Abstract
Long noncoding RNAs (lncRNAs), as well-known modulator of the epigenetic processes, have been shown to contribute to normal cellular physiological and pathological conditions such as cancer. Through the interaction with epigenetic regulators, an aberrant regulation of gene expression can be resulted due to their dysregulation, which in turn, can be involved in tumorigenesis. In the present study, we reviewed the lncRNAs’ function and mechanisms that contributed to aberrant epigenetic regulation, which is directly related to gastrointestinal cancer (GI) development and progression. Findings indicated that epigenetic alterations may involve in tumorigenesis and are valuable biomarkers in case of diagnosing, assessing of risk factors, and predicting of GI cancers. This review summarized the accumulated evidence for biological and clinical application to use lncRNAs in GI cancers, including colorectal, gastric, oral, liver, pancreatic and oesophageal cancer.
Background
LncRNA
Molecular mechanisms of lncRNAs mediated epigenetic regulation in GI cancer development, progression and metastasis. (a) lncRNAs can recruit transcription factors (TF) or bind to enhancer/promoter sites to regulate transcriptional region and gene transcription; (b) lncRNAs can recruit epigenetic regulators, including modification enzymes to modify chromatin conformation resulting in regulating gene expression.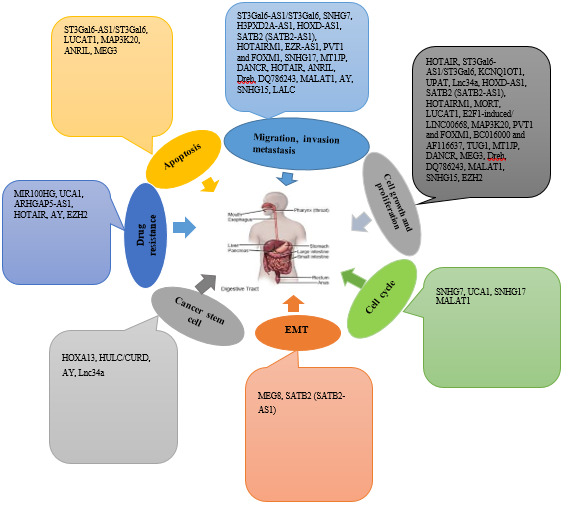
Long noncoding RNAs (lncRNAs), as transcripts that are longer than 200 nucleotides, are classified as intergenic lincRNAs, intronic ncRNAs, and sense and antisense lncRNAs. The structure of lncRNAs is similar to mRNA and may have cap structures and poly A tails. However, in most cases, they are less conserved and have relatively lower expression levels than mRNAs. It has been revealed that lncRNAs do not involve in any protein coding process to be transcribed by the RNA polymerase II (RNA Pol II) [1]. So far, 15,778 lncRNA transcripts have been recognized in human genomes that can produce about 30000 lncRNA transcripts, and it seems they are annually growing in quantity [2]. The lncRNAs can be found in specific cell types much more than protein coding RNAs (mRNAs), which can have various representations in different types of cancer. There is no exact record of the number of functional lncRNAs, yet it can be claimed that many lncRNAs have significant impacts on various biological functions, and many pathologies such as cancer can result from their dysregulation [3]. There are evidences showed that numerous lncRNAs function in tumorigenesis by involving in the regulation of cell proliferation, migration, and metastasis. The clinical studies also indicated their potentials as valuable targets for cancer therapy [4]. LncRNAs do not involve the same action mechanisms, and this related to their cellular localization and molecules interactions. Various functional mechanisms can be recognized in various cellular components, which actually have allocated to distinctive lncRNA types [4]. The epigenetic dysregulation is one of the characteristics of many cancers, yet the fact that epigenetic changes can lead to cancer development has not been indicated precisely. In the nucleus, the alteration of the epigenetic pattern of genes is frequent mode of action of lncRNAs. Notably, they act through directing epigenetic modulators to distinct loci or regulating chromatin folding and localization to mediate enhancer-promoter interactions (Fig. 1) [5].
LncRNAs basically function as two ways cis-acting and trans-acting, based on local distance to target gene transcript. In cis-acting type, the mature lncRNAs regulate the expression levels of nearby genes [6]. The function of nuclear lncRNAs is basically in cis-acting to make some changes to local gene transcription. As for the amount, cis-acting lncRNAs can frequently be regulated at a low expression level; in fact, they can be seen at only a few copies in every cell. In case of some nuclear lncRNAs function in trans-acting, the gene expression regulation is completed distance from the place they have been synthesized or positioned in diverse loci [6]. However, the detailed localization of nuclear lncRNAs and their regulatory function is so far explored. Today’s approaches toward genome-wide indicate that mutations in regulatory regions such as gene promoters and enhancers can lead to changes in chromatin structure, which result in deviations in the expression of lncRNAs [7].
lncRNAs contribute to GI cancer development and progression by involving in various cellular processes. Different modalities of lncRNA biology and their roles in epigenetic regulation are described in GI cancers. Cellular mechanisms of lncRNAs in biological processes, including cell proliferation, cell apoptosis, cell cycle, cell migration, invasion and metastasis, epithelial-mesenchymal transition (EMT), cancer stem cells and drug resistance may be mediated by epigenetic regulation.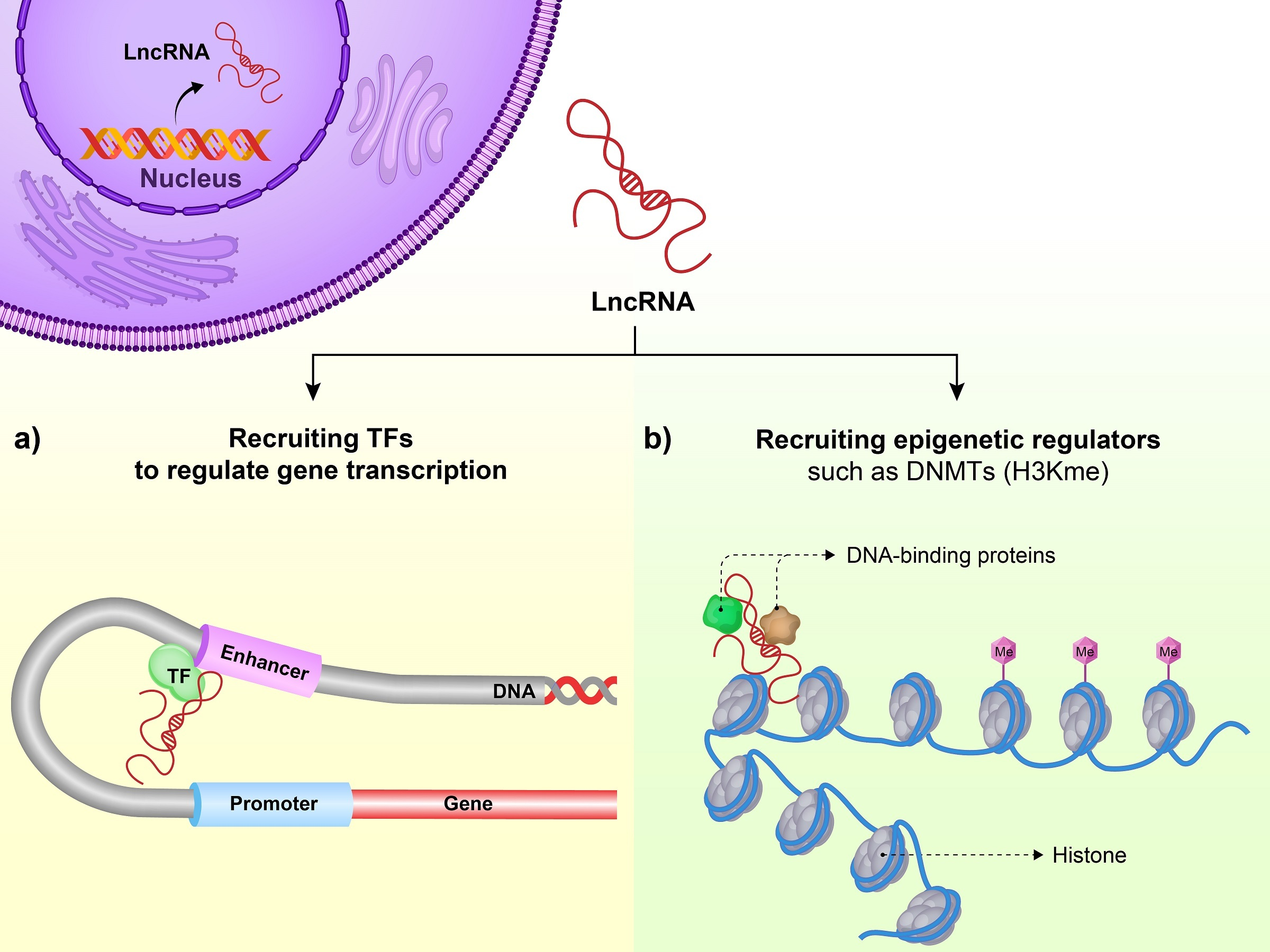
Epigenetics is described as changes in gene expression without alteration of the corresponding DNA sequences. Common epigenetic processes include DNA methylation, chromatin modifications and those mediated by non-coding RNAs specially miRNA and lncRNAs. Epigenetic alterations have frequently been revealed to contribute to both GI cancer initiation and progression. The abnormal DNA methylation has defined as the most commonly epigenetic alteration in GI cancers, including colorectal, gastric, esophageal, pancreatic, oral and liver cancers. The silencing of key tumor suppressor genes by promoter DNA hypermethylation has regularly been reported in GI cancers [8, 9]. Since aberrant DNA methylation may be involved in various GI cancers, in this review, the functions and underlying mechanisms of lncRNAs found in the related epigenetic dysregulation will be more carefully elaborated [8]. In this regard, some studies have indicated that epigenetic changes can be involved in tumorigenesis, as they can be biomarkers for diagnosis, risk assessment and prognosis of GI cancers [8, 9]. As delineated earlier, lncRNAs can epigenetically mediate various cellular processes contributing to GI tumorigenesis, including DNA methylation and chromatin modification (Fig. 2). The present review study suggests some evidence for biological and clinical applications in using lncRNAs in GI cancer types such as colorectal, gastric, oral, liver, pancreatic and esophageal cancers. Table 1 showed the lncRNAs that can regulate the GI cancers as well as their molecular functions in epigenetic dysregulation.
LncRNAs that epigenetically mediate the GI cancers and related molecular functions
DACOR1
DNMT1-associated colon cancer repressed lncRNA 1 (DACOR1), mapped on 15q22.33, has been confirmed to increase DNA methylationeither in the promoters or intergenic regions [10]. Notably, these regions are shown to be connected to hypomethylated regions in CRC. Moreover, lncRNA DACOR1 can decrease AP-1 transcription factor activity through suppressing FOS and JUN. DACOR1 also can regulate CpG-enriched sites in gene promoters and contribute to induce the activated transcription factor 2 (ATF2) and associated signaling pathways in CRC [11]. Altogether, functional studies revealed a pivotal regulatory role of lncRNA DACOR1 in DNA methylation in CRC [11]. Hence, the expression pattern of lncRNA DACOR1 might arise as an important mechanism by which DNA methylation process be involved in CRC.
HOTAIR
LncRNA homeobox transcript antisense intergenic RNA (HOTAIR) is mapped on the HOXC gene in 12q13.13 and plays an oncogenic role in CRC [12]. This lncRNA has been demonstrated to contribute to epigenetic regulation of various genes involved in cell signaling pathways by co-operating with polycomb repressive complex 2 (PRC2). LncRNA HOTAIR has also been exhibited to be overexpressed in CRC cells and clinical samples. Functional studies have revealed that knockdown of lncRNA HOTAIR can remarkably reduce growth of the cells and formation of colonies, signifying this lncRNA functions as an oncogene in CRC [13]. Moreover, lncRNA HOTAIR can bind to EZH2 in CRC cells. In a nutshell, lncRNA HOTAIR may involve in CRC tumorigenesis through interacting with EZH2 and is an useful tool for the cancer therapeutic approach [14].
ST3Gal6-AS1/ST3Gal6
LncRNA ST3GAL6-AS1 (ST3GAL6 Antisense RNA 1) is copied from the corresponding gene locating on chr3 with 18,322 bases in length [15]. LncRNA ST3Gal6-AS1 has been demonstrated to induce modification and ST3Gal6 transcription through histone methyltransferase MLL1 recruitment to the promoter of ST3Gal6. In CRC cells, lncRNA ST3Gal6-AS1 may be served as a target of transcription factor Foxo1. Indeed, ST3Gal6-AS1/ST3Gal6 axis is complicated in
SNHG7
Small nucleolar RNA host gene 7 (SNHG7), mapped on 9q34.3, has been shown to be upregulated in CRC [16]. Also, the upregulation of SNHG7 is correlated to the poor prognosis of CRC individuals and may be as a valuable clinical biomarker. Furthermore, SNHG7 can endorse CRC cell proliferation, metastasis, and mediate cell cycle. In this regard, SNHG7 and GALNT7 (Polypeptide N-Acetylgalactosaminyltransferase 7) was noticed for simultaneity expression, and in case of SNHG7 and miR-34a, a significant negative relationship was observed through endogenous RNA (ceRNA) studies. Other findings showed that SNHG7 promoted the cell growth and invasion as a striving endogenous RNA in order to modulate GALNT7 through sponging miR-34a. Furthermore, it was observed that SNHG7 could function as an oncogene to affect PI3K/Akt/mTOR axis through interacting endogenous miR-34a and GALNT7. The possibility of CRC-related SNHG7 and miR-34a implication in epigenetic-related tumorigenesis through GALNT7 suggests the possible implication of SNHG7/miR-34a/GALNT7 pathway for CRC therapy [17].
MIR100HG
MiR-100-let-7a-2-miR-125b-1 cluster host gene (MIR100HG), with 174,708 bases, is located on 11q24.1 [18]. High throughput analyses revealed that lncRNA MIR100HG was upregulated without key genetic events that are related to cetuximab resistance. MIR100HG and miR-100/125b deregulation was noticed in cetuximab-resistant CRC progressed on cetuximab. In recent years, a reciprocal feedback loop was explored in view of MIR100HG and GATA6 interaction. In this way, GATA6 (GATA Binding Protein 6) represses MIR100HG. Findings discovered that long non-coding antisense transcript of GATA6 (GATA6-AS) co-operates with the epigenetic modulator LOXL2 (Lysyl Oxidase Like 2) in order to monitor and regulate endothelial gene expression by making alterations on histone methylation. This repression has been shown by previous studies using target of GATA6 by miR-125b [19]. These studies emphasized the epigenetic mediations-related cetuximab resistance as promising clinical points for cancer management.
SH3PXD2A-AS1
SH3PXD2A (SH3 and PX Domains 2A), as a gene specific for corresponding protein with a 261,517 bases, is situated on 10q24.33. A study showed that SH3PXD2A-AS1 may promote cell proliferation and invasion and increased tumorigenesis [20]. Mechanistic examinations designated that SH3PXD2A-AS1 could epigenetically lead to a repression of P57 and KLF2 by interacting with EZH2. In the same way, results of the previous study indicated that SH3PXD2A-ASl-mediated oncogenesis could be reduced through an overexpression of P57 or KLF2. It was also in line with the results of the study that the expression of SH3PXD2A-AS1 could be negatively associated with P57 and KLF2 in CRC [21]. There is some evidence indicating that SH3PXD2A-AS1 play an oncogenic function in CRC, and have potentials for diagnosis and therapy of CRC.
UCA1
Urothelial cancer related 1 (UCA1), situated on 19p13.12, consists of 1.4 kb. An empirical research done on this field argues on the epigenetic mechanisms that are mediated through the lncRNA and can modulate cancer signalling pathways [22]. The function of lncRNA UCA1 in CRC and its function of epigenetic modulation are what being described extensively. UCA1 can be noticeably found in cancer cells and it can mediate regulation at transcriptional level when interacting with epigenetic effectors. This is being done through a specific regulation by chromatin crosstalk and miRNAs sponging [23]. In addition to this, the biological roles of UCA1 in the modulation of cell proliferation and its connection to drug resistance are subject of investigation [23]. Furthermore, it has been discovered that lncRNA UCA1 can interact with histone methyltrasferase EZH2, and in turn, promote cell proliferation and migration of cancer cells through downregulating the expression of matrix metalloproteinase-9 (MMP-9) [24]. The lncRNA UCA1 also can increase cell cycle progression and involve in tumorigenesis through binding to enhancer of EZH2 and repression of P27 transcription by histone methylation (H3K27me3) [25].
KCNQ1OT1
KCNQ1 Opposite Strand/Antisense Transcript 1 (KCNQ1OT1) is mapped on 11p15.5 with 91,671 bases in length [26]. A related study indicated that KCNQ1OT1 was remarkably upregulated in CRC cells with an overexpression of
UPAT
Ubiquitin-like with PHD and RING finger domains 1 (UHRF1) is mapped on Chr.17 [28]. The lncRNA named UPAT [ubiquitin-like plant homeodomain (PHD) is a recently more-detailed studied new gene (RING) finger domain-containing protein 1 (UHRF1) protein associated transcript] that is a requirement for the tumorigenicity and survival of tumor cells. UPAT can interact with UHRF1 and can stabilize it by intervening with its
CYP2A7, MIA and MIA-RAB4B
CYP2A7, MIA and MIA-RAB4B, with a 32 kb in length, is located on chromosome 19q13. An examination in terms of loss of heterozygosity and an allele revealed 5 common deletions [30]. Also, it has been highlighted that MIA and MIA-RAB4B, situated 35 kb upstream of the deletion, could lead to an increase in regulation in the polyps. MIA-RAB4B, RAB4B, PIM2 and TAOK1 have similar binding sites of miR-24, in their 3’UTRs. PIM2 and TAOK1, that are considered as targets of miR-24, were properly overexpressed with MIA-RAB4B in the polyps; this may imply that MIA-RAB4B acts as a competing effector. The results revealed that the 19.13 deletion interrupted chromatin structure could cause dysregulation of several genes, and lncRNA MIA-RAB4B plays a significant role in CRC through unknown genetic and epigenetic mechanisms [30].
Lnc34a
It has been recently noticed that lncRNA lnc34a is overexpressed in colon cancer stem cells (CCSCs), thereafter involves in an asymmetric division which is done through binding the miR-34a [31]. Lnc34a employees Dnmt3a through PHB2 and HDAC1 to simultaneously methylate and deacetylate the promoter of miR-34a, and thus this leads to an epigenetically silence of miR-34a expression. In this regard, Lnc34a levels can influence CCSC self-renewal and CRC cells proliferation in vivo. Lnc34a is overexpressed in advanced stage of CRC, something that can be associated with miR-34a silencing and tumor cell growth [31]. Targeting miRNAs, it is high pointed the regulatory involvedness of ncRNAs throughout genome.
HOXD-AS1
LncRNA HOXD cluster antisense RNA 1 (HOXD-AS1) exists in chromosome 1q31.2 [32]. It has been revealed that HOXD-AS locates in the nucleus and is downregulated in CRC specimens. When the HOXD-AS1 expression is at lower levels, the phenomenon is often related to reduced outcomes of CRC patients. Due to the low levels of HOXD-AS1 regulation, tumor cell growth is enhanced and CRC tumorigenesis and metastasis is promoted. Empirical studies performed on the functional aspects represented that HOXD-AS1 could inhibit HOXD3 through recruiting PRC2 to engage the H3K27me3 at the HOXD3 promoter [32]. Therefore, HOXD3 as a transcriptional activator, can increase Integrin
SNHG14/miRNA-3940-5p/NAP12
Small nucleolar RNA host gene 14 (SNHG14), with a length of 440,880 bases, is located on 15q11.2 [33]. LncRNA SNHG14, hsa-miRNA-3940-5p and NAP1L2 mRNA have some outstanding characteristics in function that are more distinctive than conventional well-known tumor markers for identification of CRC patients. LncRNA SNHG14- hsa-miR-3940-5p and NAP1L2 mRNA levels represents a high sensitivity with a quite high level of accuracy 93%. It is interesting to know that hsa-miRNA-3940-5p might act for prognosis of colorectal carcinoma [34]. In this regard, lncRNA SNHG14- hsa-miR-3940-5p - NAP1L2 mRNA can help in the controlling of CRC. The extracellular RNAs can be served as a group of minimally-invasive biomarkers for CRC diagnosis. In silico results in the recognition of NAP1L2 alteration with the distinctive epigenetic modulators showed a pattern (NAP1L2 gene, lncRNA-SNHG14 and hsa-miR-3940-5p) that can be served as promising tools for diagnosing the CRC [33].
SATB2 (SATB2-AS1)
A novel long non-coding RNA, antisense transcript of SATB2 is located on 2q33.1 [35]. The changes in the lncRNA expression levels and the antisense transcript of SATB2 (SATB2-AS1), is a pivotal cancer modulator, that has previously been discovered in the progress of CRC. SATB2-AS1 is commonly regulated in CRC cells and tissues at lower levels; patients with low SATB2-AS1 have shown a poor prognosis [36]. Regulation of SATB2-AS1 at low levels could cause cell growth and invasion, which indicates it functions as a suppressor. Noticeably, it was indicated that SATB2-AS1 can suppress CRC progression and function as a framework to bind p300. This process results in acetylation of H3K27 and H3K9 at the SATB2 promoter that regulate SATB2 at a higher level, followed by suppression of CRC. SATB2 afterwards recruits HDAC1 to the Snail promoter, through repressing Snail and moderating the movement of epithelial-to-mesenchymal transition (EMT) [35]. Considering all these together, the data reveals that SATB2-AS1 is an effective modulator of the SATB2-Snail network in CRC tumorigenesis.
HOTAIRM1
HOXA transcript antisense RNA, myeloid-specific 1 (HOTAIRM1) gene is mapped on a HOX gene cluster [37]. Recent studies have revealed that HOTAIRM1 might be aberrantly up-regulated in malignancies and was related to the grade of tumor in cancer patients [38]. Furthermore, HOTAIRM1 silencing resulted in some impacts on tumor suppression that could hinder cell proliferation. In vivo experiments have revealed that HOTAIRM1 could reduce the development of the tumor. By the way, HOTAIRM1 could function as HOXA1 gene expression regulator. HOXA1, is an oncogene, the expression levels of which could increase in the cancer cells. Functionally, HOTAIRM1 involves in the demethylation of histone H3K9 and H3K27 and could lead to a decrease in DNA methylation pattern by sequester epigenetic modifiers G9a and EZH2 (histone methyltransferases), and DNA methyltransferases (DnmTs) [37]. The inherent function of HOTAIRM1 to increase tumor cell growth through the regulation of epigenetic HOXA1 has also been shown. Such findings are the typical representation of chromatin-related gene regulation, which might benefit researchers to properly enlighten the function of lncRNAs [38].
SNHG17
LncRNA named small nucleolar RNA host gene 17 (SNHG17) is recently documented to dysregulate in some malignancies and induce carcinogenesis and metastasis. Recently, SNHG17 has been recognized for potential biological functions in CRC tumorigenesis. Dysregulation of SNHG17 has been explored to be as a valuable prognostic factor. In this regard, functional studies have displayed that SNHG17 increase cancer cell growth via mutually modulating SNHG17-Trim23-PES1 signaling and SNHG17-miR-339-5p-FOSL2-SNHG17 axis. The molecular network is considered as a distinct network including multiple signaling pathways connected to epigenetic regulation processes [39].
LALC
LZTS1 associated lncRNA in CRC (LALC), another CRC-related lncRNA, has been newly identified to mediate CRC progression and metastasis. Mechanistically studies confirmed that lnc-LALC could increase the invasion of the cancer cells via downregulation of LZTS1. Additional researches revealed that lnc-LALC might recruit DNMTs to the LZTS1 promoter through binding with enhancer of zeste homolog 2(EZH2). Thereafter, this lncRNA could induce dysregulation of LZTS1 through DNMTs-interceded DNA methylation. In total, recent findings indicated the Lnc-LALC/LZTS1 signalling pathway as an oncogenic network in CRC progression and metastasis [40].
Esophageal squamous cell carcinoma (ESCC)
MORT
LncRNA mortal obligate RNA transcript (MORT) is located on human chromosome 19 [41]. This lncRNA is known as a hominid-specific long non-coding RNA (ZNF667-AS1, Gene ID: 100128252). However, lncRNA MORT is epigenetically supressed during immortalization of tumor cells [42]. Also, it has been documented that the DNA methylation can contribute to MORT silencing in the tumors. Certainly, DNA methylation could associate with MORT silencing in a number of malignancies (cervical, oesophageal and stomach cancers). It has been indicated that MORT expression is decreased by DNA dys-methylation in numerous TCGA cancer types. Therefore, it is suggested that the MORT gene might be one of the most common epigenetic deregulations related to cancers. These discoveries provide an appropriate indication for a tumor suppressor role for MORT. In this regards, lncRNA MORT has been shown to overexpress in tumor tissues and suggest as a possible clinical therapeutic tool for ESCC [41].
LUCAT1
Lung cancer-associated transcript 1 (LUCAT1) is in 5q14.3 with a length of 11,374 base [43]. The lncRNA LUCAT1 can be linked to the tumorigenesis of ESCC. The expression of LUCAT1 has been reported to be meaningfully increased in both ESCC cells and tumor tissues, indicating as a promising target for clinical setting of ESCC such as prognosis and diagnosis. LUCAT1 knockdown causes a reduction in cell growth. This is done through decreasing DNA methylation in KYSE-30 cells. Besides, LUCAT1 siRNA causes DNMT1 protein reduction levels with no roles in transcription regulation. A research has confirmed that patients with LUCAT1 overexpression had lower survival. The clinical findings indicated that LUCAT1 could affect constancy of DNMT1 via DNA methylation, which results in promoting and metastasis of ESCC [43]. The various observations showed LUCAT1 has the potentials for diagnosis and therapy of ESCC.
EZR-AS1
Ezrin antisense RNA 1 (EZR-AS1), a lncRNA with 2,583 bases, is located on 6q25.3 [44]. A recent study revealed that antisense lncRNA EZR-AS1 could be associated with EZR expression in human ESCC [45]. Various analyses also showed that EZR-AS1 could promote cells invasion through up-regulation of EZR. Based on functional studies, antisense lncRNA EZR-AS1 might form a complex with RNA polymerase II to cause the activation of the transcription of EZR. Furthermore, EZR-AS1 might recruit SMYD3 to a binding site in a GC-rich area surrounding the EZR promoter, which can lead to the binding of SMYD3 and re-localization of H3K4me3. As a final point, the binding EZR-AS1 to SMYD3 leads to an enhancement of EZR transcription and expression. The analysis of the data indicated that antisense lncRNA EZR-AS1, a component in RNA polymerase complex and SMYD3-dependent H3K4 methylation, could up-regulate the EZR. Hence, lncRNA EZR-AS1 shows an essential function in promoting the EZR expression to increase development and invasion of some GI malignancies [44].
CASC9
LncRNA cancer susceptibility candidate 9 (CASC9) is located on chromosome 8q21.11. It has been revealed that lncRNA CASC9 is involved in proliferation, invasion, and metastasis of ESCC cells [46]. Functional studies showed that lnc-CASC9 promotes the integrin signaling pathway through stimulation of the transcriptional coactivator CREB-binding protein and H3K27 acetylation in the promoter of LAMC2 [47]. However, ECM-regulated signaling pathways and biological role of lncRNAs in tumor cell and metastasis of ESCC need to be cleared. Furthermore, clinical studies demonstrated that lncRNA CASC9 was correlated with the prognosis and metastasis of ESCC patients. These findings conclude that lncRNA CASC9 has some potential as a promising target for clinical setting of ESCC patients [48].
Gastric cancer (GC)
ARHGAP5-AS1
LncRNA antisense RNA 1 (antisense RNA 1 (head to head): Rho GTPase activating protein 5; ARHGAP5-AS1) is located on 14q12 with a 1,375 bases in length [49]. LncRNA ARHGAP5-AS1 is regulated at a higher level in chemoresistant GC cells. Meanwhile, high ARHGAP5-AS1 expression has been revealed in the literature to be linked with poor prediction of GC. It is noticeable to mention that high expression of this lncRNA is influenced by autophagy, and that SQSTM1 is in charge of moving ARHGAP5-AS1 to autophagosomes. In this regard, autophagy knockdown in the tumor cells might led to the overexpression of ARHGAP5-AS1. According to, it can increase the transcriptional level of ARHGAP5 as it can interact with ARHGAP5 promoter. ARHGAP5-AS1 can simultaneously stabilize ARHGAP5 mRNA in the cytoplasm. This is completed through engaging METTL3 to induce m
E2F1-induced/LINC00668
LncRNA LINC00668 (LINC00668), located on 18p11.31, can be noticeably found to be related to develop various malignancies. Integrated analyses have demonstrated that LINC00668 expression is considerably increase and is related to outcomes in GC [50] and significantly represses cell growth. Related studies on the phenomenon have revealed that LINC00668 can be the main transcriptional target of E2F transcription factor 1 (E2F1). LINC00668 and PRC2 are contributed to epigenetic suppression of cyclin-dependent protein kinase inhibitors (CKIs), such as p15, p16, p21, p27 and p57 that regulate the GC cell growth. Accordingly, it is suggested that E2F1-activated LINC00668, as a cell cycle effector, may enrich the relationship between lncRNA and the cell cycle regulated by E2F1 [51]. This signalling axis might be served as a prognostic and therapeutic target for therapeutic strategies in GC patients.
MAP3K20
LncRNA MAP3K20 antisense RNA 1 (MLK7-AS1), mapped on 2q31.1, has been identified in GC-related signalling pathway. LncRNA MLK7-AS1 has shown an increase in GC tissues in comparison with adjacent tissues [52]. Patients with GC who have a high level of MLK7-AS1 are less likely to survive. The phenomenon, known as loss-of-function, can reveal a knockdown of MLK7-AS1 for cell growth inhibition and apoptosis induction in HGC27 and MKN-45 cells. By the way, miR-375 has been identified as a target for MLK7-AS1. MLK7-AS1 may interact with Dnmt1 and recruit it to miR-375 promotor, hyper-methylating miR-375 promotor so that it can repress miR-375 expression. Thus, it has been determined that the suppression of MLK7-AS1 by siRNA can hinder the development of GC as it epigenetically regulates miR-375 [52]. Therefore, it can be implied that MLK7-AS1 is an effective biomarker for those who suffer from GC.
HOXA13
LncRNA HoxA transcript at the distal tip (HOTTIP), mapped on 7p15.2, functions as a transcription factor of the Homeobox (HOX) gene family. The biological functions lncRNAs-homeobox protein HOXA13 axis has been revealed to disappear in the GC derived iPS-like cells (iPSLCs) [53]. It has been demonstrated that BMP7 promoter activation occurs by the co-recruitment of HOXA13, mixed-lineage leukemia 1 lysine N-methyltransferase, WD repeat-containing protein 5. What happens completes the epigenetic alteration of the trimethylation of lysine 4 on histone H3 in tumor cells. On the other hand, HOXA13 suppressed BMP7 expression in iPSLCs through the co-recruitment of HOXA13, enhancer of zeste homolog 2, Jumonji and AT rich interactive domain 2, and lncRNA HoxA transcript antisense RNA (HOTAIR) to regulate the BMP7 promoter. The results of research on biological role of these lncRNAs in cancer cells revealed the recruitment of HOXA13-HOTTIP and HOXA13-HOTAIR to various regions in the BMP7 promoter are essential for the destiny of tumor cells [54]. Octamer-binding protein 4 and Jun dimerization protein 2 could be reprogrammed as it can stop tumorigenesis through turning BMP7 off.
PVT1 and FOXM1
Plasmacytoma variant translocation 1 (PVT1) and forkhead box protein M1 (FOXM1) are located in 8q24.21 and 12p13.33, respectively [55, 56]. The lncRNA PVT1 is reported in the related studies to upregulate in GC tissues to a large extent when it is compared to normal tissues [57]. An overexpression of PVT1 can be associated to poor prognosis in GC patients. PVT1 could power GC cell proliferation and invasion. PVT1 has been notably demonstrated to be directly bound to FOXM1 protein, which can increase post-translationally FOXM1. What is more is that PVT1 functions as a FOXM1-responsive lncRNA. Therefore, PVT1 functions as oncogenic factor in a FOXM1-mediated mode. The findings indicated that PVT1 can increase the tumorigenesis by cooperating with FOXM1 [55, 56]. Taken together, PVT1 can be considered as an appropriate predictor for GC and is suggested to be more investigated as a novel target in cancer therapy.
BC016000 and AF116637
LncRNAs BC016000 and AF116637 have been recognized as potential epigenetic effectors in carcinogenesis. Dysregulation of lncRNAs could be directly observed in the patients with GC in their body tissues. HDAC1 also plays a vital role in GC cell growth, somewhat by promoting the lncRNAs BC01600 and AF116637 expression [58]. Additionally, literature in the field has shown that BC01600 and AF116637 are modulated by HDAC1, as it has been proved by a lower expression of the lncRNAs in HDAC1knockout cancer cells. Functional studies offered enough indication on the significance of HDAC1 in the prediction of SCM1, and the correlation between HDAC1 and the transcriptional regulation of lncRNAs. Also, related research has shown that HDAC1 can be a hopeful tool for GC therapy [58].
SNHG17
LncRNA named small nucleolar RNA host gene 17 (SNHG17) is mapped on chromosome 20q11.23 [59]. Upregulation of SNHG17 in GC tissues and cells is meaningfully associated with tumor stage and metastasis [60]. The expression of plasma SNHG17 has been reported in the research to be upregulated in GC patients in comparison with their healthy control group counterparts. Functional studies on the SNHG17 showed that SNHG17 could increase GC cell growth and invasion, and inhibit apoptosis. Related studies showed that SNHG17 could be contributed to polycomb repressive complex 2. This was a necessary mediator for regulation of cyclin-dependent protein kinase inhibitors such as p15 and p57 [59]. Therefore, it can be stated that lncRNA is highly associated with the expression of cancer cell cycle and growth. Moreover, it has been indicated that SNHG17 functions as an oncogene through the activation of the enhancer of zeste homolog 2. These lines of research could bring about a novel approach toward SNHG17 which can act in GC tumorigenesis, and it might be used for as a probable biomarker for patients suffering from GC [39].
TUG1
Taurine-upregulated gene 1 (TUG1), is a lncRNA with 7.6 kb in length, is located in chromosome 22q12. Studies on the issue have revealed that upregulation of TUG1 significantly associated with advanced stages of GC [50]. Even more investigations have shown that reduction of TUG1 inhibited GC proliferation. Studies completed mechanistically have shown that TUG1 plays an outstanding function in G0/G1 arrest. TUG1 can be contributed to PRC2, and this association is essential prerequisite for epigenetic suppression of cyclin-dependent checkpoints. Therefore, it is associated with the GC cell growth regulation [50]. Therefore, it can be stated that TUG1, which is a cell proliferation regulator, might be served as a potential prognostic and therapeutics tool in case of GC.
MT1JP
LncRNA metallothionein 1 J, pseudogene (MT1JP) is placed on chromosome 16 [61]. Studies that reveal the function of such genes have shown that lncRNA MT1JP imposes a direction evident inhibition condition for proliferation, migration and invasion. Using molecular cell biology studies, it was revealed that lncRNA MT1JP has the potential to regulate FBXW7 in the process of GC development. lncRNA MT1JP also functions as a tumor suppressor and can regulate apoptosis and cell cycle in cancer cells through an epigenetic-regulated manner. The latest regulatory axis has been indicated to be induced by RNA-binding proteins Masashi 2 (MSI2) and TIA1-related protein (TIAR), and Ras-related protein Rap-1b (RAP1B) [62]. Not only the results highlight the function of lncRNA MT1JP in tumorigenesis of GC, but the findings also provide significant insights into further studying of genetic regulation mechanism in GC [61].
DANCR
LncRNA differentiation antagonizing non-protein coding RNA (DANCR) is mapped on 4q12.5 [63]. The biological function of DANCR have been extensively evaluated in GC cell phenotype [64]. The study documented that the knockdown of DANCR decrease the cell growth of GC cells although the overrepresentation of DANCR turned out to come up with opposite results. Other related studies have revealed that lncRNA-LET was a target of DANCR. By the same token, high DANCR and low lncRNA-LET were reported to be corresponding with distance metastasis and tumor stage. DANCR that is linked to EZH2 and HDAC3 in order to inhibit lncRNA-LET functioning can later effect GC migration and invasion. All in all, such findings can be an indicator of an imperative function for DANCR–lncRNA-LET in cellular function of GC, and thereafter suggest a mechanism for the regulation of lncRNA-LET [63].
EZH2
Recent functional studies have confirmed that lncRNA EZH2 could alter the promoter region of PCAT18 gene and increase methylation modification of H3K27. PCAT18 is known as a growth regulator that regulate cell proliferation in GC [65]. More molecular studies showed that PCAT18 modulates p16 by co-operating with miR-570a-3p. Together, the epigenetic regulation of histone modification of PCAT18 has a potential role GC tumorigenesis. Hence, EZH2- epigenetic modulation of lncRNA EZH2 might be indicated as an approach in GC therapy cooperating with miRNAs [65].
HOTAIR
HOX transcript antisense intergenic RNA (HOTAIR), has also been found to epigenetically affect GC tumorigenesis in several malignancies. HOTAIR could promote methylation of tumor suppressor gene pcdh10 by transcriptional regulation of DNMT1. Functional studies confirmed that HOTAIR might bind to miR-148b and DNMT1 and by inducing PCDH10 methylation promote GC carcinogenesis [66].
Oral squamous cell carcinoma (OSCC)
HOTAIR
LncRNA HOX transcript antisense intergenic RNA (HOTAIR), a lncRNA with 2.2 kbs, is located on 12q13.13 [67]. LncRNA HOTAIR functions as a distinctive structure to connect with a histone modification complexes. This lncRNA can cause a chromatin state by link histone H3K27 methylation and H3K4 demethylation to increase the tumorigenesis [68]. The HOTAIR overexpression is correlated to poor survival in numerous malignancies. The research has indicated that HOTAIR overexpression can increase OSCC in comparison with healthy tissues. HOTAIR has also shown to be correlated with metastasis in the OSCC patients with advanced stage and low histological differentiation. Therefore, the lncRNA is suggested to be more investigated for potential applications. Inhibition of HOTAIR expression by siRNA in OSCC cells could result in a low cell proliferation and growth, and it can enhance cell invasion, and thereby cause apoptosis. In this regard, the E-cadherin expression levels might be regulated through binding EZH2 and H3K27me3 with the related promoter. Biological and clinical analyses have led to the finding that HOTAIR may be one of important targets in metastatic form of OSCC [67].
CASC9
LncRNA cancer susceptibility candidate 9 (CASC9) is located on chromosome 8q21.11. The lncRNA have four transcript variants named CASC9-201, CASC9-202, CASC9-203, and CASC9-204. All various transcript variants have been revealed to involve in tumorigenesis. It has been demonstrated that lnc-CASC9 contributes to CRC tumorigenesis by targeting miR-193a and ERBB2. In this regard, CASC9 similarly may act as oncogene through the phosphorylation of SMAD3 and activation of TGF-
Pancreatic cancer (PC)
MEG3
LncRNA maternally expressed gene 3 (MEG3), a lncRNA with 1.6 kbs, is mapped on 14q32.2 [70]. MEG3 upregulation has shown to reduce the regulation of c-Met expression [71]. Mechanistically surveys such as RNA sequencing (ChIRP-Seq) revealed that Meg3 is able to bind to distinctive areas in the c-Met gene. When MEG3 does not exist, these c-Met regions have the potential to show a typical enhancer-signature histone modification. In addition to, MEG3 may act as an enhancer of zeste homolog 2 (EZH2) function, and it acts as a principal constituent of polycomb repressive complex 2 (PRC2), inhibiting c-Met. The related findings have established epigenetic mechanisms which involve in MEG3 control of c-Met. It has been reported that Meg3 exert a tumor-suppressor activity in tumor cells. This activity is due to the fact that the high expression of Meg3 in MIN6 cells can cause a suppression of cell growth and proliferation. Upregulation of MEG3 in MIN6 insulinoma causes a low level of regulation of the proto-oncogene c-Met. This lncRNA thus was established to significantly reduce cell invasion. It is interesting to say that MEG3 and c-MET expression can be changed in insulinomas using hypermethylation at the MEG3 promoter CRE-site which may simultaneously take place with decreased MEG3 expression. Moreover, in the tumor cells, DNA-demethylating medication can cause a decrease in cell proliferation and thus activate MEG3 expression [70]. In total, inactivation of c-MET via the epigenetic activation of lncRNA MEG3 could be considered as therapeutic tool for the malignancy.
MEG8
LncRNA (MEG8) is mapped on DLK1-DIO3 and 14q32.3 region [72]. An extensive study examined the biological roles of lncRNA-MEG8 in mediating EMT [73]. MEG8 lncRNA can be prompted by TGF-
HOTAIR
Epigenetic regulation has turned out to be important in tumorigenesis. Today, many studies have focused on this point and indicate that HOTAIR is expressed extensively in PC [74]. Nonetheless, the role of HOTAIR in the chemoresistance is still a controversial issue. TRAIL-resistant tumor cells have reported to have an overexpression of HOTAIR. In the same way, an overexpression of HOTAIR may inhibit the TRAIL receptor death receptor 5 (DR5). Related studies confirmed that HOTAIR modulate DR5 expression by the epigenetic regulator EZH2. It has also been confirmed that EZH2 regulates histone H3 lysine 27 trimethylation. Accordingly, the present discussion can add to the validity of the point that the upregulation of HOTAIR enhance the chemoresistance to TRAIL-mediated apoptosis through epigenetic modification of DR5 [74]. Hence, these findings support the view that modulating HOTAIR may signify an approach to control TRAIL-mediated chemoresistance.
Hepatocellular carcinoma (HCC)
ANRIL
An antisense noncoding RNA in the INK4 locus (ANRIL), with a length of 3834 bp, is located on 9p21.3 [75]. ANIL was revealed to regulate HCC tissues at a high level, and the higher expression of ANRIL could be an indicator of an important association with the clinical tumor stage. Furthermore, loss-of-function studies revealed that ANRIL could promote cell growth and migration in HCC. It has been explored that ANRIL suppresses Kruppel-like factor 2 (KLF2) epigenetically in cancer cells by binding PRC2. The findings can imply that lncRNA ANRIL can act as an indicator for cancer diagnosis and therapy [75].
MEG3
Research has shown that MEG3 expression can decrease to a large extent in human HCC cell lines when compared to healthy hepatocytes [76]. Molecular studies indicate a high expression of MEG3 in healthy liver cells with a low expression of oncogenic factors. The expression of MEG3 at a high level in HCC cells remarkably decrease cell proliferation and increase apoptosis. MEG3 promoter hypermethylation is an indicator of an inhibition to methylation with either 5-Aza-2-Deoxycytidine, or siRNA to DNMT 1 and 3b in liver cancer. Moreover, GTL2, the murine homolog of MEG3, decreases in liver cells compared to healthy control group. The findings of this study indicate that methylation-mediated regulation of the lncRNA MEG3 by miR-29a can play a major role in HCC growth [76]. These findings revealed an interaction between miRNAs and lncRNAs in the gene expression mediated by epigenetic regulation.
HULC/CURD
LncRNA highly up-regulated in liver cancer (HULC) is mapped on Chr19. Also, upregulation of long noncoding RNA cancer upregulated drug resistant (CUDR) induces carcinogenesis[77]. Research has shown that lncRNA CUDR can promote differentiation of the embryonic stem cells (ESC) through decreasing trimethylation on histone H3 twenty-seventh lysine (H3K27me3) [78]. To take it another way, one can notice that overexpression of CUDR can cause hepatocyte-like cells carcinogenesis. CUDR can upregulate HULC and
Dreh
LncRNA down-regulated expression by HBx (Dreh) has been placed on Chr17 [79]. The dysregulation of lncRNAs affected by HBx has been found. The analysis of the related data show that HBx transgenic mice display a distinctive pattern of lncRNAs in comparison with wild type mice [80]. LncRNA Dreh has the potential capacity to decrease growth, acting as a tumor suppressor. LncRNA-Dreh also has been confirmed to modification the cytoskeleton structure to decrease tumor invasion. The decreased expression of this lncRNA is significantly related to a poor survival of HCC patients. The observations exhibited a track of lncRNA-Dreh in tumor suppression in HCC tumorigenesis. Hence, it can provide a better insight into the significance of the lncRNAs by HBx. The results suggest therapeutic potentials of lncRNAs for the molecular therapy of HBV-related HCC [79].
DQ786243 (LncDQ)
LncRNA DQ786243, also known as DQ786243, is located on chromosome 8q24 [81, 82]. Relevant studies have shown LncDQ can upregulate in HCC. This can be interrelated with clinicopathological features and weak survival of cancer patients [83]. LncDQ expression has been revealed to be served as a prognostic factor for HCC. In this regard, knockdown of LncDQ can decrease cell migration. LncDQ is able to regulate the epithelial-mesenchymal transition through interaction with EZH2, in order to promote the expression of E-cadherin in HCC cells. Totally, the findings designate that LncDQ is a crucial factor in HCC progression, and might be used as biomarker for diagnosis of HCC [83].
MALAT1
Metastasis associated lung adenocarcinoma transcript 1 (MALAT1) or NEAT2 (noncoding nuclear-enriched abundant transcript 2) and is mapped on 11q13.1 [84]. LncRNA MALAT has been demonstrated to upregulate in HCC. Reduced MALAT1 has been confirmed to diminish cell cycle, cell growth and invasion of liver tumors [85]. Moreover, MALAT1 could mechanistically recruit Brahma-related gene 1 (BRG1), that is a catalytic subunit of chromatin remodelling complex switching/sucrose non-fermentable (SWI/SNF), to the promoter region of IL-6 and CXCL8. Therefore, MALAT1 can assist NF-
AY
LncRNA AY927503 (AY), is a molecule that associates with metastasis and is upregulated in HCC. Research has revealed that it is correlated with metastasis and poor survival of HCC patients [86]. AY also may enhance HCC cell migration and metastasis. In this regard, knockdown of integrin
SNHG15
Small nucleolar RNA host gene 15 (SNHG15), is mapped on 7p13 [87]. There are several evidences regarding the molecular functions of SNHG15 in some cancer types, including gastric, hepatocellular, pancreatic, colorectal and lung cancers [88]. SNHG15 is considered as an influential factor in the regulation of cell growth and invasion through numerous mechanisms. Additionally, the aberrant expression of SNHG15 has been related with some characteristics of cancer patients. Accordingly, SNHG15 may be used as a potential indicator for cancer management [88].
Crosstalk between lncRNAs and epigenetic mediators/events
The epigenetic mechanisms that are related to DNA methylation, histone modifications, and non-coding RNA dysregulation may be regarded as main features in discriminating cancers [89]. In this regard, the insights into methylation modification of chromatin elements and the emergent roles of lncRNA can help in achieving an improved insight into molecular mechanisms underlying cancers. In more recent years, methods like lncRNA profiling and epigenetic markers identification have been arguably considered to determine tumor-associated lncRNAs that are epigenetically deregulated by chromosome modifications. Therefore, an abnormal methylation changes in certain lncRNA genes could be detected as they can correlate with unfavourable consequences in a number of gastrointestinal malignancies [8]. Furthermore, the extent to which lncRNAs might mediate DNA methylation and histone alteration to modulate crucial signalling pathways in carcinogenesis [90]. The findings can imply that methylation changes in chromatin effectors mediate numerous efficient crosstalk with lncRNAs. Based on our findings, the potential roles of the lncRNA-mediated epigenetic processes and the outlining molecular mechanisms in GI cancers are more than indicated.
Conclusion and future perspectives
It is concluded that epigenetic alterations may regulate GI tumorigenesis and also, they are beneficial biomarkers in case of diagnosing and predicting of GI cancers, including colorectal, gastric, oral, liver, pancreatic and oesophageal cancers. Considering progressively application of genome-wide epigenetic analyses, it is expected that epigenetic markers are introduced in clinical settings of GI cancers such as diagnosis, prognosis and predicting response to therapies. In this regard, epigenetic-related lncRNAs in GI cancers can be reliable and promising targets and would be given more attention in the future. More investigations to improve our understanding about the molecular mechanisms of epigenetic regulation mediated by long non-coding RNAs mayprovide insights into the development of new GI cancer therapies.
Abbreviations
ANRIL: Antisense non coding RNA in the INK4 locus
ARHGAP5-AS1: ARHGAP5 Antisense RNA 1 (Head To Head)
BCLC: Barcelona Clinic Liver Cancer
BMP7: Bone morphogenetic protein 7
BRG1: Brahma-related gene 1
CARs: Chromatin associated RNAs
ChIRP-Seq: Chromatin isolation by RNA purification and sequencing
CKIs: Cyclin-dependent protein kinase inhibitors
CRNDE: Colorectal neoplasia differentially expressed
DACOR1: DNMT1-associated colon cancer repressed lncRNA 1
DANCR: Differentiation antagonizing nonprotein cod- ing RNA
Dreh: Down-regulated expression by HBx
EZR-AS: Ezrin antisense RNA 1
FBXW7: F-box and WD repeat domain- containing 7
FOXM1: Fork head box M1
GATA6-AS: GATA6 antisense RNA 1
GSE: Gene signature in the training set
Gtl2: Gene trap locus 2
H1FX: Histone 1FX
HOTAIR: HOX Transcript Antisense Intergenic RNA
HOTAIRM1: HOXA transcript antisense RNA, myel oid-specific 1
HOXD-AS1: HOXD cluster antisense RNA 1
HULC: Highly upregulated in Liver cancer
KCNQ1OT1: KCNQ1 Opposite Strand/Antisense Tran script 1
KLF2: Kruppel-like factor 2
LALC: Associated lncRNA in CRC (lnc-LALC)
LOXL2: Lysyl Oxidase Like 2
LSD 1: Lysine-specific demethylase 1
LUCAT1: Lung cancer–associated transcript 1
MALAT1: Metastasis associated lung adenocarcinoma transcript 1
MEG3: Maternally expressed 3
METTL3: methyltransferase-like 3
MIA: Mitochondrial intermembrane space import and assembly
MLK7-AS1: MAP3K20 antisense RNA 1
MORT: Mortal obligate RNA transcript
MT1JP: Metallothionein 1 J, pseudogene
NAP1L2: Nucleosome Assembly Protein 1 Like 2
NEAT2: Noncoding nuclear-enriched abundant tran script 2
PHB2: Prohibitin-2
PIM2: Proviral Integrations of Moloney virus
PNET: Pancreatic neuroendocrine tumors
PRC2: Polycomb Repressive Complex 2
PVT1: Plasmacytoma variant translocation 1
PXR: Pregnane X receptor
RAB4B: Ras-Related GTP-Binding Protein 4b
RING: Really interesting new gene
SH3PXD2A: SH3 and PX Domains 2A
SMYD3: SET and MYND Domain Containing 3
SNAI1: Snail, zinc finger transcription factor
SNHG14: Small nucleolar RNA host gene 14
SNHG15: Small nucleolar RNA host gene 15
SOGA1: Suppressor of glucose autophagy-associated
SQSTM1: Sequestosome 1
TCGA: The Cancer Genome Atlas
TFOs: Triplex-forming oligonucleotides
TUG1: Taurine-upregulated gene 1
UCA1: Urothelial cancer associated 1
UHRF1: Ubiquitin-like with PHD and RING finger domains 1
Ethics approval and consent to participate
Not applicable.
Availability of data and material
Not applicable.
Competing interests
All authors declare that they have no competing interests.
Funding
This work was supported by Grant number 13162 from Iran University of Medical Sciences.
Author contribution
Conception: A Akbari, A Talebi.
Preparation of the manuscript: A Akbari, A Talebi, S Abbasi, M Sedeghat.
Revision for important intellectual content: S.A.P Tabaeian, A.H Faghihi Kashani, N Borumandnia.
Supervision: A Akbari, A Talebi, Z Shokati Eshkiki, S.A.P Tabaeian, A.H Faghihi Kashani.
Footnotes
Acknowledgments
This work was supported by Gran number 13162 from Iran University of Medical Sciences.
