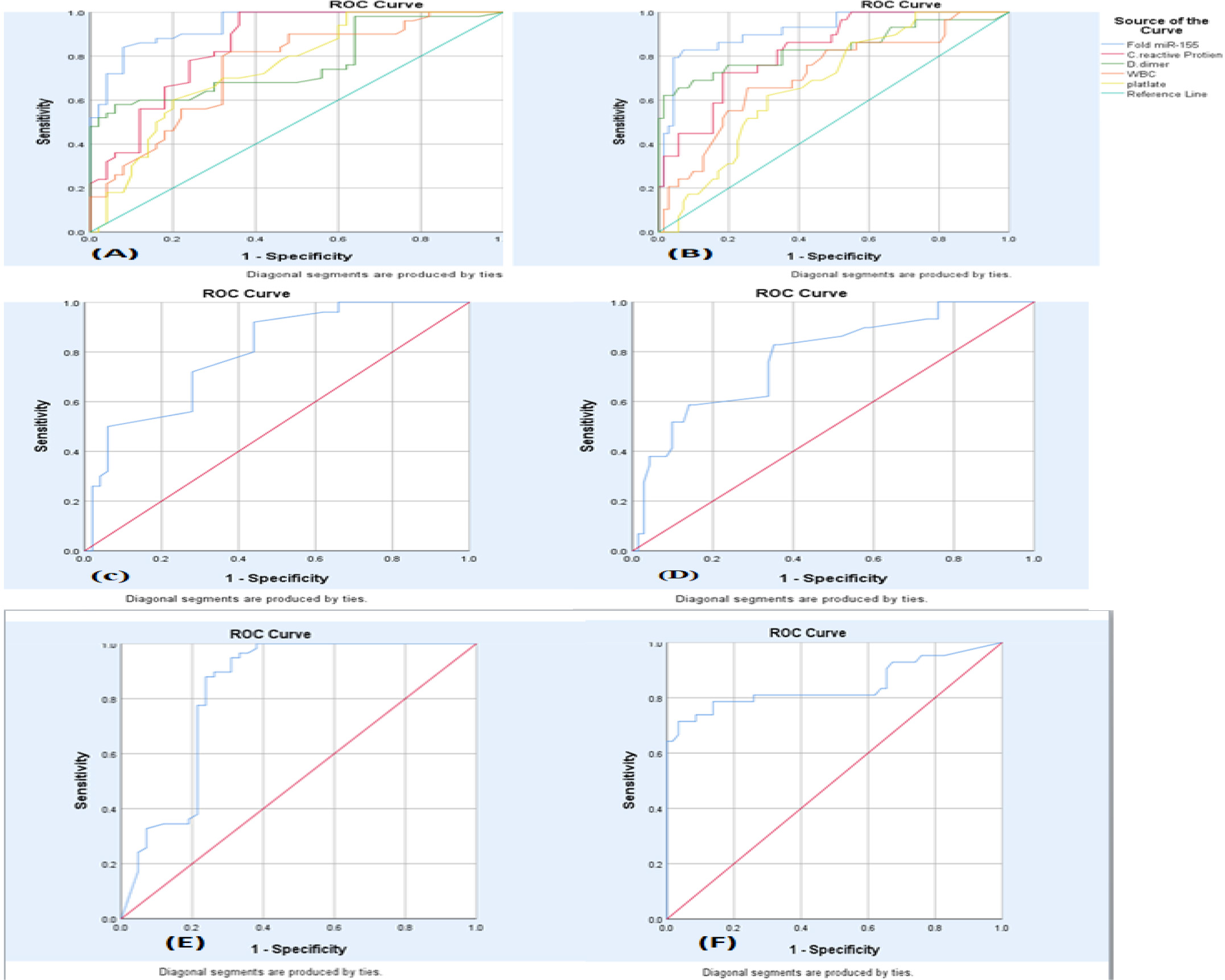
SARS-CoV-2 infection triggered a global public health crisis. This highly transmissible respiratory disease can cause mild symptoms up to severe pneumonia with potentially fatal respiratory failure. This study showed that older people were more likely to SARS-CoV-2. This study proved and this is supported by other studies that reported that individuals aged 45 – 80 with chronic conditions are more likely to die and are more likely to be hospitalized as the immune system weakens and becomes less effective in mounting an effective response against the virus [22, 23]. Furthermore, in a study similar to the research findings, there appears to be a higher number of male SARS-CoV-2 patients compared to females [24]. The findings from extensive research indicate that there is an upregulation of ACE2 expression levels in individuals who are sick, male, and older even before being infected with the SARS-CoV-2 virus, it has also been observed that certain male gender and advanced age are associated with a higher risk of developing severe SARS-CoV-2symptoms [25]. The data indicates that male patients have an over-fold expression of miR-155 compared to female patients [26].
Additionally, Smoking can weaken the immune system and damage the respiratory system, making individuals more susceptible to respiratory infections like SARS-CoV-2. Research has shown that habits such as smoking and obesity can act as risk factors for severe illness and complications in individuals with SARS-CoV-2 [27, 28].
In this study, it observed that approximately 40% of the participants were smokers, while the remaining 60% were non-smokers. One interesting finding was that the expression fold of miR-155 was higher in smokers compared to non-smokers. This implies that smoking may have a direct impact on the regulation of miR-155 expression levels. Further investigations are needed to understand the potential consequences of this higher miR-155 expression in smokers and its potential implications for health and disease.
Among the miRNAs, miRNA-155 (miR-155) has been thoroughly examined in individuals with SARS-CoV-2 that is either severe or mild. According to reports, people infected with SARS-CoV-2 may be instructed to trigger a cytokine storm or strong pro-inflammatory reactions. CRP and miR-155 exhibited a strong and positive correlation, Consequently, the molecules can control the expression of the viral and immune system proteins inside the host cells. Based on these findings, MiR-155 has been shown by researchers to have a major role in the pathophysiology and severity of SARS-CoV-2, and it may be used as a clinical biomarker for early infection severity and detection [29]. Based on these results, patients with severe conditions had the highest C-reactive protein levels, followed by those with moderate or mild conditions compared to control, where it was between (116.16
The study examined the onset and duration of SARS-CoV-2 symptoms in a group of patients. It found that for 57.1% of the patients, the onset of symptoms occurred within a span of 3 days, while for 38.6% of patients, symptoms manifested within 2 days. In terms of the duration of survival, it was observed that 65.71% of patients reached the maximum duration of 10 days, while 34.28% of patients had a longer survival period of 12 days.
The study also evaluated the relationship between miR-155 and the duration of survival in SARS-CoV-2 patients. The findings indicated no statistical significance between miR-155 and the duration of survival. This implies that miR-155 does not appear to have an impact on the length of time a patient survives with SARS-CoV-2 infection, as reflected by the ratio of 0.38 (not significant).
The inclusion of miR-155 as a parameter suggests its potential utility as a biomarker for SARS-CoV-2 diagnosis. Analyzing the sensitivity and specificity of miR-155 in conjunction with other parameters can provide a comprehensive understanding of its diagnostic performance.
To evaluate the SARS-CoV-2 disease (patient and healthy control) sensitivity and specificity of miR-155, the ROC curve was employed. The cutoff value identified was 2.34 demonstrating a sensitivity of 84% and a specificity of 92%. Furthermore, the calculation of the area under the curve (AUC) yielded a value of 0.94. The calculated (AUC) value indicates a high level of accuracy for miR-155 in detecting SARS-CoV-2 disease.
As for the detection of SARS-CoV-2 disease severity, the ROC curve was employed the cutoff value identified was 4.06, demonstrating a sensitivity of 82% and a specificity of 88%. Furthermore, the calculation of the area under the curve (AUC) yielded a value of 0.91. According to this finding, miR-155 might have a direct effect on the degree of severity and duration of COVID-19 disease. The study investigates the role of miR-155 in detecting SARS-CoV-2 disease in both vaccinated and unvaccinated individuals. This study utilizes the ROC curve analysis to determine the cutoff values for miR-155 in both groups. For the unvaccinated group, the cutoff value was identified as 3.50. This means that individuals with a miR-155 level below 3.50 were considered negative for SARS-CoV-2 disease. The analysis showed a sensitivity of 76%, indicating the proportion of true positive results, and a specificity of 86%, indicating the proportion of true negative results. The calculated area under the curve (AUC) was 0.844, suggesting a good accuracy of miR-155 in detecting the disease in unvaccinated individuals.
For the vaccinated group, the cutoff value was identified as 2.34. This means that individuals with a miR-155 level below 2.34 were considered negative for SARS-CoV-2 disease. The analysis showed a sensitivity of 74% and a specificity of 88%. The AUC value for this group was also 0.844, indicating a similar accuracy in detecting the disease among vaccinated individuals. These findings highlight the importance of vaccination and adherence to preventive measures in reducing the impact of the virus, particularly for high-risk groups. The use of miR-155 as a biomarker for detecting SARS-CoV-2 disease can provide valuable insights for effective management and early intervention.
One study revealed that it provides evidence of the effectiveness of SARS-CoV-2 vaccines in stimulating the immune response and providing protection to the Iraqi population [31], This is what our evaluation of the expression revealed miR-155 in vaccinated individuals was lower than unvaccinated based on the number of dosages taken, individuals during SARS-CoV-2 infection, suggesting that small RNAs may regulate gene expression and serve as a biomarker of infection severity, The qRT-PCR experiment conducted as part of the study revealed that the expression of miR-155 genes was elevated in SARS-CoV-2 patients compared to the healthy control group. These findings contribute to our understanding of the immune response to SARS-CoV- 2 and provide insights into potential biomarkers for evaluating infection severity. The study investigated the correlation between miR-155 expression levels and the dosage of a vaccine. The results showed a significant relationship between the two variables. The highest level of miR-155 expression was observed in individuals who were not vaccinated or had received only one dose, with a value of 8.40
MiR-155 may also be therapeutic according to studies from earlier research and the mechanism depends on their ability to target viral or cellular RNAs during infection. In recent studies, including viral infections miRNAs found in plants, animals, and some viruses like SARS-CoV-2, are crucial regulators of viral infections, suppressing gene expression during infection [35]. Also, cellular miRNAs can inhibit the translation of the viral genome to prevent virus replication [36]. Ultimately, these findings support the notion that the presence and severity of SARS-CoV-2 disease are associated with the quantity of miR-155 in the blood. Therefore, it might be a useful biomarker for SARS-CoV-2 diagnosis. Furthermore, it might be a crucial therapy medicine for those with COVID-19.
In conclusion, further research on miR-155 and its connection to the SARS-CoV-2 virus is essential for gaining insights into the underlying mechanisms of SARS-CoV-2 pathogenesis. Investigating the role of miR-155 as a potential biomarker could provide valuable information for developing effective diagnostic and therapeutic strategies for SARS-CoV-2.
This study aims to evaluate the role of miR-155 as a biomarker for the detection and monitoring of SARS-CoV-2, furthermore, C-reactive protein and miR-155 exhibited a strong and positive correlation, Also, the study found significant differences in miR-155 levels between infected patients and control groups, with patients showing a significantly up-regulated miR-155, however, further study is required to evaluate miR-155 as a potential therapeutic target.
Funding
There is no Funding.
Author contributions
Conception: Halah Kadhim and Abdulameer Ghareeb.
Interpretation or analysis of data: Halah Kadhim and Mohammed Alhilal.
Preparation of the manuscript: Halah Kadhim.
Revision for important intellectual content: Halah Kadhim.
Supervision: Abdulameer Ghareeb and Mohammed Alhilal.
Datasets/data availability statement
The data supporting the findings of this study are available within the article and/or its supplementary material.
Footnotes
Acknowledgments
The authors thank all the nurses and laboratory technicians (We especially appreciate the assistance provided by the lab personnel in the Molecular Biology Unit of the National Center for Educational Laboratories/Baghdad, Iraq) who participated in this research and assisted with blood sample collection.
Conflict of interest
No conflict of interest was declared by the authors.
