Abstract
BACKGROUND:
The role of ELAPOR1 has been evaluated in several cancers but has not been elucidated in colorectal cancer (CRC).
OBJECTIVE:
To investigate the role of ELAPOR1 in CRC.
METHODS:
In the present study, the correlation between ELAPOR1 and survival of CRC patients in TCGA-COAD-READ datasets was predicted, and the difference in ELAPOR1 expression between tumor and normal tissues was analyzed. ELAPOR1 expression in CRC tissues was measured by immunohistochemistry. Then, ELAPOR1 and ELAPOR1-shRNA plasmids were constructed and transfected into SW620 and RKO cells. The effects were assessed by CCK-8, colony formation, transwell, and wound healing assays. Transcriptome sequencing and bioinformatics analysis were performed on the genes before and after ELAPOR1 overexpression in SW620 cells; the differentially expressed genes were substantiated by real-time quantitative reverse transcription PCR.
RESULTS:
High level of ELAPOR1 is associated with favorable disease-free survival and overall survival. Compared to normal mucosa, ELAPOR1 is lower in CRC. Moreover, ELAPOR1 overexpression significantly inhibits cell proliferation and invasion in vitro in SW260 and RKO cells. Conversely, ELAPOR1-shRNA promotes CRC cell proliferation and invasion. Among the 355 differentially expressed mRNAs identified, 234 were upregulated and 121 were downregulated. Bioinformatics indicated that these genes are involved in receptor binding, plasma membrane, negative regulation of cell proliferation, as well as common cancer signaling pathways.
CONCLUSIONS:
ELAPOR1 plays an inhibitory role in CRC and may be used as a prognostic indicator and a potential target for treatment.
Introduction
Colorectal cancer (CRC) is the third most common malignancy worldwide. The result of traditional treatments is unsatisfactory with its total 5-year survival rate at 64% [1]. Although targeted therapies for solid tumors are challenging with either single drug treatment or combined with radiotherapy and chemotherapy, they might have prospects. Thus, exploring effective targets for the prognosis or treatment of CRC is an urgent requirement. The membrane proteins are considered to play a key role in the neoplastic transformation and progression, mediating intercellular communication and intracellular signal transduction [2, 3, 4]. The role of various membrane proteins in malignant tumors has been explored previously, revealing their regulatory networks in different cancers; however, their further clinical significance is yet to be unraveled.
The endosome-lysosome associated apoptosis and autophagy regulator 1 (ELAPOR1) is a transmembrane protein, encoding 1013 amino acids highly conserved across species, such as nematode, Xenopus, fish, and chicken. It is located in the plasma membrane and endoplasmic reticulum and is also related to the endosomal lysosomal compartment [5, 6]. ELAPOR1 leads to growth inhibition and apoptosis of breast cancer cells when sufficient nutrients are available, but it may protect the cancer cells from cytotoxic drugs by upregulating autophagy under stressful conditions and is related to the high metastasis of breast cancer cells [7, 8]. It also induces apoptosis in gastric cancer cells but not autophagy [8, 9]. Clinically, the role of ELAPOR1 in cancer has been evaluated in endometrial, gastric, and prostate cancers [5, 9, 10]. In pancreatic neuroendocrine tumors, ovary cancer, and CRC, only some bioinformatics analyses have shown that ELAPOR1 gene might have biological effects, albeit an in-depth investigation is lacking [6, 11, 12].
In the present study, we found that high ELAPOR1 level is associated with favorable overall survival (OS) and disease-free survival (DFS). Compared to normal mucosa, ELAPOR1was lower in CRC. The overexpression of ELAPOR1 significantly inhibited CRC cell proliferation and invasion. Conversely, its suppression by RNA interference promotes CRC cell proliferation and invasion. Transcriptome sequencing and bioinformatics analysis on the genes before and after ELAPOR1 overexpression in SW620 cells identified 355 differentially expressed mRNAs involved in receptor binding, plasma membrane, negative regulation of cell proliferation, and common cancer signaling pathways. These findings suggested that ELAPOR1 might be a prognostic indicator and a tumor suppressor in CRC.
Materials and methods
TCGA data analysis of CRC datasets
The correlation between ELAPOR1 and survival of CRC patients in The Cancer Genome Atlas (TCGA) datasets of colon adenocarcinoma and rectal adenocarcinoma was predicted using GEPIA (http://gepia.cancer-pku.cn/index.html). The cut-off value used to divide the patients into high- and low-expression groups is the median value of ELAPOR1 expression in all samples. The difference in ELAPOR1 expression between cancer and adjoining non-cancerous tissues was analyzed.
Clinical specimens and immunohistochemistry (IHC) scoring
From December 2021 to February 2022, specimens (tumor and paired normal mucosa) of 44 patients who underwent radical resection of CRC were collected from the Department of General Surgery, Jinshan Hospital, Fudan University, China. This study was approved by the clinical research ethics committee of Jinshan Hospital, Fudan University (Shanghai, China, No. JIEC2021-S57). Written informed consent was obtained from all patients before the collection of tissue samples. IHC was carried out as described previously [13]. All slides were evaluated by two investigators who were blinded to the clinicopathological characteristics of the study participants using Olympus BX43 (Tokyo, Japan) microscope. IHC scoring for ELAPOR1 expression was based on the multiplication of intensity and percentage of stained cells. The intensity was scored from 0 to 3 (0, negative; 1, weak; 2, moderate; 3, strong) and the percentage of positively stained cells was scored from 0 to 4 (1, 0–25%; 2, 26–50%; 3, 51–75%; 4, 76–100%). Three sections of each tumor were assessed for ELAPOR1 staining and the average score was calculated. Based on the IHC scores, ELAPOR1 staining patterns were classified as low-level (IHC score, 0 to 4) or high-level (IHC score, 6 to 12).
Cell culture
SW620, RKO, SW480, HT29, CACO2, HCT116 and HEK293T cells were bought from Procell Life Science and Technology Corporation (Wuhan, Hubei, China). The cells were cultured in DMEM medium (KeyGEN, Nanjing, Jiangsu, China) with 10% fetal bovine serum (Cellsera, Maitland, NSW, Australia) at 37
Plasmids and cell transfection
For overexpression, the full-length cDNA sequences of ELAPOR1 were inserted into pCMV7-3xFLAG vectors (Merck, Darmstadt, Germany) to construct pCMV7-3xFLAG-ELAPOR1 plasmids. Empty vector was used as the negative control. ELAPOR1-shRNA (NM_020775.5) targeting sequence (5’-AGAGTCTGAGTACCACTAT-3’) was designed and synthesized by EnzyArtisan (Shanghai, China). A scrambled sequence (5’-GACTTCAAATCGGGTCTAA-3’) and a nonsense sequence (5’-ACTACCGTTGTTATAGGTG-3’) were used as negative controls. These sequences were cloned into pSilencer™ 4.1-CMV (Thermo, Carlsbad, CA, USA). An equivalent of 1
Western blot analysis
The cells were lysed, and an equivalent of 30
Estimation of cell proliferation and colony formation
Cell counting kit-8 (CCK-8) assay (Dojindo, Kumamoto, Japan) was used to evaluate cell proliferation. Logarithmic phase cells were seeded in 96-well plates at the density of 2,000 cells/well. A volume of 10
Transwell and wound healing assays
Transwell chamber (Corning, New York, USA) of 8-
Transcriptome sequencing and bioinformatics analysis
Three SW620 cell samples of the ELAPOR1 group and three samples of the control group were sent toGENEWIZ (Suzhou, Jiangsu, China) for RNA extraction, quality control, library construction, and transcriptome sequencing. The sequences provided by Illumina were used for subsequent analysis. Degseq (v1.38.0) was used to screen out the differentially expressed genes (DEGs). The screening criteria for the significantly different genes were as follows:
Real-time quantitative reverse transcription PCR (qPCR) and primers
RNA Extraction Kit (Esunbio, Shanghai, China) was used to extract the total RNA of the cells, and RT Kit(Takara, Beijing, China) was used to synthesize the cDNA. The expression of ELAPOR1 and other genes was measured by fluorescence dye qPCR (Vazyme, Nanjing, Jiangsu, China). The relative gene expression was calculated using the 2
Statistical analysis
GraphPad Prism 7.0 software (GraphPad, San Diego, CA, USA) was used for data analysis. All data are expressed as mean
Prognosis analysis of CRC patients from TCGA datasets with respect to ELAPOR1 expression. Kaplan–Meier curves and log-rank tests show patients with high ELAPOR1 expression in colon cancer (A), rectal cancer (B), or both cohorts (C). CRC, colorectal cancer; TCGA, The Cancer Genome Atlas; ELAPOR1, endosome-lysosome associated apoptosis and autophagy regulator 1.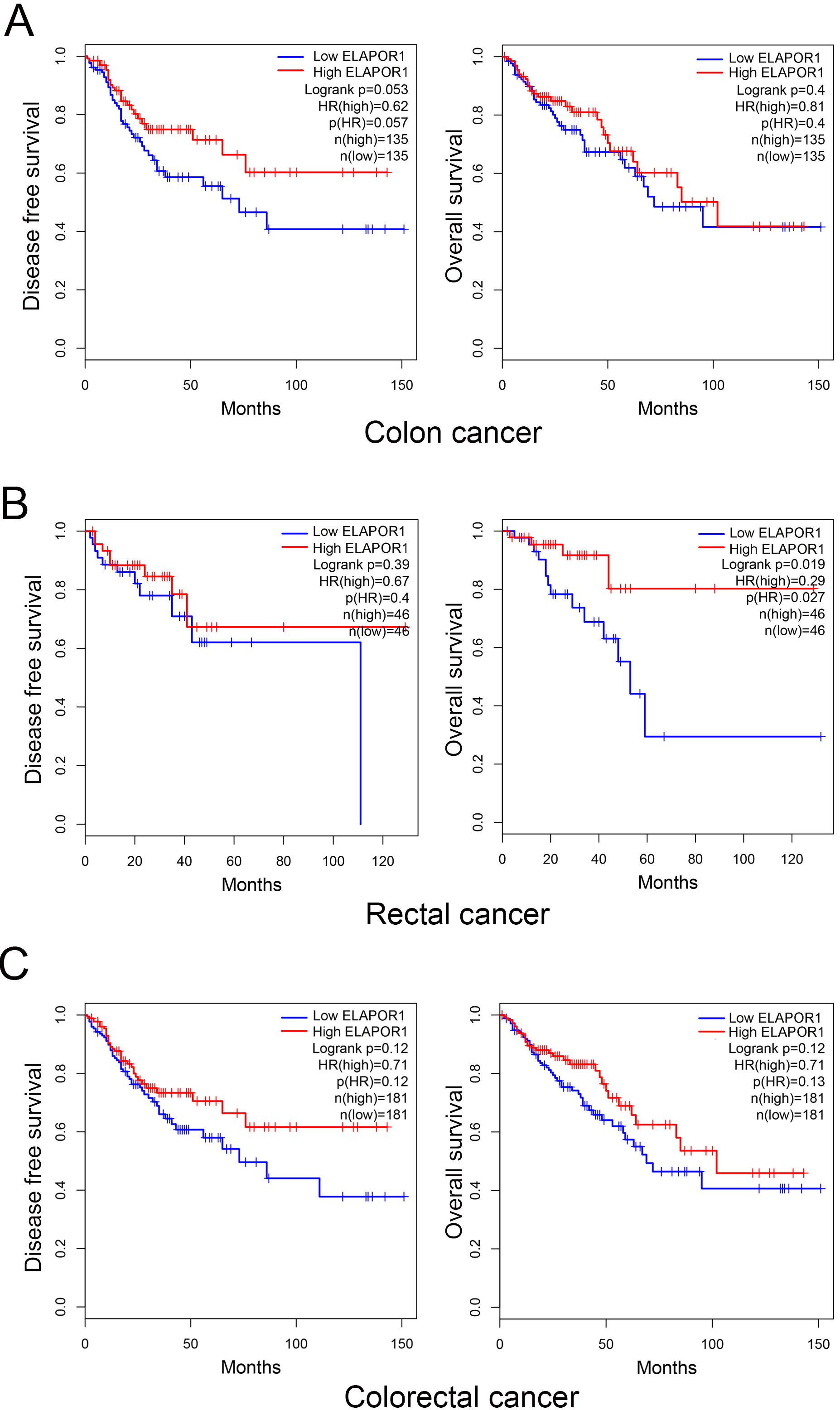
High ELAPOR1 expression is linked to a favorable prognosis in CRC patients
Kaplan-Meier curves and log-rank tests showed that patients with high ELAPOR1 expression have a favorable OS and DFS, according to GEPIA (Fig. 1). In 270 cases of colon cancer, there was an all but significant correlation between high ELAPOR1 expression level and improved DFS interval (
ELAPOR1 is downregulated in human CRC
Expression patterns of ELAPOR1 in CRC specimens. Positive ELAPOR1 expression cells were stained brown. Representative micrographs of high and low ELAPOR1 expression in normal mucosa (A–B) and tumor tissue (C–D). (E) Statistical analysis of ELAPOR1 immunostaining in CRC patients. (F) Median ELAPOR1 expression in tumor and normal tissue of CRC patients of TCGA datasets. (G) Correlation of ELAPOR1 and tumor staging in CRC patients of TCGA datasets.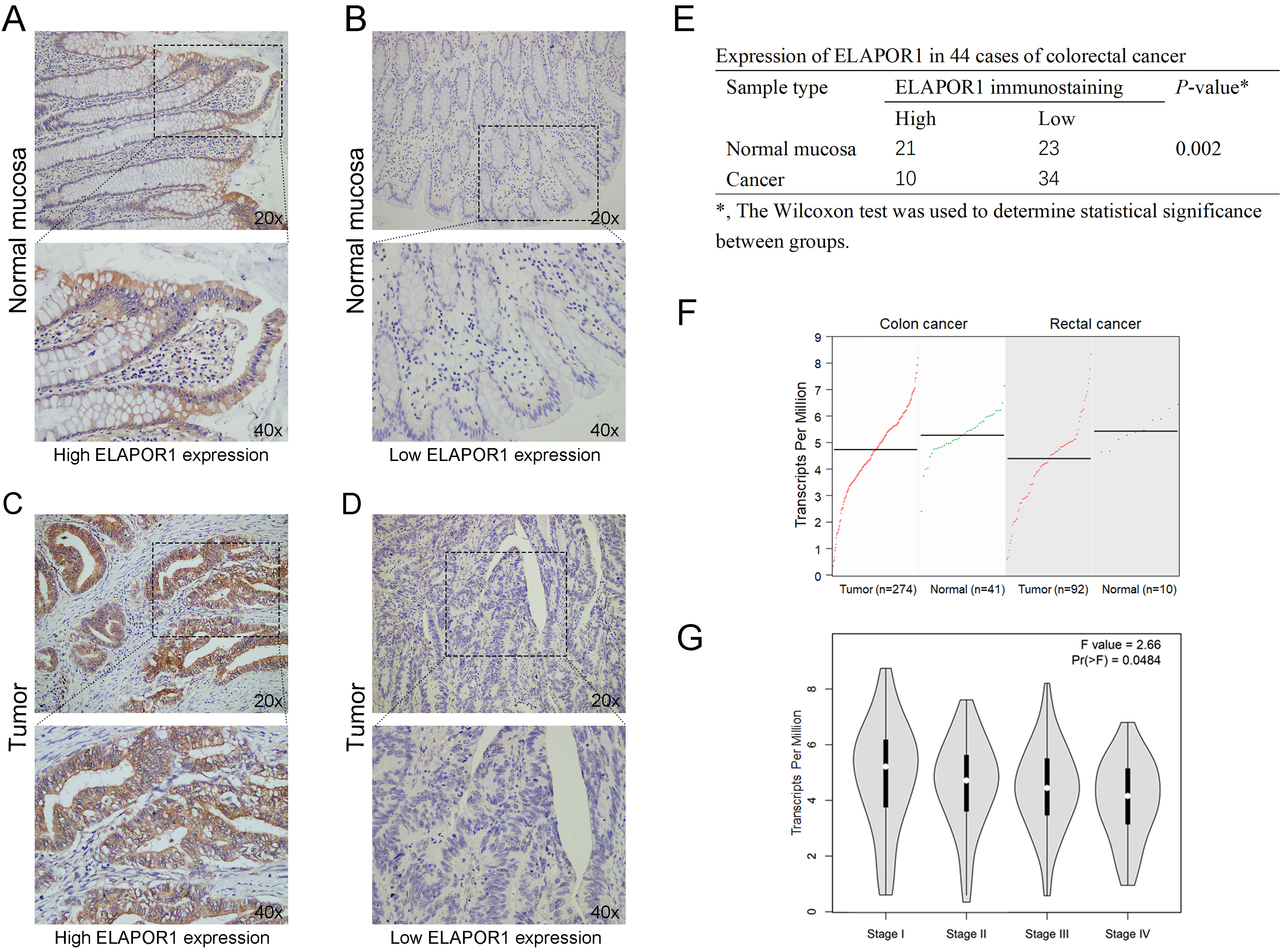
IHC was employed to detect the expression ofELAPOR1 in paraffin-embedded CRC samples (Fig. 2A–D). The results showed that ELAPOR1 is mainly expressed on the cell membrane. Compared to the adjacent normal mucosa, ELAPOR1 is lowly expressed in CRC tissues (
ELAPOR1 inhibits the proliferation and migration of CRC cells in vitro. (A) ELAPOR1 overexpression was confirmed by Western blot in the transfected SW620 and RKO cells. (B, C) Proliferation of ELAPOR1- or empty vector-transfected cells measured by CCK-8 assay and colony formation assay. (D, E) Migration of ELAPOR1- or empty vector-transfected cells evaluated by transwell and wound healing assays. OD, optical density; Scale bar, 200 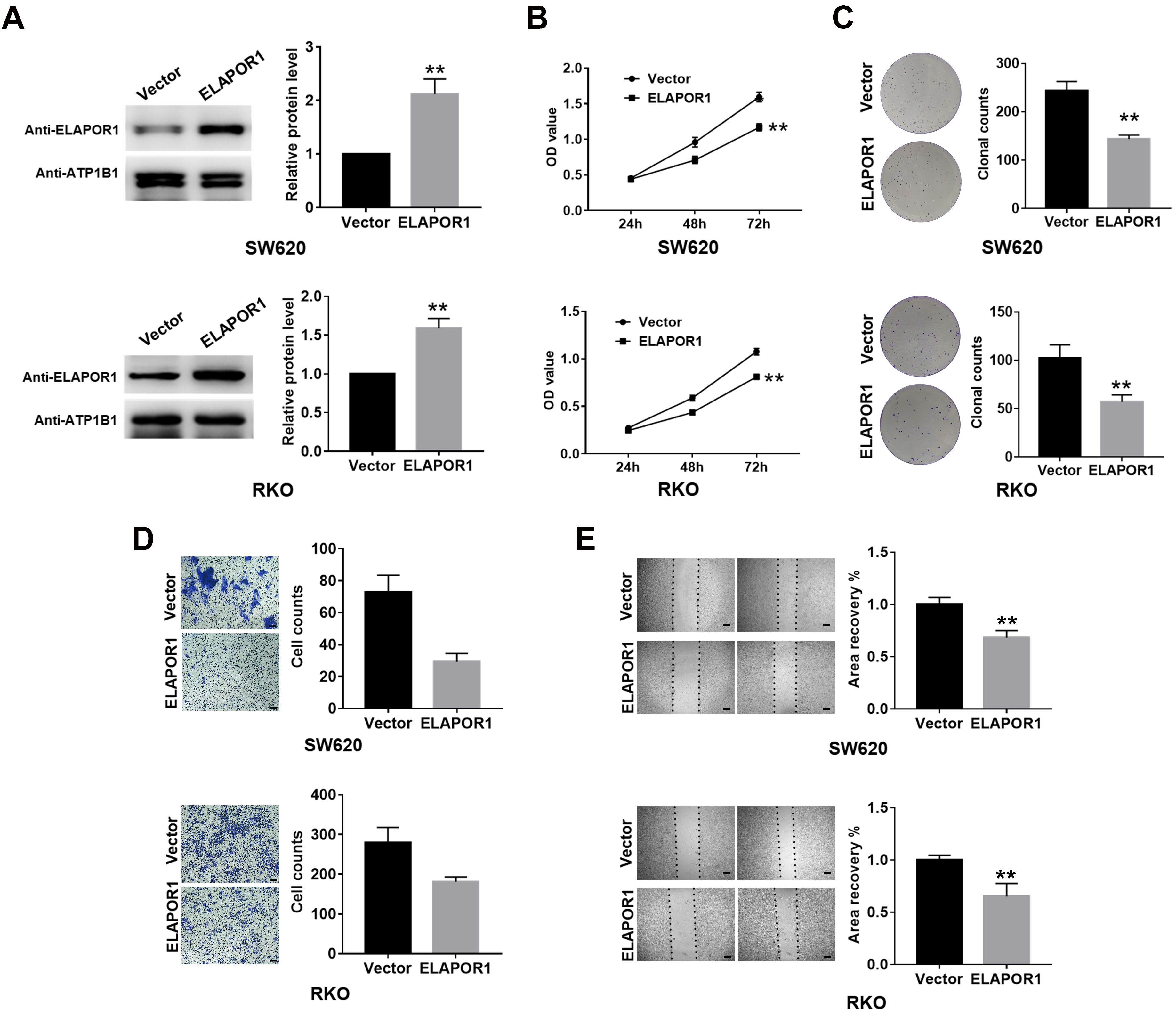
The expression of ELAPOR1 protein was confirmed by western blotting in six CRC cells including SW620, RKO, SW480, HT29, CACO2, HCT116 (Fig. S1). SW620 and RKO cells were selected for subsequent assays because they showed a relatively higher level of ELAPOR1 than the other cell lines, which means it might have a more obvious effect in cell physiological functions. We successfully transfected SW620 and RKO with ELAPOR1 plasmid or empty vector. ELAPOR1 overexpression was verified using Western blot with ELAPOR1 antibody (Fig. 3A). CCK-8 assay revealed that the number of viable SW620 and RKO cells in the ELAPOR1 group decreased significantly compared to the vector group (
ELAPOR1 knockdown promotes the proliferation and migration of CRC cells in vitro. (A) ELAPOR1 was detected by Western blot in the shRNA-transfected SW620 and RKO cells. (B, C) Proliferation of ELAPOR1-shRNA or control group measured by CCK-8 and colony formation assays. (D, E) Migration of ELAPOR1-shRNA or control group evaluated by transwell and wound healing assays. OD, optical density; Scale bar, 200 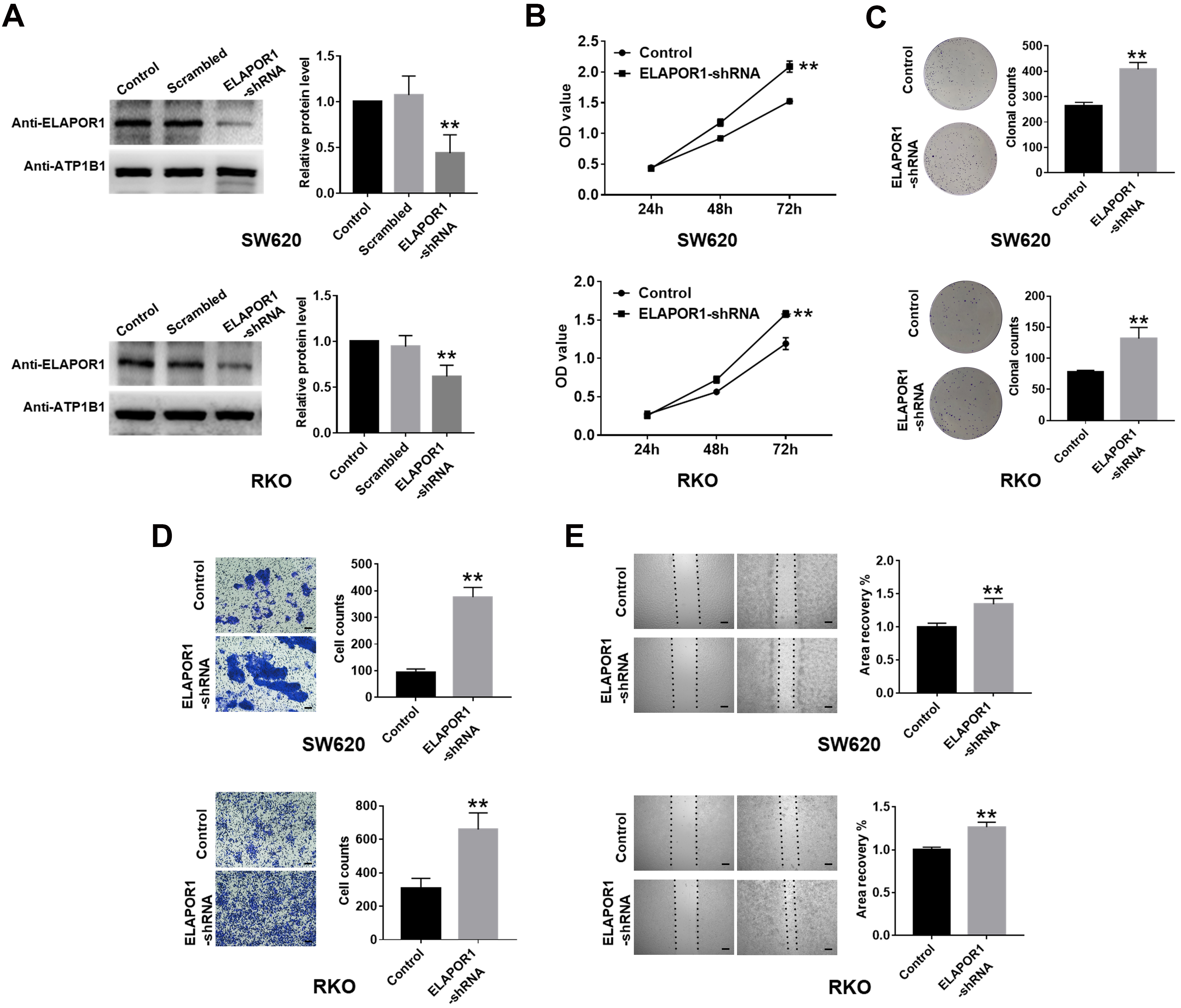
Herein, we successfully screened the appropriate ELAPOR1 gene-silencing targets and then transfected ELAPOR1-shRNA or control shRNA into SW620 and RKO cells. (Fig. 4A). CCK-8 assay revealed that the number of viable cells in the ELAPOR1-shRNA group increased significantly compared to the control group (
ELAPOR1 is associated with multiple gene sets of CRC cells
Bioinformatics analysis of DEGs related to ELAPOR1 overexpression. (A) Volcano plot showing the distribution of DEGs before and after ELAPOR1 overexpression in SW620 cells. Red color shows upregulated genes, while blue color shows downregulated genes. (B) GO shows the correlations between DEGs and predicted functions. (C) KEGG network shows the connections between the pathways and DEGs. (D) Verification of some DEGs in SW620 and RKO cells by qPCR. DEG, differentially expressed gene; 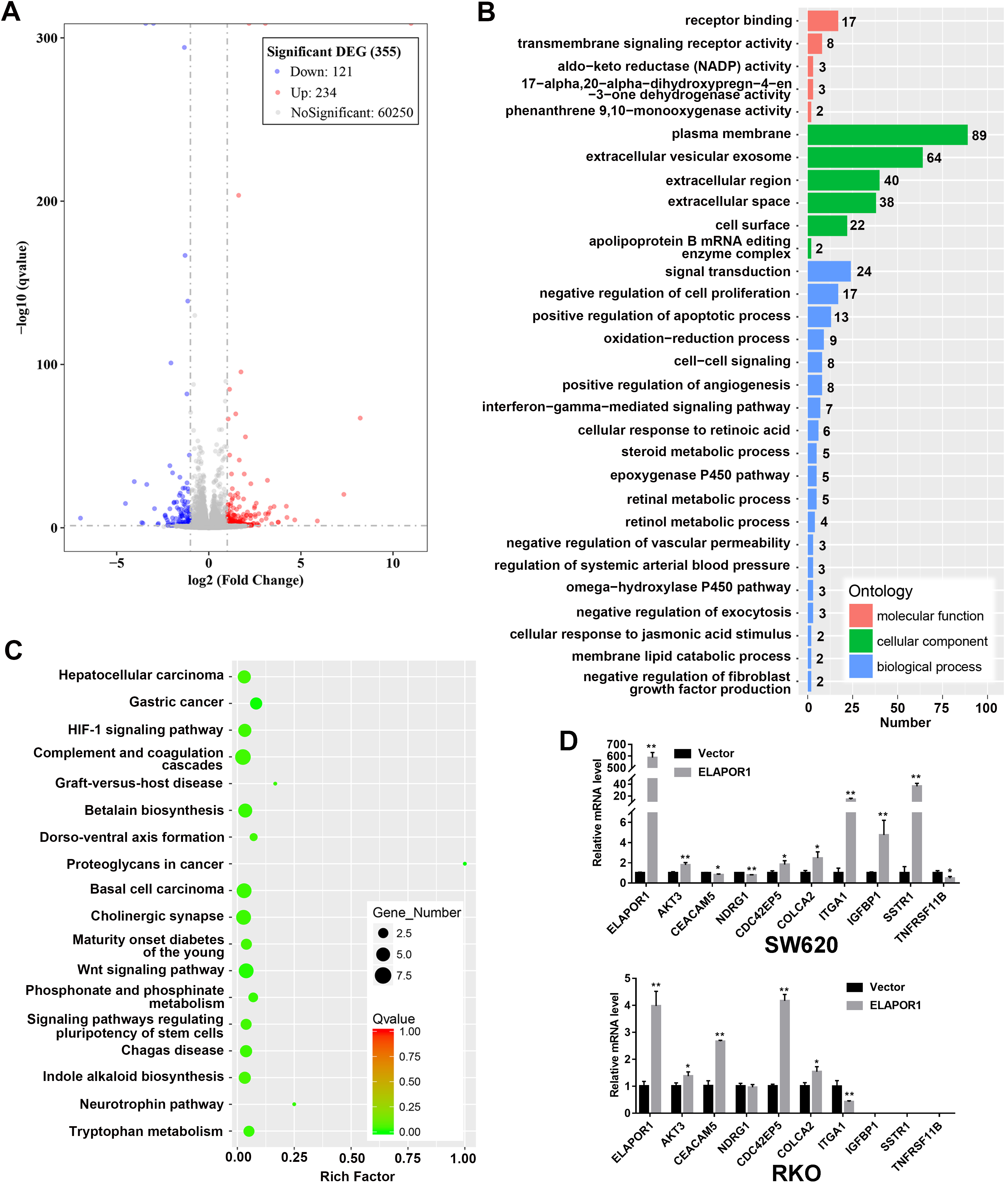
To further elucidate the potential mechanism underlying these phenomena, we conducted differential gene expression analysis using transcriptome sequencing and bioinformatics. According to the predetermined screening criteria, 355 differentially expressed mRNAs were retrieved in ELAPOR1-overexpressed SW620 cells compared to control group; among these, 234 are upregulated and 121 are downregulated (Fig. 5A, Table S2). Based on the Gene Ontology database, we found that ELAPOR1 is involved in the molecular functions of receptor binding, transmembrane signaling receptor activity, aldoketo reductase activity; cellular component of the plasma membrane, extracellular vesicular exosome; the biological process of signal transduction, negative regulation of cell proliferation, and positive regulation of apoptotic process (Fig. 5B). Based on the KEGG pathway significance enrichment analysis, we observed that these differential genes are significantly involved in common cancer signaling pathways, including WNT signaling pathway, gastric cancer, basal cell carcinoma, hepatocellular carcinoma, signaling pathways regulating pluripotency of stem cells, proteoglycans in cancer, and HIF-1 signaling pathway (Fig. 5C). In the subsequent qPCR assays on SW620 and RKO cells, the mRNA levels of AKT3, CDC42EP5, and COLCA2 in the above pathways were significantly different between ELAPOR1 and empty vector groups (Fig. 5D). Notably, the mRNA expression of CEACAM5, NDRG1, and ITGA1 showed inconsistent results in RKO cells, while that of TNFRSF11B, IGFBP1, and SSTR1 could not be detected (Fig. 5D).
In the present study, the expression of ELAPOR1 was found to be favorably correlated with the prognosis of CRC patients from TCGA datasets. It decreased gradually with the deterioration of the tumor stage. Moreover, its expression is decreased in tumor tissues compared to the paired normal mucosa in CRC patients from our cohort. These phenomena indicated that ELAPOR1 might be correlated with CRC progression. To further explore the function of ELAPOR1, we examined its expression in two CRC cell lines and observed that the overexpression inhibits the proliferation and invasion ability of the cells, while suppression promotes proliferation and enhances invasiveness. Transcriptome sequencing identified the gene sets of ELAPOR1 regulating CRC cell biology, which might involve transmembrane signaling receptor activity, negative regulation of cell proliferation, and other common cancer signaling pathways. The current study revealed that ELAPOR1 is a potential anti-oncogene in CRC. This finding is consistent with those from studies on endometrial, gastric, and prostate cancer.
Membrane-binding proteins are associated with the prognosis of many cancer patients [16, 17]. ELAPOR1 is a transmembrane protein that might play an important physiological role in tumor progression. Our immunohistochemistry is consistent with the literature, suggesting that it is mainly expressed on the membrane of CRC tissues. According to bioinformatics analysis, ELAPOR1 may be one of the genes related to malignant transformation of colorectal adenoma [12]. High expression of ELAPOR1 mRNA is also found in pancreatic neuroendocrine tumors with good prognosis [11]. Kang et al. demonstrated that low ELAPOR1 levels were associated with poor prognosis in patients with gastric cancer [9]. These findings, together with our results, suggested that as a transmembrane protein, ELAPOR1 might be a good prognostic marker for CRC. However, due to the limited number of cases, the DFS and OS of total CRC cases could not reach statistical significance. But our findings are still valuable. Since colorectal cancer has a high incidence and mortality rate, it is important to conduct high-quality observational cohort studies with a larger number of samples to establish the statistical significance of ELAPOR1 in the prognosis of CRC patients and to further investigate the association of ELAPOR1 expression levels to CRC clinicopathological characteristics such as tumor location, TNM staging, tumor budding and microsatellite status etc.
Membrane proteins are promising therapeutic targets of cancers [16, 18]. The current study demonstrated that ELAPOR1 has an inhibitory role in CRC, which is consistent with the different expression patterns between normal and malignant specimens. Kang et al. showed that ELAPOR1 activates Caspase-3 and Caspase-7 by blocking the carcinogenic activity of GRP78, reduces AKT signal, inhibits the growth, invasion, and tumorigenic activity of gastric cancer cells, and induces apoptosis, rendering it as a promising therapeutic membrane target of CRC. However, it is found that ELAPOR1 is overexpressed in some highly metastatic breast cancer and lung cancer cell lines, while it is a splice variant of its truncated protein expression in non-metastatic cancer cells and normal epithelial cells. Although in breast cancer, ELAPOR1 overexpression inhibits tumor growth, but its knockdown does not promote tumor growth; In view of its functional differences in tumor cells from different tissues, we carefully designed the overexpression and knockdown assays to demonstrate the role of ELAPOR1 in CRC cells. Noticeably, ELAPOR1 knockdown has the opposite effect on breast cancer under pressure, which is similar to TGF-
ELAPOR1 has a large molecular weight of 100 kDa. It consists of a transmembrane segment, an extracellular segment, and an intracellular peptide segment of 30 amino acids [5]. This composition is similar to the receptor tyrosine kinase, integrin, ligand, and cytokine receptors, which are various membrane-binding proteins that characterize the cells [19, 20, 21]. Our transcriptome data suggested that similar to a multifunctional molecule, ELAPOR1 may participate in the biological function of CRC cells through various pathways. Although the phenotypes of CRC cell proliferation and invasion in our study were consistent with those described previously, significant differences were noted in gene expression between different CRC cell lines (Fig. 5D), which could be ascribed to the differences between individuals.
Conclusion
In summary, the current study suggested that ELAPOR1 contributes to tumor suppression of CRC. The underlying mechanisms of ELAPOR1 activity might lead to its use in prognosis prediction or treatment of CRC patients.
Author contributions
Conception: Anzhong Huang.
Interpretation or analysis of data: Mengting Wu, Dawei Zhang, and Guangbin Wu.
Preparation of the manuscript: Anzhong Huang and Chunzhi Qin.
Revision for important intellectual content: Chunzhi Qin.
Supervision: Peilong Sun.
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-220285.
sj-docx-1-cbm-10.3233_CBM-220285.docx - Supplemental material
Supplemental material, sj-docx-1-cbm-10.3233_CBM-220285.docx
sj-xls-1-cbm-10.3233_CBM-220285.xls - Supplemental material
Supplemental material, sj-xls-1-cbm-10.3233_CBM-220285.xls
Footnotes
Acknowledgments
The study was funded by the Medical and Health Science and Technology Innovation Fund Project in Jinshan District, Shanghai, China (No. 2020-3-19).
Conflict of interest
The authors declare that they have no conflict of interest.
