Abstract
BACKGROUND:
Immunogenomics approaches to the characterization of renal cell carcinoma (RCC) have helped to better our understanding of the features of RCC immune dysfunction. However, much is still unknown with regard to specific immune interactions and their impact in the tumor microenvironment.
OBJECTIVE:
This study applied chemical complementarity scoring for the TRB complementarity determining region-3 (CDR3) amino acid sequences and cancer testis antigens (CTAs) to determine whether such complementarity correlated with survival and the expression of immune marker genes.
METHODS:
TRB recombination reads from RCC tumor samples from RNAseq files obtained from two separate databases, Moffitt Cancer Center and The Cancer Genome Atlas (TCGA), were evaluated. Chemical complementarity scores (CSs) were calculated for TRB CDR3-CTA pairs and survival assessments based on those CSs were performed.
RESULTS:
Moffitt Cancer Center and TCGA cases representing the upper 50
CONCLUSIONS:
These results indicate that TRB CDR3-CTA chemical complementarity scoring may be useful in distinguishing RCC cases with a productive, anti-tumor immune response from cases where basic immune parameter assessments are inconsistent with a productive immune response.
Keywords
Introduction
Numerous and varied genomics approaches have been applied for the analysis of renal cell carcinoma (RCC) In one approach, sequencing of primary tumors was used to track RCC evolution and to study the relationship between intratumor heterogeneity and patient outcomes, consistent with the conclusion that evolutionary subtypes determined by driver mutations or chromosomal abnormalities play a role in patterns of metastases and outcomes [1]. Many immunology approaches have focused on categorizing immune dysfunction in metastatic RCC, with worse prognosis indicated by terminally exhausted CD8+ T cells, decreased levels of pro-inflammatory macrophages, and increased levels of suppressive M2-like macrophages [2, 3]. Interestingly, worse prognosis has been found to be associated with high CD8+ T cell infiltration indicating that immune dysfunction may have a critical function in RCC tumor aggressiveness [4]. CD8+ T cell-related molecular classifications have been suggested as possible predictors of prognosis and immunotherapeutic efficiency but thus far in RCC have been suboptimal guides [2, 5]. Additionally, a predictive model for prognosis and immunotherapeutic responses based upon 14 immune-associated genes has recently been suggested as an effective alternative for predicting RCC outcomes [5]. However, overall, there remains a lack of effective markers for subdivisions of RCC that would benefit from distinct immunotherapies.
RCC is typically resistant to chemotherapy and radiotherapy, so immuno-therapeutics have become increasingly important and promising components of treatment [6].
Before the introduction of immunotherapeutics, many treatments involved targeted therapies such as tyrosine kinase inhibitors for the VEGF or mTOR signaling pathways; current therapies have improved effectiveness by incorporating immunotherapeutics such as different combinations of immune checkpoint inhibitors, e.g., PD-L1 monoclonal antibodies with or without the addition of tyrosine kinase inhibitors [7]. Clinical decision making for immunotherapy choices is complex but often based upon parameters such as overall risk classification, pathologic features such as sarcomatoid status and presence of contraindications to immune checkpoint inhibitor therapy [7]. Future therapies will likely include new immune classifications and new immunotherapy-related agents in combination with currently accepted targets, such as mTOR [8].
In this report, we explored a TRB CDR3-candidate antigen chemical complementarity scoring algorithm [9, 10], particularly suited to the big data setting, and applied the algorithm to both Moffitt Cancer Center (MCC) and TCGA case sets. We sought to identify a subset of RCC cases with active and pronounced cytotoxic immune responses, in contrast with less refined approaches that raise questions about whether successful, anti-tumor, T-cell-based cytotoxicity is in effect.
Methods
Acquisition of the immune receptor (IR) recombination reads from the renal cell carcinoma (RCC) sample RNAseq files
Kaplan-Meier (KM) analyses comparing OS probabilities for case IDs representing RCC and the upper and lower 50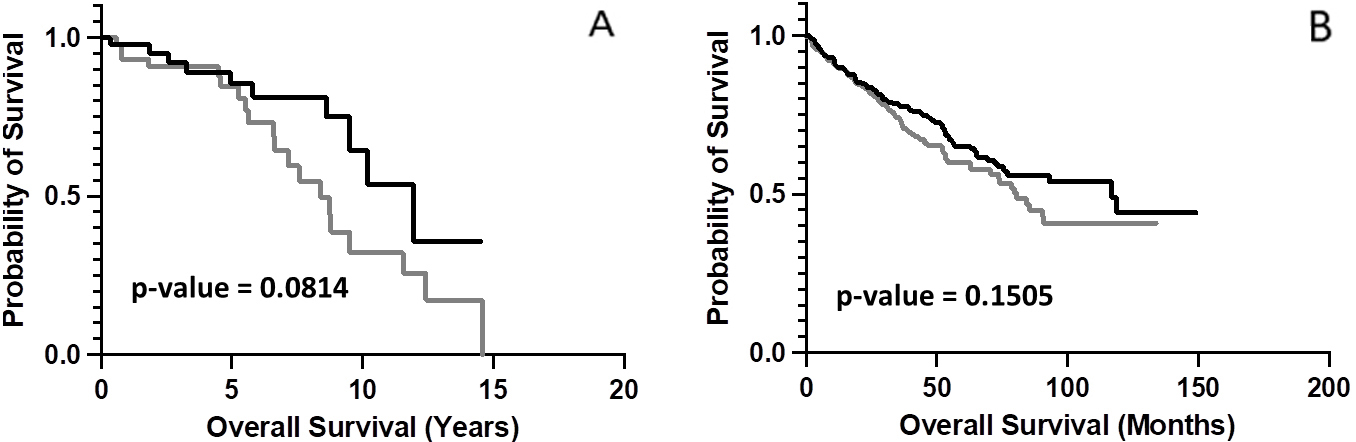
The methods for extracting the TRB recombination reads, from the RNAseq datasets, along with the benchmarking of the connection of such recombination reads to basic tumor immunology features, have been extensively described [11, 12, 13, 14]. The software for the extraction of the recombination reads, along with readme files, is available at the following links. Any one link represents the complete collection of software needed, but the three different links have various, specific conveniences as may be desired: (i)
The TRB CDR3-candidate antigen CSs were calculated according to reference [9]. The calculations were facilitated by the web tool, adaptivematch.com [16], which is publicly available and which has instructions for preparation of input files. The web tool has been extensively benchmarked in ref. [17] and further employed in refs. [16, 18]. Briefly, the chemical complementarity scoring algorithm aligns all available TRB CDR3s for a given case to a candidate antigen and engineers a stepwise approach to calculating chemical attractiveness, based on electrostatic and hydrophobic attraction (with the latter specifically based on Uversky hydropathy scores [10]). The algorithm returns the highest CS for each CDR3-candidate antigen pair. The resulting chemical CS score is referred to as Combo CS, thereby referencing the inclusion of both electrostatic and hydrophobic calculations to obtain the CS. Three example input files are available in the SOM, Tables S2–S4, with Table S2 representing TCGA-KIRC CDR3s; Table S3 representing the CTA, POTEA [19]; and Table S4 representing the TCGA-KIRC OS data publicly available via cbioportal.org. Tables S5 and S6 represent example adaptivematch.com output files.
Survival analyses
Multivariate analysis
A multivariate Cox regression analysis for OS was performed using the Statistical Package for the Social Sciences (SPSS). The input variables were CDR3-Fragment FPNTEIEEYHRPEKKSNE Combo CS upper and lower 50
Results
Survival distinctions represented by Combo CSs of TRB CDR3 AA sequences and the CTA, POTEA
To determine whether chemical complementarity between TRB CDR3 AA sequences and CTAs was associated with overall survival (OS) distinctions, we assessed Combo CSs (Methods) for the TRB CDR3 AA sequences and the AA sequences of multiple CTAs [9, 10, 16, 21, 22, 23, 24, 25]. We found that, while not meeting the conventional standard of statistical significance, Moffitt Cancer Center case IDs representing the upper 50
KM analyses comparing OS probabilities for case IDs representing the upper and lower 50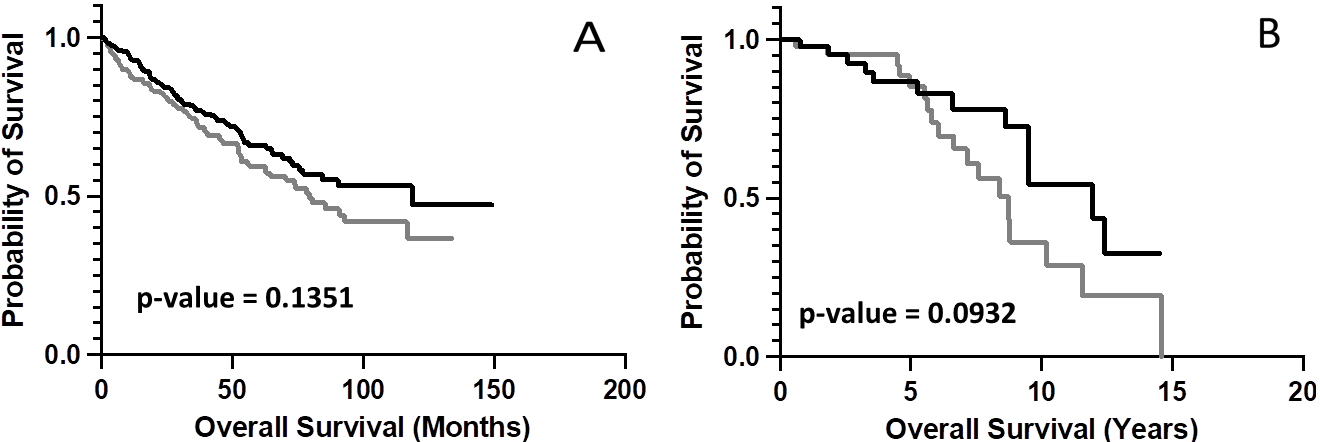
To determine whether chemical complementarity between the TRB CDR3 AA sequences and a POTEA fragment was further associated with OS distinctions, we assessed the Combo CSs for the TRB CDR3 AA sequences and the AA sequence of fragment FPNTEIEEYHRPEKKSNE, among other POTEA fragments (Table S3). Results indicated that the Moffitt Cancer Center case IDs representing the upper 50
Correlation between tumor RNA expression of immune signature genes and TRB CDR3-Fragment FPNTEIEEYHRPEKKSNE Combo CSs. Each point represents a case from the TCGA-KIRC database. (A) Scatterplot with linear regression line correlating RNA expression of granzyme A and Combo CSs (518 case IDs). (B) Scatterplot with linear regression line correlating RNA expression of granzyme B and Combo CSs (518 case IDs). (C) Scatterplot with linear regression line correlating RNA expression of interferon-gamma and Combo CSs (518 case IDs).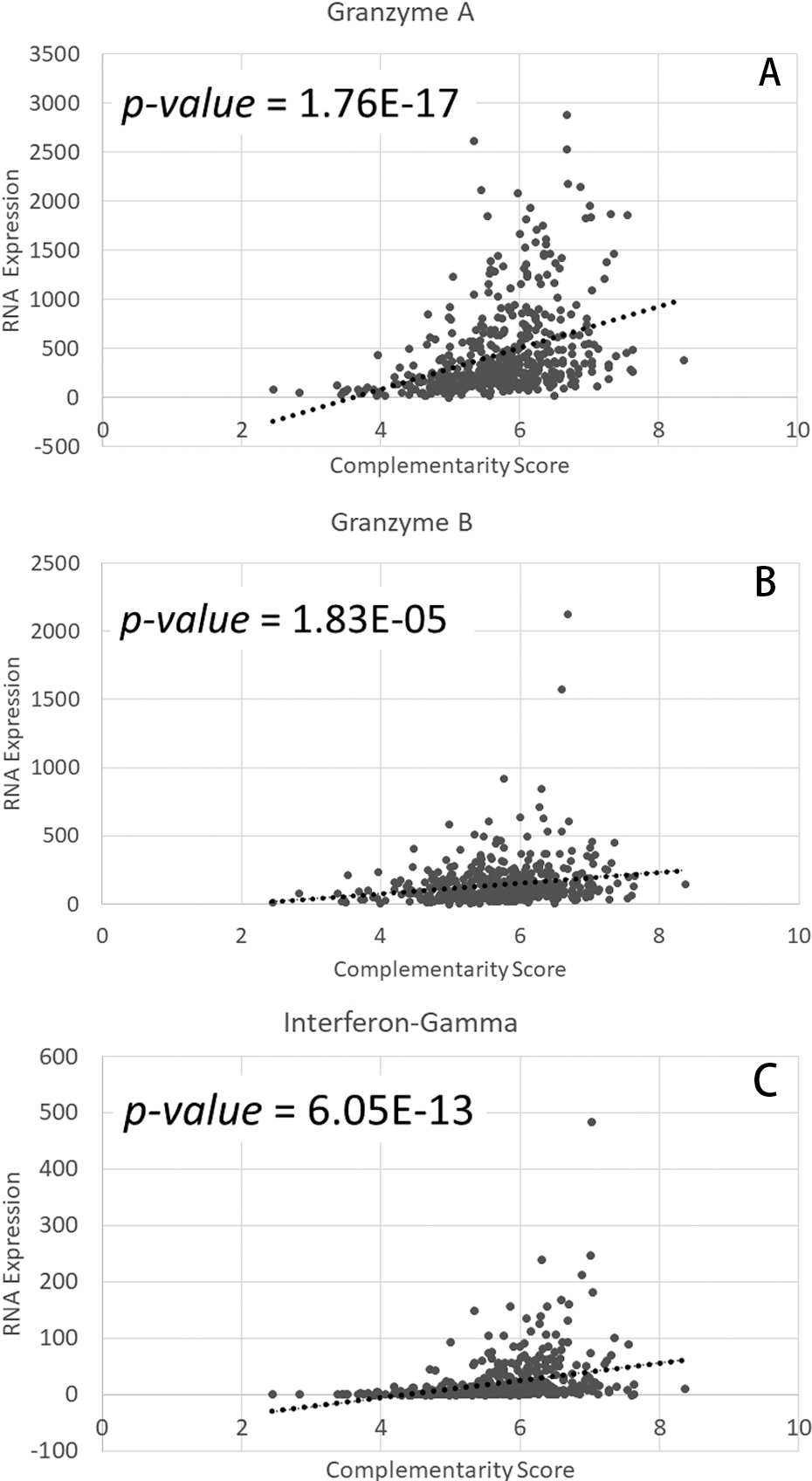
Correlation between TCGA-KIRC RNA expression of immune signature genes and Combo CSs for the TRB CDR3 AA sequences and POTEA Fragment FPNTEIEEYHRPEKKSNE (See also Table S8)
To determine whether expression of immune signature genes was associated with chemical complementarity, we assessed the correlation between TRB CDR3-fragment FPNTEIEEYHRPEKKSNE Combo CSs and the tumor RNA expression of multiple immune signature genes (Table 1). We found that TCGA-KIRC case IDs with greater tumor RNA expression of numerous immune signature genes including granzyme A, granzyme B, and interferon-gamma, among others, were associated with the higher Combo CSs (Table 1, Fig. 3). Additional data related to the correlation of the Combo CSs and immune signature gene expression is in Table S8.
Survival distinctions represented by expression of immune genes independently of TRB CDR3-POTEA chemical complementarity scoring
We determined whether RNA expression of immune signature genes in the total RCC dataset was associated with OS distinctions. Results indicated that TCGA case IDs representing the upper 50
Correlation between CPTAC RCC RNA expression of immune signature genes and Combo CSs for the TRB CDR3 AA sequences and POTEA Fragment FPNTEIEEYHRPEKKSNE (See also Table S10)
Correlation between CPTAC RCC RNA expression of immune signature genes and Combo CSs for the TRB CDR3 AA sequences and POTEA Fragment FPNTEIEEYHRPEKKSNE (See also Table S10)
We mined the CPTAC RNAseq files for TRB recombination reads using the same approach described above for the Moffitt Cancer Center dataset (Methods; Table S10). We then obtained the Combo CSs for the TRB CDR3-POTEA fragment, FPNTEIEEYHRPEKKSNE, and determined any correlations for the Combo CS value for each CPTAC RCC case and the expression of the immune signature gene set, exactly as was done for Table 1, using a Pearson’s correlation assessment. Results indicated a substantial correspondence of the direct correlation of the Combo CS values and the level of expression of genes representing the immune signature set for both the TCGA and CPTAC datasets (Table 2).
Multivariate Cox regression analysis. Age was analyzed on a continuous scale, while the upper and lower 50th percentiles for TRB CDR3-POTEA Fragment FPNTEIEEYHRPEKKSNE Combo CSs and tumor stages were treated as categorical variables. (Tumor stage I was used as a reference category)
Multivariate Cox regression analysis. Age was analyzed on a continuous scale, while the upper and lower 50th percentiles for TRB CDR3-POTEA Fragment FPNTEIEEYHRPEKKSNE Combo CSs and tumor stages were treated as categorical variables. (Tumor stage I was used as a reference category)
A multivariate Cox regression analysis was performed to detect potential variables that may be impacting OS including age and tumor stage, as well as the upper and lower 50
Discussion
This study was designed to determine whether chemical complementarity between CDR3s and CTAs could be used to subdivide RCC into groups with better or worse survival probabilities, and whether those groups could be correlated with additional features of an immune response. We focused on TRB CDR3 amino acid sequences in our approach because TRB represents the highest standard for direct contacts to an antigen in antigen binding to the T-cell receptor [27, 28, 29, 30, 31, 32, 33, 34]. We found an increased OS probability represented by cases with greater TRB CDR3-POTEA chemical complementarity, with similar results for cases with a greater TRB CDR3-POTEA Fragment FPNTEIEEYHRPEKKSNE chemical complementarity. We also identified greater RNA expression of multiple immune signature genes, notably including granzyme A, granzyme B, and interferon-gamma, in tumors with greater TRB CDR3-Fragment FPNTEIEEYHRPEKKSNE Combo CSs. One potential interpretation of these results is that improved tumor-resident, TRB CDR3 binding to the tumor antigen POTEA, or a fragment of this antigen, results in downstream immune effects that positively affect survival The association of high CS and better OS in the RCC setting is consistent with a similar study for MAGE CTAs and melanoma [17], but distinct from studies of lower grade glioma and ovarian cancer where the immune receptor-CTA CSs are inversely correlated with OS [16, 35], potentially reflecting vestigial and eventually ineffectual immune responses against the CTAs. In these latter two cases, the CTAs at issue were not expressed, further suggesting the possibility that the high CSs for the CTAs led to selection against early cancer cells expressing those CTAs.
The evidence that higher Combo CSs are related to better OS and higher expression of immune signature gene RNA becomes especially interesting in the context of RCC, a cancer that typically faces progressive levels of immune dysfunction [2]. When not taking the complementarity scoring into consideration, i.e., by assessing the entire TCGA RCC set, we found worse OS probability in cases with greater RNA expression of multiple immune signature genes. This result is consistent with a previous study that found that some immune genes, such as IFNG, correlate with greater occurrence of cytotoxic T cells, which in RCC correlates with higher stage and grade of tumors [36]. The contrast between the results from the global RCC set assessment and assessing the RCC cases by CSs indicates that complementarity scoring may be useful in distinguishing productive from non-productive or even harmful immune responses.
One recent study found that immune gene expression has shown promise in classifying solid tumors according to active favorable or unfavorable cytotoxic T cell responses, thereby identifying parameters that could potentially be used to refine current biomarkers for diagnosis and prognosis or to predict therapeutic responsiveness to treatments [37]. Another study identified a signature of five immune genes that predicted prognosis for RCC and that could potentially help in selecting patients for immune checkpoint inhibitor therapy [38]. We propose that chemical complementarity scoring could be included when searching for parameters to further improve the accuracy of molecular markers. Especially for cases of metastatic RCC, current prognostic factors have limited accuracy; it has been proposed that further refinement and validation of molecular markers should be a research priority [39].
It’s important to note that our analyses relied upon correlations, and that the mechanisms and downstream effects from TRB CDR3s binding to various RCC CTAs remains unverified by in vitro or in vivo approaches. Thus, it would be useful to extend the results indicated here to in vitro binding experiments whereby specific TRB CDR3s are tested for their affinity for the CTA, POTEA, as well as to consider animal models whereby POTEA is assessed as a potential antigen for RCC.
Conclusion
Chemical complementarity scoring, based on tumor resident, TRB CDR3s and cancer testis antigens can likely be used to distinguish productive versus harmful immune environments in renal cell carcinoma.
Disclosures
Informed Consent: Not applicable.
Registry and the Registration No. of the study/trial: Not applicable for clinical trial, but applicable for access to data, per Ethics approvals below.
Animal Studies: Not applicable
Ethics approvals
Access to TCGA-KIRC, RNAseq based TRB recombination reads was via dbGaP approval number 6300; access to the Moffitt Cancer Center RNAseq files was via protocol approvals available in reference [11]; access to the CPTAC RNAseq files was via dbGaP approval number 31752.
Author contributions
TRH: CONCEPTION; INTERPRETATION, OR AN-ALY SIS OF DATA: PREPARATION OF THE MANUSCRIPT.
VRB: INTERPRETATION OR ANALYSIS OF DATA; PREPARATION OF THE MANUSCRIPT
AC: INTERPRETATION OR ANALYSIS OF DATA: PREPARATION OF THE MANUSCRIPT.
BJM: CONCEPTION; INTERPRETATION, OR ANALYSIS OF DATA: PREPARATION OF THE MANUSCRIPT.
BIC: CONCEPTION; INTERPRETATION, OR ANALYSIS OF DATA.
MJD: INTERPRETATION OR ANALYSIS OF DATA.
JJS: INTERPRETATION OR ANALYSIS OF DATA.
GB: CONCEPTION; INTERPRETATION, OR ANALYSIS OF DATA: PREPARATION OF THE MANUSCRIPT; SUPERVISION.
Abbreviations
Supplementary data
The supplementary files are available to download from
sj-csv-1-cbm-10.3233_CBM-230047.csv - Supplemental material
Supplemental material, sj-csv-1-cbm-10.3233_CBM-230047.csv
sj-csv-2-cbm-10.3233_CBM-230047.csv - Supplemental material
Supplemental material, sj-csv-2-cbm-10.3233_CBM-230047.csv
sj-csv-3-cbm-10.3233_CBM-230047.csv - Supplemental material
Supplemental material, sj-csv-3-cbm-10.3233_CBM-230047.csv
sj-csv-4-cbm-10.3233_CBM-230047.csv - Supplemental material
Supplemental material, sj-csv-4-cbm-10.3233_CBM-230047.csv
sj-csv-5-cbm-10.3233_CBM-230047.csv - Supplemental material
Supplemental material, sj-csv-5-cbm-10.3233_CBM-230047.csv
sj-xlsx-1-cbm-10.3233_CBM-230047.xlsx - Supplemental material
Supplemental material, sj-xlsx-1-cbm-10.3233_CBM-230047.xlsx
sj-xlsx-2-cbm-10.3233_CBM-230047.xlsx - Supplemental material
Supplemental material, sj-xlsx-2-cbm-10.3233_CBM-230047.xlsx
sj-xlsx-3-cbm-10.3233_CBM-230047.xlsx - Supplemental material
Supplemental material, sj-xlsx-3-cbm-10.3233_CBM-230047.xlsx
sj-xlsx-4-cbm-10.3233_CBM-230047.xlsx - Supplemental material
Supplemental material, sj-xlsx-4-cbm-10.3233_CBM-230047.xlsx
sj-xlsx-5-cbm-10.3233_CBM-230047.xlsx - Supplemental material
Supplemental material, sj-xlsx-5-cbm-10.3233_CBM-230047.xlsx
Footnotes
Acknowledgments
The authors thank USF research computing and Ms. Corinne Walter for extensive admin support of the database approval processes. The authors thank Moffitt Collaborative Data Services for database assistance. The authors thank the taxpayers of the State of Florida.
Conflict of interest
The authors have nothing to declare.
