Abstract
BACKGROUND:
Carcinoembryonic antigen (CEA) is the only established serum biomarker for colorectal cancer (CRC). To facilitate therapy decisions and improve the overall survival of CRC patients, prognostic biomarkers are required.
OBJECTIVE:
We studied the prognostic value of five different cell free circulating DNA (fcDNA) fragments. The potential markers were ALU115, ALU247, LINE1-79, LINE1-300 and ND1-mt.
METHODS:
The copy numbers of the DNA fragments were measured in the peripheral blood serum of 268 CRC patients using qPCR, the results were compared to common and previously described markers.
RESULTS:
We found that ALU115 and ALU247 fcDNA levels correlate significantly with several clinicopathological parameters. An increased amount of ALU115 and ALU247 fcDNA fragments coincides with methylation of HPP1 (
CONCLUSIONS:
This study shows that an increased level of ALU fcDNA is an independent prognostic biomarker for advanced colorectal cancer disease.
Abbreviations
Introduction
Colorectal cancer (CRC) is the second most common cancer in Europe with almost 500,000 new cases in 2018 and is responsible for the second most cases of cancer-related deaths in men and women [13]. This high incidence, particularly in western Europe, may reflect changes in lifestyle such as an increased prevalence of obesity and great variability in diet and physical activity across all European countries [3]. Although great efforts have been undertaken to improve cancer treatment and new therapy options find their way into clinical practice, survival of patients is still poor, especially in metastatic cases [22].
CRC carcinogenesis is characterized by genetic and epigenetic changes, which can be attributed to different signaling pathways. Basically, development of colorectal cancer through mutation in or loss of the adenomatous polyposis coli gene (APC) follows the adenoma-carcinoma sequence. Subsequently, different pathways lead to cancer development. Amongst others, the most frequent pathway in 65–70% of the cases is the Chromosomal instability (CIN) – pathway, where mutations of P53 and KRAS activate different pathways in turn, such as Wnt, MYC, EGFR (epidermal growth factor receptor), TGFB (transforming growth factor beta) and Akt signaling [1, 37].
Currently, the most valuable prognostic tool for cancer patients is the UICC (Union for International Cancer Control) TNM (Tumor site and size, Lymph node involvement, Metastatic spread) staging classification [6]. Although it is the only valid marker in predicting patient outcome [4], the TNM system is surgically oriented and neither integrates tumor biology nor differentiates the metastatic stage [32]. For this reason, TNM system is not able to specify the individual risk of a given patient [38]. Biopsies taken from the tumor also cannot reflect tumor biology properly due to genetic heterogeneity of the tumor tissue [55]. Nomograms based on TNM classification, combined with different clinicopathological parameters such as age and race, were created to predict survival of CRC patients [30].
A prognostic marker for CRC, widely used in clinical practice, is carcinoembryonic antigen (CEA). CEA proved to be an independent prognostic marker in various studies and guidelines recommend measuring preoperative CEA serum levels to determine prognosis, keep patients under surveillance after curative surgery and monitor therapy in advanced diseases [11]. CEA levels vary between individuals. Therefore, CEA baseline levels must be established for each patient. Although CEA determination is also recommended in the metastatic disease [10], Wanebo et al. reported that there was no correlation between CEA level and survival in metastatic disease [56].
Cell-free DNA is circulating in the blood stream in different concentrations. Compared to normal individuals, the amount of cell free circulating DNA (fcDNA) is higher in the blood of cancer patients [2] and is also increased in metastasized disease compared to nonmetastatic patients [21, 28]. Tumor-related nucleic acids as potential biomarkers for malignancies have been intensively studied recently [16, 24]. Circulatory microRNAs are also the subject of current research and have even been demonstrated to distinguish between HCV associated fibrosis, cirrhosis, and hepatocellular carcinoma [15, 58]. Sequencing of plasma DNA revealed that fcDNA represents the complete tumor genome as it can be released from either the primary tumor, metastasis, or apoptotic circulating tumor cells. Apoptosis, necrosis and even active secretion have been discussed as possible sources of tumor DNA, but the mechanism of release of fcDNA is still incompletely understood [54].
fcDNA fragments can be measured using real-time quantitative polymerase chain reaction (qPCR), which offers a quick, cost-effective, and robust method for quantification of the fcDNA fragments of interest. Alternative approaches compared are microarray-based linear amplification or Illumina sequencing of fcDNA [14, 47].
Notably, epigenetic alterations proved to be a promising predictive and prognostic tool. We previously described the prognostic value of methylation of various genes in free circulating DNA, namely helicase-like transcription factor (HLTF) and hyperplastic polyposis 1 (HPP1) [40]. It could be demonstrated that HLTF/HPP1 methylation status corresponds with shorter survival and therefore turned out to be a prognostic marker in UICC stage IV.
In addition to DNA methylation, DNA integrity has already been discussed as diagnostic and prognostic marker for colorectal cancer [57] based on the idea that the length of nucleic acids released from cancer cells differs from the DNA of non-malignant apoptotic cells [33].
Non-LTR (long terminal repeats) retrotransposons, which include short interspersed elements (SINEs) like the ALU repetitive sequence and long interspersed elements (LINEs), represent almost one third of the whole genome [8]. ALU elements are characterized by having a cleavage site for the Arthrobacter luteus (ALU) restriction endonuclease [19]. SINEs are shorter than LINEs and contain about 300bp. These non-coding DNA elements have function in regulating transcription and gene expression [51].
European population is yet not well considered in studies concerning fcDNA as a biomarker for cancer. The prognostic and diagnostic value of ALU repeats as biomarkers has been shown for different types of cancer, such as breast cancer, non-small-cell lung cancer and colorectal cancer, and across different populations, such as American and Egyptian [12, 46, 52, 53].
LINE1 has lately been discussed not only as a biomarker but also therapeutic drug target as it seems to have a regulatory impact on gene expression in cancer cells using different pathways of carcinogenesis. Furthermore, promoter methylation, transcription, translation, and retro transposition are different activities LINE1 is said to perform in cancer cells [26, 43]. LINE1 as a prognostic and diagnostic biomarker has been investigated in different types of cancer such as colorectal cancer, gastric cancer, and breast cancer, in addition to the Japanese mostly in the American population [9, 25, 49].
Human cells contain about 200 mitochondria, which possess their own genomic material. Due to its molecular characteristics such as an inefficient DNA repair system, mitochondrial DNA (mtDNA) is more susceptible to oxidative damage than nuclear DNA. Thus, DNA mutations accumulate easily and lead to mitochondrial dysfunction, which is associated with tumorigenesis. Through activation of the Toll-like 9 receptor (TLR9) pathway, inflammatory processes are initiated, which provide a condition for cancer development [29, 36].
Furthermore, copy numbers of circulating mtDNA have been discussed as a biomarker for different types of cancer such as hepatocellular carcinoma in an Egyptian population [18], lung cancer in a Chinese population [7] or head and neck squamous cell cancer in Northeast India [27].
Using mtDNA with its simple structure and a higher copy number than nuclear DNA allows a sensitive detection and exact quantification even of very low concentrations of circulating DNA in serum and other body fluids. Thus, mtDNA is ideally suited as biomarker.
The aim of this study was to evaluate the prognostic value of cell free circulating DNA fragments in colorectal cancer patients with metastasized and non-metastasized disease. We selected five potential markers, namely ALU115, ALU247, LINE1-70, LINE1-300 and ND1-mt, which were able to discriminate between CRC patients with benign and malignant tumors and healthy individuals in blood samples of a British patient population [34] and compared the results with common tumor markers as CEA and other predictive tools, e.g. the HLTF and HPP1 methylation status [40].
Methods
Patients and serum samples
Male and female patients over the age of 18 years were recruited from the Medical Department 2, Ludwig-Maximilians-University Munich and were diagnosed by histopathological examination of biopsies taken during colonoscopy or sigmoidoscopy using the valid WHO-classification at that time. Blood serum samples from 268 patients were taken before any therapeutic interventions and processed blinded to patient data in the central laboratory using routine procedures. All subjects underwent medical treatment at the Ludwig-Maximilians-University of Munich. Exclusion criteria were pretreatment of the current cancer disease, clinical history of any other cancer and simultaneous tumors. Staging was based on postoperative pathology findings or, for inoperable patients, on diagnostic imaging, namely CT of thorax and abdomen.
All samples were centrifuged at 2,000 g at 4
DNA isolation
After thawing the serum aliquots at room temperature, genomic DNA was isolated from 100
Quantitative polymerase chain reaction
Two different fluorescence-based real-time qPCR assays were performed. The primer sequences for all markers are shown in Table S1[9, 29, 49, 53]. We confirmed in silico that these primers form neither unwanted secondary structures nor primer dimers using the software Primer Express v2.0 (Applied Biosystems). Focusing on the amplification of ALU115 and L1-79 DNA fragments, we determined an intraassay coefficient of variation (CoV) of 3.2% and the interassay CoV was 3.0% for ALU115. The corresponding values for L1-79 were 5.3% (intraassay CoV) and 2.8% (interassay CoV).
To quantify the ALU repeats, the following reaction mixture was used: 1
Real-time PCR was performed using the following conditions: 95
For each fcDNA of interest, a standard curve was generated using leucocyte DNA. A single standardised solution of leucocyte DNA was used as a standard curve reference in each qPCR run. Mean values across triplicates were used for further analysis. For each serum sample, the number of fcDNA copies was normalized to the sample volume.
Statistical analysis
Comparisons of marker values between different clinico-pathological groups were done using the Wilcoxon-Mann-Whitney test in case of two groups and the Jonckheere-Terpstra test in case of three or more groups.
Survival analysis was done with Kaplan-Meier survival curves with UICC stage 1 to 3 and UICC stage 4 being analyzed separately. For calculation of overall survival, the period between the date of diagnosis of the primary tumor and the date of death respectively end of follow-up was included. Hazard ratios (HRs) for univariate and multivariate analysis of overall survival were estimated using Cox’s regression model. All statistical analyses were performed with SAS 9.3 (SAS Institute, Cary, NC, USA).
Results
Correlation of DNA markers with clinicopathological parameters and DNA methylation status
Correlation between copy numbers/CEA concentration/methylation status and clinical features of our study population
Correlation between copy numbers/CEA concentration/methylation status and clinical features of our study population
Correlation of copy numbers with clinicopathologigal parameters.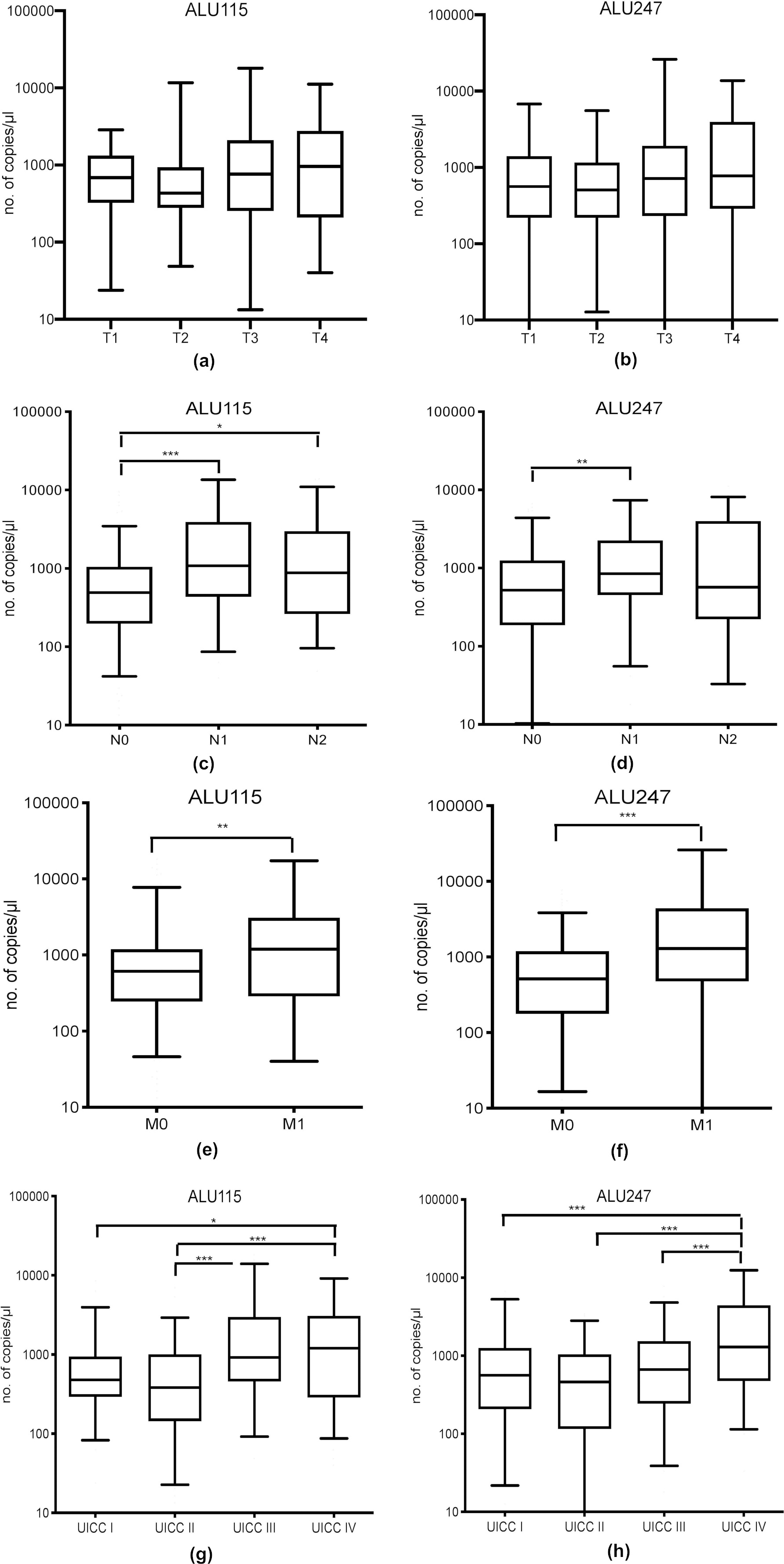
The characterization of our study population is shown in Table 1. The population consists of 142 female and 126 male patients with cancer in cecum, ascending, transverse and descending colon (46%), sigmoid colon (20%) and rectum (34%). 45% of the patients had positive regional lymph nodes. Distant metastases were present in 83 patients. UICC stage I tumors was 21%, UICC stage II 26%, UICC stage II 22% and UICC stage IV 31% of our patient population.
First, we quantified the copy numbers of each marker using qPCR. The median concentrations are shown in Table S2. Common clinicopathological parameters were analyzed for a relationship with the copy numbers of the candidate biomarkers (Table 1). The amount of ALU115 and ALU247 fcDNA showed a significant correlation with tumor size (
Overall survival in association with UICC stage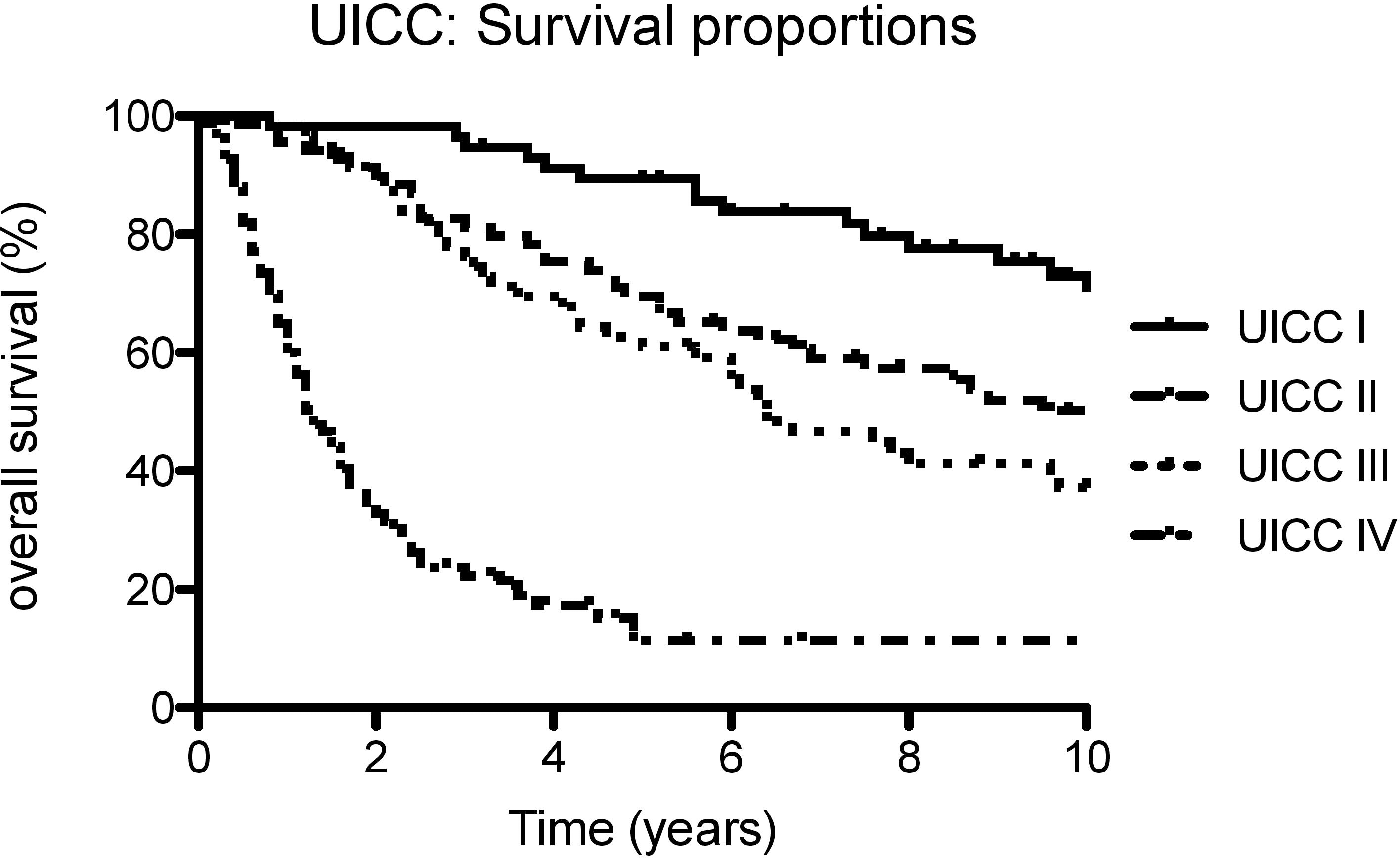
Besides the association with clinical parameters, we also analyzed the correlation of our markers with carcinoembryonic antigen (CEA) as a widely used tumor marker and the methylation status of HLTF and HPP1, which have already been demonstrated to be prognostic markers for CRC [40]. Carcinoembryonic antigen is more efficient in discriminating between good and poor prognosis when individualized cutoff values are used [23]. In this study, two different CEA levels served as cutoff values: The lower concentration (2.5 ng/ml) represents the 95
Next, we were interested in the prognostic value of raised copy numbers of ALU115, ALU247, L1-79, L1-300 and ND1-mt.
Survival proportions with combinations of ALU115 and HPP1.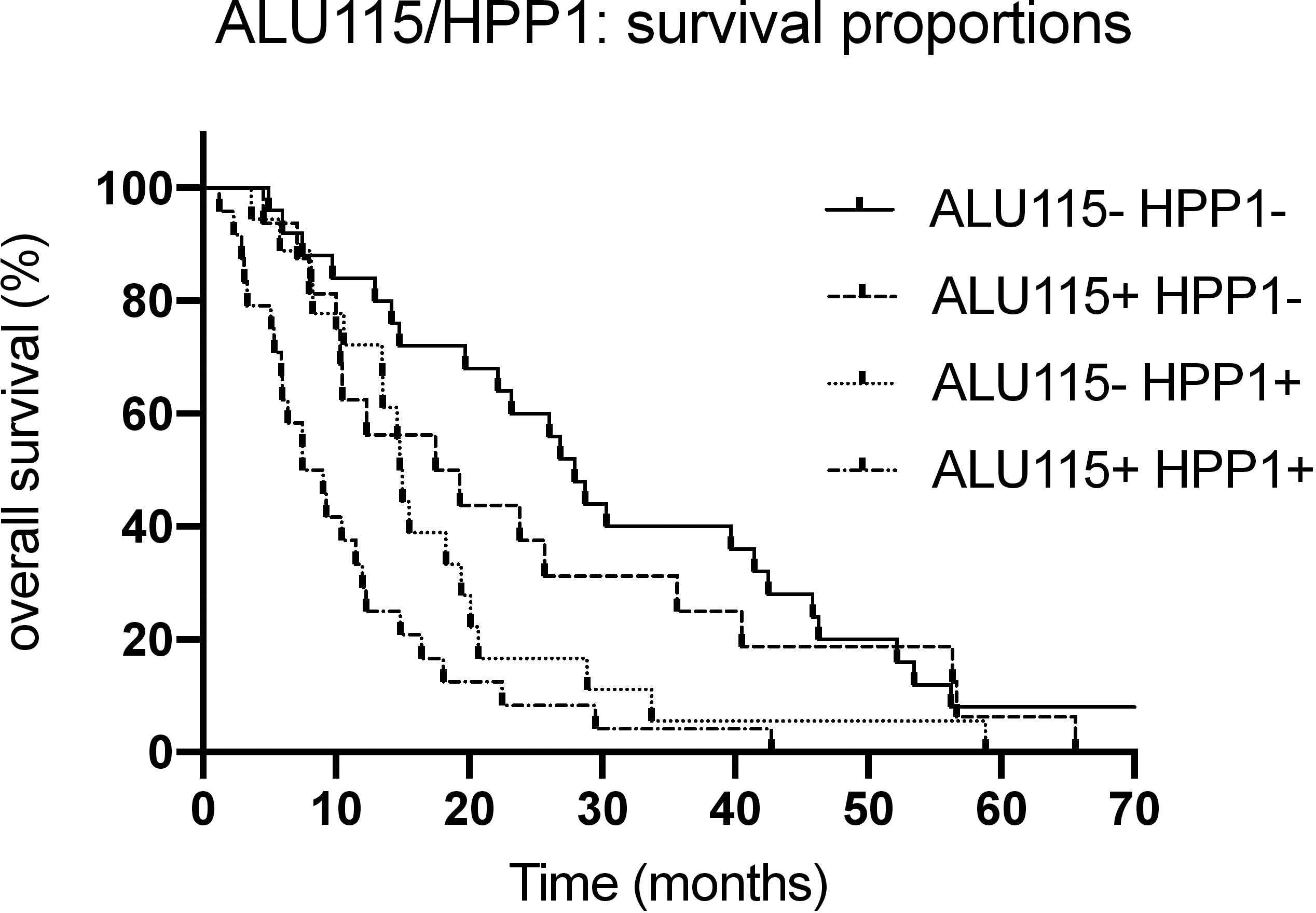
Overall survival in UICC stages I–IV
n.d.: upper limit of 95% CI could not be calculated due to insufficient number of events in this group.
Overall survival in association with our potential markers was analyzed for all patients in all UICC stages (Fig. 2). Kaplan-Meier survival statistics were carried out with the median copy number as cutoff-value. Additionally, quartiles of the marker levels were used as cutoff-values for survival analysis. The results are shown in Table 2. The amount of ALU115 fcDNA fragments in serum of cancer patients showed a highly significant correlation with overall survival with both cutoff-values (
Overall survival of patients in stage UICC IV; multivariate analysis with biomarker combinations (including HLTF and HPP1)
Second, overall survival of patients in stage UICC IV according to the five potential biomarkers was determined separately (Table 3). Determination of survival in UICC stage IV subset was done using the median copy number of each marker as cutoff value. The amount of ALU115 fragments showed a highly significant correlation with survival in UICC stage IV (
The five fcDNA sequences, the methylation markers HLTF and HPP1, which already have proved their prognostic value for CRC in former studies of our group [39], and CEA were included in a multivariate Cox model. Only ALU115 (HR
Survival statistics with combinations of both markers showed highly significant differences in survival (
Discussion
It is estimated that incidences of the 14 most common cancer types including CRC will increase in the future [42]. Furthermore, according to these projections concerning cancer incidences and cancer-related deaths in Germany, CRC will be the third most common cause of cancer-related deaths in men and women by 2030. Even though overall survival of colorectal cancer patients is improving continuously [5], among patients with metastatic disease no progress in survival can be observed [31]. Therefore, new therapeutic strategies would be needed, individually adapted to a patient’s medical conditions, his or her medical condition considering his comorbidities and commensurate to the risks of any medical treatment. To improve the overall survival of UICC stage IV patients and evaluate the individual risk of each patient avoiding under- or over-treatment, it is important to establish new prognostic tools.
In this study, we were able to identify ALU115 fcDNA as an independent prognostic marker for the survival of CRC patients. The prognostic value of ALU115 could be demonstrated in the overall survival analysis for all tumor stages. Notably, survival of patients with UICC stage IV was significantly shorter when ALU115 was above the cutoff value, which was determined by the median copy number of ALU115 fragments in the blood serum of UICC stage IV patients. Compared to CEA as a common clinical marker, ALU115 correlated stronger with clinicopathological parameters as well as survival in UICC stage IV.
In order to maximize the prognostic value, combination of markers could be a useful approach. In a previous study, we demonstrated the prognostic value of HPP1 methylation for UICC stage IV in the identical patient population [40]. Combining HPP1 and ALU115 in Kaplan Meier survival analysis, we were able to define three different risk groups. With both markers being positive, the overall survival was substantially shorter than with only one marker respectively none of both markers being positive. In summary, both markers have a similar prognostic value, but in combination, the prognostic value is even higher.
The analysis of fcDNA in serum as a non-invasive, low-risk “liquid profiling” [44] provides various information about the primary tumor characteristics. Therefore, it is a helpful instrument for understanding metastatic progression and hence a potential prognostic device for cancer patients. Here, sampling of the specimens can easily be repeated to, for example, monitor the amount of fcDNA over a longer period of time.
However, it hast to be taken into consideration that concentrations of cell-free DNA are not only higher in the blood of CRC patients. Shaban et al. evaluated the diagnostic and prognostic value of ALU fcDNA in different cancer types and found that ALU levels in different cancers (type or stage) are strongly varying in the included studies due probe processing, analysis and calculation of concentrations [45]. Levels of fcDNA can also be increased due to chronic inflammations, e.g. systemic lupus erythematosus [50], ischemic heart disease, diabetes mellitus [48] and is discussed to be influenced by psychosocial and physical stress conditions [20]. Furthermore, significant differences in ALU concentrations were found in psychiatric disorders like schizophrenia, major depressive disorder and alcohol-induced psychotic disorder [41] due to neuroinflammatory processes.
In our study population, besides sex and age, only clinicopathological data concerning CRC was recorded. Comorbidities, notably inflammatory conditions, or other types of cancer, were not captured in our study in contrast to Mead et al. [34], where patients were screened by history, patient letters and recent blood tests. However, we had no evidence that ALU concentrations were confounded by other diseases as our data were intrinsically conclusive and we had strong correlations with other markers such as CEA and HLTF/HPP1.
Further studies studying the suitability of fcDNA as a biomarker should consider the general health conditions of the study patients to exclude confounding comorbidities.
Considering the fact that our ALU markers significantly correlate with UICC stage, tumor size, nodal status and metastasis, we expect a decrease of fcDNA concentration after surgery due to a loss of tumor burden. Former studies showed inconsistent data: Bhangu et al. could detect a slight increase of fcDNA concentration after surgery, whereas ALU115 and ALU247/115 significantly and progressively decreased after surgical treatment in a study by Hao et al. [17] An increase of fcDNA markers shortly after surgery may be caused by mechanical and thermic manipulation as well as destruction of both healthy and tumor cells. So, the period of time between surgery and measurement of fcDNA concentrations seems to play an important role and should be chosen thoroughly. The changes of fcDNA concentrations in blood after different types of medical treatment, i.e., surgery, chemo- and immunotherapy, should be subject of further investigations as this question was not addressed before and new therapies are developing quickly.
In order to introduce more biomarkers to clinical practice, future studies need to consider the combination of markers and avoid selection bias. Sample proceeding should be standardized to reduce costs, accelerate analysis and generate a comparable standard. Most importantly, results need to be validated in larger studies in various populations, followed by prospective trials.
In conclusion, according to our findings, the main advantage of ALU115 fcDNA over common markers is to improve the prognostic assessment for CRC patients with metastatic disease.
Author contributions
Conception: Anzinger Isabel, Kolligs Frank T.
Interpretation or analysis of data: Anzinger Isabel, Nagel Dorothea, Ofner Andrea.
Preparation of the manuscript: Anzinger Isabel, Herbst Andreas.
Revision for important intellectual content: Philipp Alexander B., De Toni Enrico N., Holdt Lesca M., Teupser Daniel.
Supervision: Herbst Andreas, Kolligs Frank.
Footnotes
Supplementary data
Primer sequences
fcDNA marker
Forward primer
Reverse primer
ALU115
5’-CCTGAGGTCAGGAGTTCGAG-3’
5’-CCCGAGTAGCTGGGATTACA-3’
ALU247
5’-GTGGCTCACGCCTGTAATC-3’
5’-CAGGCTGGAGTGCAGTGG-3
L1-79
5’-AGGGACATGGATGAAATTGG-3’
5’-TGAGAATATGCGGTGTTTGG-3
L1-300
5’-ACACCTATTCCAAAATTGACCAC-3’
5’-TTCCCTCTACACACTGCTTTGA-3
ND1-mt
5’-CACCCAAGAACAGGGTTTGT-3’
5’-TGGCCATGGGATTGTTGTTAA-3
Median concentrations of fcDNA biomarkers/CEA; HLTF/HPP1: no. of patients with positive methylation status
ALU115
ALU247
L1-79
L1-300
ND1-mt
CEA
HLTF
HPP1
Median concentration
668/
l
675/
l
342/
l
168/
l
438/
l
3.2 ng/ml
–
–
(range)
(13–18,016)
(1.7–26,096)
(1.6–3,925)
(4.4–3,980)
(8.5–6,644)
(1.0–3,945)
–
–
No./% positive
–
–
–
–
–
–
41/15
50/19
IQR
1636
1423
605
311
666
11.1
–
–
