Abstract
OBJECTIVE:
MED subunits have been reported to be associated with various types of tumors, however, the potential role of MED7 in hepatocellular carcinoma (HCC) was still unclear. The aim of the study was to explore the role of MED7 in HCC.
METHODS:
In this study, MED7 mRNA expression levels between HCC and adjacent normal tissues were first analyzed by several public datasets. Then we utilized a tissue microarray (TMA) to investigate the clinical role of MED7 in HCC by immunohistochemistry (IHC). Meanwhile, the potential mechanisms of MED7 based on gene-gene correlation analyses were also explored.
RESULTS:
High mRNA level of MED7 correlated with advanced stage and worse grade of differentiation. IHC results showed that MED7 protein level was upregulated in HCC and associated with Edmondson grade and Microvascular invasion in 330 cases of HCC. GO (Gene Ontology) and KEGG (Kyoto Encyclopedia of Genes and Genomes) analysis revealed that MED7 co-expressed genes participate primarily in ribonucleoprotein complex biogenesis, protein targeting, mRNA processing and nucleoside triphosphate metabolic process et cetera. Further analysis also revealed that MED7 mRNA level has significant correlation with immune cells infiltration levels.
CONCLUSION:
MED7 was upregulated in HCC and correlated with progression of HCC. Meanwhile, MED7 may promote HCC through participating in multiple gene networks to influence tumorigenesis as well as immune response in HCC microenvironment.
Introduction
Hepatocellular carcinoma (HCC) is the leading cause of cancer-related deaths with 841,080 new cases and 781,631 deaths in 2018 [1]. To date, several preventive and therapeutic strategies have been significantly improved, but the five-year survival rate for advanced HCC remains low. Intrahepatic and extrahepatic metastases of HCC are the common reason for poor clinical prognosis of HCC patients [2]. Therefore, identification of the possible molecular mechanisms of malignant biological behaviors of HCC will provide therapeutic benefit for HCC.
The Mediator complex is an evolutionarily conserved RNA polymerase II (Pol II) coregulator required for the transcription of nearly all protein-coding genes [3]. Mediators control transcription in part by acting as a link between Pol II and transcription factors, which regulate their activity during initiation and elongation [4]. The destruction of mediator subunits may affect many different cell functions and fates, some of which may be related to carcinogenesis [5].
Since Med complex plays an important role in the transcription of all eukaryotic genes, alteration in its function and/or composition may have important consequences, and may lead to a variety of disease, including cancer [6]. To date, several MED subunits have been reported to be associated with various types of tumors, including HCC. Zou et al revealed that down-regulation of MED19 could significantly inhibit cell proliferation, induce delay in the G1-phase transition, resulted in tumor formation suppression in HCC in vitro and in vivo, which indicating an oncogenic role of MED19 in the HCC progression [7]. MED7, one of the Mediator subunits, was reported to play an important role in gonadal development and embryogenesis [8]. Recent studies also showed that MED7 may involve in tumorigenesis and gene expression profiling. MED7 downregulation was shown to be associated with an increased tumor risk and could therefore potentially be a marker of favorable prognosis in gastrointestinal stromal tumours [9]. Joseph et al also revealed that high MED7 mRNA and protein expression was associated with good prognostic factors in breast cancer, which may be used as a biomarker for good prognosis in invasive breast cancer, especially ER+ luminal subtypes [10]. Altogether, these findings indicate that MED7 may be used to identify the possibility of clinical outcomes, and contributed to further optimize treatment of patients. However, the potential role of MED7 in HCC was still unclear.
In this study, MED7 mRNA levels between HCC and normal tissues were first analyzed by several public datasets. Then we utilized a tissue microarray (TMA) to investigate the clinical role of MED7 protein level in HCC by immunohistochemistry (IHC). Meanwhile, the potential mechanisms of MED7 based on gene-gene correlation analyses were also explored.
Materials and methods
MED7 mRNA level in HCC based on public datasets
UALCAN (
The Gene Expression Profiling Interactive Analysis (GEPIA) database (
Analysis of MED7and immune infiltration by TIMER database
TIMER (
MED7 co-expression analysis by LinkedOmics Database
The LinkedOmics database (
Patient samples and construction of tissue microarray (TMA)
We collected 330 cases of HCC paraffinized specimens from the Department of Surgery (Zhejiang Provincial People’s Hospital, Hangzhou, China) between 2008 and 2015. All HCC participants had not received any chemotherapy or radiotherapy before operation, and the project was approved by the ethics committee of Zhejiang Provincial People’s Hospital. The HCC patients were followed up by telephone, which were acquired from patient records. The follow-up deadline was December 2016 or date of death. Patient survival time was calculated from the date of surgery to the follow-up cutoff point. Then a TMA containing these 330 HCC samples was constructed by Shanghai Outdo Biotech Company (Shanghai, China). Furthermore, key information of the patient will not be disclosed, and there is no risk to the patients, so the Ethics Committee exempted this study.
Patient and public involvement
The patients samples used in the experiment are obtained after previous operations, so the patients or the public were not involved in this study.
Immunohistochemical staining
First, the paraffin sections of TMA were baked at 60
Evaluation of Immunohistochemical staining
TMA tissues were evaluated independently by two different investigators with no prior knowledge of clinical information of patients. MED7 expression score was based on intensity score and the proportion score of positive staining tumor cells. Intensity score was recorded according to the following criterion: No staining (0); low intensity of staining (1); moderate staining (2); high intensity of staining (3). The proportion score was graded according to the following criterion: 0 (
Statistical analysis
Statistical Package for the Social Sciences (version 13.0; SPSS Inc, Chicago, IL) was used to perform all statistical analyses in the present study.
Results
MED7 mRNA level was elevated in HCC
We initially evaluated MED7 mRNA levels by UACAN. The results revealed that mRNA level of MED7 was significantly higher in HCC tissues than in adjacent normal tissues (Fig. 1). Further sub-group analysis of multiple clinic-pathological features in UALCAN database also showed elevated mRNA level of MED7 was significantly associated with stage, grade of HCC. High mRNA level of MED7 correlated with advanced stage and worse grade of differentiation.
Expression of MED7 in 330 cases of hepatocellular carcinoma tissues
Expression of MED7 in 330 cases of hepatocellular carcinoma tissues
MED7 mRNA level in HCC, as well as in subgroups of patients with HCC stratified based on grade, metastasis and stage by UALCAN. MED7 mRNA level was upregulated in HCC, and high MED7 mRNA level was associated with grade, metastasis and stage. Grade 1: Well differentiated, Grade 2: Moderately differentiated, Grade 3: Poorly differentiated, Grade 4: Undifferentiated. Stage: TNM (Tumor node metastasis) stage. N0: No regional lymph node metastasis, N1: Metastases in 1 to 3 axillary lymph nodes. 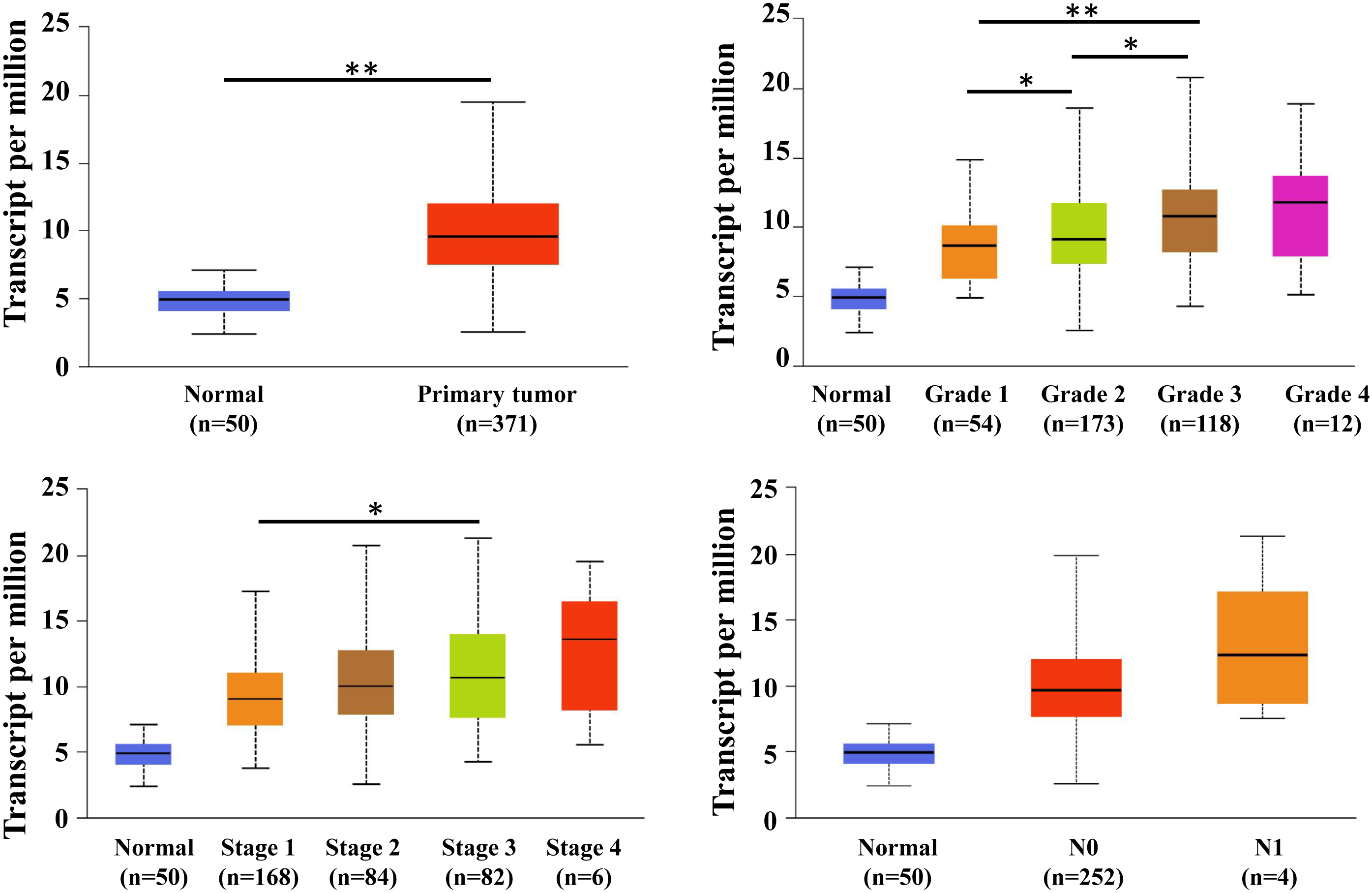
MED7 protein level was detected by Immunohistochemical staining in 330 cases of HCC. (A) Weak staining of MED7 in HCC. (B) Strong staining of MED7 in HCC. Magnification: the original magnification 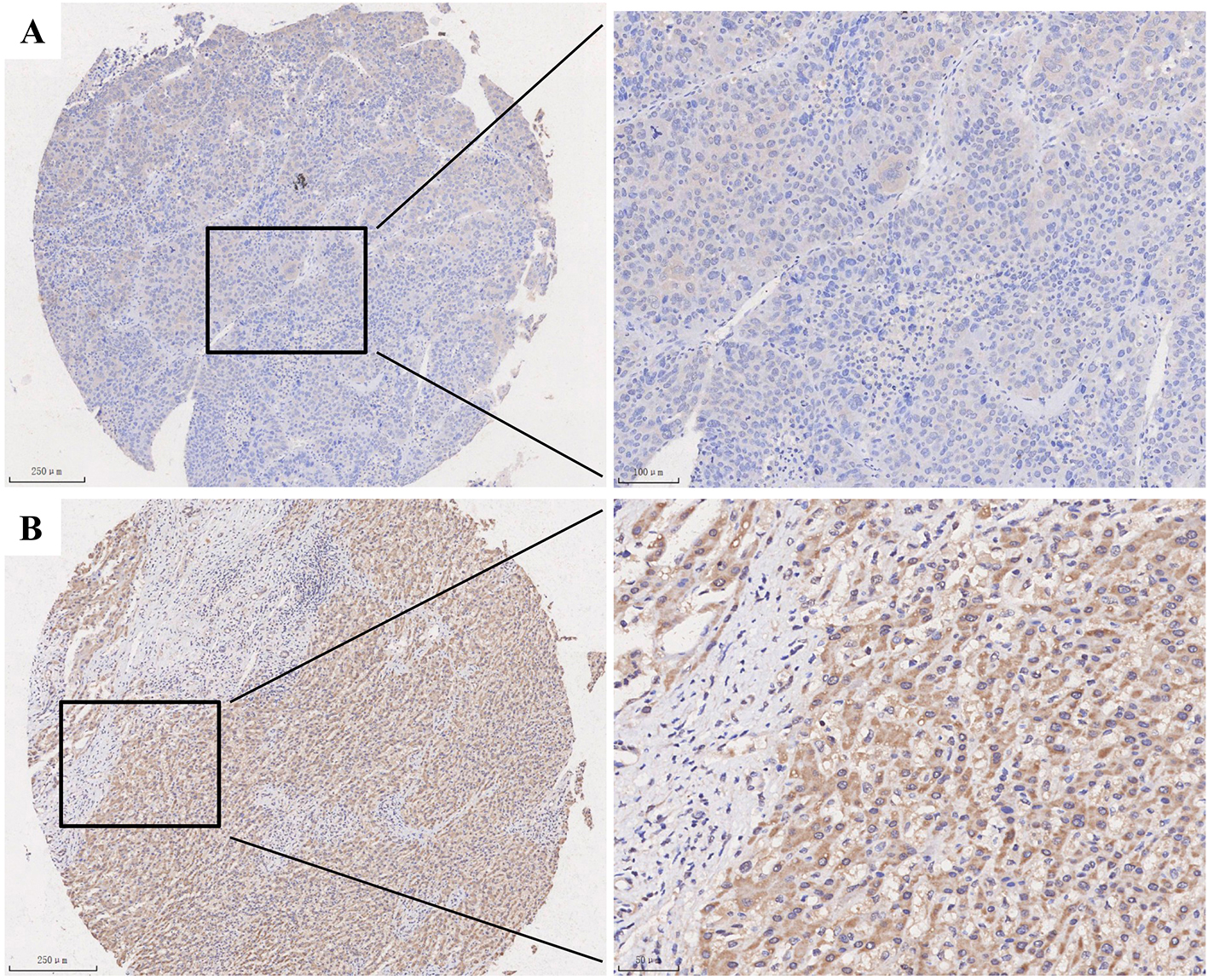
Immunohistochemistry results of 330 cases of HCC showed that MED7 was mainly located in nucleus and cytoplasm (Fig. 2). Then the correlation between MED7 expression and clinicopathological features was analyzed and showed that MED7 upregulation was closely associated with Edmondson grade and Microvascular invasion (Table 1). High expression ratio of MED7 in patients with Edmondson grade I
MED7 was correlated with prognosis of HCC
Next, we also performed Kaplan-Meier survival analysis to explore the association between MED7 expression and the survival outcomes of HCC with published database and our present study (Fig. 3). Generally, the patients with high MED7 mRNA level had significantly shorter overall survival (log-rank test,
The association between MED7 and survival outcome of HCC was detected by public database and present IHC result. Overall survival (OS) analysis detected by UALCAN, GEPIA, TIMER and in our cohort of 330 cases of HCC detected by IHC. 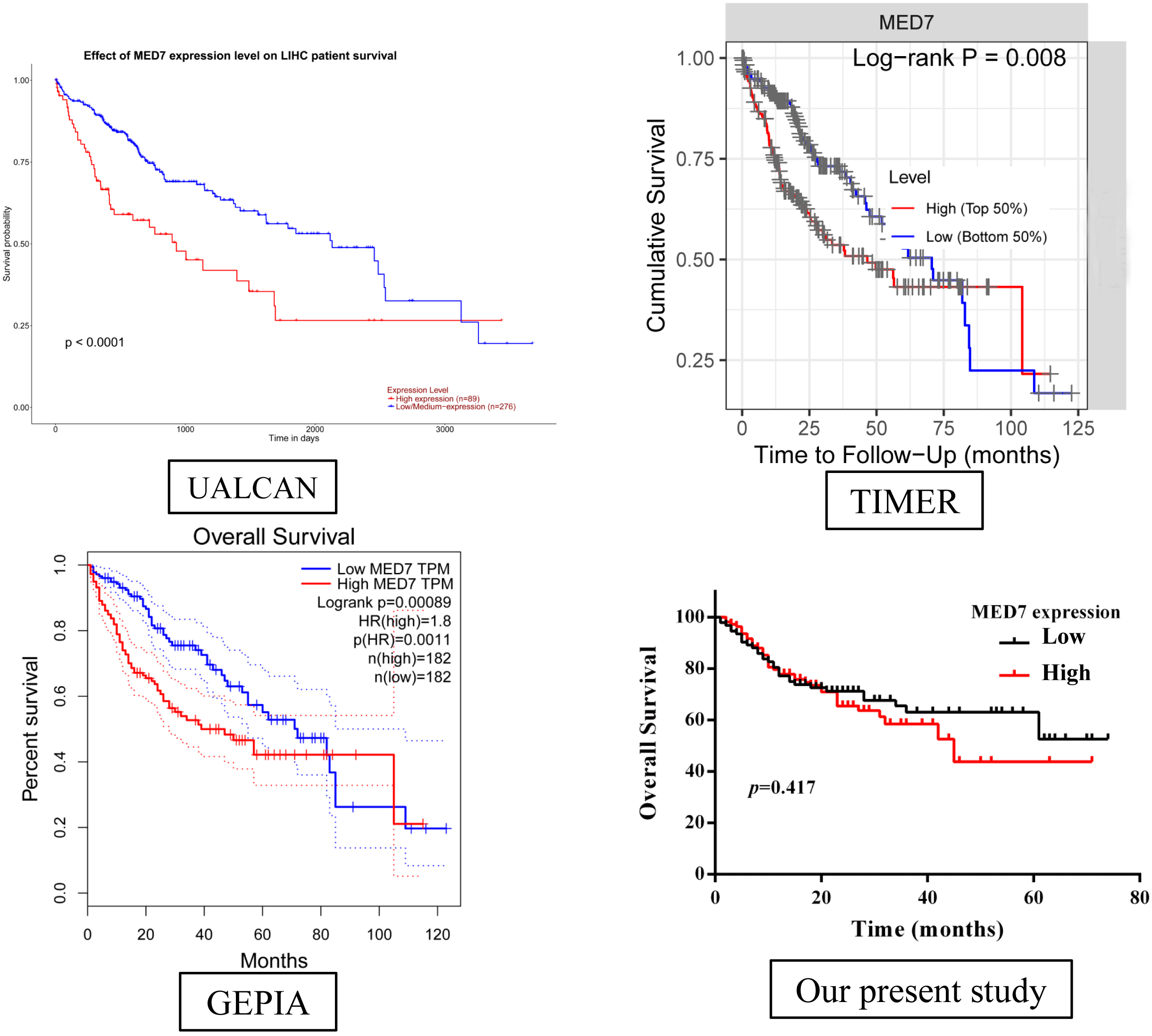
In order to gain the insight of MED7 molecular mechanism in HCC, we used function module of LinkedOmics to examine MED7 co-expression networks in HCC. Figure 4A showed that 5,102 genes (dark red dots) significant positive correlated with MED7, whereas 5,044 genes (dark green dots) negatively correlated with MED7, and the top 50 significant genes positively and negatively correlated with MED7 were shown in the heat map (Fig. 4B).
Co-expression genes of MED7 in HCC were analyzed by LinkedOmics. (A) The highly correlated genes of MED7 identified by Spearman test in HCC. (B) Top 50 genes positively and negatively correlated with MED7 mRNA level in HCC were showed by heat maps. Positively correlated genes were shown in red and negatively correlated genes were shown in blue. (C) Analysis of GO annotations and KEGG pathways of MED7 in HCC.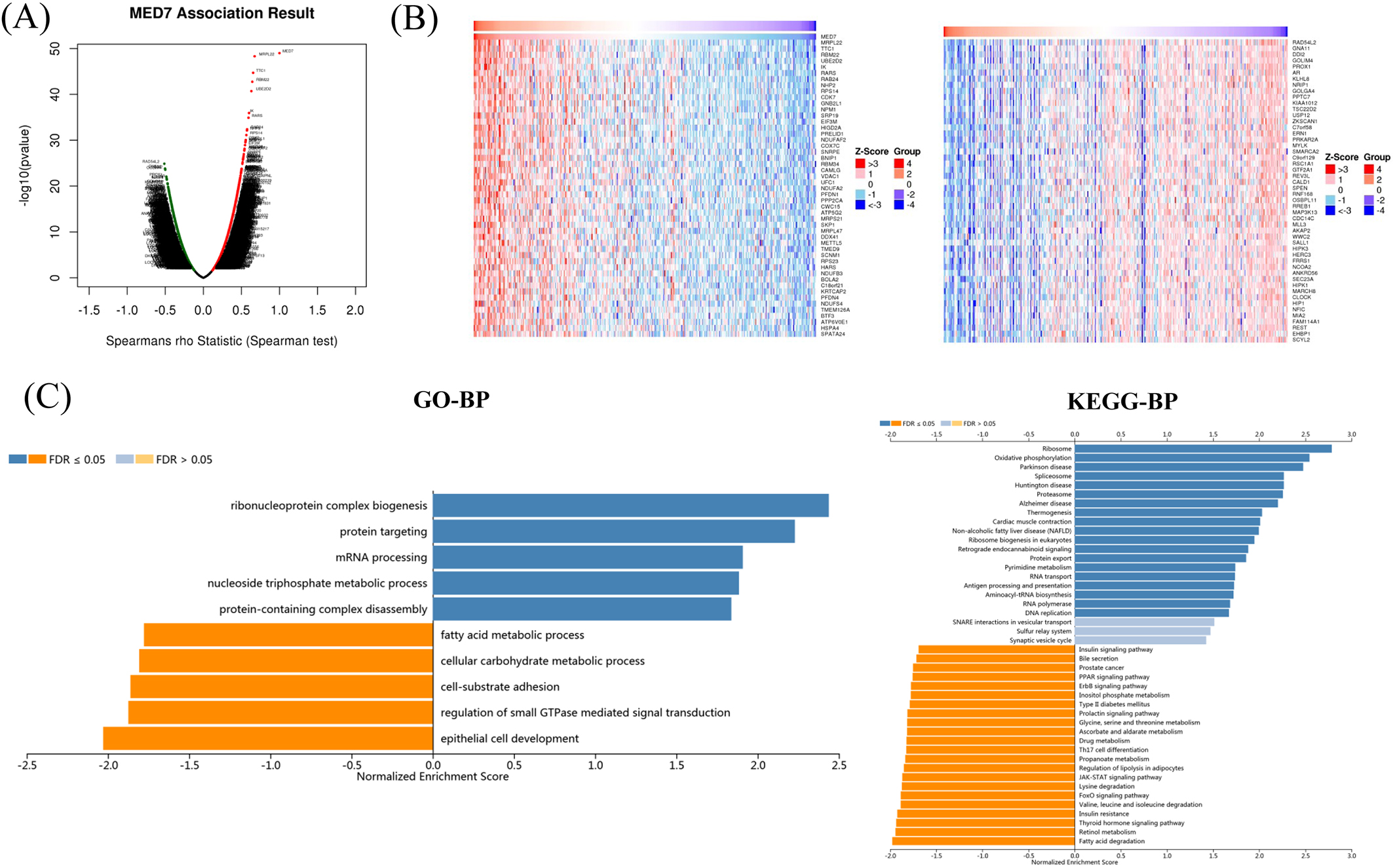
Furthermore, we also performed Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis. GO by gene set enrichment analysis (GSEA) showed that co-expressed genes of MED7 participate primarily in ribonucleoprotein complex biogenesis, protein targeting, mRNA processing and nucleoside triphosphate metabolic process et cetera, while the process like fatty acid metabolic process, cellular carbohydrate metabolic process, and cell-substrat adhesion were inhibited (Fig. 4C). Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis showed enrichment in the ribosome, oxidative phosphprylation, spliceosome, et cetera (Fig. 4C).
Next, we investigated the correlation of MED7 level with immune infiltration levels in HCC by TIMER database. As shown in Fig. 5, MED7 level has significant correlated with immune cells infiltration levels, such as B cell (
MED7 expression level was associated with immune cells infiltration in HCC. MED7 mRNA level correlated with infiltrating levels of CD8+ T cells, CD4+ T cells, macrophages, neutrophils, and dendritic cells significantly in HCC.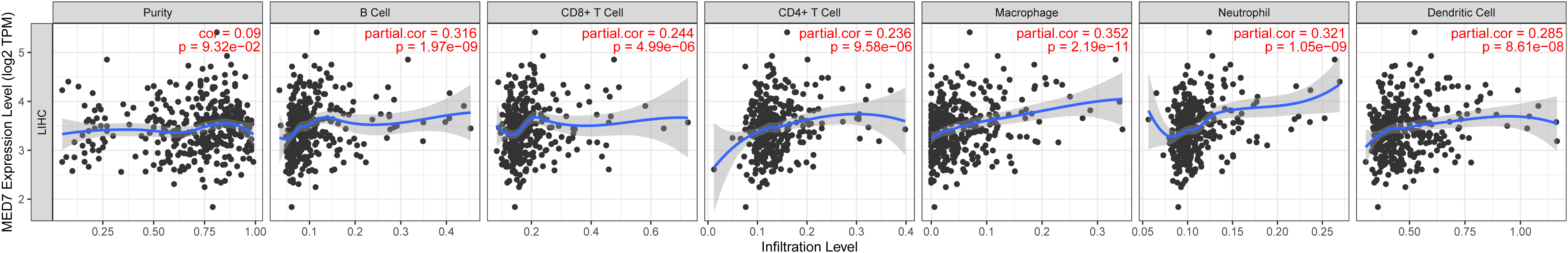
Furthermore, we also performed IHC staining of CD8 (provided as supplementary materials) and analyzed the correlation of MED7 and CD8. The results showed that the protein level of CD8 was associated with metastasis (Table 2), which suggested that CD8 may associate with the disorder of immune status in tumor microenvironment of HCC. However, further analysis showed that correlation between MED7 and CD8 was not significant (Spearmen’s rho test,
Expression of CD8 in hepatocellular carcinoma tissues
MED7, as mediator subunits, was reported to play an important role in gonadal development and embryogenesis [8]. Recently, study found that disruption of mediator subunits could affect many different cellular functions, including carcinogenesis [5]. However, the potential role of MED7 in HCC was still unclear.
In our present study, the functional relationships of MED7 with various clinicopathological variables in primary HCC were assessed. We found that high mRNA level of MED7 correlated with advanced stage, worse grade of differentiation and metastasis, and upregulation of MED7 protein was closely associated with Edmondson grade and microvascular invasion. These suggested that high expression levels of MED7 were significantly associated with poorer behaving tumour characteristics. However, Koschubs et al revealed that MED7 downregulation was significantly associated with increased risk of gastrointestinal stromal tumours [11]. Joseph et al also found that high MED7 mRNA and protein expression was associated with good prognostic factors [10]. Although MED7 is a highly conserved mediator subunit, it is not essential for viability of all species. The results revealed that the role of MED7 in cancer tissue may be diverse due to tissue-specificity. However many limitations existed in the present study including small sample size and limitation during collection, the conclusion should be made with caution.
Some studies revealed that high MED7 level was associated with good prognostic factors in breast cancer and gastrointestinal stromal tumours [9, 10]. However, in our present study, we found that patients with high MED7 mRNA level had poorer prognosis in HCC by public database, and there was no significant difference between MED7 protein level and prognosis of HCC. These findings indicate that MED7 could have a function in tumor pathogenesis with tissue specificity, which should be validated by further analysis in vitro and in vivo.
Furthermore, we also explore the molecular mechanism of MED7 in HCC by GO and KEGG analysis, and the results showed that co-expressed genes of MED7 participate primarily in ribonucleoprotein complex biogenesis, protein targeting, mRNA processing and nucleoside triphosphate metabolic process, oxidative phosphprylation, et cetera. Studies also found that Mediator plays an integral role in transcription at least in part by serving as a link between Pol II and transcription factors [4]. This indicated that MED7 level may affect multiple gene networks which are able to influence many cellular processes related to tumourigenesis in HCC. Further analysis by TIMER database tools showed that MED7 mRNA level has significant correlation with immune cells infiltration levels. However, there was no correlation between MED7 and CD8 protein level in our present study detected by IHC method. We considered that IHC staining may not reflect the whole level of immune cells comprehensively and accurately due to factors such as local sampling method and treatment process. Further study in vivo experiment will be performed in the future.
In summary, we found that MED7 was upregulated in HCC and correlated with progression of HCC. Meanwhile, MED7 may promote HCC through participating in multiple gene networks to influence tumorigenesis as well as immune response in HCC microenvironment.
Contributorship statement
Conception: Jun-Gang Zhang.
Interpretation or analysis of data: Zheng-Lin Chen, Ying-Yu Ma.
Preparation of the manuscript: Zheng-Lin Chen.
Revision for important intellectual content: Jun-Gang Zhang, Xiao-Zhou Mou.
Supervision: Xiao-Zhou Mou.
Funding
Not applicable
Data sharing statement
The data that support the findings of this study can be available from the corresponding author on reasonable request.
Ethics statement
This work was approved by ethic committee of Zhejiang provincial people’s hospital.
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-220439.
sj-pdf-1-cbm-10.3233_CBM-220439.pdf - Supplemental material
Supplemental material, sj-pdf-1-cbm-10.3233_CBM-220439.pdf
Footnotes
Conflict of interest
The authors declare no conflict of interest.
