Abstract
BACKGROUND:
The clinicopathological significance of spatial tumor-infiltrating lymphocytes (TILs) subpopulations is not well studied due to lack of high-throughput scalable methodology for studies with large human sample sizes.
OBJECTIVE:
Establishing a cyclic fluorescent multiplex immunohistochemistry (mIHC/IF) method coupled with computer-assisted high-throughput quantitative analysis to evaluate associations of six TIL markers (CD3, CD8, CD20, CD56, FOXP3, and PD-L1) with clinicopathological factors of breast cancer.
METHODS:
Our 5-plex mIHC/IF staining was shown to be reliable and highly sensitive for labeling three biomarkers per tissue section. Through repetitive cycles of 5-plex mIHC/IF staining, more than 12 biomarkers could be detected per single tissue section. Using open-source software CellProfiler, the measurement pipelines were successfully developed for high-throughput multiplex evaluation of intratumoral and stromal TILs.
RESULTS:
In analyses of 188 breast cancer samples from the Nashville Breast Health Study, high-grade tumors showed significantly increased intratumoral CD3
CONCLUSIONS:
The high- and low-grade breast cancers exhibit differential immune responses which may have clinical significance. The multiplexed imaging quantification strategies established in this study are reliable, cost-efficient and applicable in regular laboratory settings for high-throughput tissue biomarker studies, especially retrospective and population-based studies using archived paraffin tissues.
Keywords
Abbreviations
Introduction
Tumor-infiltrating lymphocytes (TILs), important components of the tumor microenvironment, represent a local immune response against cancer. Recent studies indicate that TILs have a predictive value for immunotherapy and prognostic roles for various types of neoplasms, including triple-negative and HER2-positive breast cancers [1, 2]. Common clinical application of TILs assessment is based on pathologists’ visual scoring of hematoxylin and eosin (H&E) stained tissue slides, recommended by the International Immuno-Oncology Biomarker Working Group as biomarkers in routine histopathological reporting for breast cancer patients [3, 4]. The standardized TILs assessment guidelines are of value to researchers and practicing pathologists, owing to their ease of application in clinical trials and in daily histopathology practice, with improved consistency. However, some inherent limitations cannot be fully addressed through standardization and training. For example, H&E-based visual assessment cannot evaluate intratumoral TILs (iTILs), which have cell-to-cell contact with tumor cells; cannot differentiate various TIL subpopulations, which may differentially impact tumor progression and response to treatment [5, 6]; and have some degree of inter-reader variability so that it is considered insufficiently reproducible for clinical application [7, 8]. Conventional immunohistochemistry (IHC) as a gold standard for identifying tissue biomarkers is widely used to examine TIL subsets and immune cell markers [9, 10, 11, 12]. However, IHC only labels one or a limited number of markers per tissue section; therefore, it is difficult to apply IHC for tumor immune profiling, colocalized biomarker examination and automated image quantitative assessment [13]. Due to the lack of high-throughput scalable methodology for studies with large sample sizes, the clinicopathological significance of spatial TIL subpopulations in breast cancer molecular subtypes remains to be elucidated.
Multiplex immunohistochemistry (mIHC) technologies have emerged as powerful investigative tools to circumvent the limitations of H&E-based and conventional IHC methods. mIHC technologies allow the simultaneous detection of multiple markers on a single section to provide objective quantitative data for greater insight into disease heterogeneity, molecular profiling and biological mechanisms driving diseases [13]. Studies have shown high concordance of mIHC with conventional IHC [14] and significantly higher diagnostic accuracy in predicting clinical responses to anti-PD-1/PD-L1 therapy [15]. Moreover, mIHC conserves limited tissues when study samples are of low availability, such as rare donors, small biopsy samples or limited tissue sections collected in large population-based studies. In recent years, various mIHC techniques have been developed, such as chromogenic mIHC [16], tyramide signal amplification (TSA)-based mIHC [17, 18, 19], metal-based mIHC [20, 21], and DNA barcoding-based mIHC [22]. Each mIHC platform has some advantages, but their utilization requires certain constraints, such as time-consuming, low sensitivity, high cost, requirement of dedicated instrumentation, or lack of high-throughput image analysis [13], making it impractical for most lab settings and for large population studies.
The limitations described above motivated us to conduct this study, with aims to establish a reliable and sensitive high-plex fluorescent mIHC method, to develop an automated quantitative imaging method using open-source imaging software for high-throughput evaluation of spatial distribution of immune cell markers, and to apply the quantitative mIHC/IF approach in a pilot study to evaluate associations of TILs with clinicopathological parameters of breast cancer patients. The established mIHC/IF workflow provides a robust, cost-efficient modality, which can be conducted in regular IHC settings for deep phenotyping of the tumor and its microenvironment, especially applicable for retrospective or population-based biomarker studies using formalin-fixed paraffin-embedded (FFPE) archived tissues.
Materials and methods
Specimens
The human breast cancer samples evaluated in this study were from the Nashville Breast Health Study (NBHS). The NBHS is a case-control study conducted in Nashville, Tennessee, which included 2,694 breast cancer patients diagnosed between February 1, 2001 and December 31, 2011, identified through the Tennessee State Cancer Registry and five major hospitals providing medical care for breast cancer patients. The recruitment of study participants has been described elsewhere [23, 24]. Approval for this study was obtained from institutional review boards of Vanderbilt University Medical Center and other participating institutions, and all participants provided informed consent prior to enrollment in this study. In current study, we randomly selected four tissue microarray (TMA) blocks from the NBHS tissue repository, which included tumor tissue samples from 208 patients. From each donor block, representative cancer areas of three spots in the peripheral, central and middle regions were punched with a 1 mm needle and transferred into a recipient TMA block using a semi-automated tissue microarrayer. Each TMA block contained 170 tissue cores (13X13), including 52 cases of breast cancer and 14 internal control cores (eight normal human tissues of liver, spleen, kidney, lung, lymph node, prostate, brain and placenta and six breast normal and cancer cell lines: MCF7, MCF10A, MDA-MB-231, MDA-MB-361, MDA-MB-435S, and MDA-MB-453). TMA slides were sectioned at 5
Fluorescent multiplex immunohistochemistry (mIHC/IF)
Two panels of 5-plex mIHC/IF staining protocols were developed to evaluate both TIL subpopulations and their spatial localizations in breast tumor tissues. The first panel included CD3 (for total T cells), CD8 (for cytotoxic T lymphocytes, i.e., CTLs), and forkhead box protein 3 (FOXP3, for regulatory T cells, i.e., Tregs). The second panel included CD20 (for B cells), CD56 (for NK cells), and the immune check-point molecule programmed death ligand 1 (PD-L1). Additionally, DAPI (for nuclei counterstain) and pan-cytokeratin (panCK, for tissue segmentation) were included in both panels. The primary antibodies of immune cell markers used in this study are commercially available and well validated [18, 19, 25, 26], i.e., rabbit anti-CD3 (Clone# SP7, #M3070, 1:1000 dilution. Spring Biosciences, CA, USA), rabbit anti-CD8 (#M5394, 1:500 dilution, Spring Biosciences), rabbit anti-PD-L1 (#13684S, 1:5000 dilution. Cell Signaling Technology, MA, USA), rabbit anti-CD56 (#156R-95, 1:3200 dilution. Cell Marque, CA, USA), mouse anti-CD20 (#GA604, 1:10 dilution. Agilent, CA, USA), and mouse anti-FOXP3 (#AB20034, 1:1000 dilution. Abcam, Cambridge, UK). The mIHC methods were established using tyramide signal amplification (TSA) technology, which allows for signal amplification of low abundance targets and facilitates heat-mediated removal of primary/secondary antibodies without disruption of TSA-fluorescence signals [17]. To optimize mIHC/IF staining protocols, we constructed a control TMA block with 13 human tumor and normal tissues, including breast, lung and colorectal cancers, and one each of spleen, prostate and placenta tissues. The staining methods are summarized below.
After deparaffinization and antigen retrieval using a high-pressure cooker with citrate buffer (pH 6.0), followed by blocking steps with 3% hydrogen peroxide solution and 5% normal goat serum, slides were subsequently incubated with the first primary antibody (mouse anti-FOXP3 or mouse anti-CD20) for 1 hour at room temperature (RT), Polymer-HRP goat anti-mouse secondary antibody (DAKO EnVision kits, Agilent, CA, USA) for 30 minutes at RT, and TSA-Cy5 solution (PerkinElmer Opal 3-plex Kit, NEL791001KT, 1:100 dilution, PerkinElmer, MA, USA) for 5 minutes at RT. Next, slides were incubated with the second primary antibody (rabbit anti-CD3 or rabbit anti-CD56) at 4
Fluorescence quenching and cyclic mIHC/IF
Conventional immunofluorescence imaging is typically limited to 4–6 channels, above which, crosstalk between fluorophores becomes a problem for regular laboratory settings. To increase the multiplicity of immunofluorescence imaging, various fluorescence quenching methods have been developed and reported as cyclic immunofluorescence technology [27, 28, 29]. In this study, a chemical bleaching method, based on a U.S. patent (No: US 7,741,046 B2) [30], was established using TSA-cyanine dyes in our mIHC/IF protocols. Stock solutions were 1 M sodium bicarbonate (NaHCO
Cyclic mIHC/IF staining of selected TIL markers. A. The cyclic 5-plex mIHC/IF detection scheme. B. Example images of sample breast cancers stained by 5-plex mIHC/IF of the first panel (a, CD3, CD8 and FOXP3) and second panel (b, CD20, CD56 and PD-L1). Total T cells (CD3
Measurement pipeline for spatial quantitative analysis of CD3
The mIHC stained slides were imaged using an Olympus IX-81 fluorescence fully motorized and automated multispectral slide analysis system, with filter sets specific for DAPI, GFP, CY3, CY5, and CY7. Three TMA cores per case were imaged at X 100 amplification, and exposure times were determined for each antibody stain (typically, DAPI
For the first panel of CD3, CD8 and FOXP3 stains, the five grayscale images were input into a single integrated pipeline to quantify three biomarkers, summarized in Fig. 2. Total T cells, CTLs and Tregs were defined as CD3
Statistical analysis
Mean and standard deviation were applied to describe distributions for continuous variables and percentages of each category for categorized variables. Correlations between immune cell markers in human breast cancer tissue were examined using Spearman correlation coefficients. Cox proportional hazards models were employed to evaluate associations of selected biomarkers with clinicopathologic characteristics and overall survival (OS) of breast cancer. Tests for trends were conducted by entering the order of categorized variables as continuous variables in regression models. Multivariable Cox models included adjustments for tumor-node-metastasis (TNM) stage, tumor grade, ER and PR status, and race. All analyses were performed using Statistical Analysis Software (SAS Office Analytics, version 14.3; SAS Institute Inc., Cary, NC), with a two-sided significance threshold of
Results
Establishment of cyclic mIHC/IF and automated image quantification
The basic steps of TSA-based cyclic mIHC/IF stain are illustrated in Fig. 1A. The 5-plex staining protocols for two panels of immune cell markers were successfully established, with excellent signal-to-noise ratio (Fig. 1B). Staining protocols were optimized and validated by comparing conventional standard IHC with single fluorescent staining for each biomarker to produce specific, non-interfering, and balanced labeling for three immune cell markers of interest per slide. The sensitivity of TSA-based mIHC/IF is generally 5 to 10 times higher than standard single stains for CD3, CD8, CD20, and FOXP3, but 32 and 100 times higher for CD56 and PD-L1, respectively (Supplementary Fig. 2). After one cycle of sequential 5-plex staining, we tested chemical bleaching methods to inactivate fluorescence signals in order to perform the next cycle of 5-plex staining using the same slide. We found that TSA-FITC signal could not be fully inactivated by Lin’s (activating Alex fluorophores) [28], Gerdes’s, or Riordan’s (activating cyanine fluorophores) methods [27, 29]. Therefore, we replaced TSA-FITC with CF488*TSA (Biotium), a cyanine dye equivalent to TSA-FITC, and used a simple inactivation method based on a U.S. patent [30]; all cyanine fluorophores (Cy3, Cy5, Cy7, and CF488) could be totally inactivated in 3 hours. A very weak DAPI signal might remain, which did not affect the next biomarker staining cycle. We repeated four cycles of 5-plex mIHC stains and obtained the same staining results for all biomarkers (Fig. 1C).
After image acquisition of the stained slides, positive cells within the intratumoral, stromal and total areas of breast cancer tissues were quantified by established measurement pipelines using CellProfiler. Validation sets of 50 sample images were semi-quantitatively scored by an experienced study pathologist to compare quantitative data scored using the measurement pipelines. We found that high-quality images, defined as having contrast, bright details with a dark background, and being properly focused (Fig. 1B), could be correctly quantified by the default setting of the “IdentifyPrimaryObjects” module. However, for images with unfavorable signal-to-noise ratios, the advanced setting of the “IdentifyPrimaryObjects” module, with careful threshold adjustment, was needed to correctly differentiate true positive signals from nonspecific backgrounds for all training sets of images. In this study, our established pipelines using training sets were verified until the pathologist’s visual score and automated quantitative data reached a good consistency (Supplementary Fig. 3). The verified pipelines enabled more than 1000 images to be automatically analyzed at one time. Biomarker assessment was conducted for three TMA cores per case, then averaged for further statistical analysis. Based on our experience, a workflow of high-plex, high-throughput mIHC/IF imaging quantification workflow was established and illustrated in Fig. 3.
The workflow of high-plex, high-throughput spatial quantitative analysis of tissue TIL biomarkers.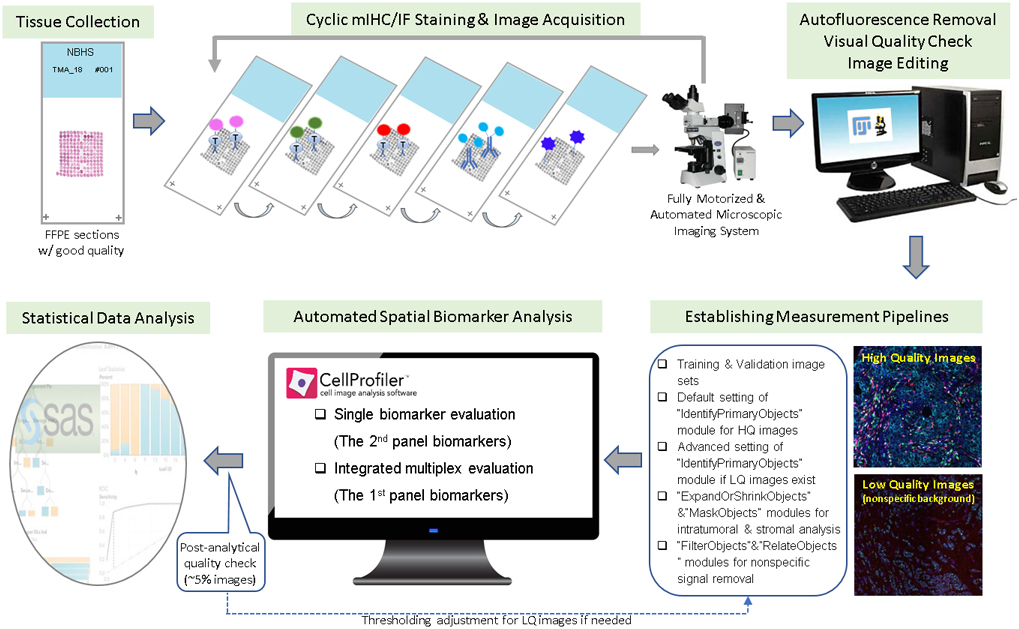
Among four TMAs, a total of 188 (89.4%) cases had evaluable tissue cores and were included in our analysis (Table 1). The mean age at diagnosis was 54 years. The majority (77.7%) were diagnosed with early-stage (0-II) disease; the remaining had advanced-stage (III-VI) disease, with a 5.5 times higher risk of death. Most patients had either grade II (33.0%) or grade III (48.4%) disease. Most tumors were ER positive (71.8%), PR positive (53.2%), and HER2 negative (75.0%). Approximately 20% of patients had triple-negative breast cancer. A total of 23 patients (12.2%) died during an average of 5.45 years of follow-up.
Characteristics of breast cancer patients in current study from the NBHS
Characteristics of breast cancer patients in current study from the NBHS
Correlation between immune cell markers in human breast cancer tissue
Correlation between immune cell markers in human breast cancer tissue
Table 2 presents correlations between immune cell markers. The positive staining of CD8 and FOXP3 were significantly correlated with CD3 (
Associations of selected biomarkers, as well as CD8
Discussion
In this study, we established a cyclic mIHC/IF method for highly multiplexed biomarker detection of FFPE tissues and developed image analysis pipelines for high-throughput automated quantification of spatial distributions of immune cell markers in tumor stromal and intratumoral compartments of breast cancer. The analyses of pilot breast cancer samples showed that higher breast cancer grades (II and III) were significantly associated with higher density of intratumoral CD3
Of several alternatives of mIHC technology and platforms, we integrated TSA, an enzyme-tyramide mediated signal amplification technology, into our mIHC/IF protocol to capitalize on the advantages of TSA, such as a much higher sensitivity than standard IHC for detection of low-abundance biomarkers, a very favorable signal-to-noise ratio, which is especially important for studies using archived FFPE tissues, the feasibility of
Association between immune cell markers and clinicopathological parameters in breast cancer patients
Footnotes
Acknowledgments
Dr. Robert Coffey’s lab at VUMC kindly provided technical support for the establishment of 5-plex fluorescent mIHC and TMA image capturing. We also thank Ms. Nabela Hamm for tissue processing of the NBHS samples, Dr. Mary Shannon Byers, Ms. Rachel Mullen, and Ms. Kathleen Harmeyer for assistance in editing and submitting this manuscript.
Conflict of interest
All the authors declare that they have no conflict of interest.
