Abstract
BACKGROUND:
The potential of microRNAs (miRNAs) as molecular tumor biomarkers for early diagnosis and prognosis in lung cancer is still unclear.
OBJECTIVE:
To analyze expression of miRNAs in A549 lung adenocarcinoma (LUAD) cells and in primary, non-malignant bronchial epithelial (BE) cells from healthy donors. To analyze the most prominently deregulated miRNAs in plasma samples of LUAD patients and healthy donors.
MATERIALS AND METHODS:
The expression of 752 miRNAs in LUAD and BE cells was assessed by RT-qPCR with mean-centering restricted normalization. The relative plasma levels of 18 miRNAs in LUAD patients and healthy donors were analyzed using RT-qPCR and normalized to miR-191-5p and miR-16-3p. Putative interactions between miRNAs and their target genes were investigated in silico.
RESULTS:
Out of 752 miRNAs, 37 miRNAs were significantly deregulated in A549 cells compared to BE cells. MiR-15b-3p, miR-148a-3p, miR-193b-3p, and miR-195-5p were significantly deregulated in plasma samples of LUAD patients compared to donors. The target genes of those four miRNAs are involved in essential mechanisms in cancer development and progression.
CONCLUSIONS:
There are substantial differences between cancer and control miRNA expression in vitro and in plasma samples of LUAD patients compared to healthy donors. Four deregulated miRNAs are promising as a diagnostic biomarker for adenocarcinoma of the lung.
Introduction
Lung cancer is a very common type of cancer. The number of deaths caused by lung cancer in 2020 in both sexes and all age groups was about 1.8 million, which is more than 18% of all cancer-related deaths, thus making lung cancer the leading cause of cancer death worldwide. Lung cancer can be divided into adenocarcinoma, squamous cell carcinoma, and neuroendocrine tumors, including small cell carcinoma and carcinoid [1]. The early detection of lung cancer plays an important role for prognosis and overall survival. Unfortunately, patients at the early stage are mostly asymptomatic; thus, lung cancer is frequently diagnosed at an advanced stage. Nowadays, for lung cancer diagnostics, different invasive methods such as bronchoscopy with endobronchial ultrasound (EBUS) or/and transbronchial needle aspiration (TBNA), CT-guided transthoracic percutaneous needle aspiration, thoracentesis of pleural effusion, or surgical procedures are used. The medical history and complaints of a patient also play an important role in this decision, since every invasive method has a risk of complications. Thus, the establishment of biomarkers for early detection of lung cancer could be crucial in diagnosing lung cancer at an earlier stage.
The known molecular tumor biomarkers have greatly changed the classification and management of lung cancer and improved the overall survival for lung cancer patients [2, 3]. New biomarkers could especially aid in the selection of individuals for low-dose computed tomography screening and management of small nodules with unclear dignity (benign/malignant) [4].
MicroRNAs (miRNAs) are small, typically 21–23 nucleotides long non-coding RNA molecules that are involved in many biological processes. MiRNAs are involved in gene regulation at post-transcriptional level. MiRNAs play an essential role in many biological processes of healthy, as well as malignant cells, via regulating mechanisms that ensure normal cell and organ activity, such as cell proliferation, differentiation, apoptosis and tumorigenesis [5]. Expression levels of miRNAs vary in different malignancies and depend on cancer type and stage [6]. Aberrant miRNA signatures in malignant cells are regulated by genetic alterations (e.g. chromosomal rearrangements, amplifications, deletions, and mutations), epigenetic changes, and transcriptional regulation [7]. Numerous studies have shown that functional miRNAs substantially contribute to the etiology and progression of different cancer entities, including lung cancer. MiRNAs exhibit tumor suppressive or promoting activities, thereby controlling initiation, progression, and metastasis of lung cancer [7, 8]. MiRNAs modulate various cellular mechanisms such as cell proliferation, motility, invasion, apoptosis, angiogenesis and immune escape, which frequently leads to development of resistance against current cancer therapies [7, 8, 9]. Several miRNAs deregulated in lung cancer have been found in extracellular vesicles (EV) secreted in body fluids, primarily in the circulation. MiRNAs are stable in plasma and serum, thus, circulating miRNA can be used as a minimally invasive biomarker for early lung cancer detection [7, 10]. Among others, Huang and coworkers have shown that EV miRNAs have a high diagnostic accuracy for non-small cell lung cancer (NSCLC) and suggested miR-21, miR-210, and miR-1290 as potential diagnostic biomarkers for NSCLC [10]. Regarding the clinical implications of aberrant miRNAs in lung cancer, miRNAs are promising as cancer subtype classifiers, diagnostic and prognostic markers, as drug-response predictor and, finally, as potential therapeutic targets [11].
In this study, we aimed to investigate in vitro expression profiles of miRNAs in A549 LUAD cells compared to primary, non-malignant bronchial epithelial (BE) cells. Furthermore, we compared the expression of miRNAs prominently deregulated in vitro in plasma samples of 18 LUAD patients and 18 healthy donors. We analyzed putative interactions and molecular functions of validated target genes of deregulated miRNAs. A detailed description and understanding of miRNA expression profiles and related molecular mechanisms can provide useful information for the search of future diagnostic and therapeutic targets in LUAD patients.
Materials and methods
Cell culture
A549 cells were purchased from ATCC (Rockville, Maryland, U.S.) and grown in DMEM/F12 medium supplemented with 10% fetal calf serum (FCS; Gibco, Paisley, UK). Primary BE cells were isolated from non-utilized human donor lungs, explanted at the Department of Surgery, Medical University of Graz, Austria. The investigations were conducted according to the Declaration of Helsinki principles and were approved by the local institutional review board. For BE cell isolation, a protocol modified after Fulscher et al. [12] was used. In brief, isolated proximal to segmental bronchi from explant lungs were cut in approximately 1–2 cm pieces, rinsed with PBS and added to a cell culture dish with DMEM/F12 medium containing 20% fetal calf serum (FCS), antibiotics (Pen/Strep), and antimycotic (Fungizone, 1:100; Gibco, Waltham, MA). Epithelial cells were scraped gently with a scalpel, resuspended, and transferred in a new cell culture dish coated with collagen type I (rat-tail collagen, 4.5 mg/ml, Sigma Aldrich, diluted 1:100 in PBS), containing DMEM/F12 and 20% FCS. After cell attachment, medium was replaced with BEGM (CC-3170, Lonza, Basel, Switzerland) containing growth factors, which was further used as a growth medium. BE cells were cultured on collagen-coated cell culture dishes and used in low passages (
Patient cohort
This study confirms to the provisions of the Declaration of Helsinki (as revised in 2013) [13]. This study was approved by the Ethics Committee of the Medical University of Graz. All participants signed written informed consent and were randomly selected from those who presented in the Pulmonology Department. The inclusion criterion for LUAD patients was a completed histology showing lung adenocarcinoma. Exclusion criteria were: i) any other active malignant disease, ii) antitumor therapy for lung cancer before the histological material was retrieved, iii) acute or chronic pulmonary infection. In all patients, lung adenocarcinoma was diagnosed by experienced interdisciplinary team of lung pathologists, radiologists and pulmonologists from the University Hospital Graz, Austria. Classification and staging were done following the newest (8
Cytokeratin immunofluorescence
BE cells were cultured in LabTek
RNA isolation from cells and quality check
The RNA isolation from cells was performed using the mirVana
Reverse transcription
For cDNA synthesis, the miRCURY LNA
SYBR-Green based RT-qPCR
The expression levels of different miRNAs in the LUAD and BE cells were detected using the miRCURY LNA
Normalization and statistical analysis of RT-qPCR data
Three replicates per sample and condition were analyzed and the results were represented as Ct-values determined with the 2
Blood collection and miRNA isolation from plasma samples
Three milliliters of full peripheral venous blood were collected from each patient and healthy donor using EDTA-anticoagulant tubes (Thermo Fisher Scientific, Vienna Austria). Immediately upon collection, blood samples were centrifuged for 15 min at 1000
For miRNA isolation from plasma samples, miRNeasy Serum/Plasma Advanced Kit (Qiagen) was used. Isolation of miRNA was performed as recommended by producer, by using 200
Mutational analyses
In all LUAD patients, the mutational status of tumor cells was determined routinely using the Ion Torrent S5 system for next generation sequencing (NGS) and two different panels. The first panel was the Ion AmpliSeq Colon and Lung Cancer Research Panel v2, containing 92 pairs of primers with an average amplicon length of 162 base pairs (bp). With this DNA-based panel following genes were analyzed: KRAS, EGFR, BRAF, PIK3CA, AKT1, ERBB2, PTEN, NRAS, STK11, MAP2K1, ALK, DDR2, CTNNB1, MET, TP53, SMAD4, FBX7, FGFR3, NOTCH1, ERBB4, FGFR1, and FGFR2. The RNA-based panel was the Archer Fusion Plex MUG Pan-TRK Lung panel v1 (custom modified for the Institute of Pathology, Medical University of Graz). This panel was used for analysis of following genes: ALK, BRAF, EGFR, FGFR1, FGFR2, FGFR3, MET, NRG1, NTRK1, NTRK2, NTRK3, RET, ROS1. PD-L1 expression was analyzed by immunohistochemistry and evaluated as percentage of positive tumor cells compared to all tumor cells.
Statistical calculations
All statistical analyses were performed by GraphPad Prism (v. 5.0). Data are represented as means
Results
miRNA expression profiles in cell samples
The experimental design of this study is shown in Fig. 1A. Isolated primary BE cells displayed a high degree of purity and nearly 100% positivity for cytokeratin (Suppl. Fig. 1A). Only RNA samples with an RNA integrity number (RIN) between 9.8–10.0 were used for further analyses (Suppl. Fig. 1B). Out of 752 miRNAs analyzed in our screening, 123 were detectable by RT-qPCR in both A549 and BE cells. The rest of analyzed miRNAs were either not expressed, or their expression level was below the detection range of our method. Based on the producer’s recommendation (Qiagen) and our own experience, all cycle threshold (Ct) values higher than 37 were considered negative.
In vitro miRNA expression profiling. (A) Workflow of the present study. MiRNAs were isolated from A549 LUAD cells and primary, non-malignant bronchial epithelial (BE) cells from three healthy donors. MiRNA expression profiles were analyzed using RT-qPCR on panels containing 752 miRNAs in total. MiRNAs which were most prominently deregulated in vitro were selected for analyses in plasma samples of LUAD patients and healthy donors. (B) Expression levels of 18 miRNAs most prominently deregulated in A549 cells compared to non-malignant BE cells (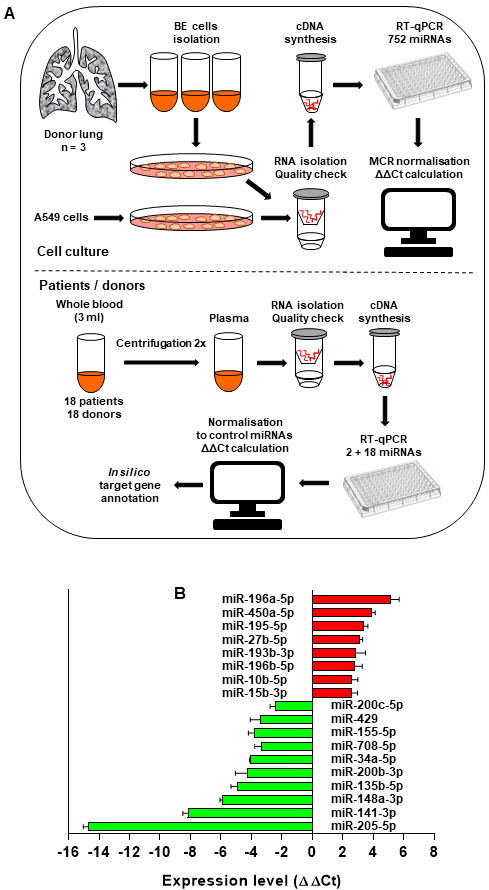
A reliable miRNA normalization is critical for data interpretation. Here, we used the mean-centering restricted (MCR) method [15]. Rather than focusing on all miRNAs, including miRNAs without detectable expression value, this method is restricted to all miRNAs that are detectable in all analyzed samples. MCR-based data normalization delivered very restrictive results. Without any cut-off, 58 miRNAs were differentially deregulated in A549 cells compared to BE cells. After setting the cut off to
Deregulated miRNAs (
In this study, plasma samples of 18 LUAD patients and 18 healthy controls were included. All patients had a confirmed lung adenocarcinoma diagnosis and did not receive any anticancer therapy before sample collection. Donors were characterized as individuals without known malignancies or acute and/or chronic infections. Clinicopathological characteristics of LUAD patients are summarized in Table 2. Following data were collected: age, sex, cancer stage, anatomical cancer localization in the lung, smoking status, indicated mutations, PD-L1 expression, and metastases status. The stage classification was based on the last (8
Clinicopathologic characteristics of LUAD patients
Clinicopathologic characteristics of LUAD patients
F, female; M, male; LLL, left lower lobe; LUL, left upper lobe; RUL, right upper lobe; R, right, L, left; n. d., no data; neg., negative; pos., positive. Multiple, at least in two organs.
Survival data of the patients
S
We analyzed the survival data of patients from our cohort approximately two years after the first diagnosis (Table 3). Seven out of 18 (39%) patients died during this time. All patients who underwent surgery (
Finding stably expressed circulating miRNAs that can be used for normalization of expression data generated in biofluids (e.g. plasma) is even more challenging than the normalization of data collected under fully controlled in vitro conditions. This is partially due to biological diversity between different patients or donors, and can be further influenced by sample collection, as well as by miRNA isolation and storage. Therefore, stably expressed miRNA were investigated in plasma samples. These results demonstrated that miR-16-3p and miR-191-5p were expressed in all samples and the expression level of each of them was very consistent, both in plasma samples from patients and from healthy donors (Fig. 2A). MiR-191-5p was expressed at average Ct
Expression profiling of selected miRNAs in plasma samples of LUAD patients and healthy donors. (A) Two control miRNAs, miR-16-3p and miR-191-5p, were chosen for data normalization based on their stable expression in all samples and in both cohorts. Expression levels of control miRNAs are presented as interleaved box and whiskers and as double gradient heat map. (B) Circulating miRNAs (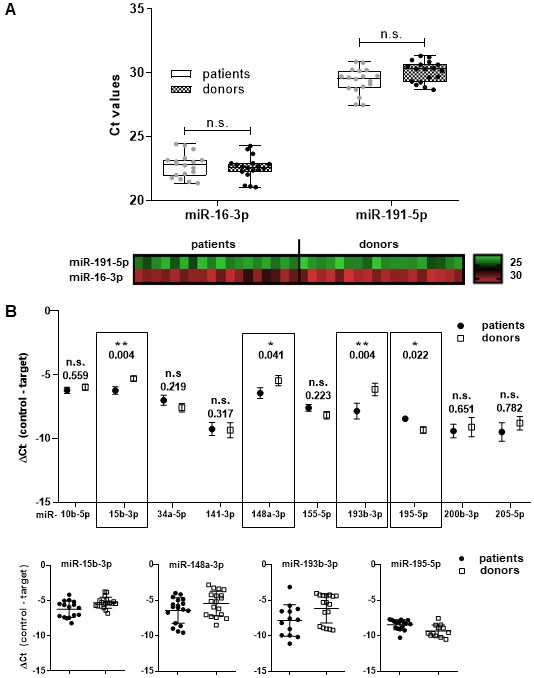
Based on the miRNA expression data from our in vitro experiments, we chose the most prominently deregulated miRNAs (
Annotation of published target genes by PANTHER overrepresentation test for pathway classification
Annotation of published target genes by PANTHER overrepresentation test for pathway classification
As a reference (Ref.), all genes in database for Homo sapiens were used and Fisher’s exact test with data correction for false discovery rate (FDR) was performed. Panther Classification System 16.0. (www.pantherdb.org; January 2021).
Regulatory network and putative interactions of differentially expressed miRNAs and their target genes and pathways. (A) Regulatory network of four deregulated miRNAs and their direct target genes validated so far. Red diamonds, yellow diamonds, and green rectangles represent up-regulated miRNA, down-regulated miRNAs, and associated target genes, respectively. For data visualization, Cytoscape (version 3.8.2) was used. (B) Based on their direct target genes, deregulated miRNAs were annotated to particular pathways by performing Panther Overrepresentation Test. The down-regulated and up-regulated miRNAs are shown in green-line and red-lined rectangles, respectively. MiR-15b-3p was not annotated to any special pathway. Analysis was done by using the Panther Classification System 16.0. (www.pantherdb.org).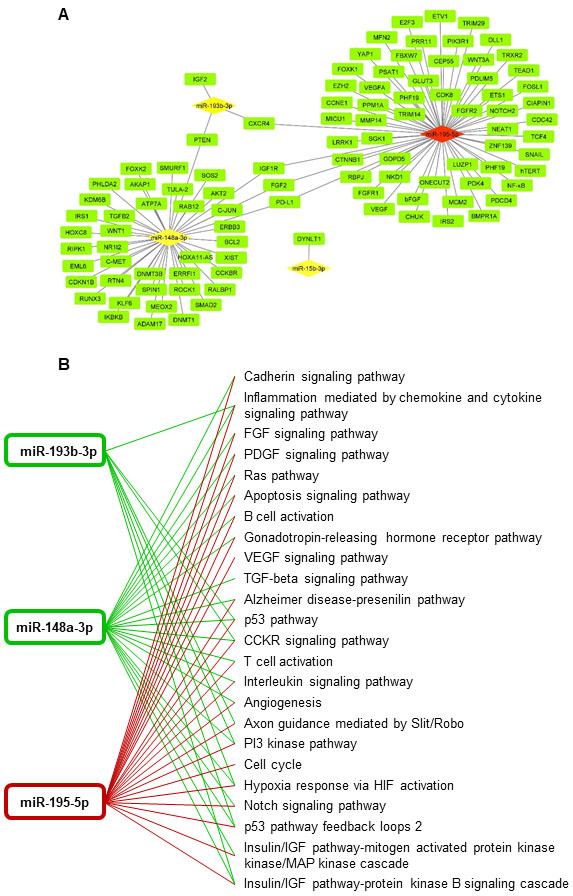
Putative interactions and molecular functions of validated target genes for deregulated miRNAs. Analysis was performed by the Panther Classification System 16.0. (www.pantherdb.org).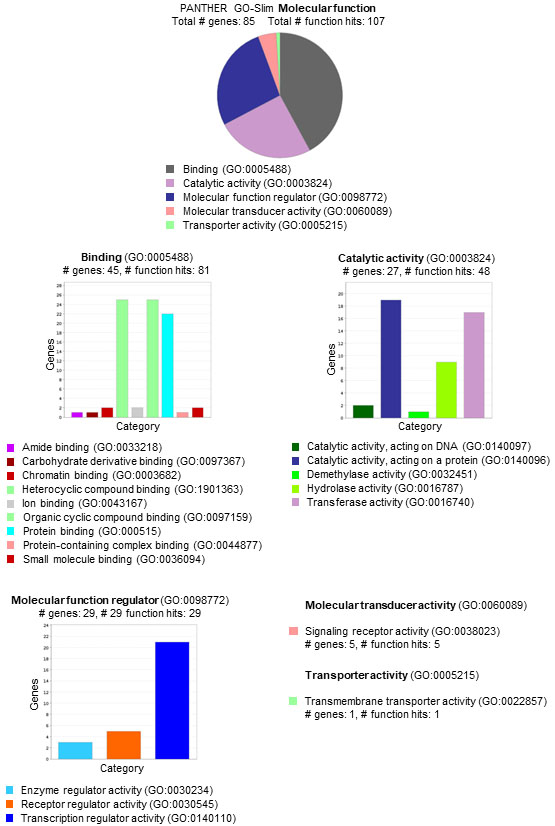
In order to shed light upon the target genes regulated by these four miRNAs (miR-15b-3p, miR148a-3p, miR-193b-3p, and miR-195-5p), we performed literature screening and regulatory network analyses for genes described so far as direct targets of these miRNAs. Upon screening (https://pubmed.ncbi.nlm.nih.gov; searching criteria: hsa-miR-X-Xp
Discussion
Early detection of LUAD is still very challenging and the need to identify specific molecular tumor biomarkers, especially in serum, plasma, or in the whole blood is essential. MiRNAs have potential as molecular tumor biomarkers for early diagnosis and prognosis and for the evaluation of treatment efficacy. Such biomarkers might even help to establish personalized therapies for LUAD patients. In the study presented herein, expression levels of miRNAs in malignant A549 LUAD cells and primary, non-malignant BE cells were compared. Based on these in vitro data, the expression levels of the most deregulated miRNAs were analyzed in plasma samples of LUAD patients and healthy controls.
To determine the expression levels and deregulation of miRNAs in different samples, RT-qPCR data must be normalized to one or more miRNAs stably expressed over all samples. In the literature, different miRNA normalization methods have been described, but there is no agreement about the one single optimal strategy. Therefore, the normalization of miRNA qPCR data requires a well-considered decision to accurately handle the obtained data [17]. For in vitro data in this study, we used the MCR method for miRNA normalization, as this method has been described as the most reliable out of seven different methods [15]. The MCR method focuses on a restricted set of miRNAs, taking into account miRNAs that are fully detected in all analyzed samples, thereby excluding the miRNAs with non-determined values (which are e.g. included in the non-restricted mean-centering method).
Based on our in vitro data, we selected a panel of 18 miRNAs with
In this study, special care has been taken to avoid haemolysis and to remove all blood cells from plasma samples. These two factors frequently lead to increased concentration of miRNAs highly expressed in red and white blood cells or in platelets. Thus, upon blood collection, two consecutive centrifugation steps were performed in order to remove any residual blood cells. Sample collection and sample preparation are still two major drawbacks in performing miRNA expression profiling in biological samples, and their standardization is urgently needed. Another big challenge is the miRNA nomenclature. It is frequently not clear from the published studies which strand of particular miRNA was investigated. This problem should not be trivialized because it is well known that 5p and 3p miRNA molecules are biologically different in terms of stability, which has an impact on their function and ability to bind to the RISC complex and, finally, on the binding to their target genes.
In this study, we detected four miRNAs differentially expressed in plasma samples of LUAD patients compared to healthy donors. Human miR-15b-3p is located at chromosome 3q25.33. To our best knowledge, there are no data about the expression of miR-15b-3p in lung cancer. Several studies have looked at either the 5p strand of miR-15b or, mostly, the active strand was not specified at all. In one study, miR-15b-5p was described as down-regulated in exosomes from plasma of squamous cell carcinoma patients compared to healthy controls [20]. In another study, miR-15b-5p was up-regulated in the serum of non-small cell lung cancer (NSCLC) patients [21]. Shi et al. described down-regulated serum expression of miR-15b in NSCLC patients in comparison to healthy donors and benign lung disease, the miR-15b expression being lower in stage III–IV patients than in stage I–II [22]. Thus, the authors suggested miR-15b, together with miR-22, as reference markers for early NSCLC diagnosis. Furthermore, miR-15b appears to play a role in chemotherapy resistance of lung cancer, as the sensitivity to cisplatin of advanced NSCLC patients on cisplatin-based chemotherapy was higher in patients with high expression of miR-15b [23]. In advanced NSCLC, serum levels of miR-15b have been shown to change pre- and post-chemotherapy, these changes being associated with chemotherapeutic response [23]. This is further supported by studies showing that miR-15b is involved in DNA-repair regulation via Wip1/ATM/Chk1/p53 signaling [24], and via expression inhibition of PEBP4 [25]. On the other side, miR-15b was found to be more highly expressed in human NSCLC tissue and cells than in normal tissue/cells, thereby controlling the viability, cell cycle, and migration via targeting TIMP2 [26]. In summary, data about miR-15b-3p expression in lung cancer are lacking, and data regarding the role of miR-15b in cancer in general and in lung cancer in special are controversial. These discrepancies may be due to the fact that miR-15b is expressed in platelets and its level depends on sample preparation. This can lead to large differences in expression of circulating miR-15b in whole blood, serum, or in plasma.
Human miR-148a-3p is located at the 7p15.2 chromosomal region and belongs to the miR-148/152 family. In our study, miR-148a-3p was down-regulated in plasma samples of LUAD patients compared to healthy donors, consistent with published data. MiR-148a-3p has been described as down-regulated in tumor tissue, serum, and plasma of NSCLC patients, frequently showing a significant correlation with tumor stage but not with survival [27, 28, 29, 30]. Regarding the molecular mechanism, it has been shown that miR-148a-3p inhibits proliferation and migration, and induces apoptosis in NSCLC cell via the PI3K/AKT signalling pathway by targeting ERBB3 and ITGA5 genes [31]. Furthermore, miR-148a-3p binds to the lncRNA HOXA11-AS, which furthermore regulates DNMTA (a central DNA methyl transferase in mammalian cells) expression and facilitates NSCLC tumorigenesis in vitro and in vivo [32]. Together, all these data indicate a tumor suppressive role of miR-148a-3p and its downregulation in plasma samples of LUAD patients indicates its direct involvement in LUAD tumorigenesis.
Human miR-193b-3p is located on chromosome 16p13.12 and, based on literature, has tumor-suppressive function in human cancers, including lung cancer [33, 34, 35]. In our study, miR-193b-3p was the most down-regulated miRNA. Thus, our data are in line with data published, including that by Hu et al., who described lower level of hsa-miR-193b-3p in NSCLC cells compared to non-malignant cells [36]. They also showed that inhibition of mir-193b in A549 cells led to increased cell proliferation, migration, and invasion. These effects were based on repression of cyclin D1 and urokinase-type plasminogen activator, suggesting the tumor suppressive role of miR-193b in NSCLC [36]. Furthermore, it has been shown that both 5p and 3p strands of miR-193b were down-regulated in metastatic LUAD cells. Ectopic expression of both miR-193b strands, based on NSCLC cell transfection with miR-193b-5p and -3p precursors, inhibited proliferation and metastatic potential of lung cancer cells [34]. In their integrative analysis of miRNAs in NSCLC, based on two publicly available datasets and validated with RT-qPCR, Zhang et al. described miR-193b-3p as up-regulated in NSCLC to a significant degree [37]. However, this data describe the expression of miR-193b-3p in NSCLC tumor tissue and not of circulating miR-193b-3p, which may be the reason for this discrepancy. Recently, miR-193b-3p has been described in plasma exosomes as down-regulated in e-cigarette smokers compared to traditional smokers [38]. Although all patients in our cohort were active or ex-smokers, explicit information about e-cigarette smoking is lacking.
Located on chromosome 17p13.1, miR-195-5p has been shown to be aberrantly expressed in various cancers such as breast cancer [39], renal carcinoma [40], colorectal cancer [41], and lung cancer [42, 43, 44, 45, 46, 47]. In our study, we showed significant up-regulation of miR-195-5p in plasma samples of LUAD patients, with a very constant expression level both in patients and in healthy donors. This is in contrast to some studies reporting down-regulation of miR-195-5p in lung cancer [44, 45, 46, 47]. However, it should be stressed that these studies analyzed miR-195-5p expression in lung cancer tissue or cell lines, whereas the expression of miR-195-5p in plasma of NSCLC patients has not been analyzed thus far. In NSCLC cell lines A549 and H1299, miR-195-5p inhibited proliferation and induced apoptosis by targeting CEP55 [42]. As further targets of miR-195-5p in lung cancer cells, CHEK1 [43], CPNE1 [44], CIAPIN1 [45], GLUT3 [46], FOXK1 [47], and PDK4 [48] have been described. Together with five other miRNAs, miR-195-5p has been also identified as smoking-related miRNA, thus perhaps being at least partially responsible for cigarette smoke-induced malignant transformation of bronchial epithelial cells [49]. Expression of miR-195-5p in lung cells can be regulated by lncRNA CASC9 [48] and by circular RNA circMYLK [46], as recently described. Altogether, these data indicate the complexity of miR-195-5p regulation, both in cancerous and in non-cancerous tissues and cells, and elicit need for further investigations.
We found five overlapping target genes (PTEN, CXCR4, IGF1R, FGF2, and PD-L1) that are directly regulated by at least two of four deregulated miRNAs. These genes have different cellular functions and are involved in several signalling pathways in non-malignant and various malignant cells. Phosphatase and tensin homologue deleted on chromosome ten (PTEN) is a tumor suppressor gene frequently lost in variety of tumors (for review see [50]). PTEN inhibits cell proliferation, survival, growth, and migration by blocking the phosphatidylinositol 3 kinase (PI3K) signaling pathway and inhibiting inter alia, the membrane recruitment and activation of AKT. Tumor-suppressive function of PTEN can be regulated by its binding partners and by its localization [50]. In addition, PTEN can be exported from the cell and can be transferred from one cell to another in form of exosomes [51] and its translational variant (PTEN-Long) has also been detected in human serum and plasma [52].
C-X-C chemokine receptor 4 (CXCR4), a 7-trans- membrane G-protein coupled chemokine receptor, is known for its numerous pleiotropic functions under physiological and pathophysiological conditions. Through interaction with its ligand CXCL12, the CXCR4 plays a role in a number of cellular processes and is expressed in over 30 types of malignant tumors (for review see [53]). CXCR4 functions in autoimmunity, immunodeficiency, and cancer are well documented and it has been identified as prognostic marker for breast, ovarian, and pancreatic adenocarcinoma [54]. It is thought that the CXCL12/CXCR4 axis has a pro-tumorigenic role in NSCLC by supporting tumor growth, enhancing angiogenesis and metastases, and promoting chemo-resistance [55]. Although it has been shown that miR-195-5p directly regulates the CXCR4 in the heart failure mice model, thereby inactivating the JAK/STAT pathway [56], in lung cancer, CXCR4 regulation by miR-193b-3p or miR-195-5p has not been described.
Insulin like growth factor 1 receptor (IGF1R), a transmembrane tetrameric receptor containing two extracellular alpha and two transmembrane beta subunits, binds insulin-like growth factors (IGFs) and insulin in a variety of cells and tissues. IGF1R is overexpressed in many tumors and plays an important role in anti-apoptosis, thus enhancing cancer cell survival and proliferation [57]. Furthermore, IGF1R activation is connected to several metastatic features of cancer cells, including tumor angiogenesis, degradation of extracellular matrix, and loss of anchorage-dependent growth [58]. However, it is not clear how deregulation of miR-148a-3p and miR-195-5p might influence the expression of IGF1R and its potential role in NSCLC development.
Fibroblast growth factor 2 (FGF2) is frequently increased in different cancer entities, including NSCLC [59, 60], acting as regulator of cancer cell proliferation, invasion, angiogenesis, and metastases [61, 62, 63]. In NSCLC cells, correlations between FGF2 expression and different miRNAs including miR-182, miR-210, miR-497-5p, and miR-152 has been shown [63, 64, 65, 66]. Regulation of FGF2 by miR-148a-3p or miR-195-5p has not been put into context with NSCLC yet and should be further investigated.
Programmed cell death ligand 1 (PD-L1) is an approved predictive biomarker for lung cancer. PD-L1 binds to PD-1 expressed on T-cells, B-cells, dendritic cells and natural killer T-cells to suppress anti-cancer immunity. PD-L1 expression is regulated by different signaling pathways, as well as by transcriptional and epigenetic factors, and miRNAs (for review see [67]). Improved survival of a subset of lung cancer patients have been observed upon immunotherapy targeted against PD-L1 and its receptor PD-1 [68]. In colorectal cancer, direct regulation of PD-L1 by miR-148a-3p has been shown [69]. In our patient cohort, showing significant down-regulation of miR-148a-3p, PD-L1 was positive (30–100% of all tumor cells) in eight (44%) patients, however, direct regulation should be experimentally validated. Altogether, pathway analysis indicates that the target genes of our four deregulated miRNAs are involved in mechanisms and pathways essential in cancer development and progression.
As a limitation of this study, our preselection of deregulated miRNAs was performed in one single LUAD cell line (A549). This cell line is widely used as an in vitro model and was chosen to provide an overview of miRNAs deregulated in malignant cells in comparison to non-malignant BE cells. In our pilot experiments (data not shown), we also compared miRNA expression profiles of H460 LUAD cells in comparison to BE cells. However, none of 10 miRNAs deregulated in A549 cells was detectable in H460 cells. Although this is a strong indication that in H460 cells, these miRNAs might be also down-regulated, it is not certain whether it is a limitation of the methods or if they are not expressed at all. Variations in miRNA expression based on different genetic background of these two LUAD cell lines are expected, since A549 cell line was established from a lung carcinomatous tissue, whereas H460 cells represent a hypotriploid human cell line derived from pleural fluid of patient with large cell lung cancer. Our study found a considerable overlap between miRNAs deregulated in A549 cells and in LUAD patient’s plasma samples, but it also opened the door for discussion, especially regarding the variations in miRNA expression between different cell models and LUAD patients.
Next, this study was retrospective and the number of patients and controls was quite small. We relied on an explorative approach, based on in vitro experiments with one adenocarcinoma cell line, and this might not be representative for all adenocarcinoma cells. We did not investigate other types of lung tumors or other malignant diseases for our detected miRNAs and, therefore, cannot assess the specificity for LUAD. Thus, further studies are needed to clarify these important issues.
Early cancer detection is essential for reduction of cancer mortality worldwide. Based on benefits for patients, low-dose computed tomography (CT) scan is recommended by different international guidelines as a screening method for early lung cancer detection, especially for high risk patients [70]. However, the CT screening has some limitations and possible drawbacks, such as additional radiation exposure. Furthermore, overdiagnosis can lead to unnecessary invasive procedures, which are connected with increased risk for patients. There are still no well-established biomarkers for early lung cancer detection, but this is under intensive investigation, with a focus on miRNAs as promising candidates. As an advantage of our approach, we focused in patient samples on miRNAs pre-selected in vitro. Patient samples are unique; functional analyses of miRNAs and their target genes are very costly and time consuming. Thus, pre-selection and focusing on specific miRNAs and target genes is a benefit to address these challenges. MiRNAs described in this study might be important molecular markers for lung adenocarcinoma, both for early diagnosis and possibly for reexamination of patients after surgery or chemotherapy. However, this is still speculative and must be verified in future prospective studies.
In conclusion, the analysis of differentially expressed miRNAs in A549 cells compared to primary, non-malignant BE cells led to the identification of four de-regulated miRNAs in the plasma of LUAD patients. We suggest miR-15b-3p, miR-148a-3p, miR-193b-3p, and miR-195-5p as potential plasma markers for LUAD. Their direct target genes play an essential role in tumor development and growth. Improved understanding of molecular mechanisms and the role of specific molecular biomarkers will lead to improved detection of early stage tumors and, ultimately, to improved biomarker-directed therapies for LUAD patients.
Author contributions
Conception: Irina Robinson, Horst Olschewski, Andelko Hrzenjak.
Interpretation or analysis of data: Irina Robinson, Alexandra Bertsch, Andelko Hrzenjak.
Preparation of the manuscript: Irina Robinson, Andelko Hrzenjak, Katharina Leithner.
Revision for important intellectual content: Irina Robinson, Andelko Hrzenjak, Katharina Leithner, Philipp Stiegler, Horst Olschewski.
Supervision: Andelko Hrzenjak.
Funding
This study was supported by the Ingrid Shaker Nessmann Cancer Research Association (grant to A.H.). The founder did not have any role in study design, data collection and analysis, interpretation of results, decision to publish, or preparation of the manuscript.
Declaration of competing interest
The authors report no declarations of interest.
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-210205.
sj-pdf-1-cbm-10.3233_CBM-210205.pdf - Supplemental material
Supplemental material, sj-pdf-1-cbm-10.3233_CBM-210205.pdf
Footnotes
Acknowledgments
We thank all patients and donors for participating in this study. We appreciate excellent technical assistance of Theodora Goessler and kind assistance of Core Facility for Molecular Biology, Center of Medical Research, Medical University of Graz.
