Abstract
Adopting open science principles can be challenging, requiring conceptual education and training in the use of new tools. This paper introduces the Workflow for Open Reproducible Code in Science (WORCS): A step-by-step procedure that researchers can follow to make a research project open and reproducible. This workflow intends to lower the threshold for adoption of open science principles. It is based on established best practices, and can be used either in parallel to, or in absence of, top-down requirements by journals, institutions, and funding bodies. To facilitate widespread adoption, the WORCS principles have been implemented in the R package
Introduction
Academia is arguably past the tipping point of a paradigm shift towards open science. Support for this transition was first motivated by several highly publicized cases of scientific fraud (Levelt et al. [20]), increasing awareness of questionable research practices (John et al. [16]), and the replication crisis (Shrout and Rodgers [36]). However, open science should not be seen as a cure (or punishment) for this crisis. As the late dr. Jonathan Tennant put it: “Open science is just good science” (Tennant [39]). Open science creates opportunities for researchers to more easily conduct reliable, cumulative, and collaborative science (Adolph et al. [3]; Nosek and Bar-Anan [24]). Open science also promotes inclusivity, because it removes barriers for participation. Capitalizing on these advances has the potential to accelerate scientific progress (see also Coyne [11]), as has been aptly demonstrated by the role of open science in the response to the coronavirus pandemic (Sondervan et al. [37]).
Many researchers are motivated to adopt current best practices for open science and enjoy these benefits. And yet, the question of how to get started can be daunting. Making the transition requires researchers to become knowledgeable about different open science challenges and solutions, and to become proficient with new and unfamiliar tools. This paper is designed to ease that transition by presenting a simple workflow, based on established best practices, that meets most requirements open science: The
This paper introduces the conceptual workflow (referred to in upper case, “WORCS”), and discusses its underlying principles and how it meets different standards for open science. The principles underlying this workflow are universal and largely platform independent. We also present a software implementation of WORCS for
The WORCS workflow constitutes a step-by-step procedure that researchers can follow to make a research project open and reproducible. These steps are elaborated in greater detail below. See Fig. 1 for a flowchart that highlights important steps in different stages of the research process. WORCS is compatible with open science requirements already implemented by journals and institutions, and will help fulfill them. It can also be used by individual authors to produce work in accordance with best practices in the absence of top-down support or guidelines (i.e., “grass roots” open science).
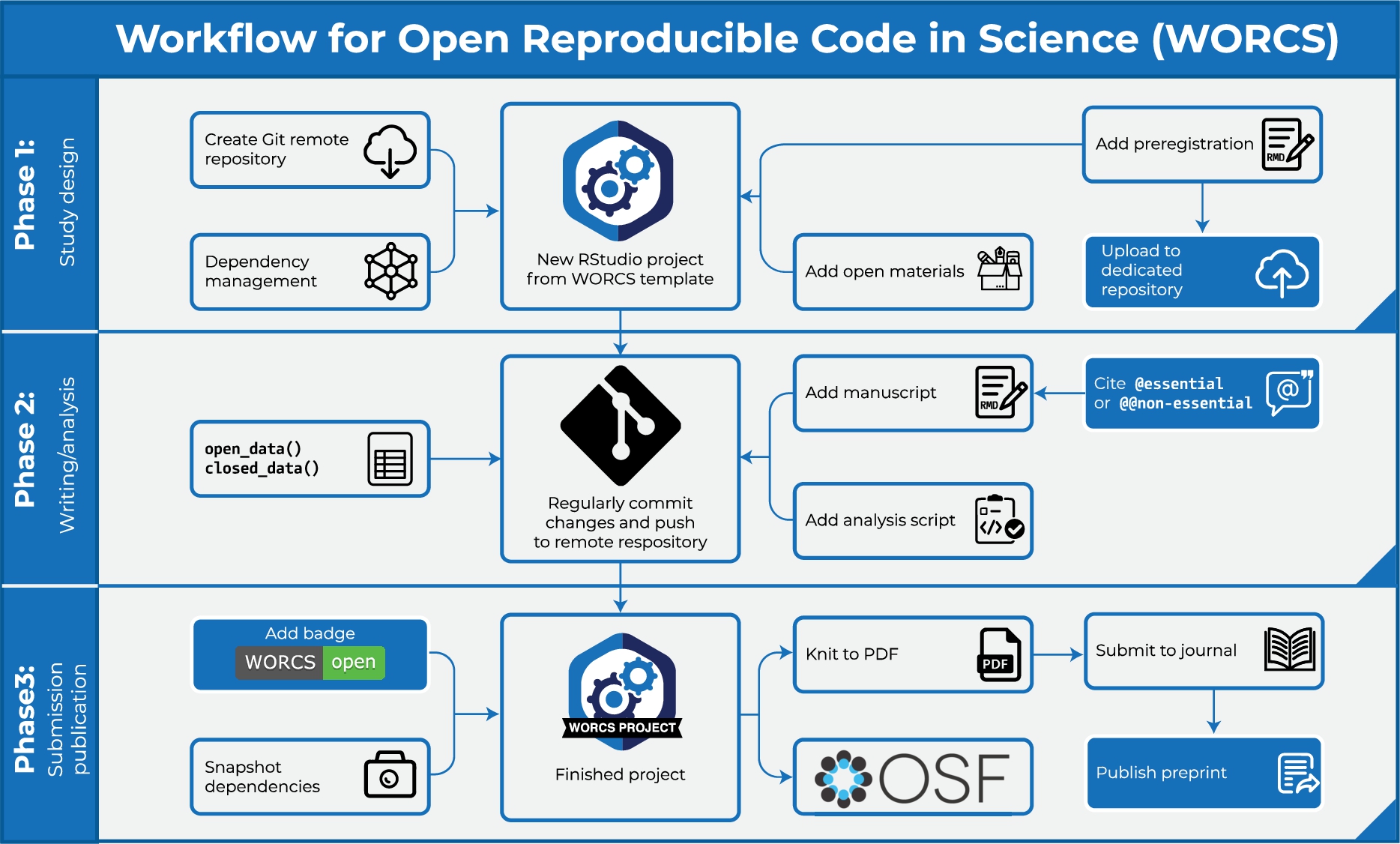
Schematic illustration of important steps in different stages of the WORCS procedure.
Although open science is advocated by many, it does not have a unitary definition. Throughout this paper, we adhere to the definition of open science as “the practice of science in such a way that others can collaborate and contribute, where research data, lab notes and other research processes are freely available, under terms that enable reuse, redistribution and reproduction of the research and its underlying data and methods” (the FOSTER Open Science Definition, as discussed in Bezjak et al. [8]). In this context, the term “science” refers to all scholarly disciplines. We further define the term
The “TOP-guidelines” are one of the most influential concrete operationalisations of general open science principles (Nosek et al. [23]). These guidelines describe eight standards for open science, the first seven of which “account for reproducibility of the reported results based on the originating data, and for sharing sufficient information to conduct an independent replication” (Nosek et al. [23]). These guidelines are: 1) Comprehensive citation of literature, data, materials, and methods; 2) sharing data, 3) sharing the code required to reproduce analyses, 4) sharing new research materials, and 5) sharing details of the design and analysis; 6) pre-registration of studies before data collection, and 7) pre-registration of the analysis plan prior to analysis. The eighth criterion is “not formally a transparency standard for authors”, but “addresses journal guidelines for consideration of independent replications for publication”. In this context, reproducibility can be defined as “re-performing the same analysis with the same code using a different analyst”, and replicability as “re-performing the experiment and collecting new data” (Patil et al. [27]). WORCS defines the goals of open science in terms of the first seven TOP-guidelines, and the workflow and its
Introducing the tools
WORCS relies solely on a set of free, open source software solutions which we will discuss before introducing the workflow.
Dynamic document generation
The first is
Although transitioning to DDG involves a slight learning curve, we strongly believe that the investment will pay off. Time saved from painstakingly copy-pasting output and manually formatting text soon outweighs the investment of switching to a new program. Moreover, human error in manually copying results is eliminated. When revisions require major changes to the analyses, all results, figures and tables are automatically updated. The flexibility in output formats also means that a manuscript can be rendered to presentation format, or even to a website or blog post. Moreover, the fact that code is evaluated each time the document is compiled requires researchers to work reproducibly, and allows reviewers and/or readers verify reproducibility simply by re-compiling the document. While writing academic papers in a programming environment might seem counter-intuitive at first, this approach is much more amenable to the needs of academics than most word processing software. It prevents mistakes, and saves time.
In the
The RMarkdown document can incorporate blocks of analysis code, which can be used to render results directly to Tables and Figures, which can be dynamically referenced in the text. Long scripts can be stored in
Version control
The second solution is
An integral part of WORCS is the
From an open science perspective, Git is particularly appealing because it retains a complete historical backlog of all commits. This can be used, for example, to verify that authors executed the steps outlined in a preregistration. Users can also compare changes between different commits, or go back to a previous version of the code (for example, after making a mistake, or to replicate a previous version of the results). Git only version controls files explicitly committed by the user. Moreover, it is possible to prevent files from being version controlled – which is useful for privacy sensitive data. A
The functionality of Git is enhanced by services such as GitHub (
The social network aspect of GitHub comes into play when a repository is made “public”: This allows other researchers to peruse the repository and see how the work was done; clone it to their own computer to replicate the original work or apply the methods to their own data; open “Issues” to ask questions or give feedback on the project, or even send a “Pull request” with suggested changes to the text or code for your consideration. Git and GitHub shine as tools for collaboration, because different people can simultaneously work on different parts of a project, and their changes can be compared and automatically merged on the website. Even on solo projects, working with Git/GitHub has many benefits: Staying organized, being able to start a new study with a clone of an old, similar repository, or splitting off an “experimental branch” to try something new, while retaining the ability to “revert” (return) to a previous state of the project, or to “merge” (incorporate) the experimental branch.
Although GitHub is the most widely used remote repository,
Dependency management
The third solution is
Many solutions exist to ensure computational reproducibility, which differ in user-friendliness and effectiveness. These solutions typically work by enveloping your research project in a distinct “environment” that only has access to programs that are explicitly installed, and maintaining a record of these programs. When choosing an appropriate solution for dependency management, there is a tradeoff between ease-of-use on the one hand, and robustness on the other. In making this tradeoff, it is important to consider that all solutions have a limited shelf life (Brown [10]). Therefore, the best approach might be to use a “good-enough” solution, and acknowledge that all code requires some maintenance if you want to reproduce it in the future.
For dependency management, the
All solutions to computational reproducibility require users to make their dependencies explicit, often by reinstalling all software for each project in a controlled environment. This can lead to long installation times and large memory requirements. Similarly,
When a project is conducted in a different software environment than
Docker instantiates a “Docker container”: An environment that behaves like a virtual computer, which can be stored like a time capsule, and identically reinstated on a user computer, or in the cloud. Although the use of Docker falls outside of the scope of the present paper, for existing users, a Docker build running
As Docker is somewhat challenging to set up for novice users, an
Text-based files are better
A key consideration when developing a research project is what filetypes to use. WORCS encourages the use of text-based files whenever possible, instead of binary files. Text-based files can be read by machines and humans alike. Binary files, such as Word (
Two additional points are worth noting: First, as Git tracks line-by-line changes, it is recommended to start each sentence on a new line. That way, the change log will indicate which specific sentence was edited, instead of replacing an entire paragraph. As RMarkdown does not parse a single line break as the end of a paragraph, this practice does not disrupt the flow of a paragraph in the rendered version. Second, the remote repository GitHub renders certain text-based filetypes for online viewing: For example,
Introducing the workflow
We provide a conceptual outline of the WORCS procedure below, based on WORCS Version 0.1.6. The conceptual workflow recommends actions to take in the different phases of a research project, in order to work openly and transparently in compliance with the TOP-guidelines and best practices for open science. Note that, although the steps are numbered for reference purposes, we acknowledge that the process of conducting research is not always linear. We refer users of the
Phase 1: Study design
Create a (Public or Private) remote repository on a “Git” hosting service When using R, initialize a new RStudio project using the WORCS template. Otherwise, clone the remote repository to your local project folder. Add a README.md file, explaining how users should interact with the project, and a LICENSE to explain users’ rights and limit your liability. This is automated by the Optional: Preregister your analysis by committing a plain-text preregistration and tag this commit with the label “preregistration”. Optional: Upload the preregistration to a dedicated preregistration server Optional: Add study materials to the repository
Phase 2: Writing and analysis
Create an executable script documenting the code required to load the raw data into a tabular format, and de-identify human subjects if applicable Save the data into a plain-text tabular format like Write the manuscript using a dynamic document generation format, with code chunks to perform the analyses. Commit every small change to the “Git” repository Use comprehensive citation
Phase 3: Submission and publication
Use dependency management to make the computational environment fully reproducible Optional: Add a WORCS-badge to your project’s README file Make a Private “Git” remote repository Public Create a project page on the Open Science Framework (OSF) and connect it to the “Git” remoterepository
Generate a Digital Object Identifier (DOI) for the OSF project
Add an open science statement to the Abstract or Author notes, which links to the “OSF” project page and/or the “Git” remote repository Render the dynamic document to PDF Optional: Publish the PDF as a preprint, and add it to the OSF project Submit the paper, and tag the commit of the submitted paper as a release of the submitted paper as a release, as in Step 4.
Notes for cautious researchers
Some researchers might want to share their work only once the paper is accepted for publication. In this case, we recommend creating a “Private” repository in Step 1, and completing Steps 13-18 upon acceptance.
The R implementation of WORCS
WORCS is, first and foremost, a conceptual workflow that could be implemented in any software environment. As of this writing, the workflow has been implemented for RStudio’s primary purpose is to create free and open-source software for data science, scientific research, and technical communication. This allows anyone with access to a computer to participate freely in a global economy that rewards data literacy; enhances the production and consumption of knowledge; and facilitates collaboration and reproducible research in science, education and industry.
Fourth,
Working with
Preparing your system
Before you can use the
How WORCS helps meet the TOP-guidelines
Comprehensive citation
The TOP-guidelines encourage comprehensive citation of literature, data, materials, methods, and software. In principle, researchers can meet this requirement by simply citing every reference used. Unfortunately, citation of data and software is less commonplace than citation of literature and materials. Crediting these resources is important, because it incentivizes data sharing and the development of open-source software, supports the open science efforts of others, and helps researchers receive credit for
To facilitate citing datasets, researchers sometimes publish
References for software are sometimes provided within the software environment; for example, in
One important impediment to comprehensive citation is the fact that print journals operate with space constraints. Print journals often discourage comprehensive citation, either actively, or passively by including the reference list in the manuscript word count. Researchers can overcome this impediment by preparing two versions of the manuscript: One version with comprehensive citations for online dissemination, and another version for print, with only the essential citations. The print version should reference the online version, so interested readers can find the comprehensive reference list. The WORCS procedure suggests uploading the online version to a preprint server. This is important because most major preprint servers – including arXiv.org and all preprint services hosted by the Open Science Framework (OSF) – are indexed by Google Scholar. This means that authors will receive credit for cited work; even if they are cited only in the online version. Moreover, preprint servers ensure that the online version will have a persistent DOI, and will remain reliably accessible, just like the print version.
Implementation in worcs
The
The
With regard to the citation of
Data sharing
Data sharing is important for computational reproducibility and secondary analysis. Computational reproducibility means that a third party can exactly recreate the results from the original data, using the published analysis code. Secondary analysis means that a third party can conduct sensitivity analyses, explore alternative explanations, or even use existing data to answer a different research question. If it is possible to share the data, these can simply be pushed to a Git remote repository, along with documentation (such as a codebook) and the analysis code. This way, others can download the entire repository and reproduce the analyses from start to finish. From an open science perspective, data sharing is always desirable. From a practical point of view, it is not always possible.
Data sharing may be impeded by several concerns. One technical concern is whether data require special storage provisions, as is often the case with “big data”. Most Git remote repositories do not support files larger than 100 MB, but it is possible to enhance a Git repository with Git Large File Storage to add remotely stored files as large as several GB. Files exceeding this size require custom solutions beyond the scope of this paper.
Sharing human participant data additionally entails legal and ethical concerns. Before sharing any human participant data, it is recommended to obtain approval from qualified ethical and legal advisory organs, guidance from Research Data Management Support, and informed consent from participants. Many legislatures require researchers to “de-identify” human subject data upon collection, by removing or deleting any sensitive personal information and contact details. However, when researchers have an imperfect understanding of the legal obligations and dispensations, they can end up over- or undersharing, and risk either placing participants at risk of being identified, or undermine the replicability of their own work and potential reuse value of their data (Phillips and Knoppers [30]). Furthermore, it is important to note that the European GDPR prohibits storing “personal data” (information which can identify a natural person whether directly or indirectly) on a server outside the EU, unless it offers an “adequate level of protection”. Although different rules may apply to pseudonimized data, there are many repositories that are GDPR compliant, such as the European servers of the Open Science Framework. Different Universities, countries, and funding bodies also have their own repositories that are complient with local legistation. In sum, data should only be shared after consult qualified legal and ethical advisory organs, obtaining informed consent from participants, and de-identifying or pseudononimizing data.
If data cannot be shared, researchers should aim to safeguard the potential for computational reproducibility and secondary analysis as much as possible. WORCS recommends two solutions to accomplish this goal. The first solution is to publish a It is theoretically possible but improbable that random changes to a file will result in the same checksum.
A second solution is to share a synthetic dataset with similar characteristics to the real data. Synthetic data mimic the level of measurement and (conditional) distributions of the real data (see Nowok et al. [25]). Sharing synthetic data allows any third party to 1) verify that the published code works, 2) debug the code, and 3) write valid code for alternative analyses. It is important to note that complex multivariate relationships present in the real data are often lost in synthetic data. Thus, findings from the real data might not be replicated in the synthetic data, and findings in the synthetic data
Note that for data synthesis to work properly, it is essential that the
When initializing a new
One issue of concern is the practice of “fixing” mistakes in the data. It is not uncommon for researchers to perform such corrections manually in the raw data, even if all other analysis steps are documented. Regardless of whether the data will be open or closed, it is important that the raw data be left unchanged as much as possible. This eliminates human error. Any alterations to the data – including processing steps and even error corrections – should be documented in the code, instead of applied to the raw data. This way, the code will be a pipeline from the raw data to the published results. Thus, for instance, if data have been entered manually in a spreadsheet, and a researcher discovers that in one of the variables, missing values were coded as
The
In
Alternatively, if the project requires data to remain closed, researchers can call
As the purpose of the
After generating tidy, shareable data, this data is then loaded in the analysis code with the function
Note that these functions can be used to store multiple data files, if necessary. As of version 0.1.2, the
Some users may intend to make their data openly available through a restricted access platform, such as an institutional repository, but not publicly through a Git remote repository. In this case, it is recommended to use
Sharing code, research materials, design and analysis
When writing a manuscript created using dynamic document generation, analysis code is embedded in the prose of the paper. Thus, the TOP-guideline of sharing analysis code can be met simply by committing the source code of the manuscript to the Git repository, and making this remote repository Public. If authors additionally use open data and a reproducible environment (as suggested in WORCS), then a third party can simply replicate all analyses by copying the entire repository from the Git hosting service, and Knitting the manuscript on their local computer.
Aside from analysis code, the TOP-guidelines also encourage sharing new research materials, and details of the study design and analysis. These goals can be accomplished by placing any such documents in the Git repository folder, committing them, and pushing to a cloud hosting service. As with any document version controlled in this way, it is advisable (but not required) to use plain text only.
Preregistration
Lindsay et al. [21] define preregistration as “creating a permanent record of your study plans before you look at the data. The plan is stored in a date-stamped, uneditable file in a secure online archive.” Two such archives are well-known in the social sciences: AsPredicted.org, and OSF.io. However, Git cloud hosting services also conform to these standards. Thus, it is possible to preregister a study simply by committing a preregistration document to the local Git repository, and pushing it to the remote repository. Subsequently taging the release as “Preregistration” on the remote repository renders it distinct from all other commits, and easily findable by people and programs. This approach is simple, quick, and robust. Moreover, it is compatible with formal preregistration through services such as AsPredicted.org or OSF.io: If the preregistration is written using dynamic document generation, as recommended in WORCS, this file can be rendered to PDF and uploaded as an attachment to the formal preregistration service.
The advantages, disadvantages, and pitfalls for preregistering different types of studies have been extensively debated elsewhere (see Lindsay et al. [21]). For example, because the practice of preregistration has historically been closely tied to experimental research, it has been a matter of some debate whether secondary data analyses can be preregistered (but see Weston et al. [43] for an excellent discussion of the topic).
When analyzing existing data, it is difficult to
Good faith preregistration efforts always improve the quality of deductive (theory-testing) research, because they avoid HARKing, ensure reliable significance tests, avoid overfitting noise in the data, and limit the number of forking paths researchers wander during data analysis (Gelman and Loken [12]).
WORCS takes the pragmatic position that, in deductive (hypothesis-testing) research, it is beneficial to plan projects before executing them, to preregister these plans, and adhere to them. All deviations from this procedure should be disclosed. Researchers should minimize exposure to the data, and disclose any prior exposure, whether direct (e.g., by computing summary statistics) or indirect (e.g., by reading papers using the same data). Similarly, one can disclose any deviations from the analysis plan to handle unforeseen contingencies, such as violations of model assumptions; or additional exploratory analyses.
WORCS recommends documenting a verbal, conceptual description of the study plans in a text-based file. Optionally, an analysis script can be preregistered to document the planned analyses. If any changes must be made to the analysis code after obtaining the data, one can refer to the conceptual description to justify the changes. The ideal preregistered analysis script consists of a complete analysis that can be evaluated once the data are obtained. This ideal is often unattainable, because the data present researchers with unanticipated challenges; e.g., assumptions are violated, or analyses work differently than expected. Some of these challenges can be avoided by simulating the data one expects to obtain, and writing the analysis syntax based on the simulated data. This topic is beyond the scope of the present paper, but many user-friendly methods for simulating data are available in most statistical programming languages. For instance,
As soon as a project is preregistered on a Public Git repository, it is visible to the world, and reviewers (both formal reviewers designated by a journal, and informal reviewers recruited by other means) can submit comments, e.g., through “Pull requests” on a Git remote repository. If the remote repository is Private, Reviewers can be invited as “Collaborators”. It is important to note that contributing to the repository in any way will void reviewers’ anonymity, as their contributions will be linked to a user name. Private remote repositories can be made public at a later date, along with their entire time-stamped history and tagged releases.
Implementation of preregistration in worcs
The
Compatibility with other standards for open science
Aside from the TOP Guidelines, the FAIR Guiding Principles for scientific data management and stewardship (Wilkinson et al. [45]) are an important standard for open science. These principles advocate that digital research objects should be Findable, Accessible, Interoperable and Reusable. Initially mainly promoted as principles for data, they are increasingly applied as a standard for other types of research output as well, most notably software (Lamprecht et al. [19]).
WORCS was not designed to comprehensively address these principles, but the workflow does facilitate meeting them. In terms of Findability, all files in GitHub projects are searchable by content and meta-data, both on the platform and through standard search engines. The workflow further recommends that users create a project page on the Open Science Framework – a recognized repository in the social sciences. When doing so, it is important to generate a Digital Object Identifier (DOI) for this project. A DOI is a persistent way to identify and connect to an object on the internet, which is crucial for findability. Additional DOIs can be generated through the OSF for specific resources, such as data sets. It is further worth noting that, optionally, GitHub repositories can be connected to Zenodo: On Zenodo, users can store a snapshot of the repository, provide metadata, and generate a DOI for the project or specific resources. When a project is connected to the OSF, as recommended, these steps may be redundant. Finally, each worcs project has a [YAML file] (
Accessibility is primarily safeguarded by hosting all project files in a public GitHub repository, which can be downloaded as a ZIP archive, and cloned or forked using the Git protocol. GitHub is committed to long term accessibility of these resources. We further encourage users to make their data accessible when possible. When data must be closed, the
With regard to interoperability, the workflow recommends using plain-text files whenever possible, which ensures that code and meta-data are human- and machine-readable. Furthermore, the
To further facilitate reusability,
Incidentally, the FAIR principles have been applied to software as well, and
Sharing all research objects, as advocated in WORCS, also provides a thorough basis for research evaluation according to the San Francisco Declaration on Research Assessment (DORA;
Discussion
In this tutorial paper, we have presented a workflow for open reproducible code in science. The workflow aims to lower the threshold for grass-roots adoption of open science principles. The workflow is supported by an
Comparing WORCS to existing solutions
There have been several previous efforts to promote grass-roots adoption of open science principles. Each of these efforts has a different scope, strengths, and limitations that set it apart from WORCS. For example, there are “signalling solutions”; guidelines to structure and incentivize disclosure about open science practices. Specifically, Aalbersberg et al. [1] suggested publishing a “TOP-statement” as supplemental material, which discloses the authors’ adherence to open science principles. Relatedly, Aczel et al. [2] developed a consensus-based Transparency Checklist that authors can complete online to generate a report. Such signalling solutions are very easy to adopt, and they address TOP-guidelines 1–7. Many journals now also offer authors the opportunity to earn “badges” for adhering to open science guidelines (Kidwell et al. [18]). These signalling solutions help structure authors’ disclosures about, and incentivize adherence to, open science practices. WORCS, too, has its own checklist that calls attention to a number of concrete items contributing to an open and reproducible research project. Users of the
A different class of solutions instead focuses on the practical issue of
One initiative that WORCS is fully compatible with, is the Scientific Paper of the Future . This organization encourages geoscientists to document data provenance and availability in public repositories, document software used, and document the analysis steps taken to derive the results. All of these goals could be met using WORCS.
Limitations
WORCS is intended to substantially reduce the threshold for adopting best practices in open and reproducible research. However, several limitations and issues for future development remain. One potential challenge is the learning curve associated with the tools outlined in this paper. Learning to work with
Another limitation is the fact that no single workflow can suit all research projects. Not all projects conform to the steps outlined here; some may skip steps, add steps, or follow similar steps in a different order. In principle, WORCS does not enforce a linear order, and allows extentensive user flexibility. A related concern is that some projects might require specialized solutions outside of the scope of this paper. For example, some projects might use very large data files, rely on data that reside on a protected server, or use proprietary software dependencies that cannot be version controlled using
Another important challenge is managing collaborations when only the lead author uses RMarkdown, and the coauthors use Word. When using the
If all collaborators are committed to using WORCS, they can Fork the repository from the lead author on GitHub, clone it to their local device, make their own changes, and send a pull request to incorporate their changes. Working this way is extremely conducive to scientific collaboration (Ram [32]). Recall that, when using Git for collaborative writing, it is recommended to insert a line break after every sentence so that the change log will indicate which specific sentence was edited, and to prevent “merge conflicts” when two authors edit the same line. The resulting document will be rendered to PDF without spurious line breaks.
Being familiar with Git remote repositories opens doors to new forms of collaboration: In the open source software community, continuous peer review and voluntary collaborative acts by strangers who are interested in a project are commonplace (see Adolph et al. [3]). This kind of collaboration is budding in scientific software development as well; for example, the lead author of this paper became a co-author on several
A final limitation is that, as WORCS is relatively new, the number of current users is still limited. We therefore do not have elaborate insights in user experiences or feedback. We strive to make the uptake of worcs as easy as possible, by means of this background paper, extensive documentation and vignettes for the
Future developments
WORCS provides a user-friendly and lightweight workflow for open, reproducible research, that meets all TOP-guidelines pertaining to reproducibility (1–7). Nevertheless, there are clear directions for future developments. Firstly, although the workflow is currently implemented only in
Conclusion
WORCS offers a workflow for open reproducible code in science. The step-by-step procedure outlined in this tutorial helps researchers make an entire research project Open and Reproducible. The accompanying
WORCS encourages and simplifies the adoption of open science principles in daily scientific work. It helps researchers make all research output created throughout the scientific process – not just manuscripts, but data, code, and methods – open and publicly assessible. This enables other researchers to reproduce results, and facilitates cumulative science by allowing others to make direct use of these research objects.
Footnotes
Acknowledgements
The lead author is supported by a NWO Veni grant (NWO grant number VI.Veni.191G.090). We acknowledge Jeroen Ooms for offering feedback and suggesting the use of the R package gert.
