Abstract
cAMP response element binding protein (CREB)-regulated transcription coactivator 2 (CRTC2), a member of the novel CRTC family of transcriptional coactivators that activates basic leucine zipper transcription factors, including CREB, is overexpressed in many carcinomas, including colon cancer. Phosphorylation of CRTC2 protein at different residues is important for its subcellular localization and activity. However, the functions of some of the serine phosphorylation sites have not been elucidated. This study aimed to investigate the effects of phosphorylation of Ser127, Ser238, and Ser245 sites of CRTC2 in colorectal cancer (CRC) cells. Recombinant lentiviral particles with a CRTC2-targeting small hairpin RNA (shRNA) sequence were transfected into CRC cells to obtained shCRTC2 cell lines. Site-directed mutagenesis of Ser127, Ser238, and Ser245 cells were constructed by transfecting CRTC2 cDNA containing S127A, S238A, and S245A mutations into shCRTC2. Cell proliferation was measured by cell counting kit-8. Cell migration and invasion were examined by transwell assay. mRNA expression was assayed by qRT-PCR, and protein expression was determined by Western blot. Our results indicate that CRTC2 is overexpressed in CRC cells. Knockdown of CRTC2 inhibits the proliferation, migration, and invasion of CRC cells. When the phosphorylation of CRTC2 Ser238 decreases due to the lack of ERK2, the phosphorylation of Ser171 site increases. The proliferation, migration and invasion of CRC cells were inhibited, the nuclear aggregation of CRTC2 in the nucleus was reduced, and the interaction between CRTC2 and CREB was weaken. It is shown that the phosphorylation of CRTC2 Ser238 is important for CREB transcriptional activity. These findings may help in the identification of potentially new targets for CRC therapy.
Introduction
Colorectal cancer (CRC) is a common malignant tumor of the digestive system. 1 The incidence and mortality rates of CRC has been increasing steadily over the past few decades, and the 5-year survival rate is about 50%. 2 Invasion and metastasis of CRC cells are the major factors that lead to the death of patients. Therefore, studying the process and mechanism of CRC invasion and metastasis will help in the identification of key functional areas and new targets for the prevention and treatment of CRC.
cAMP response element-binding protein (CREB) is an important member of the basic leucine zipper (bZIP)-containing family of transcription factors. CREB is a ubiquitous transcription factor that serves a variety of biological functions, including cellular proliferation, differentiation, and adaptive responses in the neuronal system. 3 -9 Increasing evidence suggests that CREB is involved in the transformation and immortalization of cancer cells, and is directly involved in the development of multiple cancers. 10 -12 CREB-regulated transcriptional coactivators (CRTCs), comprise a new family of transcriptional coactivators that activate bZIP transcription factors, including CREB. 13 The CRTC family consists of 3 members: CRTC1, CRTC2, and CRTC3. CRTC1 is mainly expressed in the brain, while CRTC2 and CRTC3 are abundantly expressed in most peripheral tissues. 13 Among the family members, CRTC2 acts as a coactivator of CREB and mediates multiple cAMP-dependent transcriptional pathways in metabolic tissues. Activation of CRTC2 is enhanced by the dephosphorylation of the serine 171 (Ser171) residue and subsequent nuclear translocation of CRTC2. 14 -16 Studies have shown that 28 residues in the amino terminal of CRTC2 form an α-helical structure that directly binds to the leucine zipper region of CREB. As the alpha helix of CRTC2 lacks basic amino acids, CRTC2 cannot directly bind to DNA, but promotes strong interactions between CREB and its homologous DNA (including CRE sequences) through nonspecific interactions with DNA phosphate groups. 17 CRTC2 is abundantly expressed in the liver, and the CREB pathway is activated, thereby stimulating the glycogen stored in the liver to produce glucose under fasting conditions. 18 In the liver, CREB and CRTC2 are involved in the transcriptional control of gluconeogenesis via direct activation of the gluconeogenesis-related genes and indirect activation of PGC-1α. 16 Several studies have shown that the phosphorylation of CRTC2 protein at different sites is important for its subcellular localization and activity. Under normal physiological conditions, CRTC2 is phosphorylated at Ser171 and Ser275 by salt-induced kinase 1 (SIK1) and microtubule affinity-regulated kinase 2 (MARK2), respectively. 19,20 Phosphorylation-dependent interaction with the 14-3-3 proteins subsequently causes CRTC2 to be sequestered in the cytoplasm, thus preventing CRTC2 from binding to CREB and leading to transcriptional activation of CREB. 21,22 In the liver, Ser275 phosphorylation is also regulated by extracellular glucose. 23 In mouse COS-7 cells, phosphorylation of CRTC2 at Ser307 plays an identical role as Ser171, to synergistically regulate the activity of TORC2 and block the nuclear localization of TORC2. 24
Few studies have implicated CRTC2 in the promotion and development of cancer. In prostate cancer, the expression of CRTC2 was significantly higher in patients with high Gleason scores compared to that in patients with low Gleason scores. Therefore, CRTC2 is considered to be significantly correlated with pathological outcome and postoperative survival. 25 In liver kinase B1 (LKB1)-mutant non–small cell lung cancer (NSCLC), CRCT2 is constitutively activated because of inactivation of SIKs, and promotes NSCLC progression by binding with CREB to initiate the transcription of downstream genes. 26,27 However, CRTC2 acts as a repressor gene in lymphomas and maintains the integrity of the genome by stimulating the transcription of the mismatch repair (MMR) genes. 28
We found that CRTC2 was overexpressed in CRC cells. However, its function in CRC remains unclear. As the dephosphorylation of Ser171 is very important for the nuclear localization of CRTC2, determination of the functional significance of phosphorylation of Ser127, Ser238, and Ser245 included analysis based on Ser171 to A171 mutation and investigation of the effect on CRTC2 activity. In the present study, we demonstrate that Ser238 of CRTC2 is phosphorylated by the kinase ERK2, thereby inhibiting phosphorylation of Ser171 and enhancing the transcriptional activity of CREB, to promote CRC cell proliferation and migration.
Materials and Methods
Cell Lines
Normal human colonic epithelial cell line, NCM460, was obtained from Beijing Beina Chuanglian Biotechnology Research Institute (Beijing, China). The 2 colon cancer cell lines (SW620 and T84) were obtained from the American Type Culture Collection (ATCC, Manassas, VA, USA). NCM460 cells were cultured in the Roswell Park Memorial Institute 1640 (RPMI 1640) medium, SW620 cells were cultured in L15 medium, and T84 cells were cultured in Ham’s F12 medium and Dulbecco’s modified Eagle medium (1:1 mixture). These 3 culture media were supplemented with 2.5 mM L-glutamine, 10% fetal bovine serum (ProSpec-Tany TechnoGene, Ltd., Rehovot, Israel), and 100 mg/mL penicillin/streptomycin (ProSpec-Tany TechnoGene, Ltd.), and the cells were cultured in a humidified 5% CO2 atmosphere at 37 °C.
Plasmid Construction
The human CRTC2-targeting small hairpin RNA (shRNA) sequences were acquired from Sigma-Aldrich (St. Louis, MO, USA, MISSION® shRNA TRCN0000229832 and TRCN0000229831). A Genechem-CRTC2 plasmid expressing the full-length human CRTC2 protein was purchased from Genechem (Shanghai, China), while an empty plasmid was used as a negative control. CRTC2 cDNA was cloned into the pcDNA3.1 vector (GENEray Biotechnology, Shanghai, China) to construct an overexpression plasmid pdCRTC2. Hieff MutTM Site-Directed Mutagenesis Kit (Yeasen Biotech Co., Ltd., Shanghai, China, Cat. No. 11003ES10) was used to mutate Ser171, Ser127, Ser238, and Ser245 of CRTC2 to Ala171, Ala127, Ala238, and Ala245, respectively. The sequences used for site-directed mutagenesis of Ser171, Ser127, Ser238, and Ser245 are listed in Table 1. CRTC2 cDNA with Ser171 mutated to Ala171 was cloned into the pcDNA3.1 vector to construct the plasmid pdA171. CRTC2 cDNA with Ser127 mutated to Ala127 was cloned into the pcDNA3.1 vector to construct the plasmid pdA127. Plasmids pdA238 and pdA245 were constructed in a similar manner.
The Site-Directed Sequences for Site-Directed Mutating.

Cell Transfection
Recombinant lentiviral particles with CRTC2-targeting small hairpin RNA (shRNA) sequence were generated, and CTC cells were transfected with CRTC2 or negative control recombinant lentivirus (shRNA-CRTC2 or shRNA-NC, respectively). Control siRNA (131905) and siRNAs for CDK1 (103821), AKT2 (103305), PKCa (142283), TBK (134002), ERK2 (1081), and MNK1 (111232) were purchased from Invitrogen (Carlsbad, CA, USA), and transformed into SW620 cells to obtain the corresponding siRNA cell lines. The overexpression plasmids pdCRTC2, pdA171, pdA127, pdA238, and pdA245 were transfected into the shCRTC2 cells to obtain shCRTC2 + WT, shCRTC2 + A171, shCRTC2 + A127, shCRTC2 + A238, and shCRTC2 + A245 cell lines. Transfection was performed using Oligofectamine™ (Invitrogen) according to the manufacturer’s instructions. The transfection efficiency was determined using quantitative real-time reverse transcription-polymerase chain reaction (qRT-PCR).
RNA Extraction and qRT-PCR
RNA was extracted using the QIAzol™ lysis reagent (Qiagen, Hilden, Germany) and purified using RNeasy mini kit (Qiagen). Purified RNA was treated with DNase I (New England Biolabs, Ipswich, MA, USA) to remove genomic DNA. The quality and quantity of the RNA were analyzed using a NanoDrop2000 (Thermo Fisher Scientific Inc., Waltham, MA USA), and Bioanalyzer (Agilent Technologies Inc, CA, USA. A total of 1000 ng RNA was reverse transcribed into cDNA in a final volume of 20 µL using a reverse transcription kit (Cat: RR036A; Takara, Shiga, Japan). qRT-PCR was performed with the SYBR Select Master Mix (Cat: 4472908; Applied Biosystems, Foster, CA, USA). The primers used for CRTC2, CREB, cyclin A1 (CCNA1), cyclin D1 (CCND1), B-cell leukemia 2 (BCL2), vascular endothelial growth factor (VEGF), melanoma cell adhesion molecule (MCAM), cyclin dependent kinase 1 (CDK1), AKT serine/threonine kinase 2 (AKT2), protein kinase C alpha (PKCa), TANK binding kinase 1 (TBK1), mitogen-activated protein kinase 1 (ERK2), MAPK interacting serine/threonine kinase 1 (MNK1), and β-actin (ACTB) genes are listed in Table 2. The QuantStudioTM 6 Flex Real-Time PCR System was used to perform the qRT-PCR analysis. The qRT-PCR reaction consisted of an initial denaturation step at 95 °C for 10 min, followed by 40 cycles of 93 °C for 15 s, and elongation at 60 °C for 1 min. Each sample was tested in triplicate, and the relative expression was calculated using the 2−ΔΔCt method and normalized relative to ACTB expression.
The Primers of Gene for qRT-PCR.

Western Blotting
Cells were harvested and extracts were prepared using lysis buffer (50 mM Tris-HCl [pH 7.4], 150 mM NaCl, 1%TritonX-100, 0.2% sodium dodecyl sulfate [SDS], 10 mM β-mercaptoethanol and 5% glycerol) on ice. The protein concentration in the cell extract was quantitated using a BCA Kit (KeyGEN, Pierce, USA). Cell extracts were boiled for 5 min in loading buffer, and equal amounts of protein were separated by sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE). The separated protein bands were transferred onto polyvinylidene fluoride membranes. The membranes were blocked in 5% skim milk powder in Tris-buffered saline/Tween 20 (TBS-T) for 1 h and subsequently incubated overnight at 4 °C with primary antibodies against CRTC2 (ab203187, 1:1000), CREB (ab32515, 1:1000), Cyclin A1 (ab53699, 1:1000), Cyclin D1 (ab40754, 1:3000), bcl-2 (ab32124, 1:1000), VEGF (ab32152, 1:3000), MCAM (ab134065, 1:5000), β-tubulin (ab210797, 1:1000), Lamin B1 (ab133741, 1:2000), β-actin (ab179467, 1:5000), phosph-CRTC2-Ser171 (ab203187, 1:800) or GAPDH (ab181602, 1:10,000), all purchased from Abcam (Cambridge, MA, USA). The primary antibodies to phospho-CRTC2-Ser127 (1:500), phospho-CRTC2-Ser238 (1:1000), and phospho-CRTC2-Ser245 (1:700) were from Beijing GeneX Health Co., Ltd (Beijing, China). After washing with TBS-T 3 times, the membrane was incubated with a goat anti-rabbit secondary antibody (ab7090, Abcam, 1:10,000) at 37 °C for 2 h. All experiments were independently repeated at least 3 times.
In Vitro Proliferation, Migration, and Invasion Assays
Cell proliferation was examined using Cell Count Kit-8 (CCK-8) assay. Cells were plated at a density of 1000 cells/well in 96-well plates, and cultured at 37 °C in a humidified 5% CO2 atmosphere. Prior to analysis of proliferation, 10 µL of CCK8 reagent (Beijing Solarbio Science & Technology, Beijing, China) was added to each well and incubated at 37 °C for an additional 1.5 h. The optical density was measured at 450 nm using an ELx-800 universal microplate reader (BioTek, CA, USA). The cell number was analyzed and a cell growth curve was generated. Each experiment was repeated 3 times. Cell migration and invasion assays were performed in 6.5-mm Transwells (#3422, Corning, NY, USA). Cells (2 × 105) suspended in 100 µL of serum-free medium were added to the upper chamber, and the lower chamber was filled with complete medium with 10% serum. The cells were allowed to migrate following incubation at 37 °C for 48 h. After removing the non-migrated cells, the membranes were fixed in methanol and stained with 0.05% crystal violet. The migrated cells were photographed and quantified in 5 random fields per membrane. Each sample was assayed in triplicate. For the invasion assay, the Transwells were first coated with Matrigel (BD Biosciences, NY, USA) and cells were allowed to invade for 72 h.
Statistical Analysis
All statistical analyses were carried out using the SPSS 18.0 statistical software package. All values are depicted as the mean ± standard deviation. All data were statistically analyzed using one-way analysis of variance (ANOVA) with the Bonferroni correction. The Mann-Whitney U test was used for statistical analysis of 2 groups of independent data. P < 0.05 was considered statistically significant.
Results
Overexpression of CRTC2 in CRC Cells
The expression of CRTC2 and its associated transcription factor, CREB, was analyzed in normal intestinal epithelial cell line, NCM460, and CRC cell lines, SW620 and T84, using western blotting and qPCR. The results showed that both mRNA and protein levels of CRTC2 and CREB were significantly higher in the CRC cells than those in NCM460 (Figure 1). The mRNA levels of CRTC2 and CREB were 3–6 times and 4–6 times higher, in the 2 CRC cell lines compared to NCM460 cells, respectively (Figure 1A). Consistent with the mRNA level, CRTC2 and CREB protein levels were also higher in the 2 CRC lines (Figure 1B).

CRTC2 and CREB are overexpressed in CRC cells. (A) mRNA and, (B) protein levels of CRTC2 and CREB in the 2 CRC cell lines, SW620 and T84, were higher than those in the normal intestinal epithelial cells, NCM460. mRNA expression of CRTC2 and CREB in the cell lines was assessed using RT-qPCR. β-actin was used a reference gene (n = 3).
Knockdown of CRTC2 Affects Proliferation, Migration, and Invasion of CRC Cells
An shRNA lentiviral vector that specifically targeted CRTC2 was constructed and used to infect SW620 and T84 cells, after packaging the virus. The results showed that the relative level of CRTC2 mRNA and protein in the 2 shRNA cell lines was significantly (p < 0.01) reduced. The protein and mRNA levels in shCRTC2 group were 5%–20% (Figure 2A) and 10–20% (Figure 2B), respectively, of that in the shCtrl group according to western blotting and qRT-PCR analyses. Analysis of cell proliferation using CCK-8 assay showed that following the downregulation of CRTC2, the proliferation ability of SW620 and T84 cells was significantly reduced (Figure 2C). After 72 h, the number of shCRTC2 cells was only approximately half of that in the control group. Transwell assay was performed to analyze the effect of CRTC2 on cell invasion and migration. As shown in Figure 2D, migration and invasion of cells in the shRNA group of SW620 and T84 cell lines were significantly inhibited. Western blot analysis showed that the level of EMT-related protein, E-cadherin, was upregulated while that of vimentin was downregulated (Figure 2E). These results indicate that CRTC2 plays an important role in the proliferation, invasion, and migration of CRC cells.

Knockdown of CRTC2 affected the proliferation, migration, and invasion of CRC cells. (A) Protein and, (B) mRNA expression of CRTC2 was downregulated in the shCRTC2 cell lines. (C) Growth curve showing the cell count of SW620 and T84 cells. (D) Transwell assay, and relative migration and invasion activity of shCtrl and shCRTC2 cell lines. (E) Analysis of expression of EMT-associated proteins. Compared to the shCtrl cells, the shCRTC2 cells showed increased expression of E-cadherin, and decreased expression of vimentin. The Kolmogorov-Smirnov test was used for statistical analysis in panels 2C and 2D.
Phosphorylation of the Ser238 site of CRTC2 Inhibits Phosphorylation of Ser171 and Promotes the Proliferation, Invasion, and Migration of CRC Cells
Studies have shown that different phosphorylation sites of CRTC2 protein are involved in the execution of different functions in cells; for example, the phosphorylation and dephosphorylation of Ser171 are very important for the function of CRTC2. 15,16,21 Dephosphorylation of Ser171 promotes the translocation of CRTC2 into the nucleus. 22 However, there are no reports on the functional relevance of the other known CRTC2 phosphorylation sites including Ser127, Ser238, and Ser245. We mutated Ser127, Ser238, Ser245, and Ser171 phosphorylation sites to Ala in SW620 cells and the phosphorylation levels of the derived cell lines were analyzed using western blot. shCRTC2 + WT cell lines were shCRTC2 cells containing CRTC2 cDNA; shCRTC2 + A171 cell lines were shCRTC2 cells containing CRTC2 cDNA with Ser171 mutated to Ala171; shCRTC2 + A127, shCRTC2 + A238, and shCRTC2 + A245 cell lines were shCRTC2 cells containing CRTC2 cDNA with Ser127, Ser238, or Ser245 mutated to Ala127, A238, or Ala245, respectively. Western blot analysis showed that the mutants were successfully constructed. Unexpectedly, we found a negative correlation between the phosphorylation levels of Ser171 and Ser238 (Figure 3A). In the shCRTC2 + A171 cell line, the phosphorylation level of Ser238 was significantly higher than that in the shCtrl cell line, and the phosphorylation level of Ser171 was markedly increased in the CRTC2 + A238 cell line. The effects of these site-directed mutations on proliferation, invasion, and migration of SW620 cells were then analyzed (Figure 3B, 3C, and 3D). The results showed that shCRTC2 + A238 cells were significantly (p < 0.05) inhibited, while shCRTC2 + A171 cells were slightly promoted, although not significantly, which was probably due to the low original phosphorylation level of Ser171 in shCtrl cells.

Phosphorylation of CRTC2 at Ser238 residue affected the migration, invasion, and proliferation of CRC cells. (A) Western blot analysis of the phosphorylation sites of CRTC2. Following mutation of the phosphorylation site is, the degree of phosphorylation of the corresponding site was significantly reduced. Phosphorylation level of Ser238 was significantly higher in shCRTC2 + A171 cell line, than that in shCtrl cell line. Similarly, the phosphorylation level of Ser171 was markedly higher in CRTC2 + A238 cell line. (B) Transwell assay, and (C) relative migration and invasion activity of shCtrl, shCRTC2, shCRTC2 + WT, shCRTC2 + A171, shCRTC2 + A127, shCRTC2 + A238, and shCRTC2 + A245 cell lines. (D) Growth curve showing cell count of shCtrl, shCRTC2, shCRTC2 + WT, shCRTC2 + A171, shCRTC2 + A127, shCRTC2 + A238, and shCRTC2 + A245 cell lines. For details about the cell lines, see the methods section.
Phosphorylation of Ser238 Site May Inhibit the Phosphorylation of Ser171
Studies have shown that Ser171 phosphorylation can inhibit CRTC2 from entering the nucleus, thus preventing it from binding to CREB and stimulating the transcriptional activity of CREB. To verify the effect of CRTC2 Ser238 phosphorylation at the Ser171 site, the distribution of CRTC2 in the cytoplasmic and nuclear compartments was studied. Although the mRNA and protein levels of CRTC2 were not significantly affected compared to those in shCtrl, the nuclear distribution of CRTC2 was affected and the nuclear aggregation of CRTC2 in shCRTC2 + A238 cells was significantly decreased compared to that in shCtrl and shCRTC2 + A171 (Figure 4A). We evaluated the interaction of CRTC2 and CREB in shCRTC2 + A171 and shCRTC2 + A238 cell lines, simultaneously. The interaction between CRTC2 and CREB was distinctly observed in CRTC2 + A171 cells, while in the shCRTC2 + A238 cells, the interaction was significantly weak (Figure 4B). The expression of the downstream genes of CREB (Cyclin A1, Cyclin D1, Bcl-2, VEGFA, and MCAM) was detected subsequently, and both the mRNA (Figure 4C) and protein (Figure 4D) levels were significantly downregulated in CRTC2 + A238 cells. These results indicate that phosphorylation of Ser238 and Ser171 antagonize each other, thus showing a negative correlation. Phosphorylation of Ser238 promotes the interaction of CRTC2 and CREB, thereby enhancing the transcriptional activity of CREB.
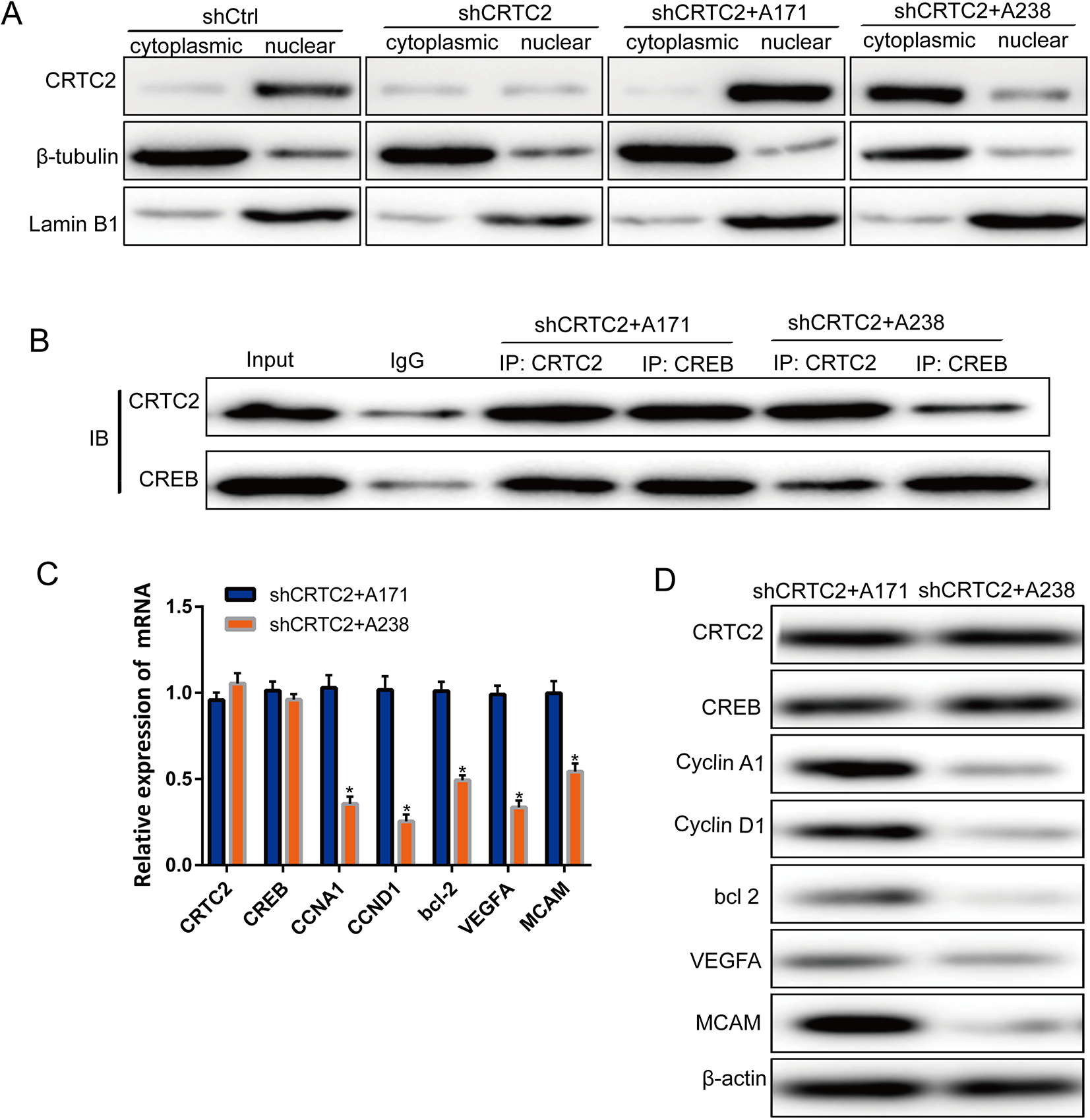
Phosphorylation of CRTC2 at Ser238 residue affected the transcriptional activation of CREB. (A) Protein expression of CRTC2 in cytoplasmic and nuclear fractions of shCtrl, shCRTC2, shCRTC2 + A171, and shCRTC2 + A238 cell lines. The nuclear distribution of CRTC2 increased significantly in shCRTC2 + A171 cell line, while it was reduced in shCRTC2 + A238 cell lines. (B) The interaction between CRTC2 and CREB, detected by co-immunoprecipitation in shCRTC2 + A171 and shCRTC2 + A238 cells. An interaction was detected between CRTC2 and CREB in shCRTC2 + A171 cells. (C) mRNA and (D) protein expression downstream of CREB were downregulated in the shCRTC2 + A238 cells compared to shCRTC2 + A171 cells. The Kolmogorov-Smirnov test was used for statistical analysis in Figure 4C.
Ser238 Site of CRTC2 Is Phosphorylated by the Kinase ERK2
The kinases that can phosphorylate the Ser238 site of CRTC2 were predicted using the online tools, PHOSPHNET (http://www.phosphonet.ca/default.aspx) and GPS 5.0-Kinase-specific Phosphorylation Site Prediction (http://gps.biocuckoo.cn/wsresult.php?p=5). The analysis revealed that CRTC2 Ser238 site may be phosphorylated by the phosphokinases (PKs) CDK1, AKT2, PKCa, TBK, ERK2, or MNK1. We used siRNA to knockdown the expression of these PKs (Figure 5A), and found that the knockdown of AKT2 and ERK2 significantly affected the phosphorylation of CRTC2 Ser238. The protein expression of CRTC2 in siERK2 was comparable to that in NC, while it was clearly reduced in siAKT2 (Figure 5B). The reduced phosphorylation of CRTC2 Ser238 in the siATK2 cell line may be due to the suppression of CRTC2 protein expression. The results indicate that ERK2 may be a kinase that phosphorylates CRTC2 Ser238.

ERK2 phosphorylated CRTC2 at the Ser238 residue. mRNA (A) and protein (B) expression of PKs that may phosphorylate CRTC2 at the Ser238 site, analyzed using qPCR and western blotting. (C) pCRTC2-Ser238 and CRTC2 protein expression in NC siRNA and TF siRNA cell lines.
Discussion
CREB is an important transcriptional factor that is located downstream of cell surface receptors and mitogens, and is very important for normal and tumor hematopoiesis. 29 Overexpression of CREB promotes proliferation and survival of cancer cells. 30 CREB activity is regulated by a group of co-activating factors located in the cytoplasm. This group of activators are called CRTCs (also known as TORCs), which serve as sensors in regulation of CREB activity. This family of proteins includes 3 members, CRTC1, CRTC2, and CRTC3, all of which have been shown to be strong activators of transcription. 31 Following stimulation by extracellular stimulatory signals (such as cAMP, calcium, and hormones), these coactivators bind to CREB. Once activated, CRTCs are transported into the nucleus, bind to CREB, and activate the transcriptional activity of CREB. 32 As a coactivator of CREB, CRTC2 plays an important role in hepatic glucose metabolism. 22,33 CREB is upregulated in many cancer cells, suggesting that it is closely associated with the occurrence of tumors. We found that CRTC2 mRNA and protein were highly expressed in both the CRC lines, SW620 and T84. To elucidate the role of CRTC2 in CRC progression, CRTC2 was downregulated in SW620 and T84 cell lines, and the proliferation, invasion, and migration of shCRTC2 cells were analyzed. We found that the proliferation, migration, and invasion abilities of SW620 and T84 cells were significantly decreased following the downregulation of CRTC2.
Some studies have shown that the phosphorylation of CRTC2 may play a crucial role in its activity. CRTC2 localizes to the nucleus, and interacts with CREB to initiate the transcription of downstream genes following dephosphorylation of the Ser171 residue. 34 CRTC2 Ser171 is phosphorylated by insulin through SIK2, which causes CRTC2 to translocate from the nucleus to the cytoplasm and undergo degradation through the proteasome pathway. 35 Ser171, Ser275, and Ser369 phosphorylation accelerates the interaction between CRTC2 and 14-3-3 proteins. Ser171 and Ser307 have been shown to regulate CRTC2 in hepatocytes. 14,16,24 As a critical factor in LKB1-deficient NSCLC, CRTC2 becomes constitutively activated in LKB1-mutant NSCLC because it remains unphosphorylated due to inactivation of SIKs, 21,36 thereby promoting tumor growth. 27 In addition to these known phosphorylation sites of CRTC2 (including Ser171), there are other phosphorylation sites in CRTC2, including Ser127, Ser238, and Ser245, whose specific functions in cancer cells remain unknown. 23 We analyzed the function of CRTC2 Ser127, Ser238, and Ser245 phosphorylation and found that Ser171 phosphorylation was inhibited by Ser238 phosphorylation. The localization of CRTC2 was altered and nuclear aggregation of CRTC2 was significantly reduced in shCRTC2 + A238 cells. The proliferation, invasion, and migration of the shCRTC2 + A238 cells were significantly inhibited, and the interaction of CRTC2 and CREB in shCRTC2 + A238 cells was significantly weakened. The downstream genes of CREB were downregulated at both the mRNA and protein levels in the shCRTC2 + A238 cells, thereby indicating that phosphorylation of CRTC2 Ser238 was important for CRTC2 to bind CREB and to activate CREB-mediated transcriptional activity. The CREB binding domain (CBD) peptide (amino acids 18–55) in CRTC2 undergoes a coil-helix transition and is important for its interaction with CREB. 17,37,38 As shown in Figure 6, we speculate that the coil-helix transition of CRTC2 may be stabilized by Ser238 phosphorylation in the 3-D structure. SIK2 is an AMP-activated protein that can phosphorylate the Ser171, Ser275, and Ser307 sites of CRTC2, 14,39 while the Ser275 site of CRTC2 is also phosphorylated by MARK2 23 and calcineurin (Cn). 14 Subsequently, the kinases that phosphorylate the Ser238 site were predicted and knockdown studies revealed that downregulation of ERK2 reduced phosphorylation of Ser238.
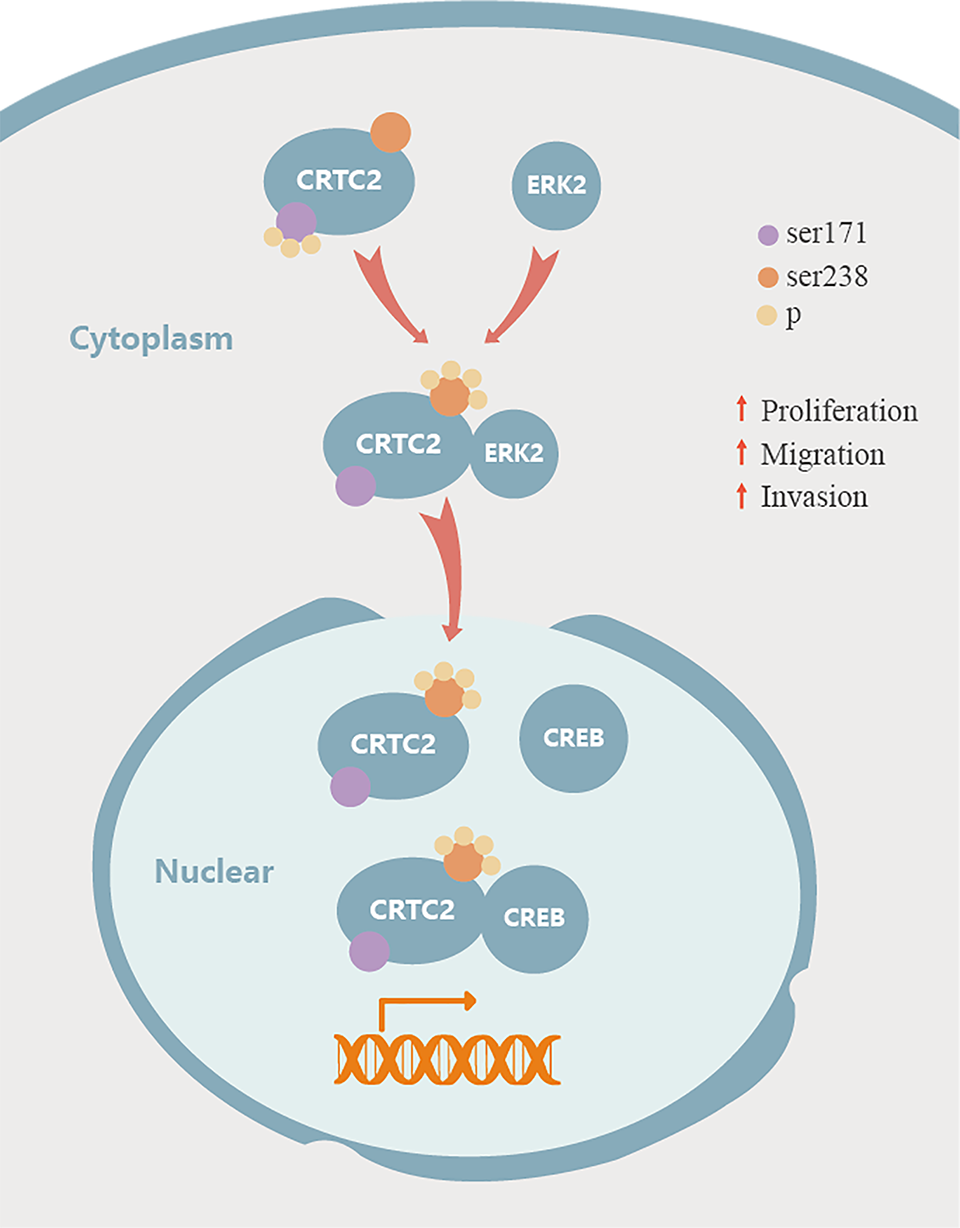
Schematic diagram of CRTC2 in CRC. CRTC2 is phosphorylated at Ser238 residue by ERK2 and inhibits the phosphorylation of Ser171, to promote CRTC2 and CREB interaction.
Conclusion
The results of this study demonstrate that CRTC2 is overexpressed in CRC cells, and downregulation of CRTC2 expression inhibits the proliferation, migration, and invasion of CRC cells. The abilities of proliferation, migration, and invasion are correlated with phosphorylation of CRTC2 at Ser238. Further, we found that ERK2 might be a kinase that phosphorylated CRTC2 on Ser238. These findings contribute to a better understanding of CRC development and may help identify potential new targets for CRC therapy.
Footnotes
Abbreviations
Authors’ Note
Our study did not require an ethical board approval because it did not contain human or animal trials.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: National key R & D Program “Research on Prevention and Control of Major chronic Infectious Diseases” key special project (2019YFC1315705), China Cancer Foundation Beijing Hope Marathon Special Fund (LC2017L07), Medical and Health Science, Technology Innovation Project of the Chinese Academy of Medical Sciences (2017-12M-1-006) Beijing Nova Program of Science and Technology under grant (Z191100001119055).
