Abstract
BACKGROUND:
There is a lack of effective biomarkers that predict immunotherapy efficacy in clear cell renal cell carcinoma(KIRC).
OBJECTIVE:
We aimed to identify biomarkers that would predict the efficacy of KIRC treatment with immune checkpoint inhibitors (ICIs).
METHODS:
Cohort data of KIRC patients with somatic mutations, mRNA expression and survival data from The Cancer Genome Atlas (TCGA) database and immunotherapy cohort and Genomics of Drug Sensitivity in Cancer (GDSC) database were analyzed and divided into interleukin 3 (IL3) pathway-related genes high expression (IL3-High) and IL3 pathway-related genes low expression (IL3-Low) groups according to pathway expression status to assess the relationship between the IL3 pathway-related genes activation status and the prognosis of KIRC patients treated with ICIs. The data were validated by immunohistochemistry experiments, and possible mechanisms of action were explored at the level of gene mutation landscape, immune microenvironment characteristics, transcriptome and copy number variation(CNV) characteristics
RESULTS:
The IL3 pathway-related genes was an independent predictor of the efficacy of ICIs in KIRC patients, and the IL3-High group had a longer overall survival (OS); KIRC patients in the IL3-High group had increased levels of chemokines, cytolysis, immune checkpoint gene expression and abundant immunity. The IL3-Low group had poor immune cell infiltration and significant downregulation of complement activation, cytophagy, B-cell activation, and humoral immune response pathways. The high group was more sensitive to targeted drugs of some signaling pathways, and its efficacy in combining these drugs with immunity has been predicted in the published literature.
CONCLUSION:
The IL3 pathway-related genes can be used as a predictor of the efficacy of ICIs in KIRC. The IL3 pathway-related genes may affect the therapeutic efficacy of ICIs by affecting the expression of immune-related molecules, immune cell infiltration, and the level of immune response pathways.
Keywords
Introduction
Kidney cancer is one of the top ten most common cancers in both men and women, accounting for 3.7% of all new cancer cases[1]. KIRC is the most common form of kidney cancer, with a prevalence of up to 85 percent; it is more prevalent in males than in females (ratio of 1.7:1), and the majority of patients are older, with a mean age of 64 years[2]. Although the 5-year relative survival rate at the time of diagnosis has improved, the overall prognosis is still poor, especially for patients with high staging[3]. In recent years, the discovery of immune checkpoint molecules including programmed cell death-1 (PD-1), programmed death-ligand 1 (PD-L1), and cytotoxic T lymphocyte-associated protein 4 (CTLA-4), has led to the development of ICIs for treating late-stage KIRC, which has ushered in a new era in the management of advanced cases. The overall response rate of KIRC with ICIs is approximately 40%, indicating that most patients do not benefit from ICIs treatment[4]. Therefore, identifying biomarkers to assess immunotherapy efficacy is particularly important for predicting therapeutic response. Recent studies have shown that PD-L1 expression, tumor mutation load (TMB), microsatellite instability, mismatch repair gene defects, special gene mutations (e.g., EPHA5)[5], the tumor immune microenvironment, gene expression profiles (GEPs), and antigen presentation defects may be predictive markers of ICI efficacy[6]. However, certain predictors still have limitations. For example, PD-L1 immunohistochemical measurements are more subjective, and some markers have no obvious cutoff values (e.g. EPHA5). Therefore, finding new markers with clearer cutoff values that can predict the treatment response to ICIs is the focus of our study.
The activation or inhibition of specific pathways is associated with cancer immunotherapy efficacy. For example, the accumulation of DNA damage response (DDR) pathway mutations[7] and the inhibition of the folate pathway[8] are used as biomarkers for predicting immunotherapy efficacy. However, no pathways to serve as biomarkers have been reported in KIRC that predict the efficacy of ICIs.
We screened the IL3 pathway-related genes as a predictive pathway for the efficacy of KIRC immunotherapy through bioinformatics and analysis and validated the immunopredictive role of the IL3 pathway-related genes in other cancer types. IL3 is a cytokine belonging to the interleukin family[9]. Intracellular signal transduction initiated by the IL3 pathway[10] depends on the receptor subunits IL3-R
However, the relationship between the IL3 pathway-related genes and prognosis in KIRC patients receiving ICI treatment is unclear. In this study, we further explored the possible molecular mechanism focusing on how IL3 pathway-related genes levels affect the therapeutic efficacy of ICIs.
Methods and materials
IL3 pathway-related genes activation and prognosis in KIRC patients receiving immunotherapy
To assess the relationship between IL3 pathway-related genes activation and ICI efficacy in KIRC patients, data from the Miao cohort[18] and the IMvigor210 cohort[19], in which patients were receiving ICIs (anti-PD1/anti-PD-L1 or combined antiCTLA-4), were collected. The “TCGAbiolinks”[20] package was used to download the cancer genome map of the renal cell carcinoma (TCGA-KIRC) cohort, which included mRNA expression profile, somatic mutation, CNV, and clinical prognosis data. The unit of gene expression was pancancer standardized log2 (the number of thousand base fragments per million mapping fragments [FPKM]
Mutation analysis
Mutation data from the IMvigor210 and TCGA-KIRC cohorts were used to analyze correlations between IL3 pathway-related genes activation, mutations in other genes, and other clinical features. The R package “ComplexHeatmap”[28] was used to display mutations (the top 20 genes with the highest mutation rates were selected) and clinical features from the IMvigor210 and TCGA-KIRC cohorts. The “Maftools”[29] package was used to identify sites with significant mutations at different levels of IL3 pathway-related genes activation.
Correlation analysis of the immune microenvironment characteristics
Immune cell infiltration status and immune-related gene expression were assessed in the TCGA-KIRC cohort with the R package “IOBR”[30] using the CIBERSORT, MCP, xCell, TIMER methods. The expression levels of these genes were quantified as log2 (FPKM
Copy number variation analysis
Gistic2.0[32] was used to analyze the data using the platform GenePattern (
Drug sensitivity analysis
It has now been shown that the use of large-scale drug sensitivity profiles in conventional cancer cell line models to identify clinically relevant biomarkers[33, 34]] that can be used for treatment provides an opportunity to design effective clinical trials and combination therapeutic strategies.
Here, we downloaded drug sensitivity information for 198 drugs from GDSC[35], cell line half-maximal inhibitory concentration (IC50) data, and 19 sets of KIRC cell line transcriptomics data (Supplementary Table 2) and entered the downloaded expression matrix as a training set using the R package “oncoPredict”[36]. Data from the TCGA-KIRC cohort were selected, grouped according to IL3 pathway-related genes status, and then used as the expression matrix for which predictions were made. The training set and the pharmacoresensitivity data matrix were combined and analyzed using the calcPhenotype[37] function in R to derive pharmacoresensitivity IC50 information for the KIRC cell lines according to the IL3-High and IL3-Low groups.
Genome enrichment analysis
Gene expression data from the TCGA-KIRC cohort were analyzed by the “edgeR”[38] package. The “clusterProfiler”[39] package was used for gene annotation enrichment analysis. This included Gene Ontology (GO), Kyoto Encyclopedia of Genes and Genomes (KEGG), and Reactome pathway analyses.
Immunohistochemistry
Tumor tissues were collected from six KIRC patients who had been treated with ICIs and were diagnosed at Zhujiang Hospital of Southern Medical University between May 2014 to January 2022. Primary antibodies against ERK1/2, STAT5, CD4, CD8, PD-1 and IL3RA were purchased from Wanleibio, and secondary antibodies were purchased from Guangzhou Yongjin Biotechnology Co. Hematoxylin and eosin (H&E)stained slides from each tumor before ICI treatment were examined, and the slides with the most abundant viable renal cell carcinoma cells were selected from each tumor. Formalinfixed and paraffin-embedded samples corresponding to the selected slides were collected from the archives of Zhujiang Hospital of Southern Medical University, and the tissue specimens were cut into 4
Statistical analysis
The Mann-Whitney U test was used to assess the relationships between IL3 pathway-related genes activation and TMB, immune gene[40] expression, and immune cell infiltration. Fisher’s exact test and the chi-square test were used to evaluate the association between the mutation frequency among the top 20 genes and IL3 pathway-related genes activation. The Kaplan-Meier method, log-rank test, and univariate and multivariate Cox proportional hazards regression analyses were used to analyze the effect of IL3 pathway-related genes activation on clinical prognosis. Spearman correlation analysis was used to further assess the association between the IL3 pathway-related genes enrichment score and immune cell infiltration score.
Results
IL3 pathway-related genes activation is a predictive biomarker for ICI response in KIRC patients
Univariate survival analysis showed that the activation status of the IL3, DNA repair, protein secretion, and IGFI pathways was associated with prognosis in ICI patients and that differences in prognosis were statistically significant (Fig. 1A;
The IL3 pathway-related genes is associated with prognosis in KIRC patients receiving ICIs. A) Results of the univariate Cox analysis in the Miao and IMvigor210 cohorts; B) Results of the multivariate Cox analysis (
To further verify the predictive function of IL3 pathway-related genes activation and TMB on OS in patients receiving ICIs, a Kaplan-Meier curve was used to assess the combined prognostic effectiveness of IL3 pathway-related genes activation and TMB status in predicting OS in each cohort. In the IMvigor210 cohort, patients who were grouped as IL3-High and TMB-High had a longer OS than IL3-Low and TMB-Low patients (Fig. 1F, Figure S2B, group 1 vs. group 4,
To determine whether IL3 pathway-related genes activation status can be used as a biomarker to predict ICI efficacy in other cancers, Kaplan-Meier analyses were performed for the GSE91061 ICI cohort for melanoma and the GSE176307 ICI cohort for bladder cancer. These analyses revealed that OS was longer in the IL3-High group than in the IL3-Low group (Fig S3,
We analyzed the distribution of sex, ethnicity, and TNM stage in the IMvigor210 and TCGA-KIRC cohorts and found no differences between the IL3-High and IL3-Low groups (Fig. 2A and B). Next, we compared mutational data from patients in the TCGA-KIRC cohort who were not treated with ICIs and those in the IMvigor210 cohort who received ICIs. According to IL3 pathway-related genes activation status, differences in the mutation status for the top 20 genes with the highest mutation frequencies were compared (all
Association between IL3 pathway-related genes activation, gene mutations, and clinical features. A) The top 20 most frequently mutated genes in the TCGA-KIRC cohort. Genes were ranked according to mutation frequency in KIRC patients. Change type, IL3 pathway-related genes status, sex, OS time, clinical TNM stage, and ethnicity were annotated. Significant differences in mutations or clinical features are highlighted by asterisks (significance calculated using Fisher’s exact test, 
To explore the immune features in TCGA-KIRC cohort patients and the activation of the IL3 pathway-related genes, the relative expression of immunerelated genes (chemokines/cytolytic activity/immune checkpoint) in the IL3-High and IL3-Low groups were compared and determined to be higher in the IL3-High group (
Immunogenicity and immune cell infiltration patterns based on IL3 pathway-related genes activation status. A) Mean differences in mRNA expression of immune-related genes between IL3-High and IL3-Low tumors in the TCGA-KIRC cohort. The Y-axis indicates the immunolabel, or gene name. The filled numbers in the graph represent the statistics, the number fill color is LogFC, where FC 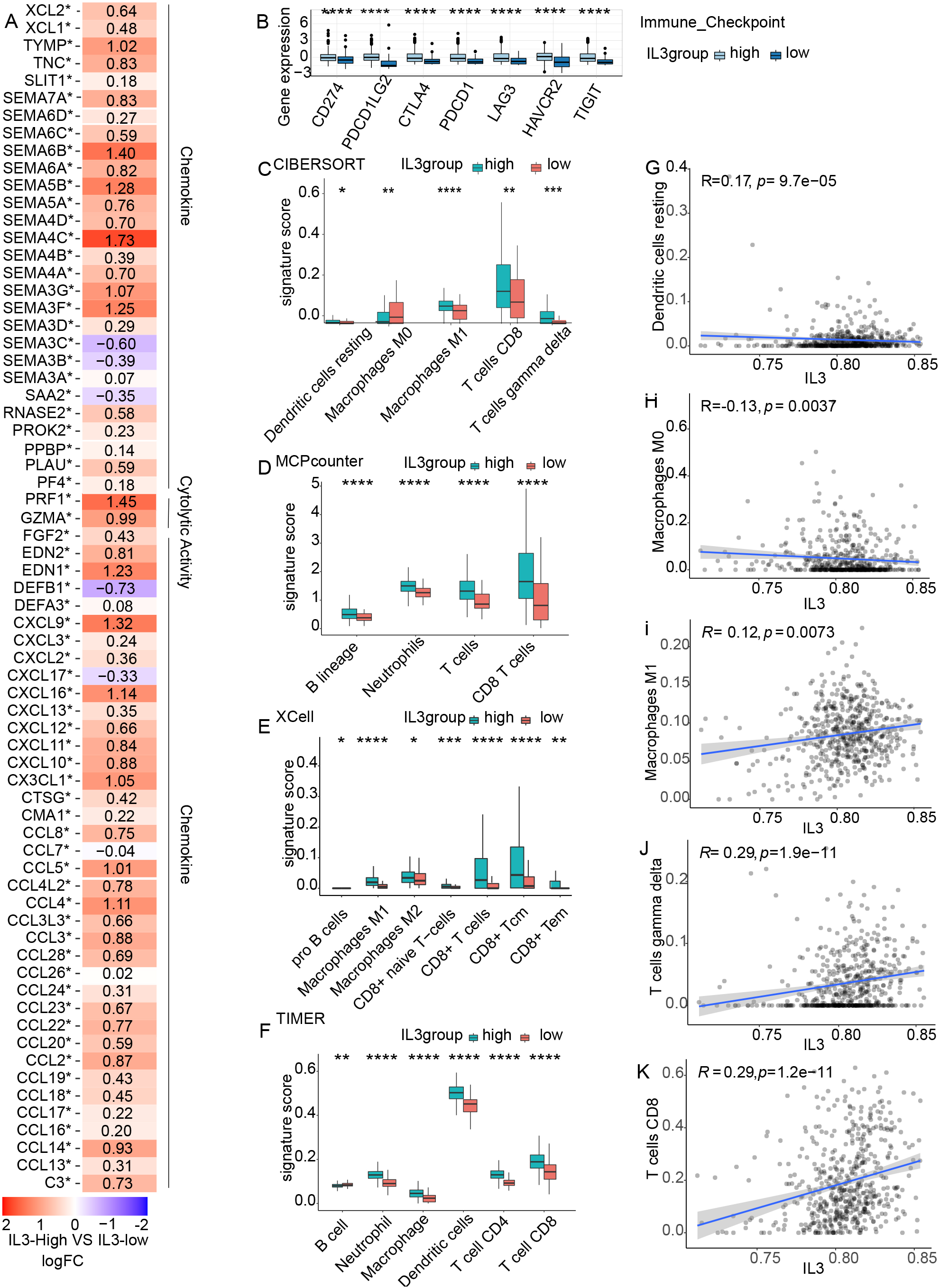
Transcriptomic data from patients in the TCGA-KIRC and GSE167573 cohorts who did not receive ICIs were analyzed by GSEA, and associations between IL3 pathway-related genes activation and transcriptome characteristics were assessed. In both cohorts, 22 pathways were significantly enriched in the IL3-Low group (Fig. 4A and B, Fig. S4,
Analysis of transcriptome signatures and copy number variations based on IL3 pathway-related genes activation status. Differences in pathway activity of GSEA scores between the IL3-High and IL3-Low tumors from patients in A) the TCGA-KIRC cohort and B) the GSE167573 (KIRC) dataset. Enrichment results showed a significant correlation between IL3-High and IL3-Low tumors. The orange bar indicates that the enrichment score (ES) of the pathway is less than 0. All transcripts were ranked by log2 (fold change) between IL3-High and IL3-Low tumors. Each run was performed using 1000 permutations. C) and D) Copy number variation in IL3-High and IL3-Low tumors in the TCGA-KIRC cohort, where red represents copy number fragment gain, blue represents copy number fragment loss, and the mutation site has been marked.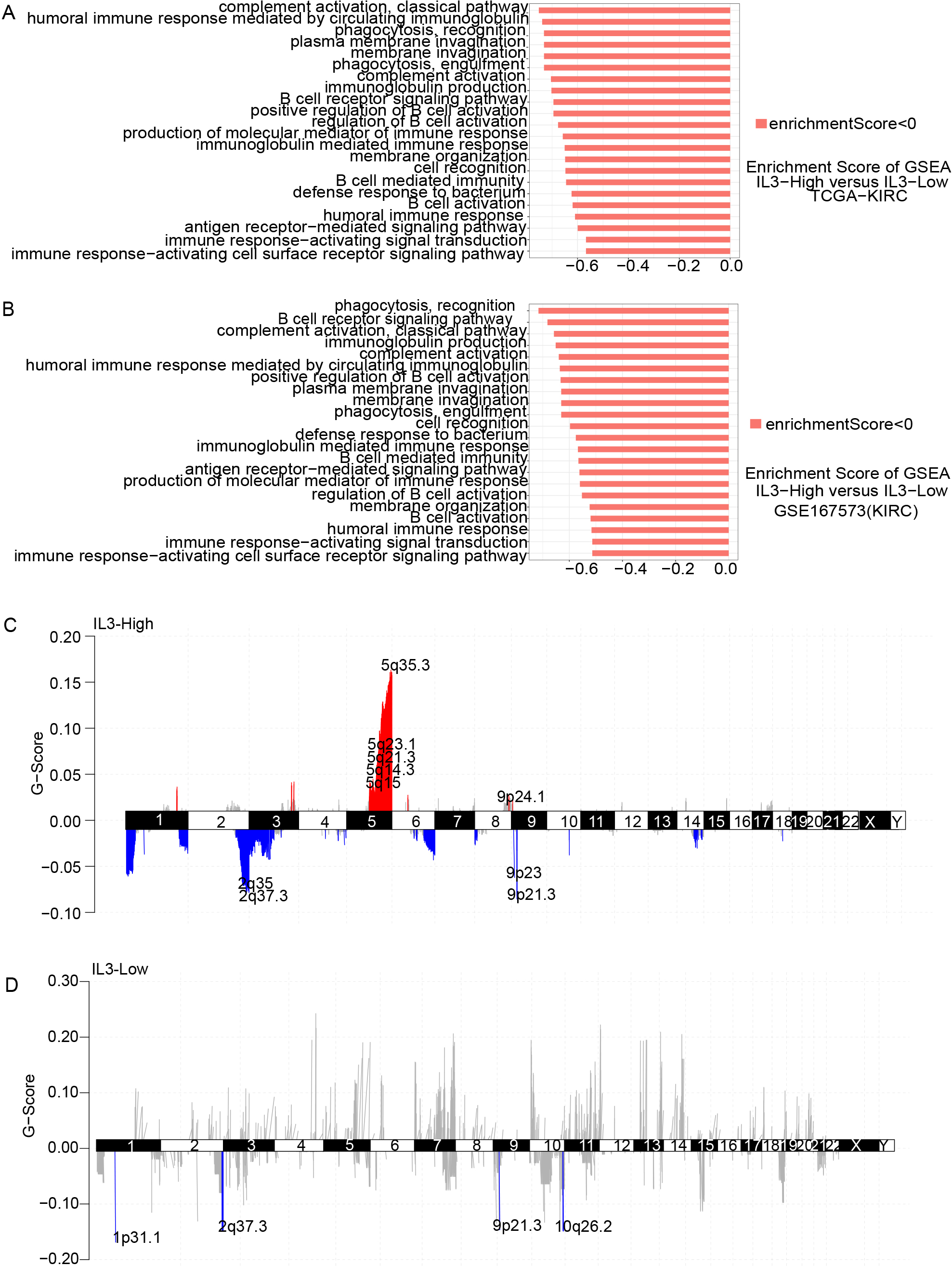
We also analyzed CNV data from the TCGA-KIRC cohort (Fig. 4C, 4D). Increased copy numbers of 5q35.3, 5q23.1, 5q21.3, 5q15, 5q14, and 9p24.1 and decreased copy numbers of 2q35, 2q37.3, 9p23, and 9p21.3 were observed in the IL3-High group, and decreased copy numbers of 1p31.1, 2q37.3, 9p21.3, and 10q26.2 were observed in the IL3-Low group. The distribution and peak of amplified/deleted regions were significantly higher in the IL3-High group than in the IL3-Low group.
According to the analysis of 198 anticancer drugs found in the GDSC, the IC50 values of 15 drugs were observed to increase in IL3-Low KIRC cell lines, indicating lower sensitivity than in the IL3-High KIRC cell lines (all
Drug sensitivity results for the KIRC cell line in the GDSC database. Box plots showing the IC50 of IL3-High and IL3-Low KIRC cell lines against targeted drugs in the GDSC database. The X-axis represents IL3 expression groups, and the Y-axis represents the drug IC50 score, measured in M. Differences were assessed by the Mann–Whitney U test, ns 
Immunohistochemical analysis of the IL3 pathway-related molecules ERK1/2, STAT5, CD4, CD8, PD-1 and IL3RA in local immunotherapy samples. A) Expression of ERK1/2, STAT5, CD4, CD8, PD-1 and IL3RA in immune-responsive samples; B) Expression of ERK1/2, STAT5, CD4, CD8, PD-1 and IL3RA in nonimmune-responsive samples; microscope magnification 
We performed immunohistochemical assays of the protein expression profiles of IL3 pathway-related molecules versus immune infiltration-related molecules in six KIRC immunotherapy samples (Fig. 6, Table 1). The results showed that the IL3 pathway-related molecules ERK1/2, IL3RA, and STAT5 and the immune infiltration-related molecules CD4 and CD8 were most highly expressed in the ICI-responsive (partial response, PR) KIRC patient samples, whereas they were weakly expressed or not expressed in the ICI-naïve (progressive disease, PD; stable disease, SD) patient samples. The PD-1 molecule was not expressed in most of the samples.
Results of immunohistochemical experiments in KIRC patients receiving immunotherapy
Results of immunohistochemical experiments in KIRC patients receiving immunotherapy
Response: efficacy of treatment with immune checkpoint inhibitors; SD: stable disease; PD: progressive disease; PR: partial response. Patient refers to a tumor sample from a patient with renal clear cell carcinoma that received immunotherapy.
In recent years, ICIs have shown promising prospects for the treatment of KIRC. However, most patients with KIRC receiving ICIs do not launch an effective antitumor immune response, and there is a lack of accurate biomarkers for assessing ICI efficacy in KIRC. In the present study, we found that IL3 pathway-related genes activation can be used as a predictor of response to ICIs in patients with KIRC. Compared with IL3 pathway-related genes low expression patients, IL3 pathway-related genes high expression patients showed significantly better OS. Based on IL3 pathway-related genes activation, we explored the possible factors that may influence ICI effectiveness in KIRC patients. Our study explores the mechanisms that may influence immune efficacy through the genetic, immune microenvironment, and copy number variation characteristics levels.
At the genetic level, as derived from the analysis, the frequency of VHL, ERBB3, FGFR3, mTOR and CDKN2A gene mutations was significantly higher in the IL3 pathway-related genes high expression group; however, the mutation frequencies of FGFR3, mTOR, and CDKN2A were higher in the IL3 pathway-related genes low expression group. It has been reported that VHL mutations can regulate the innate immune pathway of tumor promotion and antitumor response[41]. ERBB3 induces the infiltration of CD4+ T cells, CD8+ T cells, and neutrophils[42]. FGFR3 can reduce PD-L1 protein levels by affecting ubiquitination[43], and FGFR3 mutations may be associated with poor T-cell infiltration[44]. mTOR is a poor prognostic factor in KIRC patients[45], and the deletion of CDKN2A can eliminate innate and immune checkpoint blockade (ICB)-induced cancer immune control[46]. The above results suggest that high expression of the IL3 pathway-related genes is more likely to promote antitumor innate immune pathways and enhance immune cell infiltration to enhance ICI efficacy, while low expression of the IL3 pathway-related genes is more likely to affect PD-L1 expression levels and may also cause poor immune cell infiltration, imbalance in cell cycle regulation, and enhanced immune escape. The combination of these factors makes the outcome of patients with low IL3 pathway-related genes expression even worse.
At the immune microenvironment level, research showed that in the IL3 pathway-related genes high expression group, the tumor immune microenvironment (TIME) had high immune cell infiltration (including CD4+ T cells, CD8+ T cells, dendritic cells, MI macrophages, and M2 macrophages), and the expression of immunerelated genes (such as immune checkpoints and chemokines) was significantly upregulated. In addition, immune responserelated pathways were significantly downregulated in IL3 pathway-related genes low expression tumors. According to the status of immune cells in the TIME, tumor immune infiltration patterns can be roughly divided into three types: immune inflammatory type, immune exclusion type, and immune desert type. The immunoinflammatory type is characterized by the presence of CD8+ and CD4+ T cells in the tumor parenchyma and is accompanied by the expression of immune checkpoint molecules. It is suggested that these tumors have the greatest antitumor immune responses to ICIs[47]. The IL3 pathway-related genes high expression immune microenvironment is inflammatory and may result in a better antitumor immune response. T-cell recruitment of chemokines, such as CCL5, CXCL10, and CXCL11, may benefit patients with T-cell inflammatory tumors receiving checkpoint blockade therapy[48, 49, 50]. B-cell activation, humoral immune response, and phagocytosis-related pathways were significantly downregulated in IL3 pathway-related genes low expression, which may partially explain the poor prognosis of KIRC patients with low IL3 pathway-related genes expression after treatment with ICIs. In terms of in vitro experiments in mice it has been reported in the literature that by increasing IL3/anti-IL3 antibody complexes, basophils can be induced and the infiltration of tumor-specific CD8+ T cells can be enhanced[51]; the results of in vitro experiments in another study point to the fact that by enforcing the expression of IL3 in tumor cells, monocytes can be differentiated into tolerant dendritic cells, decreasing the frequency of regulatory T cells and induction of immunogenic dendritic cells in tumors, thus improving host immune surveillance[52]. In terms of human cellular experiments cellular experiments on human peripheral blood revealed that IL3 enhances the proliferation and survival of B and T cells and increases the release of proinflammatory cytokines from B and T cells[53]; IL3 enhances IFNGR2 (one of the strands of the IFNGR heterodimeric complex) and CD274, thus augmenting IFN-
At the copy number variation level, the results showed that increased copy numbers of 5q35.3 and 9p24.1 and decreased copy numbers of 2q35 were observed in the IL3 pathway-related genes high expression group, while decreased copy numbers of 9p21.3 were observed in the IL3 pathway-related genes low expression group. Studies have reported that amplification of chromosomal region 5q31-5q35.3 is strongly associated with OS in ovarian cancer, and SOX30 located in this chromosomal region has been identified as a new important tumor suppressor[58]. In relapsed/refractory classical Hodgkin’s lymphoma, 9p24.1 gene alterations were associated with a significantly higher objective response rate to anti-PD-1 monotherapy[59]. In some cancers, the 2q33-q36 deletion is associated with tumor suppression[60]. 9p21.3 homozygous deletion (9p21 deletion), one of the most frequent genomic defects occurring in approximately 13% of all cancers, results in a ’cold’ tumor immunophenotype characterized by a reduced abundance of tumor-infiltrating leucocytes (TILs), particularly T/B/NK cells, reduced PD-L1 and the activation of immunosuppressive signaling[61]. In summary, chromosomal copy number variants in the IL3 pathway-related genes high expression group tend to cause tumor suppression, enhance T-cell infiltration and strengthen the efficacy of anti-PD-1 therapy. chromosomal variants in the IL3 pathway-related genes low expression group may cause a reduced abundance of tumor infiltrates, decreased PD-L1 activity rates, and immunosuppression.
We found that the IL3 pathway-related genes high expression group had significantly increased sensitivity to drugs targeting ERK1/2, PARP1/2, WEE1, SYK, VEGFR, and mTOR. Reports have shown that ERK inhibitors, PARP1/2 inhibitors, and WEE1 inhibitors can enhance the efficacy of PD-1 inhibitors for the treatment of certain cancers[62, 63, 64]. In KIRC, the efficacy of ICIs combined with VEGFR inhibitors was higher than that of VEGFR inhibitors alone[65], while the second-line use of VEGFR inhibitors after progression on first-line ICIs was observed to prolong the OS of patients[66]. The combination of mTOR inhibitors and anti-PD-1 synergistically inhibited tumor growth and induced immunogenic cell death of tumor cells[67]. Inhibition of SYK kinase activity affects a variety of innate and adaptive immune responses[68]. There is potential to combine these compounds with immunotherapy in IL3-High KIRC patients to further improve patient prognosis. These compounds may be effective in combination with ICIs for KIRC patients suffering from the IL3 pathway-related genes high expression group, which offers a potential new approach to targeted drugs in combination with ICIs for anti-tumour therapy.
Although PD-1 is used as a conventional antitumor detection target, its high expression for ICIs is usually considered to have a better prognosis. The results of several recent studies of PD-1 monoclonal antibodies in combination with chemotherapy have demonstrated that PD-1 monoclonal antibodies in combination with chemotherapy treatment can provide a significant survival benefit to tumor patients over chemotherapy alone, even if PD-1 detection is negative or if there is no condition to detect PD-1[-]. In the current trial, we found that PD-1, although also an immune infiltration-related molecule, was largely unexpressed in most samples. In this regard, we suggest that the IL3 pathway-related genes may serve as a superior biomarker for identifying the efficacy of ICIs when PD-1 expression is negative.
Our study has the following limitations: due to the limited number of KIRC cancer patients receiving ICIs, the relationship between IL3 pathway-related genes activation and the prognosis of ICIs was tested only in the IMvigor210 and Miao cohorts. Beyond this, this study lacks further in vitro or in vivo experimental validation of the expression of proteins belonging to the target pathway in cell lines and the possible correlation between their expression and IL3 pathway-related genes high/low expression status. Finally, the IL3 pathway as a predictive factor of efficacy in PD-1 negative KIRC patients, needs further study.
Conclusions
Our findings suggest that IL3 pathway-related genes activation may be a novel prognostic marker for KIRC patients receiving ICI treatment. In addition, our study found that IL3 pathway-related genes high expression patients have an inflammatory immune microenvironment and the upregulated immune checkpoint molecules and T-cell recruitment chemokines. The above findings can provide a better reference for the clinical selection of ICIs to treat KIRC patients and provide ideas on whether to enhance ICI efficacy and reverse immune resistance through IL3 pathway-related genes activation (for example, TLR4 agonist, IL18).
Author contributions
Conception was performed by Shuang Hou. Interpretation or analysis of data was performed by Shuang Hou, Tianqi Gu Ying Shi, Jiarong Yao and Yushan Huang. Preparation of the manuscript was performed by Shuang Hou. Revision for important intellectual content was performed by Manming Cao, Anqi Lin, and Jian Zhang. Supervision by Weiliang Zhu. All authors have read and approved the final manuscript.
Funding
This work was supported by the Natural Science Foundation of Guangdong Province (Grant Nos. 2018A030313846 and 2021A1515012593) and the National Natural Science Foundation of China (Grant Nos. 81802257, 81871859, 81772457, 82172750 and 82172811).
Competing interests
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Data availability
The code and raw data used in this study are available online at
Other relevant GSE datasets are available online at the following addresses: GSE91061:
GSE176307:
GSE167573:
Ethical approval
All data used in this study were anonymous. The families of six KIRC patients from whom tumor tissue was obtained were contacted by telephone (four patients had died and two survived). Verbal informed consent was obtained from each patient. The process of this study followed the 1964 Helsinki Declaration guidelines and was reviewed and approved by the Medical Ethics Committee of Zhujiang Hospital, Southern Medical University (accessible online at zjyyll.wetrial.com, approval No. 2020-YW-053-01). The ethical approval letter and the address of the ethics committee can be found in Supplementary 3.
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-230226.
sj-pdf-1-cbm-10.3233_CBM-230226.pdf - Supplemental material
Supplemental material, sj-pdf-1-cbm-10.3233_CBM-230226.pdf
sj-pdf-2-cbm-10.3233_CBM-230226.pdf - Supplemental material
Supplemental material, sj-pdf-2-cbm-10.3233_CBM-230226.pdf
sj-xls-1-cbm-10.3233_CBM-230226.xls - Supplemental material
Supplemental material, sj-xls-1-cbm-10.3233_CBM-230226.xls
sj-xls-2-cbm-10.3233_CBM-230226.xls - Supplemental material
Supplemental material, sj-xls-2-cbm-10.3233_CBM-230226.xls
sj-xls-3-cbm-10.3233_CBM-230226.xls - Supplemental material
Supplemental material, sj-xls-3-cbm-10.3233_CBM-230226.xls
Footnotes
Acknowledgments
The authors would like to acknowledge the Zhujiang Hospital of Southern Medical University for providing equipment and technical support for immunohistochemical experiments.
