Abstract
BACKGROUND:
Pan-TRK inhibitors Entrectinib and Larotrectinib have been recently approved as tumor-agnostic therapies in NTRK1-2-3 rearranged patients and there is therefore an urgent need to identify reliable and accessible biomarkers for capturing NTRK fusions in the real-world practice.
OBJECTIVE:
We aim to assess the analytical validity of the recently released pan-TRK assay (Ventana), running a head-to-head comparison between immunohistochemistry and Archer FusionPlex Lung Panel (ArcherDX) that is designed to detect key fusions in 13 genes, also including NTRK1-3.
METHODS:
Pan-TRK IHC and NGS analysis were conducted on a retrospective/prospective cohort of 124 cancer patients (carcinomas, 93 cases; soft tissue sarcomas, 19; primary central nervous system tumours, 10; and neuroblastomas, 2). FISH data were available in most of the IHC/NGS discordant cases.
RESULTS:
A comparison between IHC and NGS results was carried out in 117 cases: among 30 pan-TRK positive cases, NTRK rearrangement by NGS was found in 11 (37%), while one of the 87 (1.1%) pan-TRK negative cases (a case of NSCLC) showed a TPM3-NRTK1 rearrangement by NGS. Accordingly, sensitivity and specificity of IHC in predicting NTRK status were 91.7% and 81.9%, respectively, while negative (NPV) and positive predictive value (PPV) were 98.8% and 36.7%, respectively.
CONCLUSIONS:
These data lead to suggest that IHC with VENTANA pan-TRK antibody can be a reliable screening tool for the identification of patients potentially bearing NTRK rearranged tumours.
Introduction
The neurotrophic receptor tyrosine kinase (NTRK) genes NTRK1, NTRK2 and NTRK3 encode for the tropomyosin receptor kinases TRKA, TRKB and TRKC, respectively, which are involved in central and peripheral nervous system development [1, 2]. NTRK gene fusions are characterized by a chromosomal rearrangement of the 3’ region of the NTRK gene with the 5’ region of a variety of different partners, most commonly TPR and TMP3 for NTRK1 and ETV6 for NTRK3 eventually leading to a constitutive activation of the intracellular tyrosine kinase domain of the TRK receptor [3, 4, 5, 6]. In clinical samples, NTRK rearrangements may be detected by different methods, including DNA and RNA next generation sequencing (NGS), fluorescence in situ hybridization (FISH) and immunohistochemistry (IHC) [7, 8]. The prevalence of NTRK gene fusion is heterogeneously distributed across cancer types, occurring at a high frequency (
Larotrectinib and Entrectinib are selective TRK inhibitors reported to achieve 79% and 63.5% overall response rate in children and adult patients with different types of metastatic cancer harbouring NTRK gene fusion, with a median duration of response of 34 and 12.9 months, respectively [11, 12]. Based on these data, the two compounds received an accelerated approval by FDA and EMA as cancer agnostic drugs for tumours with a fusion of the NTRK gene [13]. There is therefore an urgent need to identify reliable and accessible biomarkers for capturing NTRK fusions in the real-world practice, outside clinical trials setting. Foundation One has been approved by FDA as a companion test for Entrectinib [14], while in Europe the testing rules are more heterogeneous: for example, in Italy NTRK inhibitors have recently been approved for patients with NTRK fusion positive tumours as per with FISH, reverse transcriptase-polymerase chain reaction or NGS analysis, and NGS confirmation is needed only for tumour types with a low NTRK fusion prevalence [15]. In this heterogeneous scenario, ESMO recently released a diagnostic algorithm for leveraging NTRK-positive cancer patients’ detection in low-prevalence settings, in which IHC should be used frontline and confirmed by FISH and/or NGS [16, 17]. Moreover, there are some discordances in the usefulness of IHC in detect NTRK-fusion positive cancers: a large meta-analysis of 1329 cases revealed a false-negative rate of 18% [18].
In the present study, we aim to validate ESMO approach in the real-world practice by a comprehensive head-to-head comparison of IHC, FISH and RNA-based NGS in a retrospective/prospective cohort of 124 real-world patients.
Materials and methods
Patients
The study population is represented by 124 consecutive patients for which NTRK gene fusion was analysed for diagnostic and/or treatment purposes, including: i) cases for which RNA-based NGS tests are routinely performed according to INT standard diagnostic procedures (i.e. stage III-IV NSCLC); ii) cases for which the clinician ordered the test for NTRK fusion identification for therapeutic purposes; iii) cases for which the pathologist carried out the analysis for diagnostic purposes, such as salivary gland secretory carcinomas and spindle cell sarcomas. Tumour types were as follows: carcinomas (93 cases), soft tissue sarcomas (STS, 19), primary central nervous system tumours (PCNST, 10) and neuroblastomas (NB, 2). The study was approved by our Ethical Review, and was performed in accordance with the Declaration of Helsinki. All the patient tumours were evaluated by IHC and RNA-based NGS. FISH was selectively carried out in cases yielding discordant results to fully characterize NTRK gene status.
IHC
Immunohistochemical analysis was performed using the VENTANA pan-TRK (EPR17341) (Ventana Medical Systems, Tucson, AZ, USA) rabbit monoclonal primary antibody directed against the C-terminal region of TRK A, B and C proteins. This region is known to be conserved across wild-type and chimeric fusion proteins [8]. Immunostaining was carried out on a BenchMark Ultra Platform (Ventana Medical Systems), using the OptiviewDAB Detection Kit (Ventana Medical Systems) according to the manufacturer’s instructions. We applied a procedure previously validated in our laboratory by comparing IHC/FISH/NGS data in a small series of cases not included in the present analysis (unpublished data). Briefly, IHC was performed on freshly cut 3
Molecular analysis
Molecular analyses were performed using the Archer FusionPlex Lung Panel (ArcherDX), that is designed to detect key fusions in 13 genes, also including NTRK1-3. Five
The same raw data were also evaluated by ARRIBA informatic tools. Among the fusion transcripts detected by ARRIBA tool only those with high level of confidence were considered for subsequent analysis (as detailed in [20]).
Representative examples of VENTANA pan-TRK immunoreactivity (IR) . Panel A shows a salivary gland secretory carcinoma with an ETV6-NTRK3 rearrangement showing a nuclear IR pattern (40x, case 3, Table 1); panels B and C show typical examples of cytoplasmic IR pattern in a papillary thyroid carcinoma (B, 40x, case 1) and in a NTRK-rearranged spindle cell neoplasm (C, 40x, case no. 13) bearing TPR-NTRK1, and TPM3-NTRK1 rearrangement, respectively. Panel D shows a combined cytoplasmic and membranous pattern in another case of NTRK-rearranged spindle cell neoplasm bearing TPR-NTRK1 rearrangement (40x, case 16).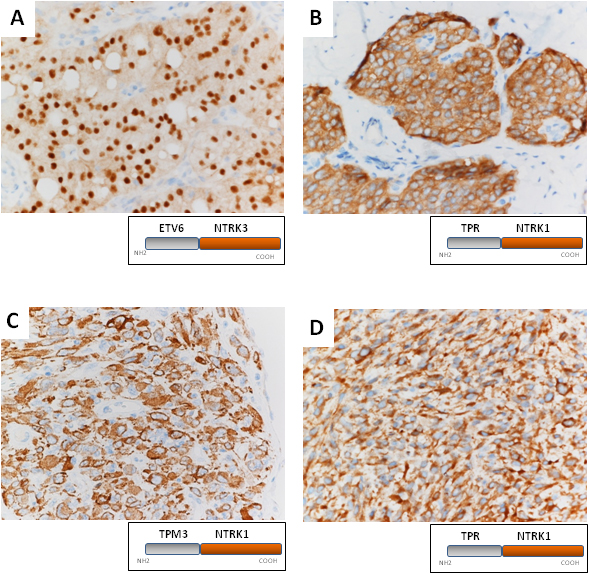
FISH experiments were carried out on FFPE samples, using the following break-apart probes: spectrum green labeled RP11-1038N13 and spectrum orange labeled RP11-180B22 for 3’ and 5’ NTRK1, respectively; spectrum green labeled RP11-244M18 and spectrum Orange labeled RP11-117C17 for 3’ and 5’ NTRK2, respectively; spectrum green labeled RP11-258I14 and spectrum Orange labeled RP11-121M4 for 5’ NTRK3, respectively. Dual color dual fusion FISH probes were used to characterize ETV6-NTRK3 fusion (spectrum orange labeled RP11-75N23/RP11-418C2 for ETV6 and spectrum green labeled RP11-258I14/RP11-121M4/RP11-46G5 for NTRK3) and TPM3-NTRK1 fusion (spectrum orange labeled RP11-737P18/RP11-1043P24 for TPM3 and spectrum green labeled RP11-1038N13/RP11-180B22 for NTRK1). BAC clones were obtained from C.H.O.R.I. (bac-pac resources. Children’s Hospital Oakland, Ca), labelled by means of nick translation (Nick Translation Reagent Kit, Abbott Molecular) in accordance with the manufacturer’s instructions, and validated on normal metaphase spreads. At least 50 tumor nuclei were scored in each case using a Leica DM 6000B (Wetzlar, Germany) microscope at 100x magnification with immersion oil and the appropriate fluorescence filters. The images were captured using Cytovision software (v. 7.0, Leica).
Comparison between pan-TRK ICH and NGS data
Comparison between pan-TRK ICH and NGS data
Abbreviations. NE, not evaluable; NOS, not otherwise specified; WT, wild type; C, cytoplasmic; N, nuclear; M, membranous; 1+, mild; 2+, moderate; 3+, high staining intensity.
IHC
Overall, 34 out of the 124 cases (27%) included in the present series showed nuclear, membranous or cytoplasmatic NTRK immunoreactivity beyond the threshold of 1% (Fig. 1), with a mean and median prevalence of 66% and 75% (interquartile range 35%–100%), respectively. Detailed histological diagnosis and IHC patterns of TRK-immunoreactive cases are shown in Table 1. NTRK IR was found in 10 of 93 carcinomas (108%), 14 of 19 STS (737%), 10 of 12 nervous system tumors (833%), including 10 PCNST cases and 2 NB cases; mean and median prevalence were respectively 555% and 60% in carcinomas (interquartile range 225%–90%), 829% and 95% in STS (interquartile range 725%–100%), 55% and 60% in nervous system tumors (interquartile range 225%–80%).
NTRK NGS analysis
Archer FusionPlex analysis was performed in all the 124 cases included in this analysis. Seven cases (5.6%) were not evaluable for low-quality RNA. A fusion transcript involving one of the NTRK genes was observed in 12 (10.2%, 8 STS and 4 carcinomas) out of the 117 cases successfully analysed by NGS. The fusion patterns were as follows: NTRK3/ETV6 (5 cases, 41.7%), NTRK1/TPR (3 cases, 25%), NTRK1/TPM3 (3 cases, 25%) and NTRK3/TFG (1 case, 8.3%) (Table 1). Out of the 105 NGS NTRK-negative cases, 11 (11/117, 9.4%) showed different rearrangements: ALK fusions with EML or KIFSC genes in 5 cases (4 non-small cell lung cancer – NSCLC – and 1 PCNS); RETNCOA4 and RET-KIF5B fusions in one NSCLC and one follicular thyroid carcinoma, respectively; BRAF-KIA1549 fusion in one PCNST; FLI1-EWSR1 in one STS; and MET exon 14 skipping variants in 3 NSCLC (data not shown).
Correlation between pan-TRK IHC and NGS results
Correlation between pan-TRK IHC and NGS results
Concordance rate and predictive values of pan-TRK immunohistochemistry as related to NGS results, according to tumor type
A comparison between IHC and NGS results was carried out in 117 cases, as detailed in Tables 2 and 3. NTRK rearrangement was found in 11 of the 30 (37%) pan-TRK positive cases, and in one of the 87 (1.1%) pan-TRK negative cases (a case of NSCLC bearing a TPM3-NRTK1 rearrangement). In particular, all the 5 cases carrying the ETV6-NTRK3 fusion transcript showed a moderate/strong nuclear NTRK staining in 60–100% of the tumour cells, while the remaining patterns of rearrangement were associated with a cytoplasmic or combined cytoplasmic/membranous IR (Fig. 1). The concordance between IHC and NGS was independent of prevalence, intensity and pattern of IR (supplementary table 1).
According to these metrics, sensitivity and specificity of IHC in predicting NTRK status were 91.7% and 81.9%, respectively, while negative (NPV) and positive predictive value (PPV) were 98.8% and 36.7%, respectively (Table 3). Interestingly, IHC NPV was high (from 98.8% to 100%) across the different histotypes investigated (Table 3). The concordance rate was higher in carcinomas (82/87
Panel A-C shows a sarcoma NOS (case no. 21 of Table 1) of the parotid with a pan-TRK IHC moderate cytoplasmic positivity in 50% of neoplastic cells, NTRK gene wild type by NGS (not shown) and NTRK3 gene amplification by FISH analysis (in green clustering signals for NTRK3 gene and in orange signals for ETV6 gene); Panel A: H&E, magnification: 10x; Panel B; pan-TRK IHC, magnification; 40x; Panel C: NTRK3/ETV FISH analysis, magnification; 100x, oil immersion. Panel D-F shows an endometrial stromal sarcoma (case no. 24 of Table 1) with a pan-TRK IHC weak cytoplasmic positivity in 90% of neoplastic cells, NTRK gene wild type by NGS (not shown) and YWHAE gene translocation by FISH analysis (ZytoLight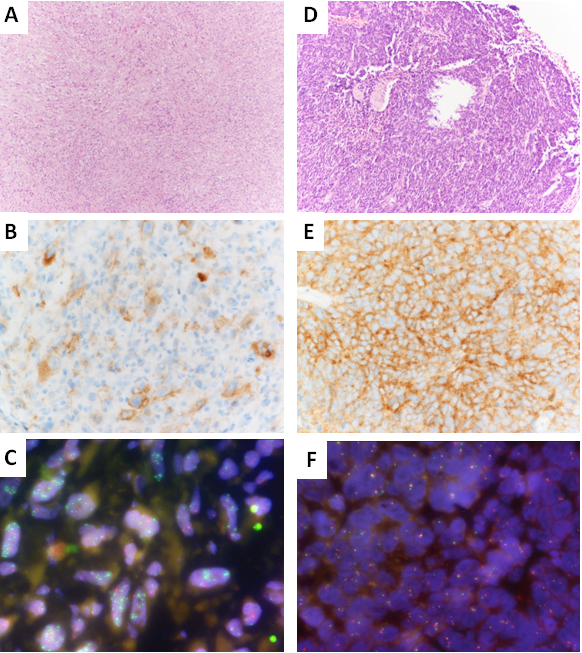
FISH analyses were performed in 14 out of 20 cases (13 IHC+/NGS-, 1 IHC-/NGS+) with IHC/NGS discordant results, in which enough tissue for further analyses was available. None of the 13 IHC+/NGS- cases proved to have a NTRK gene rearrangement by FISH; one of them (an undifferentiated pleomorphic sarcoma of the neck in a 24-years old female patient) bore NTRK3 gene amplification by FISH (Fig. 2A–C).
Finally, the unique IHC-/NGS+ case of our series (a NSCLC with a TPM3-NRTK1 NGS fusion) did not show any signal of gene rearrangement by FISH. In detail, this case showed 27 unique start site and 64% of breakpoint spanning reads whereas the three other cases with TPM3-NTRK1 fusion and positive IHC showed a mean of 196 unique start site and 80% of breakpoint spanning reads. Of note, tumour cellularity was 15% in the IHC negative case (versus a mean of 70% in other three cases). Moreover, in this case, TMP3-NTRK1 fusion call were confirmed by two different pipelines (ARRIBA and Archer Dx; [20]).
TRK inhibitors are effective in patients with tumors harboring NTRK fusions, irrespective of their site of origin, histological diagnosis, or pattern of rearrangement [11, 21, 22]. NTRK fusions can be detected by different methods, including IHC, FISH and NGS [7, 8, 23]. To date, different diagnostic algorithms for the identification of NTRK rearrangements have been proposed: the ESMO guidelines [16] recommend upfront molecular assessment (FISH, PCR and/or NGS) in tumors with a known high prevalence of NTRK fusions; for tumors with a low prevalence of NTRK rearrangements, the guidelines suggest using IHC as a screening method when NGS technology is unavailable, and confirming IHC positive cases by molecular assays, preferably including RNA testing, including Archer assay. A similar approach was proposed by Penaut-Lorca et al. [7], whereas others suggest using NGS upfront, irrespective of the histological subtype [24, 25].
Assessing the reliability of IHC as a screening tool for identifying potential NTRK-positive tumors to be confirmed by second-level molecular analyses is therefore mandatory for implementing the use of NTRK inhibitors in the clinical practice, considering that the application of comprehensive genomic profiling is still limited in pathology labs: in Italy, for example, where NTRK inhibitors have been recently approved with a generic indication for NTRK rearranged tumors, the distribution of NGS technology is still heterogeneous and mostly relying to midsize (up to 50 genes) DNA-based panels, usually not including NTRK genes [26]. To investigate the role of IHC in predicting the occurrence of NTRK gene fusions, we carried out a comprehensive head-to-head comparison between IHC and RNA-based NGS in a large monoinstitutional retrospective/prospective case series. We found that sensitivity and specificity of IHC in predicting NGS results were 91.7% and 81.9%, respectively, with a NPV of 98,8%. These results are in line with previous studies: Solomon et al. [8] reported a sensitivity and specificity of pan-TRK IHC of 87.9% and 81.1%, respectively, with a NPV of 97.0%, in a retrospective series previously analyzed by DNA NGS with the hybrid DNA/RNA MSK Impact panel. Likewise, Koopman et al. [27] evaluated the concordance of IHC and NGS in 327 tumour samples, reporting for IHC a sensitivity and specificity of 77% and 84%, respectively. NTRK rearrangement occurs at a very low frequency in the commonest tumours: in this perspective, the high negative predictive value identified in our study, firstly comparing VENTANA pan-TRK antibody with an RNA-based NGS approach, provide evidence that IHC can be reliably used as a screening tool. Of note, our approach RNA-based NGS is a fast, sensitive and specific method to detect known and novel gene fusions even in FFPE specimens with low concentration and low-quality RNA. A possible downside of the proposed workflow is represented by the low positive predictive value of IHC, likely due to the potential detection of wild-type protein expression, especially in tumours with neural and myogenic differentiation [28, 29]. This will give rise to additional NGS analyses in a fraction of IHC false positive cases (19 out of 117 patients in the present series, 16.2%), possibly impacting on the cost-efficacy of IHC screening: nevertheless, it has to be underlined that tumor profiling would anyhow be valuable for the identification in these patients of alternative biomarkers actionable with on-label drugs or in clinical trials.
The interpretation of IHC results can actually be particularly cumbersome and may at least in part explain the low positive predictive value observed in our series (36.7%). Our findings are consistent with previous reports, which have reported a PPV ranging from 49.2% to 10.4%, with even lower specificity (20.8%) observed in gliomas, possibly due to nonspecific background staining within the normal tissue. Some authors previously discussed the potential reduction of IHC specificity in cases with cytoplasmatic immunoreactivity, especially in particular settings, such as sarcomas, gliomas or salivary gland tumors [8]. Nevertheless, given the high frequency of the cytoplasmic staining pattern among NGS positive samples in our and other case series [30], we believe these cases should be submitted to NGS analysis to reduce the risk of possible false negatives.
Some authors suggest that cytoplasmic immunoreactivity could be a potential marker of IHC false-positive results [8], but as it was the most frequent staining pattern of NTRK rearrangement in this and other [30] series, we suggest that compartmentalization of the IHC signal cannot prevent further NGS analysis for validation.
We found one case with TPM3-NRTK1 fusion detected by NGS, with an acceptable number of unique start sites (n. 27, according to [31]) and 64% of breakpoint spanning, comparable with other cases with the same fusion detected, which resulted negative in IHC and FISH, highlighting the relevance of multidisciplinary discussion in routine laboratory practice.
The 2020 WHO Classification of soft tissue and bone tumors [32] has identified the “NTRK-rearranged spindle cell neoplasm” as a molecular defined emerging entity, with frequent co-expression of CD34 and S-100 by IHC. Consequently in our case series, 5 spindle cell sarcomas have been classified as “NTRK-rearranged spindle cell neoplasm”. In detail, all but one case had at least 70% of pan-TRK IHC positive tumour cells with a cytoplasmic staining pattern and rearrangements involving NTRK1. The remaining case showed pan-TRK IHC nuclear positivity in 100% of the neoplastic cells associated with an ETV6-NTRK3 fusion, consistently with prevalence of NTRK1 and NTRK3 rearrangements reported in literature [32].
Along this line, a recent study on 196 glioblastoma patients [33] reported 7 cases with NTRK fusion by FISH (4 of them confirmed by RNA sequencing), with inconclusive IHC results despite different algorithms of evaluation, leading the authors to suggest performing FISH or NGS upfront in this setting.
On the other hand, our analysis suggests potential mechanisms of NTRK overexpression alternative to gene rearrangement, highlighting the relevance of IHC: for instance, we presented a case (#21 in Table 1), which was pan-TRK IHC positive and negative by NGS, yet exhibiting substantial NTRK3 amplification when analysed with FISH (shown in Fig. 2A–C). Similarly, in case #24, which is also pan-TRK positive and NGS negative, the FISH panel used for endometrial stromal sarcoma routinely identified a YWHAE translocation (shown in Fig. 2D–F), known to be associated with NTRK3 mRNA overexpression [34].
It has to be underlined that our study that the primary endpoint of our study was the evaluation of the role of IHC in predicting NTRK genes rearrangement. As a consequence, our study population was selected in order to have a sufficient number of events to evaluate the predictive value of IHC testing in a clinical practice setting, and was not intended to provide information related to the prevalence of NTRK rearrangement in the overall population.
Collectively, our data prompt us to propose the following algorithm to reliably detect NTRK gene rearrangement in the real-world practice:
For tumours with a low prevalence of NTRK fusions, for which ESMO guidelines do not suggest carrying out comprehensive genomic profiling upfront (i.e., breast and gastric cancer), IHC should be used as a screening tool: IHC positive ( For tumours with a low prevalence of NTRK fusions, for which ESMO guidelines suggest carrying out comprehensive genomic profiling for the detection of multiple actionable biomarkers (i.e., lung cancer, cholangiocarcinoma, prostate cancer, colon cancer), hybrid RNA-DNA NGS panels including NTRK genes should be used upfront. In cases in which pre-analytical issues (i.e., insufficient tissue available, low RNA quality) hamper reliable NGS analysis, IHC could be used as a surrogated diagnostic tool, IHC positive cases to be confirmed by FISH. For rare histotypes with a known high prevalence of NTRK fusions (for example, infantile sarcomas, secretory carcinomas), NGS or FISH can be performed upfront. For primary CNS tumour and neuroblastomas, NGS or FISH should be performed upfront, in consideration of the low reliability of IHC in theses settings.
This approach, as aforementioned, rises from ESMO guidelines (17), with a specifical focus on our clinical practice. This integrated approach is supported by data from a Canadian national clinical diagnostic testing program cohort in which the majority of patients were screened with IHC, NGS upfront reserved mainly in CNS and pathognomonic tumours [35]. Moreover, a Chinese expert consensus recommended the use of IHC in cancer with rare occurrence of NTRK rearrangements, which do not routinely undergo comprehensive DNA or RNA based NGS [36]. As exposed before, unfortunately this panels are not available in a significant proportion of small-to-medium size centres.
In conclusion, we provided evidence that IHC with VENTANA pan-TRK antibody can be reliably used as a screening tool for the identification of patients potentially bearing NTRK rearranged tumours, and proposed a combined IHC/NGS approach to be integrated within the routine diagnostic workflow of pathology labs.
Funding
ROCHE Diagnostics SpA supported the study by providing the VENTANA pan-TRK antibody (EPR17341).
Author contribution
Conception: A.V, D.L., E.C., A.B., and G.P.
Interpretation or analysis of data: C.C.V., D.V.T., A.G., E.C., F.P., E.T., I.C., A.B., and G.D.
Preparation of the manuscript: A.V, D.L., C.C.V., and G.P.
Revision for important intellectual content: A.V., D.L., L.A., and G.P.
Supervision: A.G., and G.P.
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-220357.
sj-docx-1-cbm-10.3233_CBM-220357.docx - Supplemental material
Supplemental material, sj-docx-1-cbm-10.3233_CBM-220357.docx
