Abstract
BACKGROUND:
Pancreatic ductal adenocarcinoma (PDA) is one of the major human health challenges with minimal therapeutic benefits due to its late detection, and de novo – and acquired chemotherapy resistance.
OBJECTIVE:
In this work we unravel the potential pro-survival role of RAB25 in pancreatic cancer chemotherapy resistance and aim to identify if RAB25 is a prognostic marker of patients’ survival in PDA.
METHODS:
We used RNA sequencing, shRNA mediated gene knockdown, BioGRID open repository of CRISPR screens (ORCS), GEPIA, kmplot.com, and cBioPortal.org databases to identify the role of RAB25 in PDA cell proliferation, chemotherapy response, expression in tumour versus normal tissues, and overall patients’ survival.
RESULTS:
RNA sequencing show Rab25 to be one of the top upregulated genes in gemcitabine resistance mouse PDA cells. Knockdown of Rab25 in these cells enhanced gemcitabine toxicity. In addition, re-analysis of previously published CRISPR/Cas9 data confirm RAB25 to be responsible for chemotherapy resistance in KRAS
CONCLUSION:
RAB25 expression is a prognostic marker for patient’s survival and gemcitabine resistance in PDA.
Introduction
Pancreatic cancer is a major human health challenge and the third leading cause of cancer-related deaths in the USA [1]. The death rate of pancreatic cancer varies among different populations and is reported to be
Despite being the standard of care for pancreatic cancer for decades gemcitabine treatment produces minimal benefits in PDA [6, 7, 8], largely due to de novo or rapid development of resistance [9, 10, 11]. Several factors contribute to gemcitabine resistance including tumor microenvironment, deregulation of nucleoside transporters/enzymes, epithelial to mesenchymal transition, and gene expression regulation by the miRNAs [11]. Despite the persistent resistance of PDA against gemcitabine, it is still widely used for its treatment largely due to lack of alternate effective treatment options. Thus, there is an urgent need to identify novel drug targets or biomarkers that could be effective against pancreatic cancer or predict the chemotherapy response accurately.
RAB25 (Ras-related protein Rab-25), is a small GTPase from the Rab protein family involved in vesicle transport, cell proliferation, signal transduction, cell motility and protein transport [12, 13]. In addition, RAB25 mediates tumorigenesis, cellular migration, metastasis, and chemoresistance [14, 15]. Depending on the context RAB25 may be an oncogene [14, 15, 16] or a tumour suppressor [17, 18, 19]. For example, in pancreatic cancer, RAB25 acts as an oncogene or tumour suppressor depending on the presence and absence of chloride intra cellular channel protein 3 (CLIC3) respectively [20]. Collectively, CLIC3 and RAB25 are responsible for recycling of integrin
In this study we generated gemcitabine resistant mPDA cells and performed whole transcriptomics analysis with a goal to identify novel proteins and pathways that could dictate gemcitabine resistant (GemR) in PDA cells. We used multiple approaches to narrow down on 10 most upregulated genes in GemR cells. Strikingly, we find that RAB25 protein which was previously not reported to be involved in gemcitabine resistance, is not only required for gemcitabine resistance, but it is also a prognostic marker for poor patients survival in PDA. Our results highlight the importance of targeting RAB25 to overcome gemcitabine resistance and use of RAB25 expression as a biomarker for stratifying gemcitabine responders from non-responders in PDA.
Methods and materials
Mouse husbandry
Mice carrying conditional alleles for Kras
Generation of mouse KPC cell lines
Small to medium sized palpable tumours from KPC (KRas
Generation of stable mPDA cell lines
2 to 3 shRNA targeting each shortlisted candidate genes were ordered from Sigma-Aldrich on an pLKO.1 background including a control non-targeting scrambled shRNA plasmid (shpLKO.1). The lentivirus preparation and stable cell line generation was done as before [26]. For Rab25, two independent shRNA, shRab25#1 and shRab25#2 were used to generate KPC-GemR stable cell lines.
Cell viability assays
Cell proliferation data in Fig. 3A was performed via quantification of cell density by taking live cell images at regular intervals of 3 hour up to 76 hours, by Incucyte FLR device equipped with 20x objective (Essen Biosciences). Cell viability was determined by either counting the viable cells in a 500
RNA sequencing analysis
RNA sequencing was performed as before [24]. Total RNA was isolated using Qiagen’s RNAesy kit and following the kit’s protocol. RNA was converted into strand-specific cDNA libraries using the KAPA mRNA HyperPrep kit (Roche) according to the manufacturer’s instructions. Sequencing was performed on Illumina HiSeq 4000 platform with single ended 75 bp reads with and 25–30 million reads per sample. Trimmomatic/0.36-Java-1.7.0_80 was used for adaptor trimming, with parameters “LEADING:3 TRAILING:3 SLIDINGWINDOW:4:20 MIN-LEN:36”. The RSEM package (v.1.2.31)54 in conjunction with the STAR alignment algorithm (v.2.5.2a) 55 was used for the read mapping and gene-level quantification with respect to mouse Ensembl GRCm38 – release 89. The parameters used were: – star-output-genome-bam – forward-prob 0. Differential expression analysis was performed with the DESeq2 package (v.1.20.0) 56 within the R programming environment (https://www.r-project.org/; v.3.5.1), where the significance threshold for the identification of differentially expressed transcripts was set to an adjusted
Pathway analysis of significantly deregulated genes in mPDA GemR cells
To cluster differentially regulated genes based on their common functionality, a set of genes was analyzed for pathway analysis using Ingenuity Pathway Analysis (IPA). The differentially expressed genes (DEG) that are adjusted for false discovery rate (FDR) with a corrected
Meta analysis of RAB25 expression, genetic alterations, and patients’ survival datasets
The online database Gene Expression Profiling Interactive Analysis (GEPIA) (http://gepia.cancer-pku.cn/index.html) was used to visualize the expression of RAB25 in various tumors versus their matched normal tissues. GEPIA is an interactive web tool that analyses RNA sequencing expression of 9,736 tumors and 8,587 normal samples from TCGA and the GTEx projects [28]. Analysis was conducted using the ANOVA statistical method with a log2FC cutoff of 2 and a
The cBioPortal for Cancer genomics (http://www.cbioportal.org/) was used to analyse the genetic alterations associated with in different cancer types in the TCGA pipeline. The keyword “RAB25” was searched within the cBioPortal database to obtain a cross-cancer summary and identify the genetic alterations (amplification, missense mutations, deep deletion and CNV) of RAB25 associated with pancreatic cancer.
Patient prognoses were evaluated by Kaplan-Meier survival curves of Pancreatic ductal adenocarcinoma patients with low or high expression of RAB25 using the Kaplan-Meier plotter (http://kmplot.com) and the GEPIA platforms. The database includes RNA-seq information based on TCGA and GEO datasets
Re-analysis of CRISPR/Cas9 screen data
The CRISPR/Cas9 screen data were collected from BioGRID Open Repository of CRISPR Screens (ORCS) (https://orcs.thebiogrid.org/), which is a publicly available scientific online resource. The keyword “RAB25” was searched within the database and 22 hits were identified out of 1099 screens. We selected screens with the phenotype “response to chemicals” and a
Generation of gemcitabine resistant mouse PDA cell lines. A, Scheme of functional genomics screen to identify mediators of gemcitabine resistance. B, mPDA cell viability as judged by counting of viable cells after 72 hours 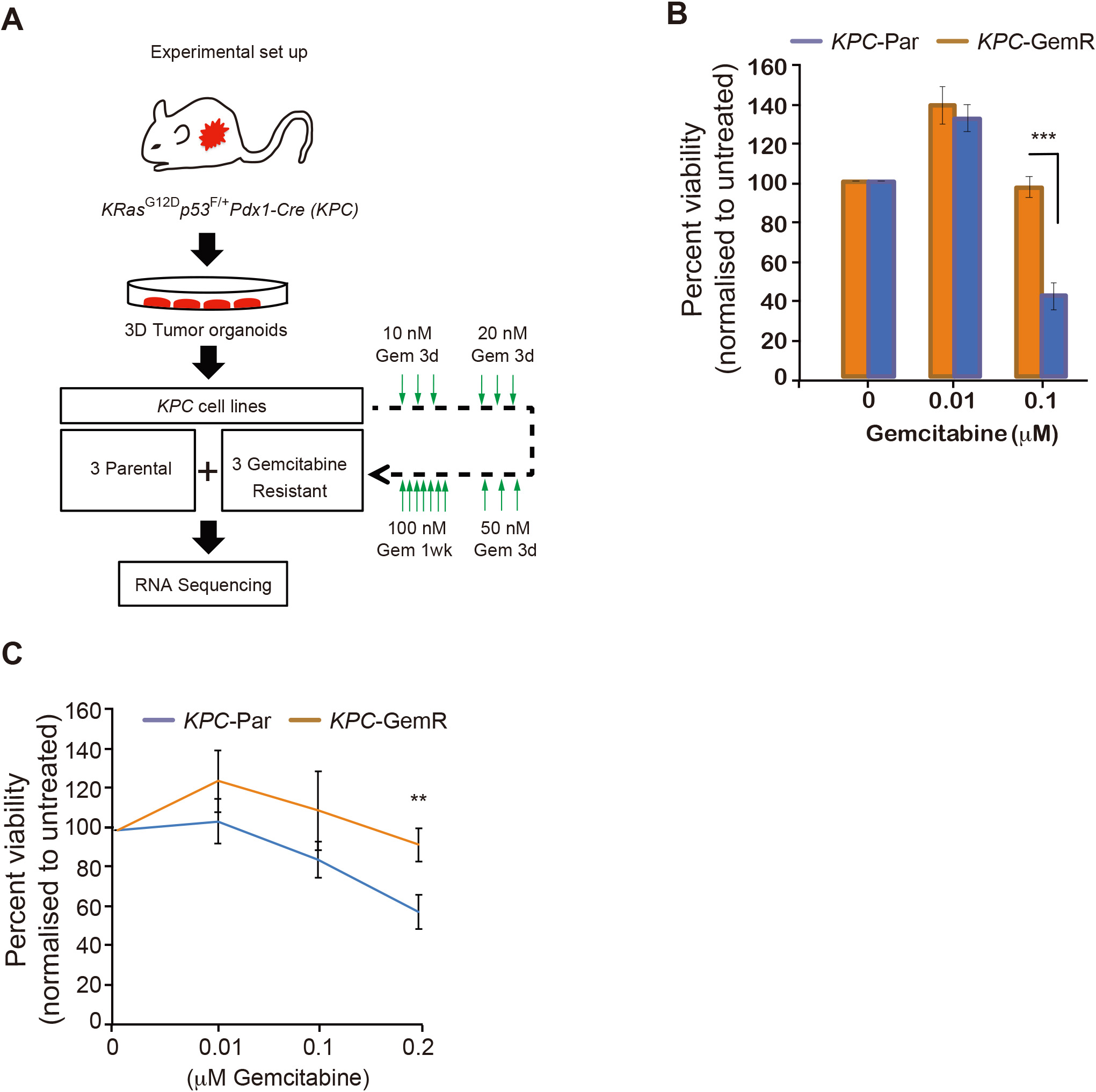
RNA-sequencing of KPC-Par versus KPC-GemR cells. A, Heat map showing changes at mRNA levels of 79 upregulated and 279 downregulated genes in gemcitabine resistant mPDA cells. B, Molecular and cellular functional pathway analysis generated by IPA. Ca’–Ch’, Immunohistochemistry of pancreatic adenocarcinoma tissues obtained from The Human Protein Atlas (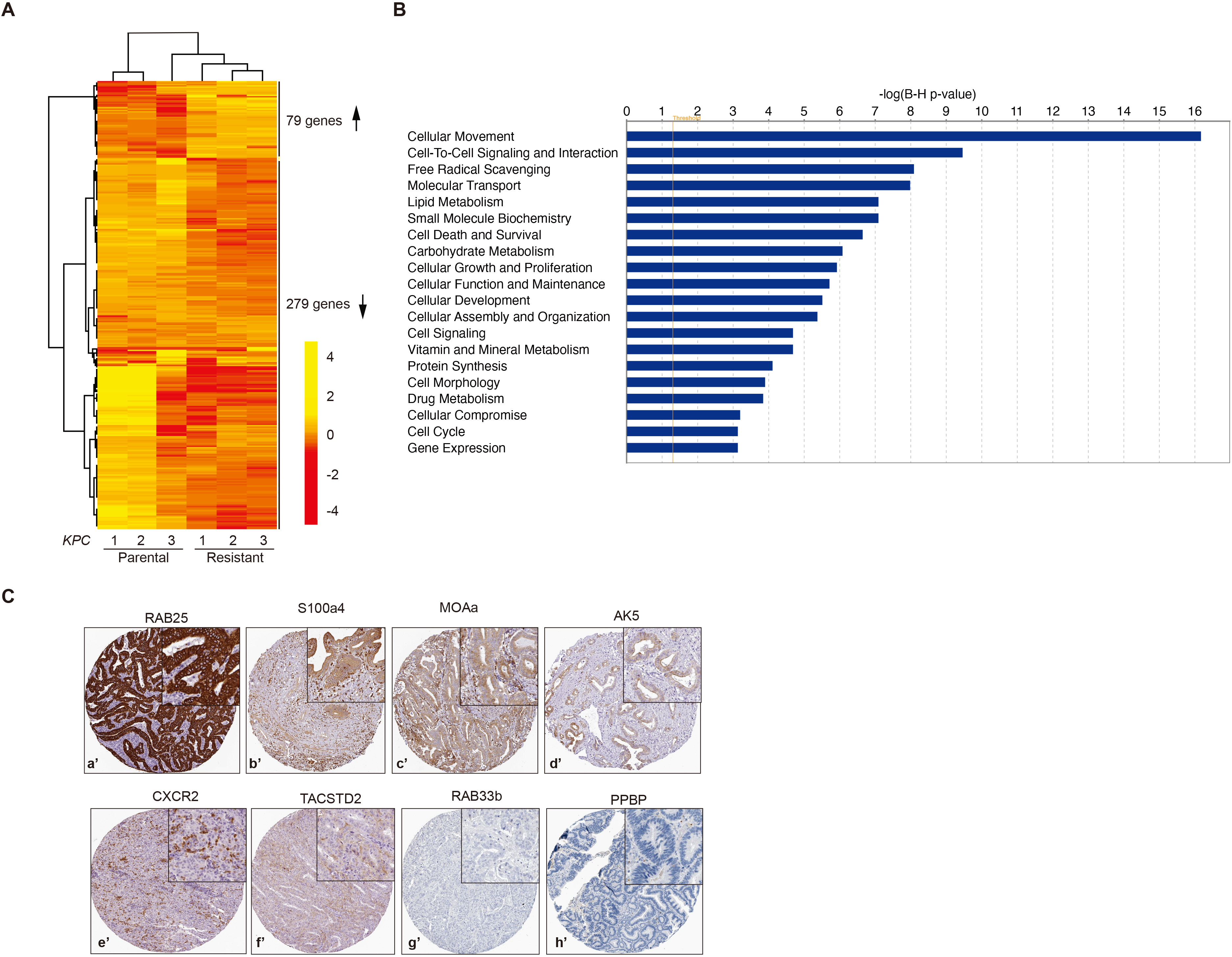
Establishment of mPDA GemR cell lines
To understand the mechanisms of gemcitabine resistance in pancreatic cancer, we first harvested small-to-medium-sized tumours from the pancreas of KPC mice. To prevent the confounding factors such as immune and stromal cells affecting our studies, we derived 3 independent primary organoid tumour cell lines (Fig. 1A) (KPC-Par). To model gemcitabine resistance in PDA we treated these low passage primary cell lines with gradually increasing doses of gemcitabine and derived 3 gemcitabine resistant mouse PDA (KPC-GemR) cell lines (Fig. 1A). Indeed, mPDA tumour cells develop resistance to gemcitabine and provide a model to study mechanisms of acquired gemcitabine resistance (Fig. 1B and C).
RNA-sequencing revealed diverse set of genes deregulated in KPC-GemR cells
Next, we performed a whole transcriptome comparison of low passage KPC-Par and KPC-GemR cells and found 79 genes significantly up-regulated while 279 genes downregulated in resistant cells (Fig. 2A). Interestingly, we found downregulation of Ent4 – one of the four members of the equilibrative nucleotide transporter (ENT) family involved in gemcitabine uptake, suggesting that gemcitabine uptake might be affected in those cells [29]. In addition, S100 Calcium binding protein 4 (S100a4) – a proposed prognostic marker for gemcitabine resistance in PDA [30] – is also upregulated in resistant mPDA cell lines. Thus, our data suggest that pro-survival mechanisms of chemotherapy resistance are conserved between murine and human PDA. We performed ingenuity pathway analysis to uncover the molecular and cellular pathways deregulated in KPC-GemR cells (Fig. 2A). The top 10 most deregulated pathways in KPC-GemR cells include cell movement, growth and proliferation, and drug metabolism (Fig. 2B). Importantly we made a shortlist of 10 druggable genes (proteins with enzymatic activity) (Table 1), that were significantly upregulated in KPC-GemR cells, and reason that one or more of those candidate proteins might be involved in gemcitabine resistance in those cells. We first confirmed the pancreatic adenocarcinoma tissue expression of our shortlisted targets by human protein atlas (Fig. 2C). We find various degrees of expression of our target proteins in human PDA ranging from tumour cell specific expression to stromal or no expression at all (Fig. 2C). Four proteins showed tumour specific expression in the order of strongest to weakest including RAB25, S100a4, MOAa, and AK5 (Fig. 2Ca’–Cd’). Two proteins CXCR2 and TACSTD2, showed strong stromal expression although weak to moderate expression was detected in tumour cells, (Fig. 2Ce’ and Cf’), while two proteins, PPBP and RAB33b were undetected (Fig. 2C-g’ and C-h’). Tumour expression data for two of our shortlisted proteins (ILR5a and GLT1D1) was not available with the human protein atlas.
List of selected DEG in gemcitabine resistant mPDA cells
List of selected DEG in gemcitabine resistant mPDA cells
RAB25 mediates cell survival in response to gemcitabine. A, Cell viability shown as percent confluency for stable mPDA cells with indicated shRNA expression, shpLKO.1 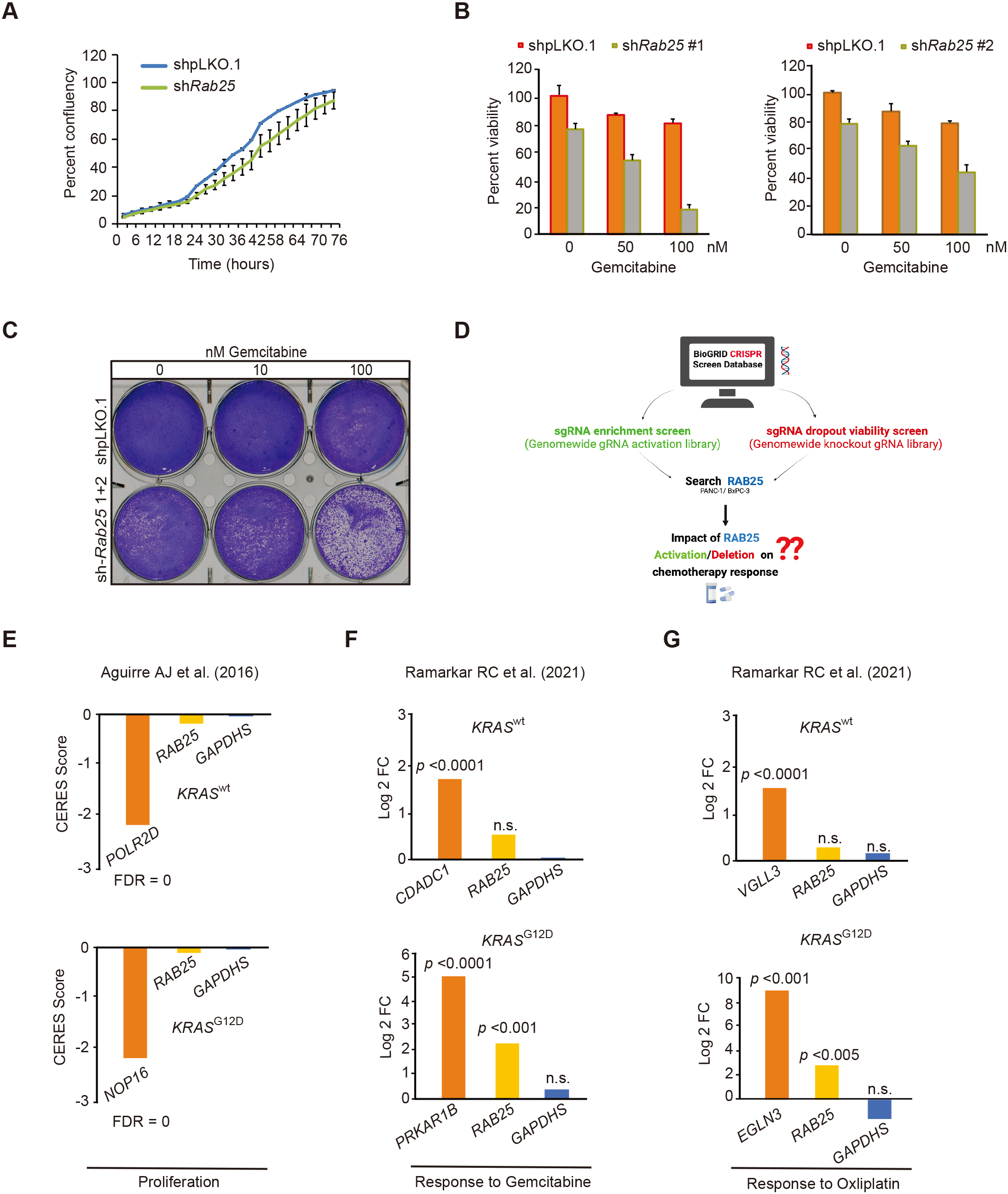
Genetic aberrations of Rab25 in different cancer types and survival rate. A, Alteration frequency of RAB25 gene in different cancers, including pancreatic cancer obtained from cBioPortal (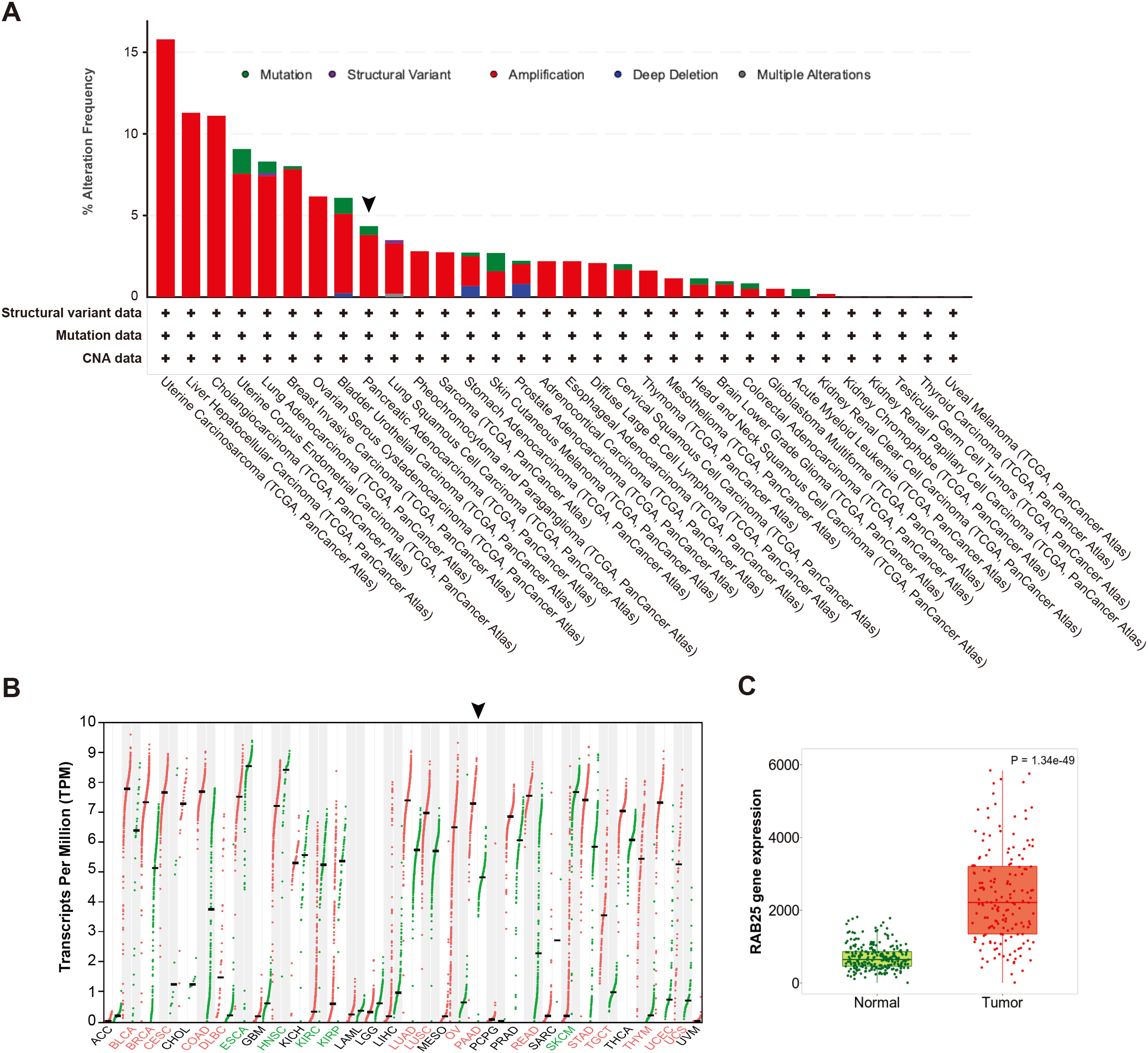
Nevertheless, we generated stable KPC-GemR cell lines expressing at least two different lentivirus mediated shRNA targeting each of our shortlisted genes and treated those cells with gemcitabine to monitor the viability of those cells in response to gemcitabine (data not shown). Interestingly, knockdown of Rab25 despite having minimal effect on KPC-GemR cell proliferation, strongly sensitized KPC-GemR cells to gemcitabine (Fig. 3A–C). To understand if RAB25 plays a similar pro-survival role in human PDA, we used publicly available datasets from pooled CRISPR/Cas9 viability screens performed in PDA cell lines (Fig. 3D). As indicated by the results from CRISPR/Cas9 viability screen (CERES Score
RAB25 is amplified across multiple human cancers
We next determine the mutational spectrum of RAB25 across multiple human cancers through cBioPortal.org platform using publicly available TCGA datasets. Strikingly, RAB25 was amplified in multiple human cancers including up to 5% PDA (Fig. 4A), suggesting an oncogenic role for RAB25. Consistent with that, the RAB25 gene expression was increased
RAB25 genomic alterations in pancreatic cancer. A, Data showing alteration frequency of RAB25 gene in pancreatic cancer as obtained from cBioPortal (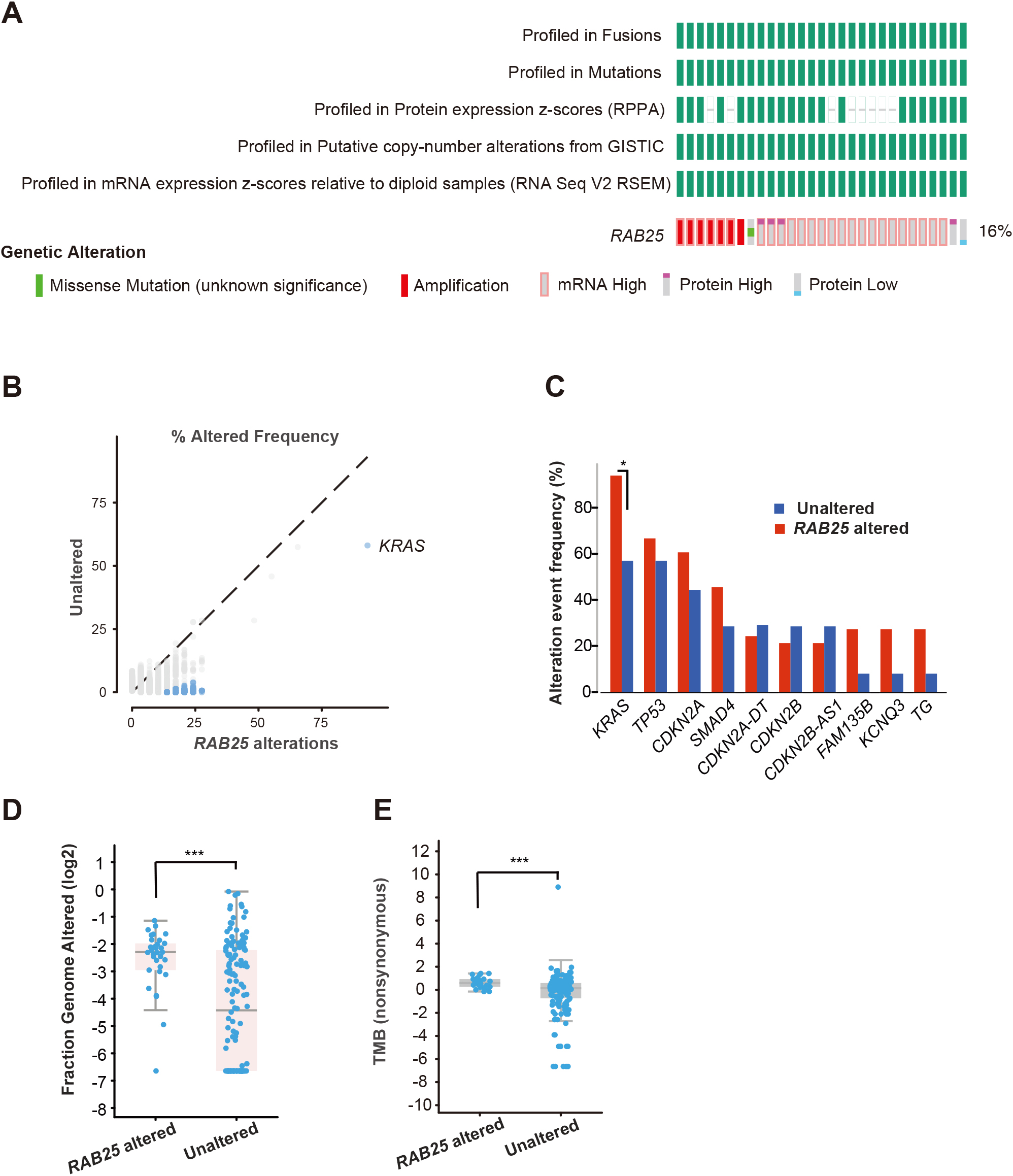
Higher expression of RAB25 is associated with poor patients’ survival in PDA cohorts. A, Overall survival of PDA patients with RAB25 high versus low expression cohorts as evaluated by kmplot.com survival analysis. B, Overall survival of PDA patients in RAB25 high versus low expression cohorts from GEPIA platform. C, Schematic showing possible mechanisms of RAB25 pro-survival activities in KRAS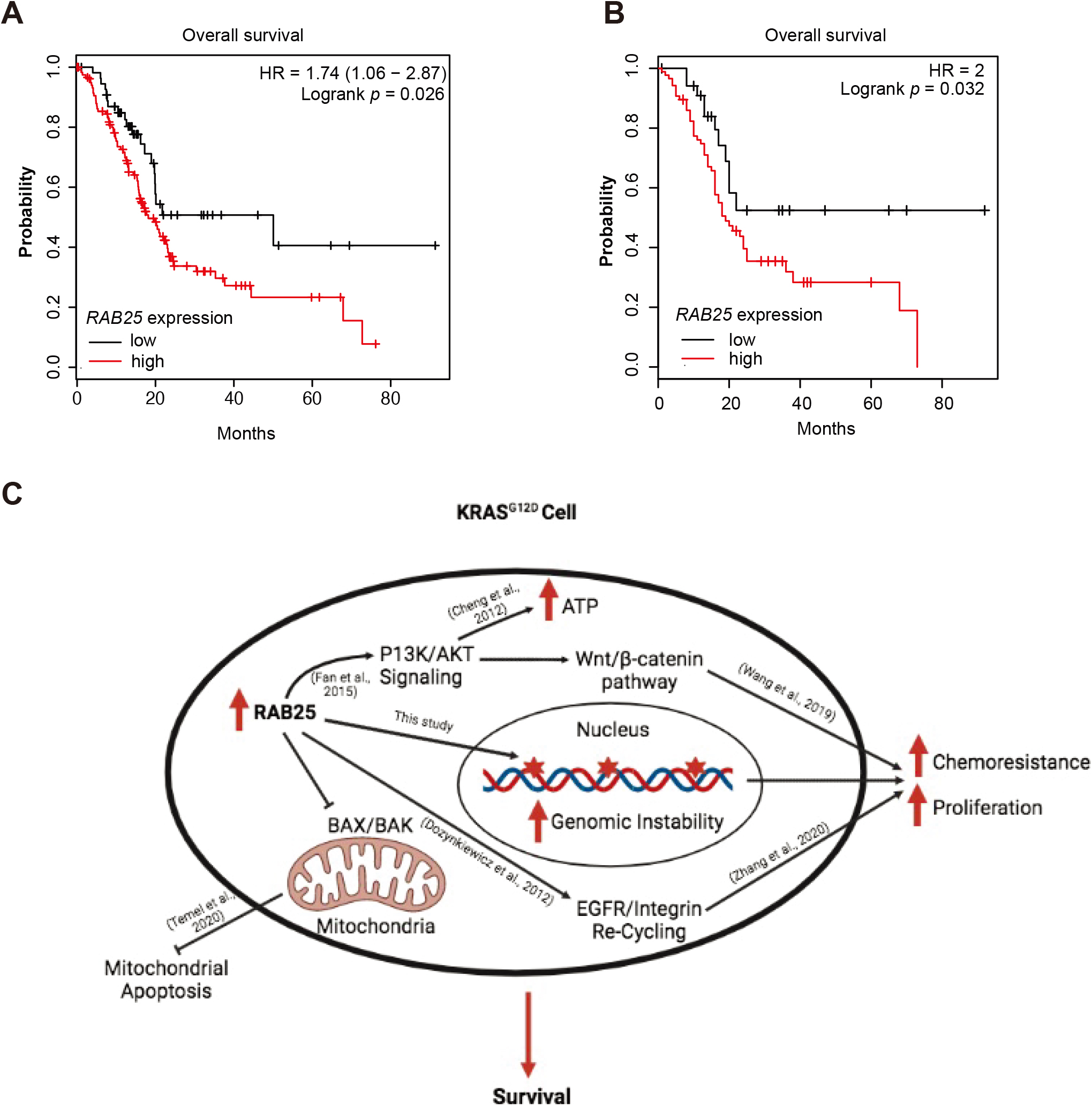
Next, we aim to detect the genetic alteration frequency of RAB25 specifically in pancreatic cancer. We use cBioPortal platform to visualize the TCGA pancreatic cancer dataset of
Finally, we used two independent publicly available datasets to check whether RAB25 expression correlates with patients’ survival in PDA. As expected, higher expression of RAB25 is associated with significantly reduced survival in patients cohorts from both the datasets (Fig. 6A and B).
Discussion
Pancreatic cancer continues to be one of the most difficult to treat cancers which is in sharp contrast to other cancers including breast, colon, and lung cancer, and myeloid malignancies where the overall patient survival has significantly improved over the last few decades. The frontline chemotherapy gemcitabine shows minimal benefits against PDA [32]. Thus, there is an urgent need to identify novel drug targets and prognostic markers of clinical relevance that can help tackle the menace of pancreatic cancer more efficiently. In this study we identify RAB25 to be associated with chemotherapy resistance in pancreatic cancer cell lines and with genomic instability and poor patients’ survival.
Interestingly we show that mPDA cells rapidly develop resistance to increasing doses of gemcitabine in a long-term culture. This resistance is associated with dramatic upregulation of genes from multiple cellular signalling pathways including cell survival, cell death, and drug metabolism. One of the highest upregulated gene RAB25, is very strongly expressed in human PDA, leading us to believe that RAB25 must be involved in PDA pathophysiology. However, RAB25 is not essential for survival of neither mPDA nor human PDA cell lines but rather its inhibition sensitized those cells to gemcitabine. These results highlight the possibility that RAB25 is specifically required for survival of pancreatic cancer cells in response to chemotherapy.
Strikingly, by reanalysing the published data from the CRISPR/Cas9 viability screens in response to chemotherapy drugs gemcitabine and oxaliplatin in PDA cell lines, we find that RAB25 sgRNA was significantly enriched only in KRAS
Whereas our study shed some important light on the role of RAB25 in pancreatic cancer, the exact molecular mechanism of how RAB25 helps PDA cells evade gemcitabine induced cell death is merely speculative. There could be several possibilities that may explain RAB25’s anti-chemotherapy activities in PDA (Fig. 6C). For example, RAB25 overexpression is associated with aggressive phenotypes of ovarian, breast, and pancreatic cancer [33]. Some of those aggressive phenotypes can be explained by CLIC3/RAB25 mediated recycling and activation of integrin/EGFR signalling [20]. Additionally, RAB25 is associated with chemotherapy and radiotherapy resistance in multiple cancers [33], most likely due to the inhibition of mitochondrial apoptotic pathway [34]. Noticeably, RAB25 helps cancer cells encounter bioenergetic stress by mTOR activation and production of ATP [35]. Tumours in PDA are largely hypoxic and under continuous nutritional and hypoxic stress due to poor vascularization [36]. Thus, RAB25 upregulation may help those tumours by providing prosurvival signalling and maintaining the ATP levels for cellular growth and signalling (Fig. 6C). This is further highlighted by strong upregulation of RAB25 in tumour versus normal pancreatic cancer tissues as shown in our analyses.
Recent advances in the field of stapled peptide technology for targeting difficult to target small molecules like RAB25 has generated a lot of interest for clinical manifestation. Particularly, targeting RAB25 in multiple cancer cell lines using a RAB25 specific peptide showed context dependent anti-tumour effects [37]. Our study highlights the importance of targeting RAB25 in combination with chemotherapy in human PDA cell lines and in autochthonous mice models of PDA and provides evidence that RAB25 might be an interesting therapeutic target in addressing chemotherapy resistance in highly aggressive otherwise difficult to treat PDA. Finally, our results indicate that high RAB25 expression in human PDA is strongly associated with poor clinical outcome and stratification of PDA patients based on RAB25 expression can identify gemcitabine responders and help identify alternate treatment plans for non-responders.
Footnotes
Acknowledgments
We would like to thank Advanced Sequencing, and Bioinformatics and Biostatistics Core facilities of the Francis Crick Institute for RNA sequencing and data analysis. This work was supported by the Francis Crick Institute (U.K) and Hamad bin Khalifa University (Qatar). O.M.K. was funded by an EMBO long-term postdoctoral fellowship award (ALT-549-213), a Swedish International postdoctoral fellowship award (VR-537-2013-359), and by an intramural grant from Hamad Bin Khalifa University (Qatar Foundation, Qatar).
Author contributions
Conception: OK designed the study. OK performed mouse experiments and generated the RNA sequencing data.
Interpretation or analysis of data: AK analysed ORCS, GEPIA, and TCGA datasets. AK performed IPA analysis on the RNA sequencing data. OK analysed survival curves from the TCGA datasets.
Preparation of the manuscript: OK, AK, and SA assembled figures and wrote manuscripts. AB provided important feedback and final approval for submission.
Revision for important intellectual content: N/A.
Supervision: AK and SA are graduate students under supervision of OK.
