Abstract
BACKGROUND:
Lung adenocarcinoma (LUAD) has a high incidence and poor prognosis, and multiple circRNAs (circRNAs) have been found to regulate LUAD.
OBJECTIVE:
This study focuses on the effect and mechanism of hsa_circ_0070661 in LUAD.
METHODS:
LUAD tissues and para-cancerous tissues were collected from 38 patients diagnosed with LUAD in our hospital. Hsa_circ_0070661, miR-556-5p and TEK Receptor Tyrosine Kinase (TEK) levels were evaluated using western blotting and RT-qPCR, and the targeting relationship was detected by luciferase reporter and RIP assays. Cell migration, viability, apoptosis-related proteins, (Bcl-2 and Bax) and tumor growth in vivo were assessed by Transwell, CCK-8, western blotting and xenograft assays, respectively.
RESULTS:
Results indicated downregulation of hsa_circ_0070661 and TEK in LUAD cell lines and tissues, whereas upregulation of miR-556-5p. Hsa_circ_0070661 upregulation restrained the viability, migration and tumor growth of LUAD cells, and promoted apoptosis. Hsa_circ_0070661 could directly target miR-556-5p to upregulate TEK expression in LUAD. MiR-556-5p upregulation promoted the malignant phenotypes of LUAD cells and reversed the anti-cancer effect of hsa_circ_0070661 overexpression, while TEK upregulation inhibited LUAD progression and somewhat eradicated the cancer-promoting effect of miR-556-5p upregulation.
CONCLUSIONS:
Hsa_circ_0070661 sponges miR-556-5p to inhibit LUAD development via regulating TEK, providing a promising molecular target for LUAD clinical therapy.
Introduction
Lung cancer is the highly widespread cancer, accounting for more than 18% of global cancer-related deaths [1, 2]. Lung adenocarcinoma (LUAD) is the most common histological type of non-small cell lung cancer (NSCLC), a major category of lung cancer that accounts for 85% of all lung cancers, accounting for approximately 40% of all lung cancers [3, 4]. A complete diagnostic system consisting of mediastinoscopy, sputum cytology, radionuclides, bronchoscopy, and X-rays can identify patients with LUAD to a large extent [5]. However, approximately 70% of patients are already in stage III-IV at the time of diagnosis of LUAD, resulting in a 5-year survival rate of less than 20% [6, 7]. In addition, traditional chemoradiotherapy has a tendency to fail in the treatment of LUAD due to the development of drug resistance [2]. To better understand the pathophysiology of LUAD and enhance its treatment efficacy, it may be useful to identify efficient molecular targets.
A unique endogenous RNA called circular RNAs (circRNAs) is created by a non-classical reverse splicing process [8]. Its lengthy half-life and resistance to ribonuclease digestion are caused by the ability of its downstream splice donor sites to covalently bind to the upstream splice acceptor sites [8, 9]. CircRNA has been shown to be a rich and conserved RNA and is generally expressed in a cell type, complex tissue or stage-specific manner [10, 11]. Current research has supported circRNA’s crucial biological roles, particularly as a predictive biomarker for human cancer [12, 13, 14]. For example, hsa_circ_0000792 is highly expressed in LUAD and has significant diagnostic accuracy for LUAD [15]. High circXPO1 expression was related with poor survival of LUAD [16]. Hsa_circ_0070661 is located on chr4 :110580166-110608872, and its related gene symbol is CCDC109B. There is no research to prove its effect on human diseases. Therefore, the hsa_circ_0070661 function and molecular mechanism in LUAD need to be preliminary investigated.
MicroRNAs (miRNAs) are short non-coding RNAs of about 18–25 nucleotide lengths that play a wide and important role in biological process regulation together with apoptosis, migration, proliferation, and differentiation [17]. MiRNAs have been found to be sponges of circRNAs and interact with seed sequences on the 3’-untranslated region (3’ UTR) of the target mRNAs [18]. Notably, a growing number of molecular mechanisms based on circRNA-miRNA-mRNA have been shown to play an significant regulatory role in LUAD. For instance, hsa_circ_0008274 overexpression inhibits LUAD cells invasion and proliferation by down-regulating miR-578 and activating HMGA2 [19]. CircFBXW7 repress the migration and proliferation of LUAD cell lines by controlling miR-942-5p/BARX2 axis [20]. MiR-556-5p has been widely reported to participate in the development and incidence of human malignant tumors, inhibits breast cancer (BC) [21], colorectal cancer [22] and meningioma [23], while promotes prostate cancer [24]. Importantly, Feng et al. [25] suggested that miR-556-5p was heavily expressed in NSCLC tissues, and its knockdown reduced cancer tumorigenesis and cell viability, and persuaded apoptosis. In this paper, the binding of hsa_circ_0070661 to miR-556-5p was found through the target prediction website. Therefore, we examined the association among miR-556-5p and hsa_circ_0070661 in LUAD.
TEK receptor Tyrosine kinase (TEK) also known as TIE2 is an endothelial tyrosine kinase receptor of vascular growth factor responsible for angiogenesis [26]. According to previous studies, TEK could regulate endothelial cell proliferation, migration, survival and inflammation by binding to Angiopoietin [27, 28]. Due to the malignant proliferation and migration characteristics of tumor cells, the role of TEK in cancer attracts many researchers’ attention. For instance, Yang et al. identified that TEK knockdown interacted with SIRT7 could reduce adriamycin-dependent breast cancer cell malignancy [29]. TEK expression was found to be downregulated in papillary thyroid carcinoma, and its knockdown also could decrease the proliferation and migration of papillary thyroid carcinoma cells [30]. However, TEK was identified as an anticancer in LUAD, which could be regulated by miR-486-5p [31] or miR-19a-3p [32]. The previous study on TEK in LUAD suggested that TEK could be regulated by different miRNAs to play the anticancer role in LUAD. This study found that TEK was a target gene of miR-556-5p. So far, the relationship between TEK and miR-556-5p in LUAD is still not be explored.
Here, we inspected the expression of hsa_circ_ 0070661 in LUAD as well as its effect LUAD cells biological process of in vitro as well as in vivo. Additionally, we focused on the specific regulatory mechanism of hsa_circ_0070661 acting on miR-556-5p/TEK Receptor Tyrosine Kinase (TEK) axis. The findings of this study should offer useful tools for LUAD diagnosis and therapy.
Materials and methods
Sample Tissues
The characteristics of all patients with LUAD
The characteristics of all patients with LUAD
This study included 38 cases of LUAD patients treated in our hospital between January 2021 and January 2022 LUAD tissue and corresponding para-cancerous tissue were collected during operation and stored in liquid nitrogen. The inclusion criteria were as follows: 1) All patients were diagnosed with LUAD by three pathologists; 2) All patients did not diagnose with other diseases; 3) All patients or their families signed informed consent forms. The exclusion criteria were as follows: 1) The patients had LUAD history; 2) The patients underwent radiation, chemotherapy or other treatments; 3) The characteristics of some patients were lost. Table 1 listed the characteristics of all patients. All participants and their families signed informed consent forms after being informed of the study. Additionally, this study adhered to the Declaration of Helsinki and was carried out with endorsement from our hospital’s Ethics Committee.
H1650 and A549 two cell lines for LUAD and BEAS-2B (the human bronchial epithelial cell line) acquired from ATCC (USA), and one LUAD cell line (PC9) acquired from Sigma-Aldrich (USA) were incubated in DMEM (Thermo Fisher Scientific, USA) containing 10% FBS (HyClone, USA) and 1% penicillin/streptomycin (Gibco, USA). Cells were cultured at 37
Cell transfection
H1650 and A549 cells were selected to establish stable transfection cell lines. Overexpression vector loading hsa_circ_0070661 and TEK (circ-OE and TEK-OE; 2
RT-qPCR assay
Real-time PCR primer synthesis list
Real-time PCR primer synthesis list
Total RNA isolated by RNAiso Plus (TAKARA, Japan) and miRNA extracted by miRNEasy EFPE Kit (BioTeke, China) were treated with ultraviolet spectrophotometer and agaLUADose gel electrophoresis to determine their concentration, purity and integrity. The transcription was then performed by means of the TaqMan reverse transcription kit (ABI, USA) and the TaqMan miRNA reverse transcription Kit (ABI). RT-qPCR was shown by means of Hieff qPCR SYBR Green Master Mix kit (Yeasen, China) on ABI 7500 Rapid RT-PCR system (ABI). The RT-qPCR conditions were as follows: 95
Hsa_circ_0070661 levels in subcellular site of H1650 and A549 cells was located via a PARI kit (Norgenbiotek, Canada) and quantified by RT-qPCR. Briefly, subcellular localization was performed in 1
Total RNA in H1650 and A549 cells acquired as described in RT-qPCR assay was utilized for RNase R tolerance test. In brief, qualified total RNA was divided into RNase R group and control group, treated with or without RNase R (3 U/
CCK-8 assay
H1650 and A549 cells after transfection for 48 h were transferred to a 96-well plate (4
Cell migration assay
H1650 and A549 cells after transfection for 48 h were suspended in serum-free medium (5
Western blotting
Total cell protein was lysed with RIPA buffer (Thermo Scientific, USA), and protein concentration were quantified by BCA kit (Thermo Scientific). The proteins were then separated in equal quantities using 10% SDS-PAGE and transferred to a PVDF membrane (Millipore, USA), which was sealed in 5% skim milk at 37
Xenograft tumor model
Ten BALB/c nude mice, aged 5 weeks and weighing 18–22 g, were purchased from Hunan SJA Laboratory Animal (China). A549 cells in circ-OE or empty vector groups were prepared into cell suspension, and each nude mouse was subcutaneously injected with 200
Luciferase reporter assay
CircInteractome (
RIP assay
This experiment was conducted using a Magna RIP (RNA immunoprecipitation) RNA-binding protein immunoprecipitation kit (Millipore) under strict kit instructions. Magnetic beads were incubated with anti-Ago2 or IgG antibody (Millipore) in a rotator at 25
Statistical analysis
SPSS (version 20.0, IBM, USA) was utilized for the statistical analysis of the data. Continuous data from all experiments performed in triplicates were in the format of mean
Results
Hsa_circ_0070661 was lowly expressed in LUAD
Hsa_circ_0070661 was lowly expressed in LUAD. (A) Relative levels of hsa_circ_0070661 in LUAD tumors and adjacent normal tissue were assessed via RT-qPCR. (B) Relative levels of hsa_circ_0070661 in different tumor stages were assessed via RT-qPCR. (C) Relative hsa_circ_0070661 expression in a human bronchial epithelial cell line (BEAS-2B) and LUAD cell lines (PC9, H1650 and A549). 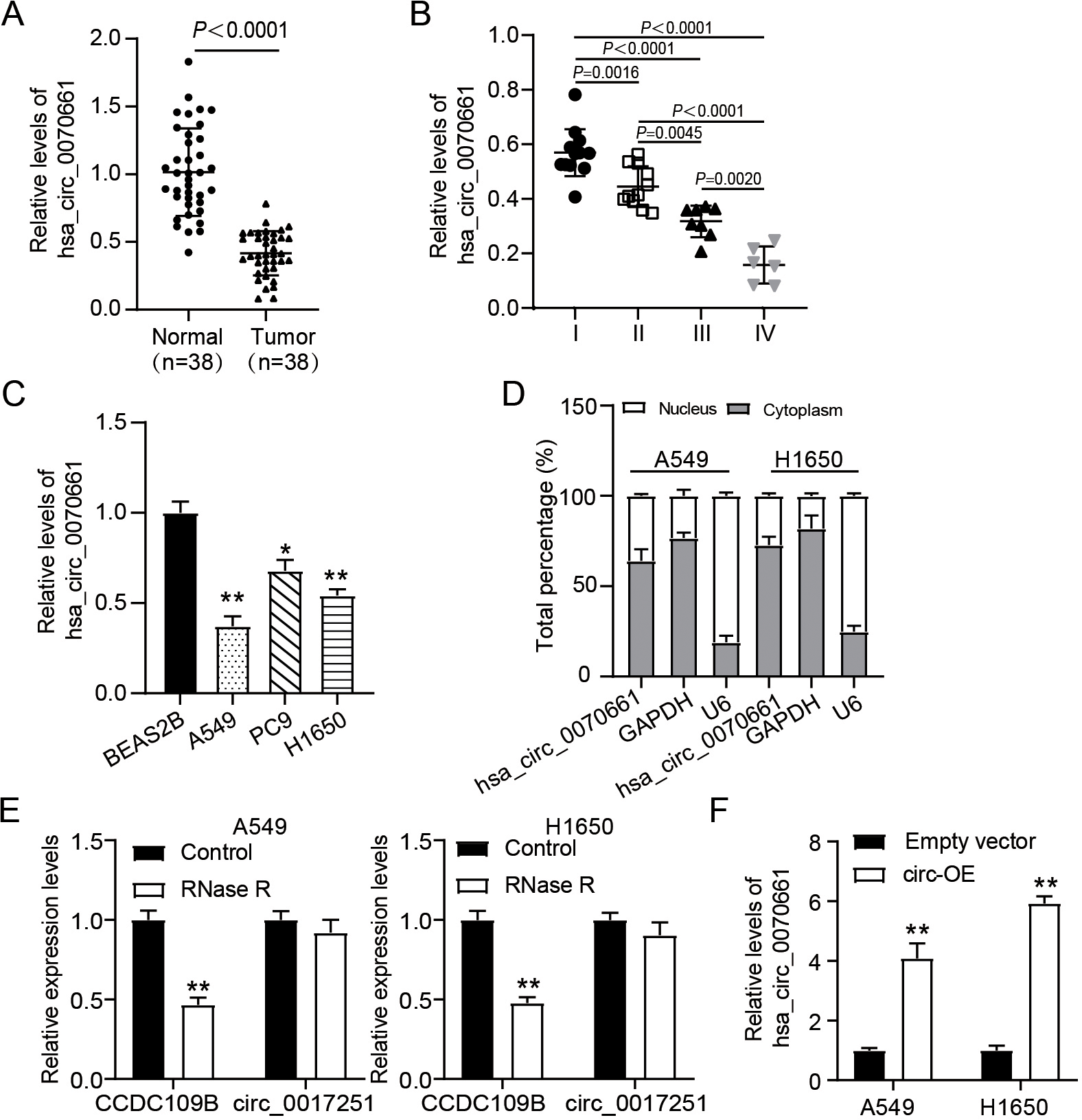
To evaluate the abnormal hsa_circ_0070661 expression in LUAD, RT-qPCR was performed. The results showed that the expression level of hsa_circ_0070661 in LUAD tissues was 60% lower than that in adjacent tissues (Fig. 1A), and the expression level of hsa_circ_0070661 was reduced with the increase of tumor stage (Fig. 1B). Similarly, compared with BEAS-2B cells, hsa_circ_0070661 was uniformly down-regulated in all three kinds of LUAD cell lines (H1650, PC9 and A549), and H1650 and A549 showed lower hsa_circ_0070661 expression (Fig. 1C). In addition, to determine the subcellular localization of hsa_circ_0070661, we also examined its relative abundance in the nucleus and cytoplasm. It was found that the proportion of hsa_circ_0070661 in the cytoplasm of H1650 and A549 cells was greater in the nucleus (Fig. 1D). This suggests that hsa_circ_0070661 is mainly located in the cytoplasm. In addition, an RNase R tolerance test was performed, and observed that hsa_circ_0070661 remained after RNase R treatment (Fig. 1E).
Hsa_circ_0070661 up-regulation suppressed the migration, proliferation, and tumor growth of LUAD cells while facilitated apoptosis. (A) Effect of Empty vector and circ-OE transfection on the viability of H1650 and A549 cells, as determined by CCK-8 assay. 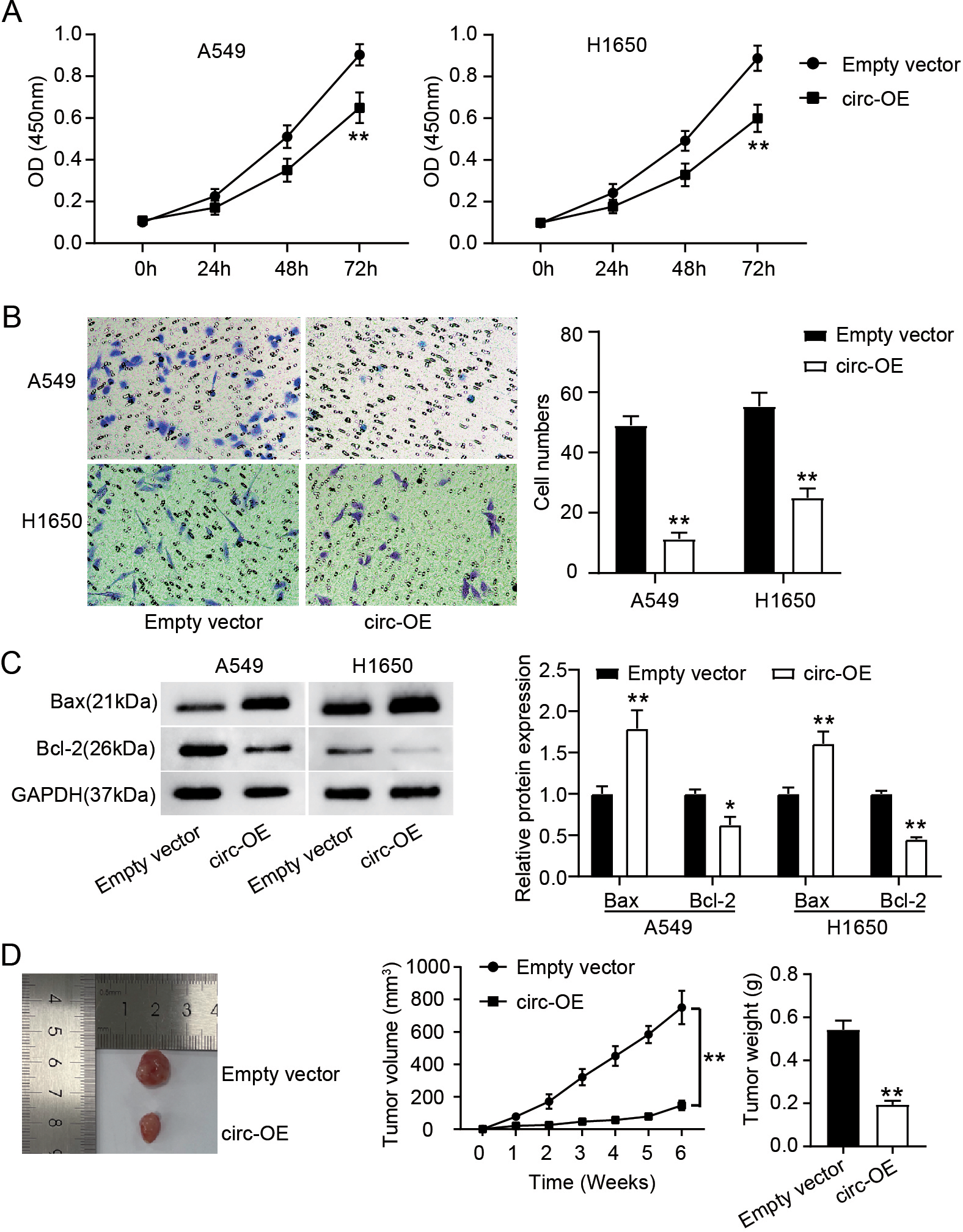
In addition, overexpression vector of hsa_circ_0070661 was transfected into LUAD cells. As shown in Fig. 1F, the expression level of hsa_circ_0070661 in LUAD cells in the circ-OE group was up-regulated by more than 4-fold compared with the empty vector group. These results suggest that hsa_circ_0070661 is down-regulated in LUAD and may play a role in the cytoplasm through the circular structure.
Hsa_circ_0070661 sponged miR-556-5p. (A) Speculated binding site of miR-556-5p within hsa_circ_0070661 was predicted by circInteractome database (https://circinteractome.nia.nih.gov/index.html). (B) Luciferase activity was estimated in H1650 and A549 cells carrying miR-NC/miR-556-5p mimic and hsa_circ_0070661 WT/MUT. 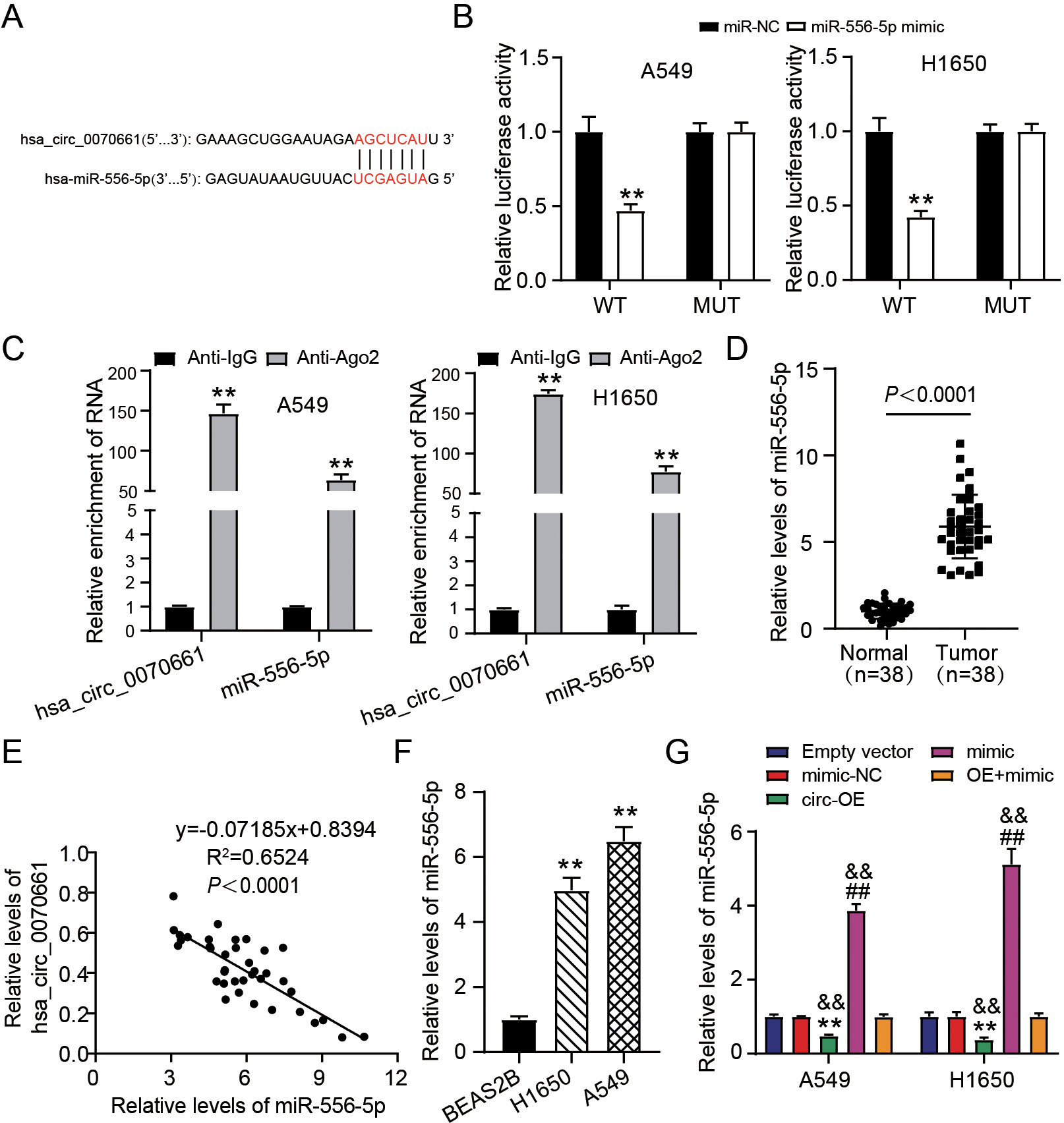
Next, we investigate the role of hsa_circ_0070661 in LUAD cells, the changes of LUAD cells viability and migration were analyzed by CCK-8 and Transwell assay. We observed that the viability of H1650 and A549 cells in the circ-OE group was about 35% and 45% of that in the empty vector group, respectively (Fig. 2A). Furthermore, hsa_circ_0070661 overexpression reduced the migration level of H1650 and A549 cells by 80% and 55%, respectively (Fig. 2B). Moreover, western blotting showed an obvious up-regulation of Bax and down-regulation of Bcl-2 protein levels in the circ-OE group than that in the empty vector group (Fig. 2C). Additionally, we investigated the impact of hsa_circ_0070661 on the progress of LUAD cells in vivo through xenograft tumor in nude mice. Six weeks after the model was established, the tumor size in the circ-OE group was significantly smaller than in the empty vector group, and tumor growth curve and tumor weighing showed that the weight and volume of tumor in the circ-OE group were lessen as compare to the empty vector group (Fig. 2D). These results suggest that up-regulation of hsa_circ_0070661 could destroy the malignant phenotype of LUAD cells in vitro and in vivo.
Hsa_circ_0070661 blocked the malignant behavior of LUAD cells through targeting miR-556-5p. H1650 and A549 cells were transfected with Empty vector, circ-OE, mimic-NC, mimic and OE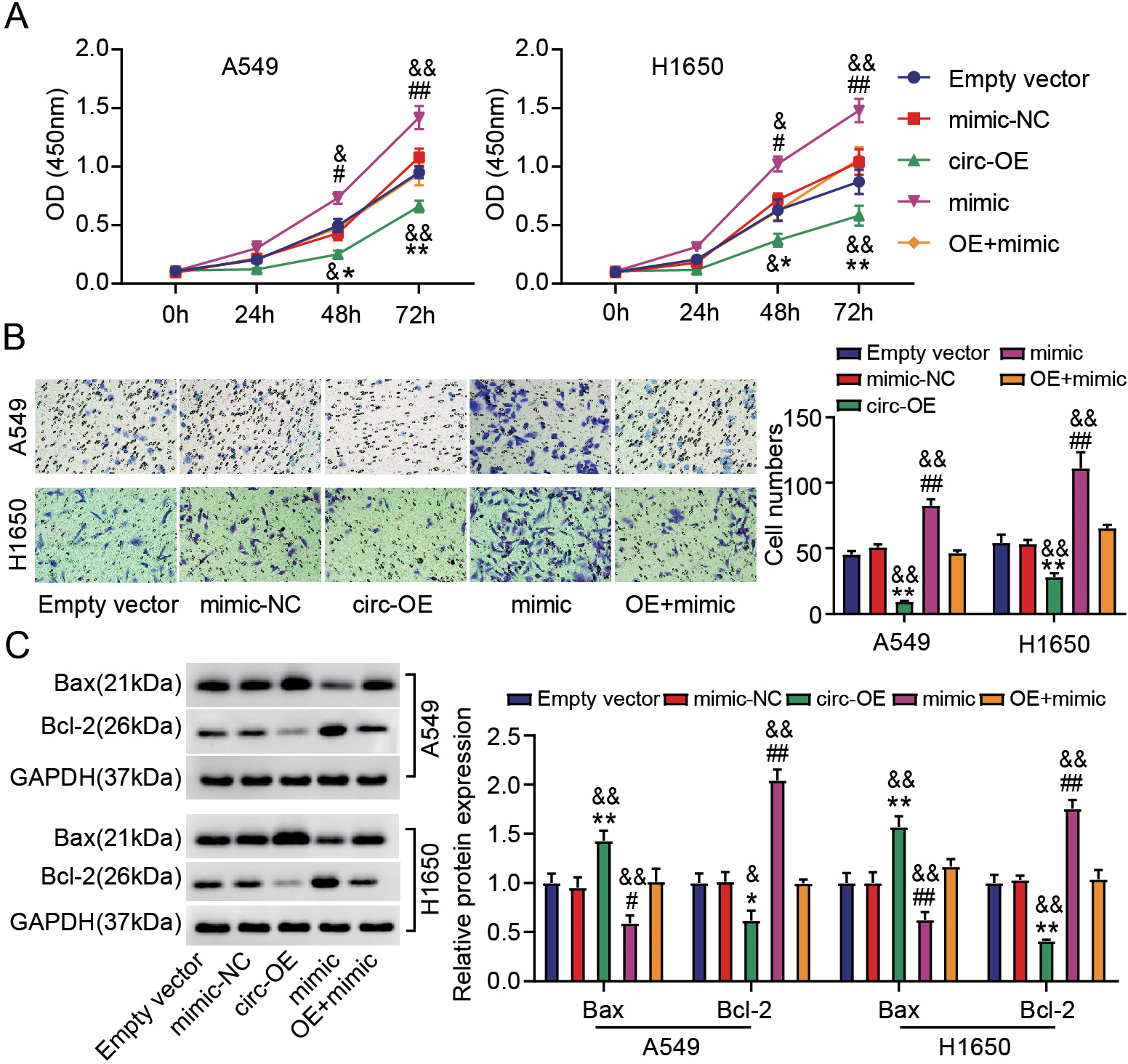
MiR-556-5p targeted TEK. (A) Binding site of miR-556-5p within TEK, predicted using TargetScan database (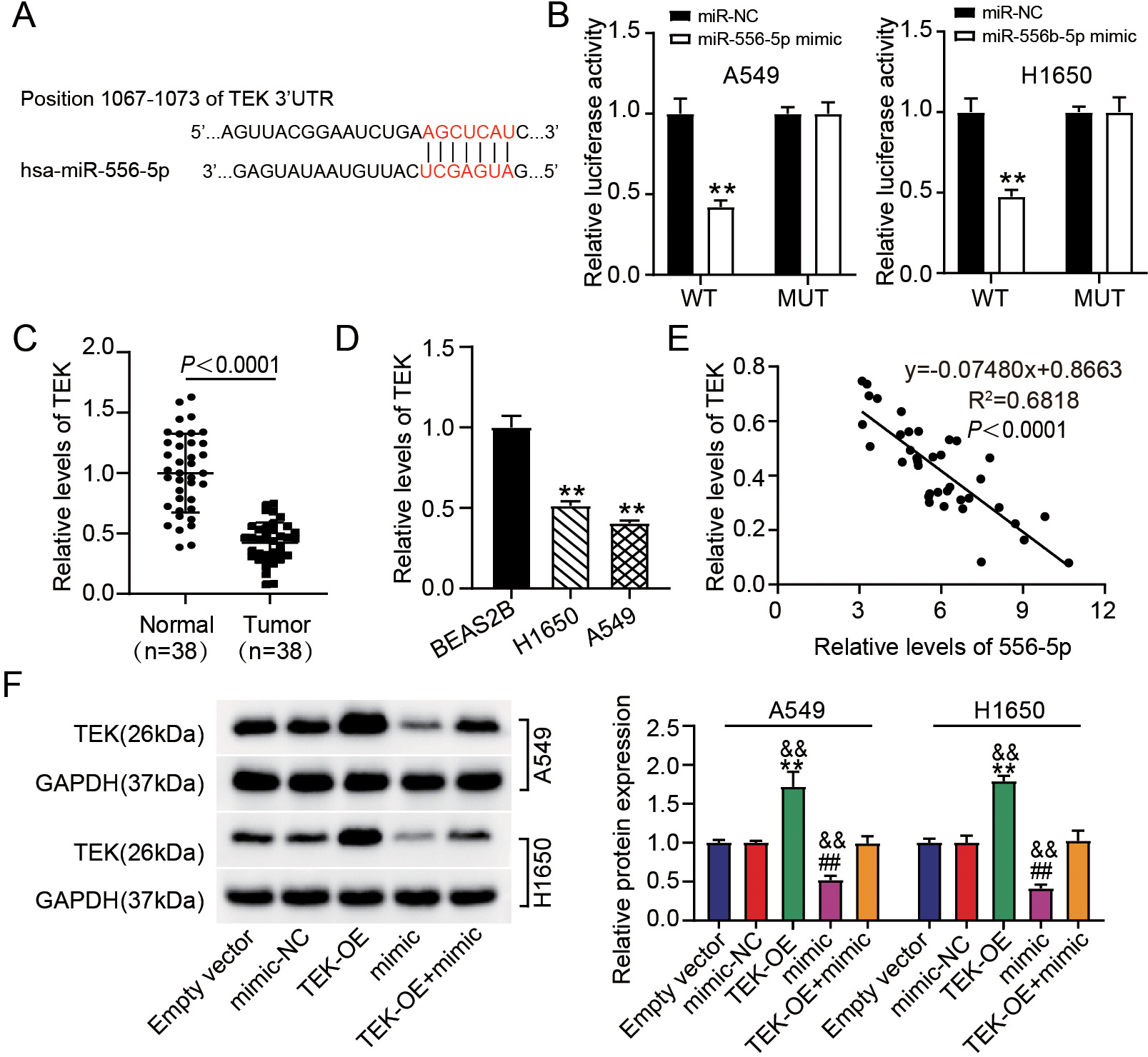
Based on the LUAD inhibitory properties of hsa_ circ_0070661 revealed above, we continued to explore the downstream miRNA of hsa_circ_0070661. As shown in Fig. 3A, the binding sites for hsa_circ _0070661 and miR-556-5p were predicted by circInteractome. To verify their binding, we executed a luciferase reporter assay. The luciferase activity of wild type hsa_circ_0070661 (WT) vectors could be inhibited by miR-556-5p mimic, but the luciferase activity of MUT vectors showed no change (Fig. 3B). Next, we used RIP to verify whether hsa_circ_0070661 and miR-556-5p could bind Ago2 protein. According to the data, Ago2 antibodies enriched hsa_circ_0070661 and miR-556-5p levels (Fig. 3C). Based on these data, we hypothesize that hsa_circ_0070661 directly bound to miR-556-5p. Then, we performed RT-qPCR and Pearson correlation analysis in clinical sample of LUAD patients. The outcomes presented that miR-556-5p expression in LUAD was up-regulated by 6.5-fold and negatively associated with the expression of hsa_circ_0070661 (Fig. 3D and E). In addition, miR-556-5p levels in H1650 and A549 increased 5.0 and 7.0 times, respectively, compared to BEAS-2B (Fig. 3F). Subsequently, we examined the regulation impact of miR-556-5p by hsa_circ_0070661 at the cellular level. The level of miR-556-5p in hsa_circ_0070661 over-expressed LUAD cells was 20% of the empty vector group, and the up-regulation of hsa_circ_0070661 reversed miR-556-5p mimic induced up-regulation of miR-556-5p (Fig. 3G). These outcomes specified that miR-556-5p was over-expressed in LUAD and its up-regulation was caused by hsa_circ_0070661.
Next, biological function experiments was performed to verify the role of miR-556-5p in LUAD and its interaction with hsa_circ_0070661 in vitro. Further analysis by Transwell assay and CCK-8 assay exhibited that miR-556-5p up-regulation increased viability and migration of LUAD cells by about 1.5 and 1.6-fold, respectively, while hsa_circ_0070661 overexpression attenuated this effect (Fig. 4A and B). In addition, western blotting displayed that miR-556-5p overexpression of LUAD cells decreased Bax protein and increased Bcl-2 protein, but this effect could also be reversed by transfection of circ-OE (Fig. 4C). In conclusion, miR-556-5p accelerates the malignant behavior of LUAD cells and mediates the anticancer impact of hsa_circ_0070661.
MiR-556-5p targeted TEK
High TEK expression inhibited the malignant behaviors of LUAD cells and reversed the tumorigenic effects of miR-556-5p upregulation. H1650 and A549 cells were transfected with Empty vector, TEK-OE, mimic-NC, mimic and TEK-OE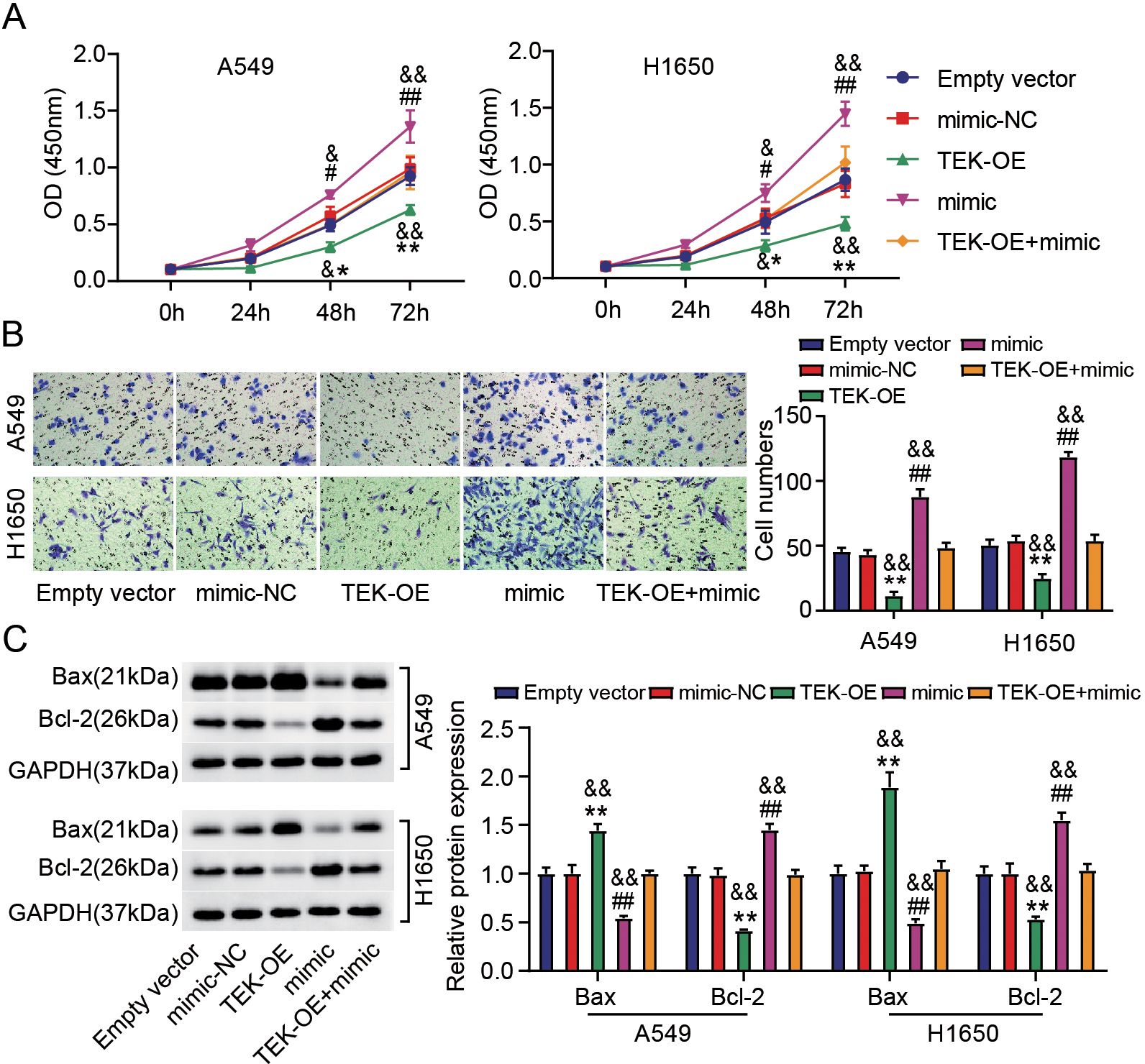
We additionally refined the downstream target of miR-556-5p. Based on the TargetScan database prediction, we found that TEK sequences have miR-556-5p binding site (Fig. 5A). Luciferase analysis showed that the luciferase activity decreased by more than 55% in LUAD cells containing TEK-WT vector and miR-556-5p mimic, while in LUAD cells containing TEK-MUT vector and miR-556-5p mimic, there was no significant change in luciferase activity (Fig. 5B). Moreover, the mean level of TEK mRNA in LUAD tumors was almost 50% lower in normal tissues (Fig. 5C). In H1650 and A549 cells, TEK mRNA levels were 50% and 40% of those in BEAS-2B cells, respectively (Fig. 5D). Moreover, the expression of TEK in LUAD tumors was negatively associated with miR-556-5p levels (Fig. 5E). To confirm the association among miR-556-5p and TEK in LUAD, cells were transfected with either TEK-OE or miR-556-5p mimic. According to the quantitative result of TEK protein in cells, the level of TEK protein increased more than 1.5 times when TEK was overexpressed, however, these changes might be reversed by mean of introducing miR-556-5p mimic (Fig. 5F). In conclusion, miR-556-5p could target and negatively regulate TEK.
We continued to study the impacts of miR-556-5p and TEK on LUAD cell behavior in vitro. The results of CCK-8 assay and Transwell assays uncovered that transfection of TEK-OE slowed LUAD cell proliferation and migration, but miR-556-5p upregulation reversed this effect (Fig. 6A and B). In addition, high levels of TEK could increase Bax protein levels and decrease Bcl-2 protein levels in LUAD cells, while this could also be reversed by miR-556-5p mimic (Fig. 6C). These results suggest that miR-556-5p modulates TEK to achieve its promoting-cancer function in LUAD cells.
Discussion
This paper investigated the impact of hsa_circ _0070661 on the generation and development of LUAD. Our analysis presented that hsa_circ_0070661 was under-expressed in LUAD, and its overexpression could inhibit the migration and survival of LUAD cells, encourage cell apoptosis, and obstruct the formation of tumors in vivo. Moreover, we exhibited that hsa_circ_0070661 sponged miR-556-5p, and miR-556-5p targeted TEK.
Accumulated literatures have indicated that circRNA possesses the ability to regulate the malignant actions of LUAD cells. For example, high circXPO1 expression helps the survival, invasion and migration of LUAD cells in vivo as well as in vitro, and inhibits apoptosis [16]. Silencing circ-ENO1 inhibits the progression of LUAD, which is specifically manifested as delaying glycolysis of LUAD cells, inhibiting proliferation, metastasis and epithelial-mesenchymal transition (EMT), and inducing apoptosis [36]. Hsa_circ_0005576 promotes LUAD cell proliferation and osimertinib resistance [37]. There have been no reports on hsa_circ_0070661. Here, we first time study the mechanism and role of hsa_circ_0070661 in LUAD. Similar to various circRNA involved in LUAD development, we exhibited that hsa_circ_0070661 expression was down-regulated in LUAD. Over-expression of hsa_circ_0070661could suppress the LUAD cell growth and migration and induce cell apoptosis. These results suggest that hsa_circ_0070661 plays a role of tumor suppressor in LUAD.
Since circRNA has the effect of sponge miRNA, we continue to explore the downstream of miRNA of hsa_circ_0070661. In this study, the target miRNA of hsa_circ_0070661, miR-556-5p, was screened out by bioinformatics tools. MiR-556-5p plays different roles in different tumors. Overexpression of miR-556-5p promotes the growth of cancer cells in prostate cancer [24]. In breast cancer, miR-556-5p knockdown promotes cancer cell invasion and migration and EMT [21]. Our suspection was that this is due to the heterogeneity of the tumor. Furthermore, previous one report found that miR-556-5p plays a carcinogenic role in NSCLC, and its downregulation can effectively improve the sensitivity of NSCLC to cisplatin [25]. Consistent with the results reported by Feng et al. [25], our study observed miR-556-5p overexpression in LUAD. Furthermore, up-regulation of miR-556-5p promoted LUAD migration and viability, as well as inhibited apoptosis. The mechanism by which miR-556-5p is regulated by circRNA has also been discovered. MiR-556-5p is absorbed by circ-LAMP1 sponge to inhibit the growth and metastasis of cholangiocarcinoma [38], and is suppressed by circSNHG1 sponge to inhibit meningioma tumorigenesis [23]. By targeting analysis, we ascertained that miR-556-5p was the downstream miRNA of hsa_circ_0070661, and was negatively regulated by hsa_circ_0070661. In addition, miR-556-5p overexpression partially eliminates the inhibitory effect of hsa_circ_0070661 upregulation on the malignant actions of LUAD. miR-556-5p was discovered to promote LUAD progression and mediates the antitumor effect of hsa_circ_0070661.
TEK receptor Tyrosine kinase (TEK) is an endothelial tyrosine kinase receptor of vascular growth factor responsible for angiogenesis and angiogenesis [26]. Studies have shown that TEK mutates in disease, resulting in abnormal endothelial cell growth, elongated, overlapping cells, and reduced extracellular fibronectin [39]. Reports indicate that TEK is participated in the development of cancer. For instance, TEK enhances invasion and metastasis of breast cancer cells as an oncogenic factor [29]. As a tumor suppressor in clear cell renal cell carcinoma, TEK downregulation predicts the poor prognosis of cancer patients [40]. In LUAD, TEK was identified as crucial for an anticancer, which significantly inhibiting the proliferation, migration as well as adhesion of cancer cells [41]. Consistent with the above analysis, this study found that TEK was down-regulated in LUAD and that its overexpression inhibited the malignant behavior of cancer cells. Furthermore, TEK was found to be the target of miR-556-5p, reversing the promoting-cancer effects of miR-556-5p on LUAD cell growth. In short, TEK suppressed LUAD cell progression via targeting miR-556-5p.
In this study, we have systematically explained the mechanism of hsa_circ_0070661/miR-556-5p/TEK axis in LUAD progression, however, there are still some limitations in this study. Firstly, this study showed that the hsa_circ_0070661 expression was different in A549, PC9 and H1650 cells. As we known, A549 is K-Ras mutant positive cell line, whereas PC9 and H1650 are BRAF mutant positive cell line. Therefore, whether the effects of hsa_circ_0070661 on different LUAD cells induced by different mutant are not same is worthy to explore. Due to the clinical samples collected from recently years, the prognosis is well so that the prognostic correlation of hsa_circ_0070661 needs to be analyzed by following up with these patients. In addition, the downstream signaling pathways that could be regulated by TEK remain to be further studied.
Conclusion
The effects of hsa_circ_0070661 on LUAD progression and its interaction with miR-556-5p/TEK were discussed for the first time. We found that hsa_circ_0070661 sponged miR-556-5p and released TEK to prevent the migration, tumor growth as well as proliferation of LUAD, and encourage cell apoptosis. To our knowledge, this research deepens the knowledge of LUAD pathogenesis, offers valued biomarkers for the LUAD diagnosis, and provides a favorable molecular target for LUAD clinical treatment.
Funding
Not applicable.
Ethics approval
The study was approved by the Wuhan No. 1 Hospital Ethics Committee (China). The processing of clinical tissue samples strictly conforms to the ethical standards of the Helsinki Declaration. A documented informed consent has been signed by each patient.
The Animal Care Committee of Wuhan No. 1 Hospital approved this animal experiment, which was conducted in agreement with the ARRIVE criteria.
Approval of participation
All patients have written informed consent.
Publication consent
Participants approved the publishing with their permission.
Availability of data and material
This article contains all of the data that were created or examined during this investigation.
Author contributions
Conception: YPC and YYW.
Interpretation or analysis of data: YPC and YYW.
Preparation of the manuscript: YYW.
Revision for important intellectual content: YPC.
Supervision: YPC.
Footnotes
Acknowledgments
None.
Conflict of interest
The author states that there are no conflicts of interest.
