Abstract
Background
Pathological angiogenesis is crucial for tumor progression, thus targeting neovascularization is regarded as an effective strategy for cancer therapy. Vascular endothelial growth factor (VEGF), a specific pro-vascular endothelial regulator, contributes to aberrant tumor angiogenesis.
Objective
To identify sequence polymorphisms of VEGF gene and the effects on breast cancer.
Methods
Protein-DNA binding was validated by EMSA and ChIP assay. Gene expression levels were detected by qPCR and western blot. The CCK-8, wound healing and transwell assays were used to assess proliferation, migration, and invasion. Tube formation, CAM, ELISA and IHC assays were performed to evaluate tumor angiogenesis.
Results
A novel 18-bp indel mutation of the VEGF promoter was detected in breast cancer cases, and the deletion allele (DD) presented dominant distribution in patients comparing to the insert type (II). Further analysis revealed that the 18-bp deletion eliminated the recognition sites of GA binding protein alpha (GABPα), which was confirmed by binding experiments. Functionally, the GABPα expression is decreased in breast cancer tissues, and acts as a tumor suppressor to inhibit proliferation, migration, invasion and angiogenesis of breast cancer cells, accompanied by accelerated tumor cell apoptosis. In addition, consistent regulatory roles were investigated in mouse models in response to GABPα overexpression or knockdown as well. Mechanistically, we revealed that GABPα inhibited breast cancer progression and angiogenesis by downregulating VEGF transcription via the 18-bp promoter sequences.
Conclusions
Our findings provide insights into angiogenic targeted strategy aiming at GABPα-VEGF axis in clinical diagnosis and therapy of breast cancer.
Introduction
Breast cancer is a highly heterogeneous disease and has exceeded lung cancer as the world's most epidemic solid tumor based on the 2020 World Health Organization (WHO) announcement. 1 According to the pathological nidus and genic profiles, breast cancer was defined as multiple clinical subtypes originating from luminal or basal epithelial cells in the mammary duct, including luminal A, luminal B, human epidermal growth factor receptor 2 (HER2) overexpression, and basal-like triple-negative breast cancer (TNBC). 2 The available treatments for breast cancer as surgery, endocrine therapy, radiotherapy and chemotherapy, and targeted therapy do not significantly alleviate the 5-year survival, prognosis and relapse. 3 The current medical plight drives a pressing need to systematically clarify the pathogenesis and tumorigenesis for excavating novel regiments and markers in breast cancer immunotargeted therapy.
GA binding protein alpha (GABPα), a subunit of GABP transcription factors belonging to E26 transformation specific (ETS) family, forms a heterotetrameric complex with its two partners GABPB1 or GABPB2 to adjust target gene expression by regulating DNA binding and transcription.
4
GABPα plays a critical role in the oncogenesis and deterioration of various human diseases including endometrial carcinoma, thyroid carcinoma, bladder cancer, hepatocellular carcinoma and other malignancies.5–8 It was previously reported that depletion of GABPα inhibited telomerase expression, while enhanced tumor metastasis and stemness of renal cell carcinoma by directly targeting transforming growth factor beta receptor type 2 (TGFBR2).
9
Consistently, the
Sequence mutations and polymorphisms of promoter region can significantly alter DNA binding and transcriptomic activity, subsequently contribute to the pathological features of mutation-bearing diseases.
13
For instance, three single nucleotide polymorphisms (SNPs) within the
Material and methods
Tissue samples of patients and ethical statement
A total of 218 breast cancer patients aged from 30 to 70 years old enrolled in this study at People's Hospital Affiliated to Shandong First Medical University between 2016 and 2023. Blood samples from 230 healthy subjects were collected from the population undergoing routine health checkups at the Hospital of Wuhan University from March 2016 to June 2017. Their paraffin-embedded breast tumor tissues and paired adjacent tissues were collected for genomic DNA extraction and immunohistochemistry assay. All patients received no radiotherapy and chemotherapy before surgery. This study was approved by the Ethics Committee at People's Hospital Affiliated to Shandong First Medical University, and the written informed consents were obtained for sample collection.
DNA polymorphism detection
The genomic DNA of tissue or blood samples from all participants was isolated by phenol-chloroform method, and quantified using NanoDrop One (ThermoFisher, Waltham, MA). According to the promoter sequences harboring 18-bp indel polymorphism of
Cell lines and cell culture
All cells (MDA-MB-231, MCF-7, MCF-10A, HUVECs, HEK293 T) used in this study were purchased from the American type culture collection (ATCC; Manassas, VA, USA). The human breast cancer cell lines (MDA-MB-231, MCF-7) and HEK293 T cell were cultured in Dulbecco's modified Eagle's medium (DMEM; Gibco, USA). The non-tumorigenic human breast cell line MCF-10A was cultured in Mammary Epithelial Cell Medium (MEpiCM; ScienCell, USA) with mammary epithelial cell growth supplement (MEpiCGS). Human umbilical vein endothelial cells (HUVECs) were cultured in Endothelial Cell Medium (ECM; ScienCell, USA). All cell culture mediums were supplemented with 10% fetal bovine serum, 100 U/mL penicillin and 100 µg/mL streptomycin, and the cells were maintained in a humidified incubator at 37 °C with 5% CO2.
RNA extraction and quantitative real-time PCR (qPCR) analysis
Total RNA was isolated from cultured cells or tumor tissues using the Ultrapure RNA Kit (CWBIO), and the extracted RNA was reversely transcribed into cDNA using the PrimeScript RT Reagent Kit (Takara). Quantitative real-time PCR (qPCR) assay was performed to detect the transcriptional levels of various gene expressions. According to the manufacturer's instructions, the qPCR experiments were carried out with Hieff qPCR SYBR Green Master Mix (YEASEN, Shanghai, China) on a CFX-96 real-time PCR system (Bio-Rad Laboratories Inc.) using an activation start cycle for 30 s at 95°C, followed by 39 cycles of 10 s at 95°C and 30 s at 60°C and 30 s at 68°C. The GAPDH was used as an internal control for normalization, and gene expression was calculated by the 2−ΔΔCt formula. The primer sequences used in this study are provided in Table S1. Each sample was repeated in triplicate.
Western blot
The total protein was isolated from cultured cells using RIPA lysis buffer (Beyotime) with 1% PMSF (Beyotime), and the consistence was quantified using the BCA protein concentration determination kit (Beyotime) according to the instructions. The proteins (50 μg) were fractionated by SDS-PAGE gel electrophoresis electroblotted onto polyvinylide ne fluoride film (PVDF; Millipore). The film with protein were washed with TBST, blocked with 5% skim milk in TBS containing 0.1% Tween 20 (TBST) for 2 h at room temperature, and examined with primary antibodies against GABPα (Cat No. sc-28311, 1:1000 dilution, Santa Cruz Biotechnology Inc), VEGF-A (Cat No. BA0407, 1:500 dilution, BOSTER) and GAPDH (Cat No. CSB-MA000071M1 m, 1:1000 dilution, CUSABIO) at 4°C overnight. The films were then washed three times with TBST for 8 min, incubated with the IgG-hydrogen peroxide (HRP) secondary antibody (Cat No. A19056, 1:10000, ABclonal) for 1 h at room temperature. The protein blots were uncovered to ECL luminol reagent (Monad) using the BIO-RAD Gel imaging system, analyzed using Image lab software.
Plasmids construction and lentivirus infection
To overexpression the
Luciferase reporter assay
According to the 18-bp indel polymorphism within
Flow cytometry
Flow cytometry assay was used to detect the apoptosis of MDA-MB-231 cells under different treatment using FITC Annexin V Apoptosis Detection Kit (BD Biosciences, CA, USA). Cells were cultured in 6-well plates for 12 h, and then pCDH-GABPα or sh-GABPα or control plasmid was transfected into MDA-MB-231 cells. After 24 h infection, cells were collected and washed with PBS for three times. The cells were resuspended with 100 μL 1× binding buffer, and then were incubated with 5 μL PI and 5 μL FITC Annexin V for 15 min at room temperature in dark. Finally, the dispersive cells were detected by NovoCyte flow cytometry.
Cell viability and clone formation assay
Cell Counting Kit-8 (CCK-8) assay (Beyotime, China) was used to detect the cell viability of breast cancer cells. The cells of different treatment were cultured in a 96-well plate for 96 h, and 10 μL CCK-8 solution and 100 μL of DMEM (without FBS) were added to each well and incubated for 2 h in 37°C, 5% CO2 humidified incubator at every 24 h. Afterward, the optical density (OD) at 450/620 nm was measured using a microplate reader (Molecular Devices, Sunnyvale, CA, USA). Each treatment was performed in six replicates. In addition, a total of 1000 treated cells per well were seeded in 6-well plates and cultured for 10 days. Cells were washed with PBS and fixed with 4% paraformaldehyde for 60 min, and stained with 1% crystal violet staining solution for 30 min.
Wound healing migration assay
The transfected cells were cultured until 100% confluence in 6-well plate, and a sterile pipette tip (10 μL) was used to scratch the cell monolayer to generate uniformed lines. The deciduous cells were washed with PBS, and continue to culture in FBS-free medium for 72 h at 37 °C, 5% CO2 in humid environment. The cell migration process was measured by phase-contrast imaging at every 24 h. Each group was performed independently in triplicate. The width of the wound was analyzed using Image J software.
Transwell invasion assay
Cell invasion assay was carried out in the 24-well transwell chambers with 8.0 μm pore size. The treated breast cancer cells were resuspended in 200 μL serum-free medium (1 × 105 per well) and seeded into the upper chamber coated with 20 μL Matrigel (BD Biosciences, CA, USA), and 600 μL DMEM containing 10% FBS was added to the bottom well. After 24 h of invasion, the detained cells were removed from the membrane with a cotton swab, and the invaded cells on chamber were fixed with 4% paraformaldehyde for 20 min and then stained with crystal violet. The stained cells were imaged and counted from at least six views. The invasion assay of each group was repeated three times.
Tube formation assay
A total of 2 × 105 MDA-MB-231 cells with GABPα overexpression or depletion for each well were plated in the 6-well plate, and cultured for 24 h in serum-free medium, and the supernatants were collected as conditioned medium (CM). After Matrigel (BD Biosciences, Bedford, MA, USA) dissolved at 4 °C overnight, the pre-cooled 96-well plate was polymerized by 50 μL Matrigel per well at 37°C for 30 min. HUVEC cells were digested and seeded on the layer of Matrigel according to the amount of 3 × 104 cells/well, and were incubated in 100 μL CM at 37°C for 2–4 h. The capillary tubes were quantified based on the images that randomly chosen from bright-field microscope. The tubule structure network was quantified using Image J Angiogenesis Analyzer program.
Chick chorioallantonic membrane (CAM) assay
Day-3 fertilized chicken eggs (presented from Huazhong Agricultural University) were chosen to conduct the CAM assay. Firstly, the eggs were incubated in a 37°C incubator with 40%∼60% relative humidity for 4 days. A window with 1 cm diameter was opened on the 7-day-old chicken embryo eggshell. To expose the CAM, the window was closed with a piece of steriled scotch tape and incubated for one day to adapt the change. Then, a qualitative filter paper in 0.5 cm diameter was treated with 100 μL conditioned medium (CM) and were placed in the middle of the blood vessels. After closing the window with sterile tape, the chicken eggs were incubated at 37°C with constant humidity for 72 h. The CAMs were fixed with stationary solution (methanol: acetone = 1:1) for 15 min. Finally, the CAMs were collected and photographed using a digital camera (Panasonic, Osaka, Japan), and the angiogenesis were assessed by analyzing the number of newly formed vessels.
Electrophoretic Mobility Shift Assay (EMSA)
For EMSA assay, the nuclear extracts were isolated from HUVEC cells with Nuclear and Cytoplasmic Extraction Reagents (Beyotime, China). Based on the 18-bp indel polymorphism within
Chromatin immunoprecipitation (ChIP) assay
The ChIP experiment was carried out using ChIP assay kit (Cell Signaling Technology, Beverly, MA, USA) according to the manufacturer's protocol. Briefly, cells were cross-linked with 1% formaldehyde for 10 min and terminated with 0.125 M glycine. Then, the cells were lysed with SDS lysis buffer containing protease inhibitor and sonicated on ice to produce 200–500 bp fragments. The cross-linked complex containing soluble chromatin was immunoprecipitated overnight at 4°C with anti-GABPα antibody (Santa Cruz Biotechnology Inc) or isotype control rabbit IgG antibody (Cell Signaling Technology, Beverly, MA, USA) bound to protein A/G agarose beads. Finally, the precipitated chromatin DNA was purified using spin columns and analyzed by real-time quantitative PCR to amplify the specific sequences (PCR product containing II genotype or DD genotype). The primer sequences are provided in Table S1.
Enzyme Linked Immunosorbent Assay (ELISA)
The VEGF-A levels in the supernatant of transfected MDA-MB-231 cells were measured using Human VEGF-A quantitative ELISA Kit (R&D Systems) according to the manufacturer's protocol. Then, the detection signals were quantified under a 450 nm wavelength. Each experiment was performed in triplicate.
Immunohistochemistry (IHC) assay
The immunohistochemistry (IHC) experiment was performed in clinical tissue sample or xenograft tumor tissues of mice using the ready-to-use Ultra Sensitive s-p immunohistochemistry kit (MXB, China) according to the manual instructions. The paraffin-embedded tissues were cut into 5 μm of slides, and deparaffinized with xylene, absolute ethanol, 95% ethanol, ddH2O. Then, the slides were repaired with sodium citrate solution and incubated with primary antibodys overnight at 4 °C. After incubation, sections were stained with DAB and counterstained with hematoxylin, finally, sealed by neutral resin. The associated primary antibodies used in IHC assay are included: GABPα (1:100 dilution, Santa Cruz Biotechnology Inc), VEGF-A (1:100 dilution, BOSTER), CD31 (1:100 dilution, CUSABIO).
Xenograft nude mice model
All mice used in this study were treated according to institutional guidelines approved by Animal Care and Use Committee of Wuhan University of Science and Technology. A total of 25 female BALB/c nude mice (4 weeks old) were used to establish the xenograft mice models, which were designed for five groups: untreated group, GABPα overexpression group, GABPα knockdown group and their respective controls. MDA-MB-231 cells stably GABPα overexpression or knockdown were cultured for injection. After trypsin digestion, the cells (3.0 × 106) were suspended with 100 μL PBS and mixed at a 1:1 ratio with 100 μL Matrigel basement membrane matrix (BD) for each group. The cell mixture was subcutaneously injected into the flank of BALB/c nude mice. Body weight and tumor volume were measured every three days. At 30 days post injection, the mice were anesthetized using diethyl ether and sacrificed by cervical dislocation. The tumors were dissected, weighed, and used for molecular detection and IHC analysis.
Statistical analysis
All statistical analysis in this study was performed using GraphPad Prism software (version 8.0.1). Data are presented as mean ± standard error of the mean (SEM). The differences between/among experimental groups were compared and analyzed using student's t-test or one-way ANOVA. The
Results
A novel 18-bp indel mutation of VEGF promoter was identified in breast cancer populations
A variety of mutations within

Identification and genotyping of an 18-bp indel mutation within
GABPα inhibited the transcriptional activity of VEGF gene by binding to the 18-bp indel
Substantial previous studies reported that the majority of sequence mutations within the core target sites of gene promoter might lead to transcriptional activity alteration and therefore partially or completely abolish related phenotypes.20,21 In this study, to assess whether the 18-bp indel disrupted binding sites of potential transcription factors, we predicted a series of critical regulatory factors within the indel locus and its adjacent sequences, including TATA box, SP1 or RBPJ binding sites. Surprisingly, GABPα binding site was identified in the insertion/insertion genotype (II), while it disappeared for DD genotype (Figure 2(a)). These

GABPα regulates the promoter activity of
To further confirm whether GABPα participate in the regulation of VEGF transcription through the 18-bp indel locus, we conducted the ChIP-qPCR assay in breast cancer cells. The primer pair spanning the indel mutation was designed and synthesized to amplify the product containing II or DD genotype. As shown in Figure 2(c), chromatin precipitation with GABPα antibody showed a significant high enrichment in the II group (
GABPα acts as a tumor suppressor in breast cancer
To investigate the functional role of GABPα in tumor progression, firstly, the expression pattern of GABPα in pan-cancers was analyzed using the Cancer Genome Atlas (TCGA) datasets. Importantly, we observed that the transcripts of GABPα were promoted in several kinds of primary tumors, including AML, esophageal cancer, pancreas cancer and gastric carcinoma, but the majority of other tumor presented a remarkably decrease of GABPα expression comparing to normal subjects, such as breast cancer, colon cancer and prostate cancer (Figure 3(a)). Specially, the comparable expression was conducted in paired or non-paired breast cancer and control samples. The results showed that a significant reduction of GABPα level was displayed in breast tissues (Figure 3(b) and (c)). In addition, similar patterns were found shown in Figure S1(a) to (c), which revealed that low GABPα expression was significantly correlated with N stage, M stage, and pathology stage comparing to the normal group. A Kaplan-Meier analysis indicated that the breast cancer patients with a high GABPα level exhibited a significant elevated overall survival rate than the individuals with a low GABPα level (HR = 0.65, Figure 3(d)). Based on the effect of GABPα on VEGF transcriptional activity, we hypothesized that there is a regulatory relation between GABPα and VEGF. Expression correlation analysis revealed that GABPα expression was negatively associated with the VEGF levels (R = −0.451, GSE29431, Figure 3(e)). The protein levels detected in the tissue sections using immunohistochemistry assay confirmed the bioinformatic results (Figure 3(f)). In addition, we conducted a qPCR assay to detect mRNA levels in distinct cell lines. In contrast to the normal mammary epithelial cells MCF-10A, the expression of GABPα depleted, while the VEGF increased in the breast cancer cell lines (MCF7 and MDA-MB-231) (Figure 3(g) and (h)). Thus, we speculated that GABPα may act as a tumor suppressor in breast cancer and participate in VEGF-involved biological process.

GABPα expression is negatively associated with breast cancer progression. (a) Differences of
Effects of GABPα on proliferation and apoptosis of breast cancer cells
Considering the suppressive role of GABPα in breast cancer, we hypothesized that GABPα may regulate the tumor pathophysiological process. First, we constructed the lentivirus vector to screen stable MDA-MB-231 cells with GABPα overexpression or depletion, which was validated by measuring the mRNA and protein levels (Figure 4(a) and (b)). The plate clonogenesis assay was established to assess the effects of GABPα on proliferation of breast cancer cells, and the results showed that the GABPα-overexpressed MDA-MB-231 cells displayed inhibitory clonal colonies compared to control lentivirus treated cells (Figure 4(c)), while depletion of GABPα presented the opposite results (Figure 4(d)). In addition, CCK-8 assay also revealed marked inhibition of cell viability in the GABPα overexpression group (Figure 4(e) and (f)), that was consistent with the role observed in the colony formation experiment. The results were further confirmed by examining the expression of proliferation related genes including Cyclin D1, c-myc and PCNA, which were significantly decreased/increased in response to overexpression/silence of GABPα (Figure 4(g) and (h)). Next, to determine whether GABPα is involved in breast cancer cell apoptosis, we carried out flow cytometry assay by annexin V-FITC/PI staining. The results showed that a high percentage of cell death and apoptosis was identified in MDA-MB-231 cells with GABPα overexpression, while the repressed apoptotic feature was detected in silencing GABPα group (Figure 5(a) and (b)). We also detected several key molecules related to cell apoptosis, as shown in Figure 5(c) and (d), a decreased expression of Bcl-2 but increased levels of Bax, Caspase-3 and Caspase-9 occurred in GABPα overexpressed MDA-MB-231 cells, and corresponding results were found in cells silencing GABPα. Collectively, these data suggested that GABPα inhibited proliferation and promoted apoptosis of breast cancer cells.
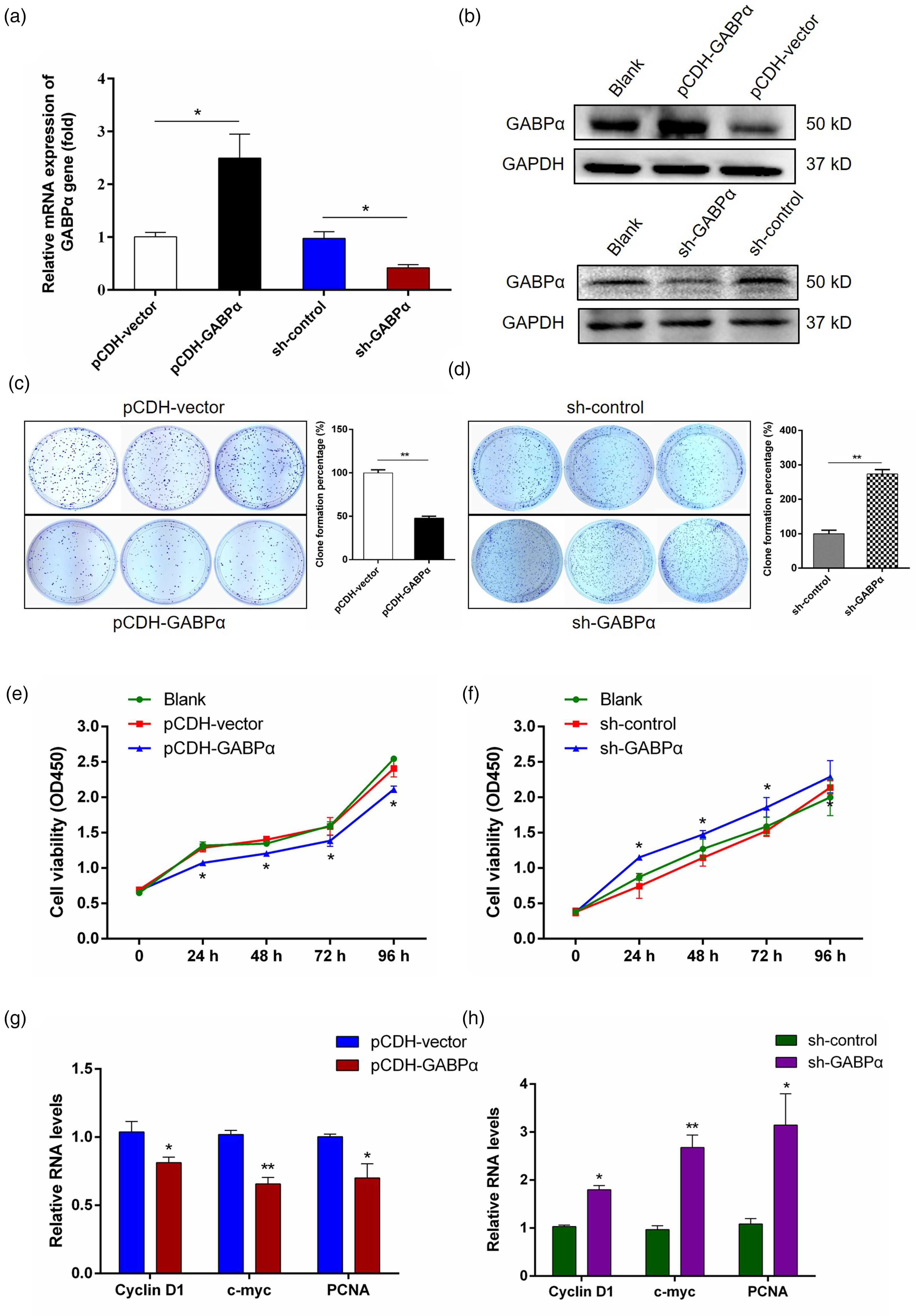
GABPα inhibited the proliferation of breast cancer cells. (a) Stable GABPα overexpression or knockdown was detected by qPCR assay. (b) Stable GABPα expression was validated by Western blot. Blank represented untreated group. (c-d) The effects of GABPα overexpression or depletion on cell proliferation were assessed by colony formation assay. (e-f) The proliferation of MDA-MB-231 cells was measured by CCK-8 assay (OD450 absorbance) when GABPα was overexpressed or repressed. (g-h) The mRNA expression of cell proliferation marker genes (Cyclin D1, c-myc, PCNA) was compared between GABPα overexpression and knockdown groups. Data are expressed as means ± SEM from at least three experiments. *

Effects of GABPα on cell apoptosis of breast cancer cells. (a) Cell apoptosis was detected by flow cytometry in MDA-MB-231 cells with GABPα overexpression or knockdown. (b) The quantized results of (a). (c-d) Expression differences of apoptosis involving genes in stable MDA-MB-231 cells. Data are expressed as means ± SEM from at least three experiments. *
Effects of GABPα on migration and invasion of breast cancer cells
Given the inhibitory role of GABPα in tumor cell proliferation, we then investigated whether GABPα could influence cell migration and invasion

GABPα inhibited migration and invasion of breast cancer cells. (a) The effects of GABPα overexpression on cell migration in MDA-MB-231 cells by wound healing assay. (b) GABPα knockdown promoted MDA-MB-231 cell migration. (c-d) The wound closure rate of breast cancer cells in replicates were quantified using Image J. (e-f) Cell invasion was demonstrated in MDA-MB-231 cells with GABPα overexpression or depletion by transwell assay. Data are expressed as means ± SEM from at least three experiments. *
GABPα inhibited angiogenesis of breast cancer cells by regulating VEGF
Given the regulatory role of GABPα in transcriptional activity of
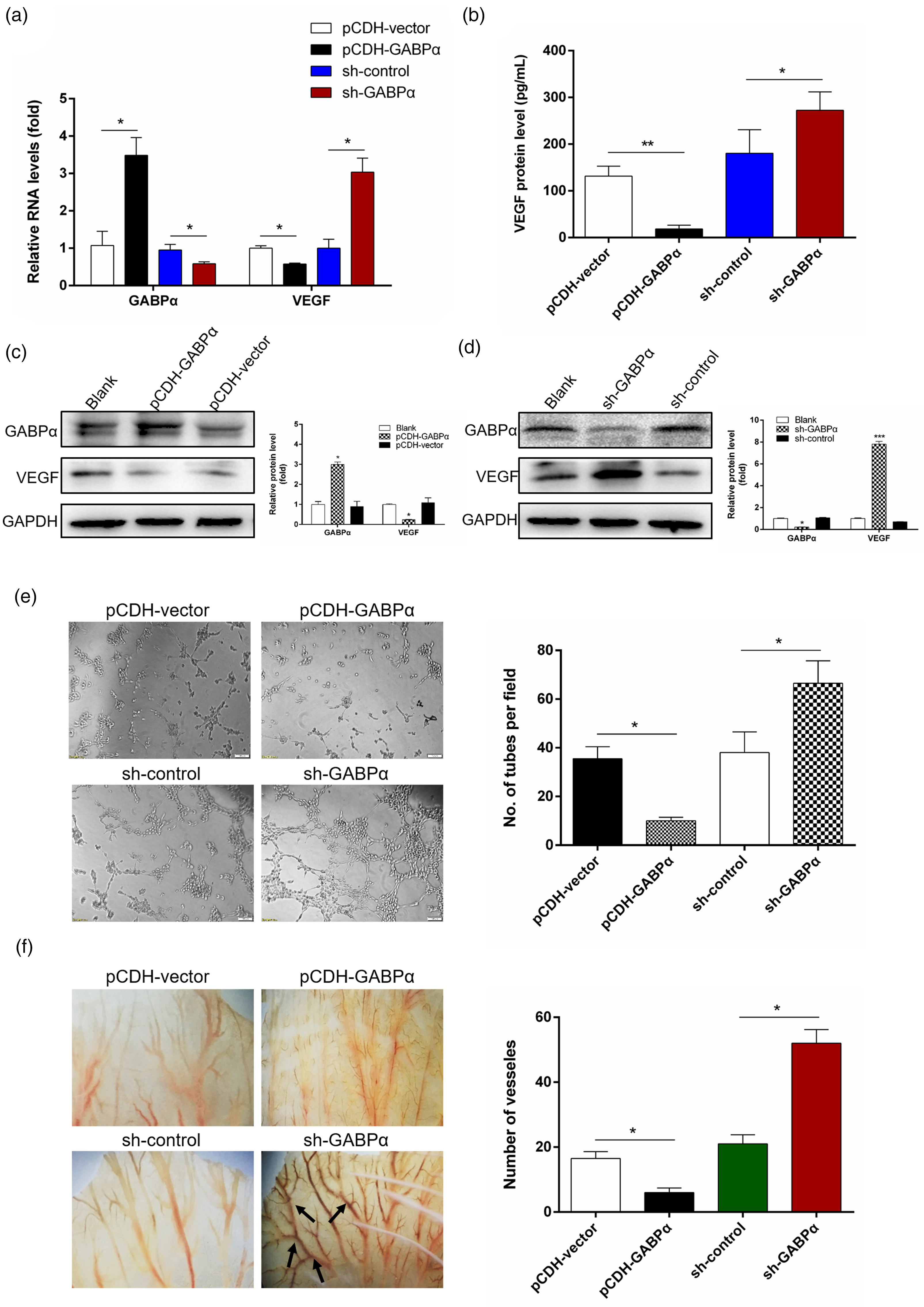
GABPα inhibited angiogenesis of breast cancer cells by regulating VEGF. (a) The mRNA expression of GABPα and VEGF in MDA-MB-231 cells with GABPα overexpression or depletion. (b) The secretory VEGF protein levels in supernatant in stable MDA-MB-231 cells with GABPα overexpression or depletion by ELISA. (c-d) The protein level of GABPα and VEGF in MDA-MB-231 cells with GABPα overexpression or depletion. (e) Effects of conditional medium from GABPα overexpressed or depleted MDA-MB-231 cells on tube formation in HUVECs. Quantification of tubes was displayed as bar graph. (f) Effects of conditional medium from GABPα overexpressed or depleted MDA-MB-231 cells on neovascularization by CAM assay. Bar graph showed the quantified blood vessels. Data are expressed as means ± SEM from at least three experiments. *
The effects of GABPα on breast cancer progression and angiogenesis in mice
In light of the anticancerous function

Effects of GABPα on tumor progression and angiogenesis in MDA-MB-231 tumor-bearing mice. (a-b) The body weight variations were reported in each treated mice group during the periods of subcutaneous tumor growth. (c) Representative image of tumor tissues in GABPα overexpression and the comparing control treated mice at the termination of the
Discussion
Though the diagnosis and treatment of breast cancer are remarkably advanced, the benefits of current therapy do not fulfill the positive expectation to reduce the morbidity and mortality. Thus, to identify the critical regulator or target and to elucidate the molecular mechanisms of carcinogenic process are paramount importance for breast cancer therapy. Microvascular architecture of tumor lesions is characterized as a hostile hallmark of cancer progression,
23
also being a fundamental synergy event for cancer invasion and metastasis.24,25 VEGF, a homodimeric glycoprotein, can induce new blood vessels of tumor by stimulating endothelial cell proliferation and migration.26,27 In the present study, our unprecedented discovery revealed that a novel 18-bp indel existed in the promoter region of
As a member of ETS family, GABPα is characterized by a conserved DNA binding domain that recognizes the
Malignant solid tumors are highly heterogeneous tissues composed of neoplastic cells as well as surrounding stromal cells, such as endothelial cells, immunosuppressive cells and perivascular mural cells, forming the complex tumor microenvironment.
35
Angiogenesis has been considered as a rate-accelerating event in tumor growth and metastatic dissemination, and the process of neovasculature involves incorporation of endothelial cells into sprouting blood vessels.36,37 Thus, an increasing area of cancer therapy focuses targeting angiogenesis process and proangiogenic factors to block tumor progression. VEGF family plays a crucial role in pathologic angiogenesis through regulating endothelial cell survival, migration, and differentiation.
38
Overexpression of VEGF has been reported to facilitate the maintenance of vasculature networks in several tumor types, such as breast cancer,
39
gastric carcinoma
40
and colorectal carcinoma.
41
In recent years, accumulated evidences demonstrate that many mutations existed in the
Gene transcription is launched by the specific binding of key upstream transcriptional factor to target gene promoter regions, thus sequence variants within the binding sites may lead to either elimination of the interaction effects, or alteration of regulatory efficiency, finally cause aberrant phenotypes, ranging from physiological dysfunction to disease predisposition.45,46 For instance, E2F1 transcription factor binding was significantly decreased due to an SNP locus located in TERT promoter, which was associated with poor prognoses of hepatocellular carcinoma.
47
Zhang et al.
48
revealed that the T allele of a point mutation (−6C > T) in Mtor gene promoter created a consensus recognition element of MZF1 factor, and the regulatory roles linked to plasmacytoma risk in mice. Moreover, the 405 G > C SNP locus of VEGF promoter was correlated with aerodigestive carcinogenesis by altering the MZF1-binding specificity.
49
Of note, our study first reported a novel 18-bp indel mutation in the VEGF promoter, and the DD genotype lead to eliminate the binding sites of GABPα protein, contributing to the progression and angiogenesis of breast cancer. Furthermore, the EMSA and ChIP assays were used to confirm the combination and interaction of GABPα with VEGF promoter sequence containing the 18-bp indel. Similarly, a previous study reported the negative effect of GABPalpha/beta on
Conclusions
To conclude, in the present study, we highlighted a novel 18-bp indel mutation, existed in the promoter of
Supplemental Material
sj-docx-1-cbm-10.3233_CBM-230541 - Supplemental material for GABPα inhibits tumor progression and angiogenesis via a novel 18-bp indel within VEGF promoter in breast cancer
Supplemental material, sj-docx-1-cbm-10.3233_CBM-230541 for GABPα inhibits tumor progression and angiogenesis via a novel 18-bp indel within
Footnotes
Abbreviations
Acknowledgements
The authors would like to thank Dr Yuan Cao at The Medicine & Sciences Analysis Center of Wuhan University of Science and Technology (Wuhan, China) for their help with immunohistochemistry imaging and analysis.
Ethics approval and consent to participate
This study was approved by the Ethics Committee at People's Hospital Affiliated to Shandong First Medical University, and the written informed consents were obtained for sample collection.
Consent for publication
Not applicable.
Author contributions
Conception: author Hui Guo, Yue Han, Qiaoyi Zhou, and Yao Xu.
Interpretation or analysis of data: author Jihua Chen, Zhenrong Wang and Fan Li.
Preparation of the manuscript: all authors.
Revision for important intellectual content: author Hui Guo, Mengyi Wang, and Xiaotong Deng.
Supervision: author Yao Xu.
Funding
This study was supported by the Foundation of Hubei Province Supporting Enterprise Technology Innovation Development (2021BAB126), Wuhan East Lake High-tech Zone “JieBangGuaShuai” Project (2022KJB113), Foundation of Wuhan University of Science and Technology (2016xz036).
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Availability of data and materials
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Supplemental material
Supplemental material for this article is available online.
