Abstract
BACKGROUND:
Globally, breast cancer (BC) has become one of the most prevalent malignancies and the leading cause of tumor-related deaths among women. Dysregulation of the cell cycle is a well-known hallmark of cancer development and metastasis. CDKs are essential components of the cell-cycle regulatory system with aberrant expression in a variety of cancers, including BC. In the development of targeted cancer treatment, reestablishing the regulation of the cell cycle by modulation of CDKs has emerged as a promising approach.
METHODS:
Herein, we used a bioinformatic approach to assess the expression pattern, prognostic and diagnostic importance, and clinical relevance of CDKs in BC. Additionally, we conducted a functional enrichment analysis of deregulated CDKs using the STRING and KEGG databases to delineate the role of CDKs in breast tumorigenesis.
RESULTS:
Gene expression analysis revealed substantial deregulation of CDKs in BC, with CDK1, CDK11A, and CDK18 showing a fold change of
CONCLUSION:
The study’s multimodal analytical methodologies imply that modulating CDKs for BC treatment is a promising approach.
Introduction
Breast cancer (BC) is one of the most common malignancies as well as the leading cause of tumor-related mortality among women globally. In the United States alone, 1700 people will die from cancer each day, with breast, lung, and colorectal cancers accounting for the majority of fatalities in women [1]. BC incidence rates are rising in transitional nations such as South America, Africa, and high-income Asian countries (Japan and the Republic of Korea) due to substantial changes in lifestyle, social, and constructed environments brought about by developing economies [2]. In addition, there has been a rapid rise in BC mortalities in Sub-Saharan African countries, largely due to the underdeveloped health infrastructure and circumstances that result in poor survival [2]. BC is a highly heterogeneous solid tumor with more than 20 distinct subgroups differing in appearance, genetics, and clinical behavior. Based on gene expression, BC has been categorized into luminal A, luminal B, HER2-enriched, and triple-negative breast cancer (TNBC) [3]. TNBC accounts for approximately 20% of all BC cases and is defined by the lack of estrogen (ER) and progesterone (PR) receptors, as well as the lack of amplification or overexpression of human epidermal growth factor receptor 2 (HER2) [3, 4]. Despite advancements in diagnostics and treatment modalities, BC continues to be a problem due to low treatment response and the development of recurring and metastatic cancers [5].
Several novel drugs targeting crucial pathways in cellular development and regulation are being developed and tested in combination treatment to improve the effectiveness of therapeutic regimens and extend the survival of cancer patients [6]. The cell cycle and subsequent cellular development are controlled by a highly coordinated cell cycle control system involving cyclin-dependent kinases (CDKs) and cyclins [7, 8]. Interactions amongst cyclin-dependent kinases (CDKs) and cyclins are critical in maintaining the regulation of cell cycle progression [8]. Loss of control over the cell cycle, leads to unrestricted growth, a classic hallmark of cancer. CDK modulation to reestablish cell cycle control has thus been a promising alternative in the development of targeted cancer therapy [9]. In the present study, we evaluated the expression of CDKs in clinical samples of BC patients by utilizing TCGA BRCA datasets available on the UCSC XENA and Gepia2 Web portals. We report here that the expression of CDKs is significantly deregulated in BC. In addition, deregulated CDK expression was found to affect both overall survival (OS) and relapse free survival (RFS) in patients with BC. Enrichment studies showed that CDKs are essential components of neoplastic pathways and that modulating CDKs in combination with conventional therapies will be a promising approach to treat BC patients.
Materials and methods
Identification of differentially expressed CDKs in breast cancer
The UCSC XENA online database was utilized to analyze the expression pattern of CDK in breast cancer and a heatmap was generated [10]. The Gepia2 database (
Kaplan-Meier plotter
The Kaplan-Meier plotter (
ROC plotter analysis
To evaluate the correlation between expression levels of the highly deregulated CDKs, viz., CDK1, CDK11A, and CDK18 with sensitivity to endocrine therapy and HER2 directed therapies, the ROC Plotter portal was accessed [14]. The ROC Plotter (
bc-GenEXminer
Breast cancer Gene-Expression Miner V4.5 (bC-GenEXminer v4.5) (
Log2 fold change of CDKs in breast cancer
Log2 fold change of CDKs in breast cancer
For gene-gene interaction analysis, we utilized the online portal GENEMANIA (
Functional enrichment of highly deregulated CDKs
The Mayaan lab online database (
CDKs exploration in single cell sequencing database (TME)
The Tumor Immune Single-cell Hub (TISCH) was first processed to visualize significantly deregulated CDK expression across several datasets in BC tissues at the single-cell level [23]. We investigated expression of CDK1, CDK11A, and CDK18 in primary and metastatic breast cancer datasets to determine the relationship between CDK expression patterns and tumor progression.
Results
Expression of CDKs is deregulated in BC
The expression pattern of CDKs in BC was examined using UCSC XENA. CDK expression analysis in the TCGA BrCa dataset revealed significant CDK deregulation in BC. Figure 1. Further analysis using the Gepia2 database revealed the log2 fold change of different CDKs in BC Table 1. CDK1 was found to be highly upregulated, with a log2 fold change of 2.84 and a p-value of 2.82E-184, whereas CDK11A and CDK18 were found to be downregulated, with log2 fold changes of
mRNA expression pattern of CDKs in breast cancer patients. Heat map displaying the expression patterns of CDKs. XENA was used to extract CDK expression. Comparing expression of CDKs amongst GTEx (normal breast, 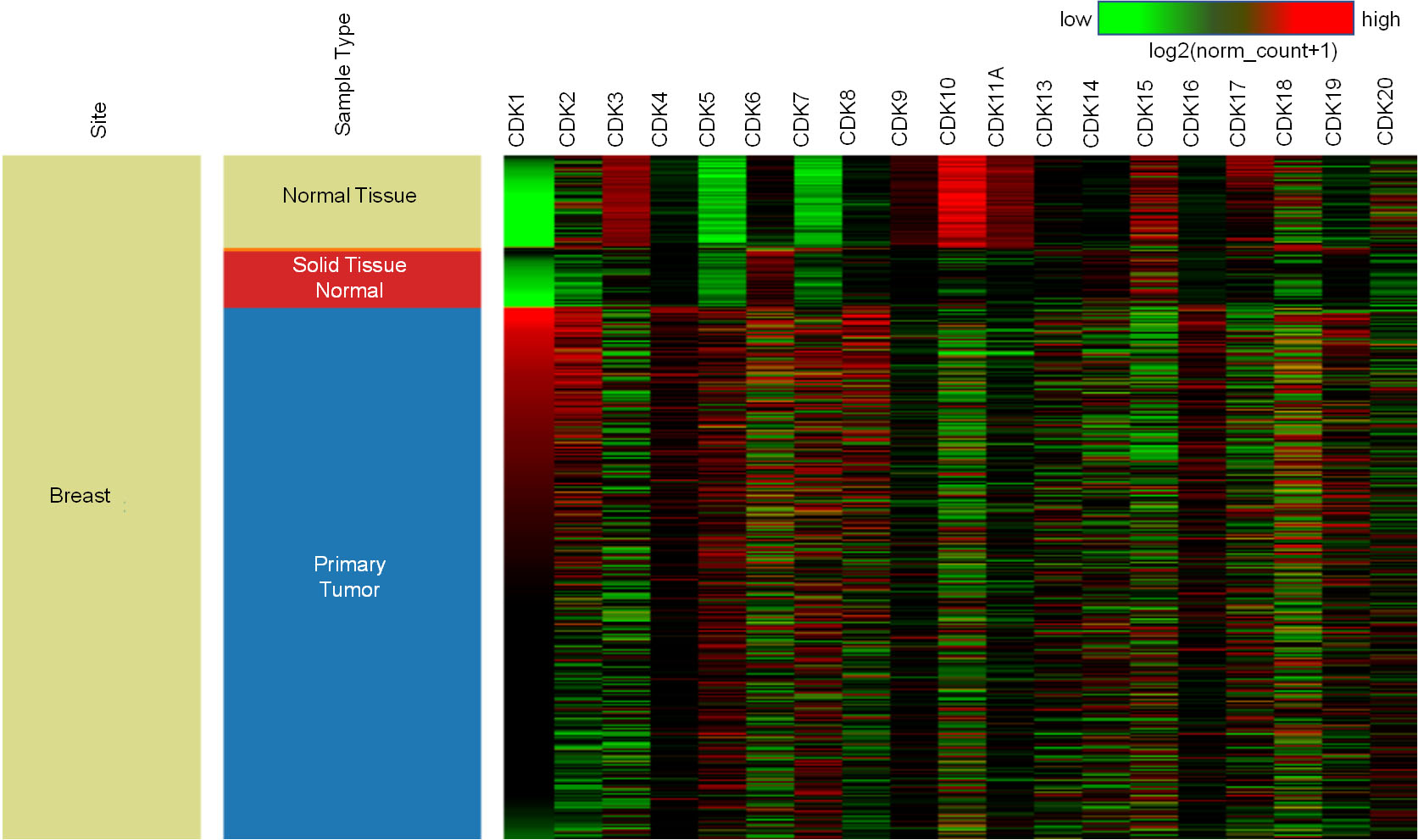
Expression pattern of CDKs in Breast cancer. A. Box Plots of the expression pattern of CDKs analyzed using TCGA BRCA and GTEx datasets using UCSC XENA. B. Expression profile of highly deregulated CDK1, C, CDK11A, and D, CDK18 in BC using Gepia2 database 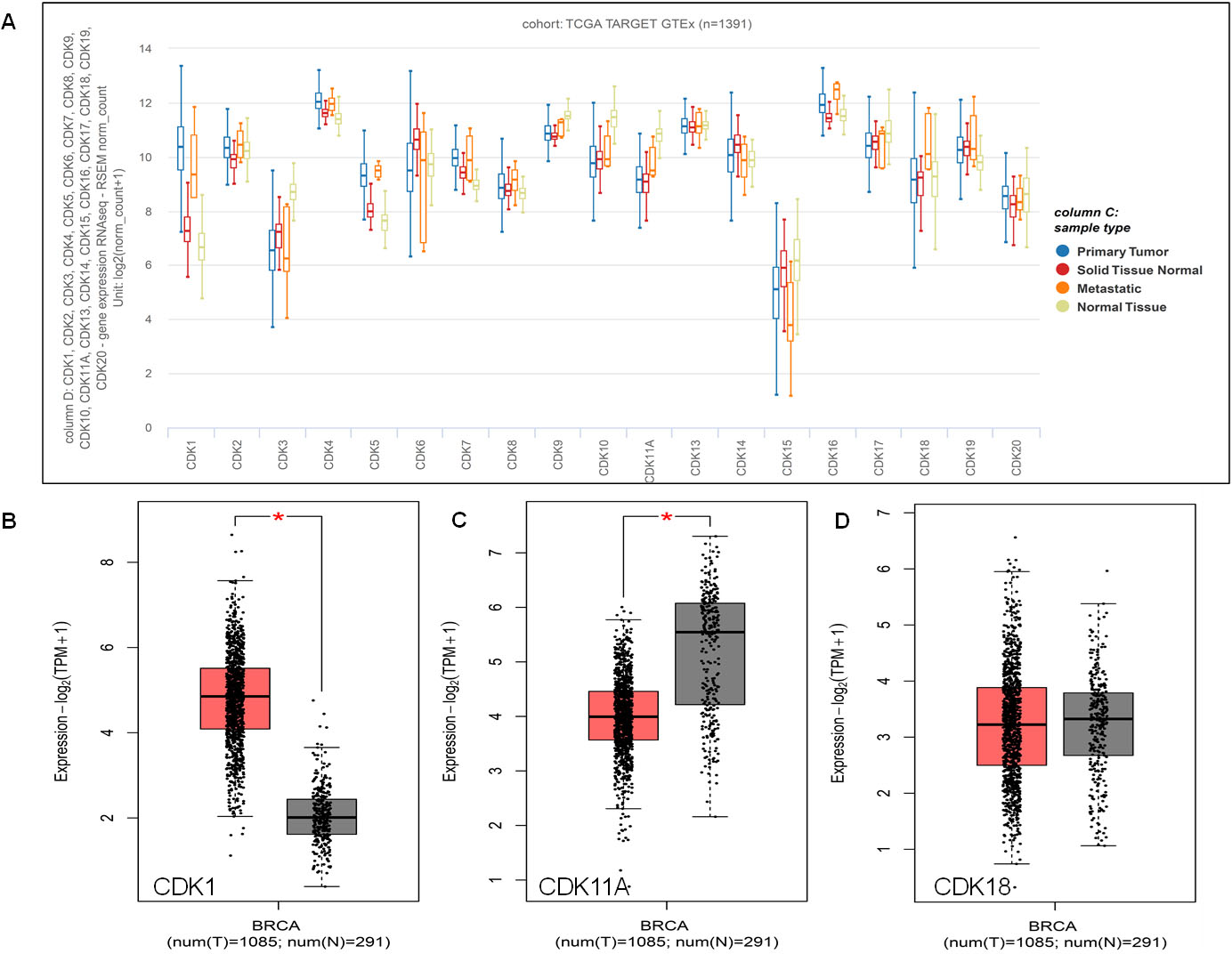
Expression pattern of highly deregulated CDKs in BC subtypes, ethnicity, and age groups. A. CDK1 showed high expression in TNBC patients with high statistical difference, while as CDK18 showed significant deregulated expression between in luminal and TNBC groups but lacked difference in HER2 enriched BC patients. CDK11A lacked statistical significance in expression with BC subtypes. B. CDK1 and CDK18 showed low median expression in Caucasian women, with CDK18 highly expressed in African-American women (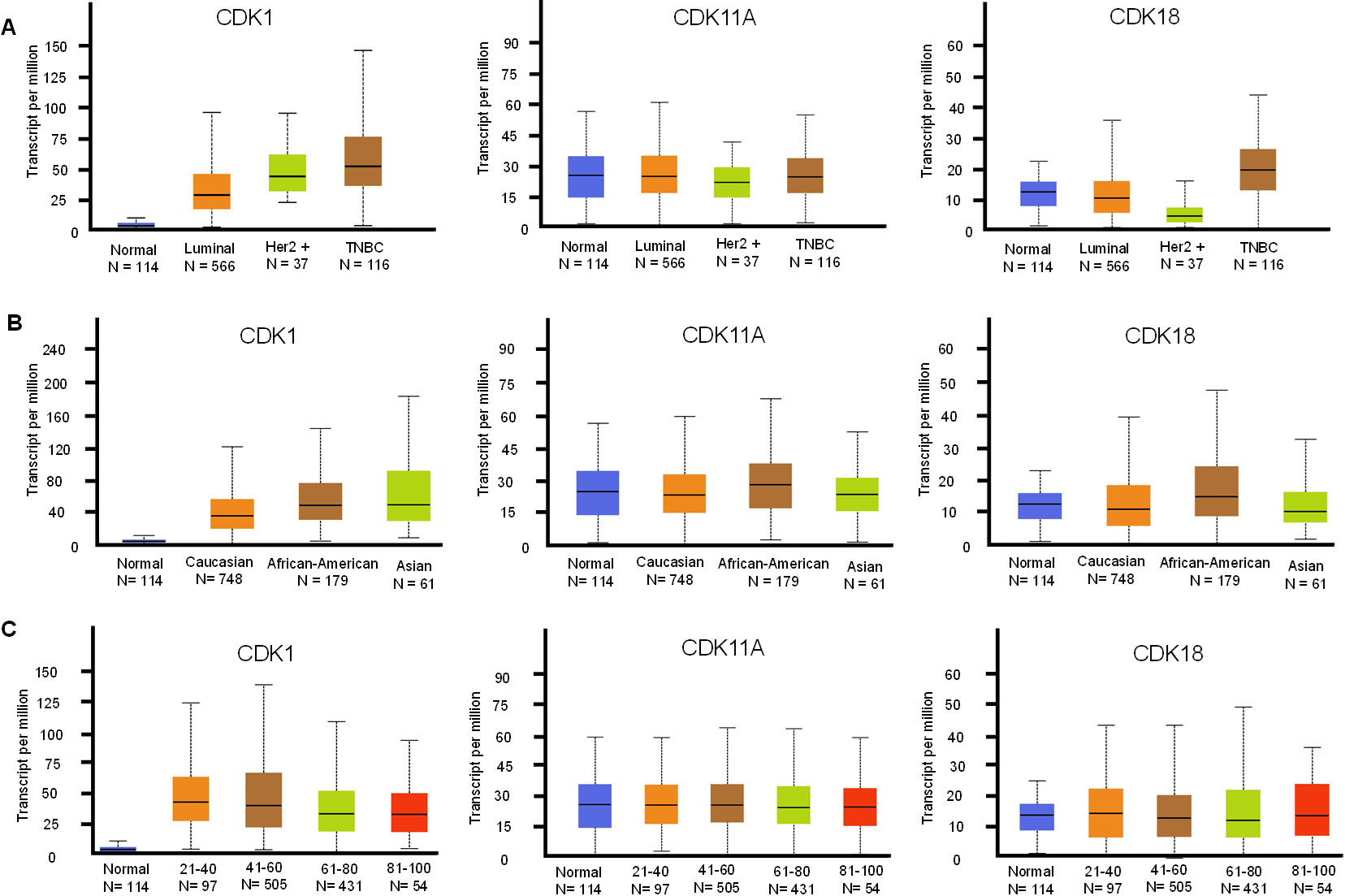
Additionally, we explored the expression pattern of deregulated CDKs in TISCH database, a single sequence database focusing on tumor microenvironment. We performed expression pattern analysis in two BrCa dataset with primary and metastatic tumors. The study revealed that CDK1 is upregulated in metastatic tumors compared to primary tumors. Moreover, CDK1 expression showed heterogeneity in expression pattern across the metastatic tumor cell population. CDK11A and CDK18 showed low expression, however stromal cells in the TME of primary tumor showed high expression of CDK11A compared to CDK18 Fig. 4.
Expression pattern of deregulated CDKs in primary and metastatic tumors using single cell RNA-seq database TISCH.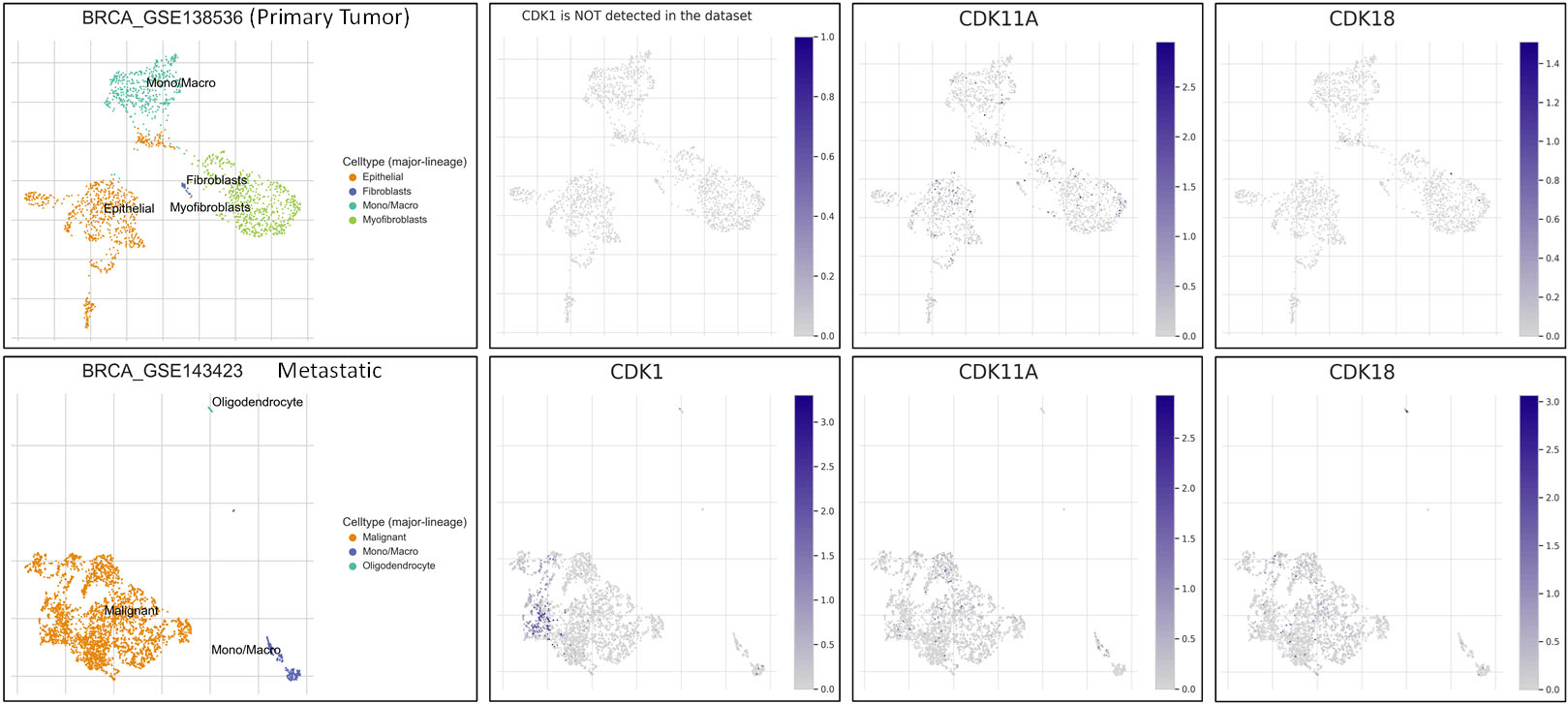
The KM Plotter, which contains gene expression information as well as the survival probability of breast cancer patients, was used to investigate the survival probability of highly deregulated CDKs such as CDK1, CDK11A, and CDK18. BC patients were divided into two cohorts based on median expression, with the two groups termed as low and high expression groups. The relapse-free survival (RFS) analysis was performed on 4929 BC patients for 300 months, and it was found that BC patients with low CDK1 expression have better RFS than patients with higher CDK1 mRNA levels, with a hazard ratio of 1.68 and a log-rank p-value of 1e-16 (Fig. 5a). Furthermore, BC patients with high CDK11A and CDK18 expression had better RFS survival, with HR values of 0.69 and 0.8, respectively. The correlation between overall survival (OS) and CDK expression was investigated in 1879 BC patients, and it was discovered that BC patients with high mRNA levels of the CDK1 gene have poor overall survival compared to patients with low survival, with a hazard ratio (HR) of 1.44 and a log-rank p-value of 0.00016 (Fig. 5b). Low expression of CDK11A and CDK18, on the other hand, is associated with a worse OS in BC patients, with HRs of 0.76 and 0.84, respectively.
Expression of CDKs correlates with survival outcomes. A. Correlation between expression of CDKs with relapse-free survival (RFS) in BC patients. B. Correlation between expression of CDKs with overall survival in BC patients.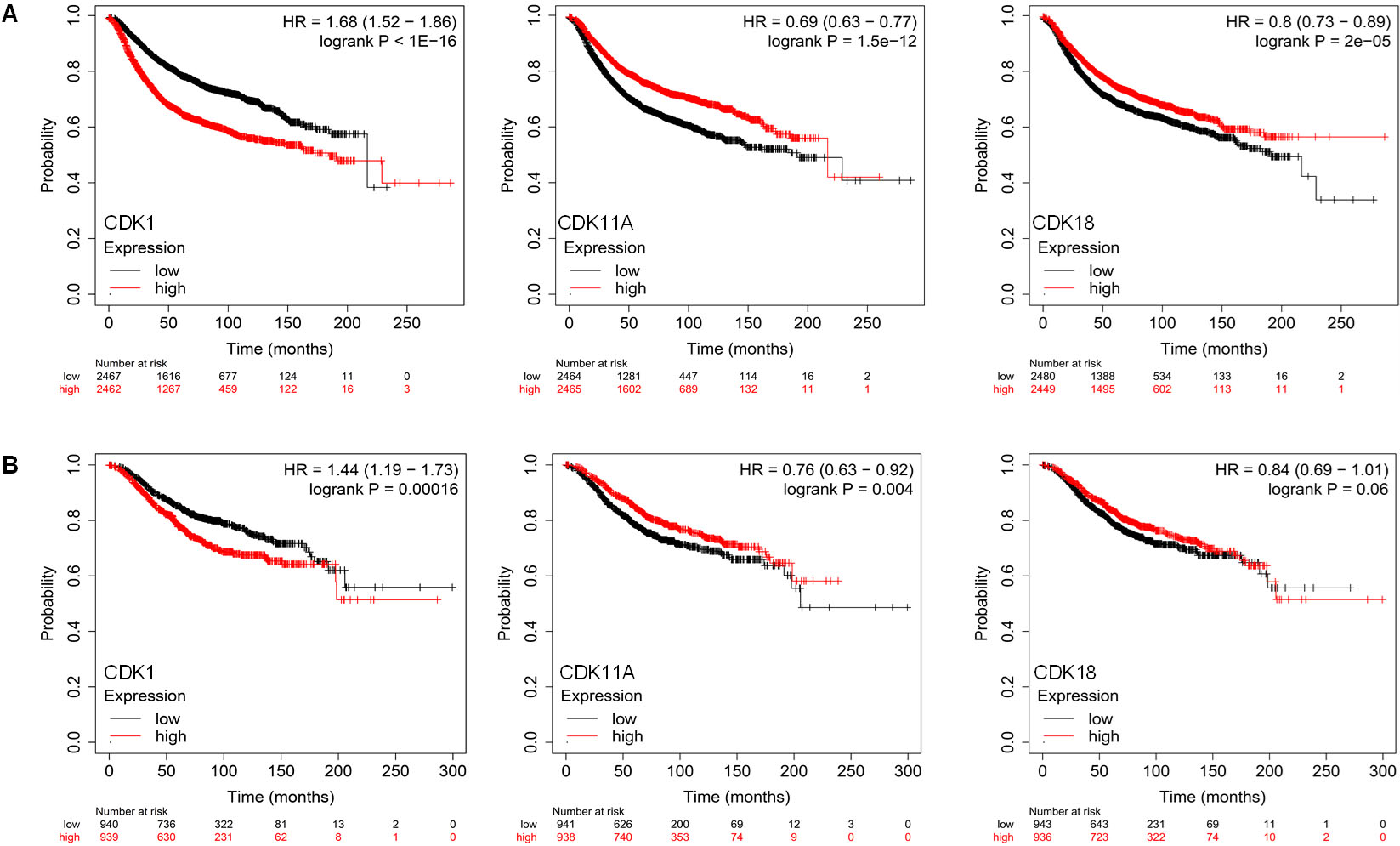
Diagnostic analysis of deregulated CDKs in BC receiving endocrine therapy. A. ROC plot for the Pathological complete response of breast cancer patients receiving endocrine therapy. B. ROC plot for RFS of breast cancer patients receiving endocrine therapy.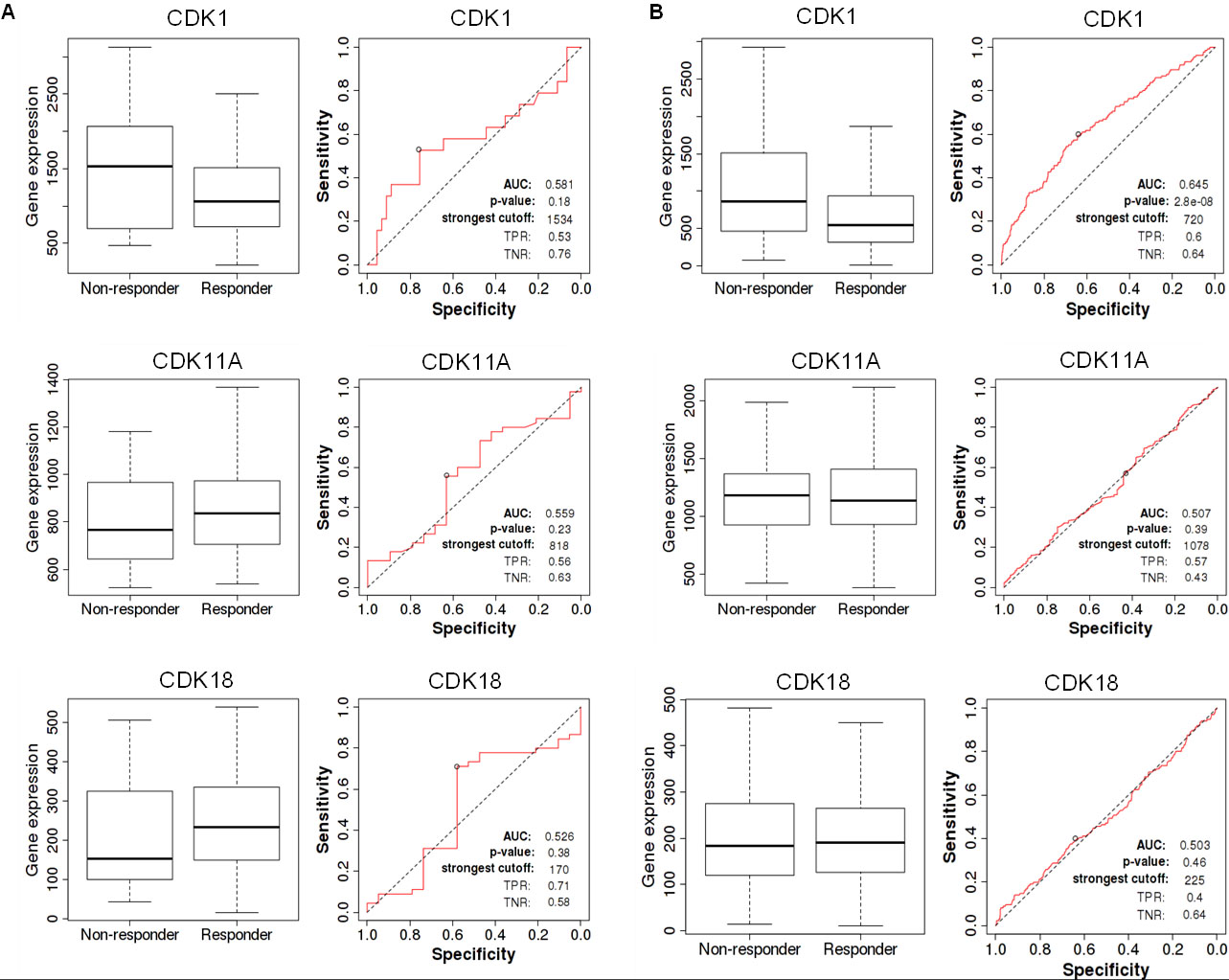
Receiver operating characteristic (ROC) curve analysis revealed several highly predictive values for deregulated CDKs. CDK1 was shown to be an extremely sensitive biomarker for individuals receiving endocrine therapy. On the other hand, CDK11A and CDK18 exhibited limited specificity and sensitivity as biomarkers in patients taking endocrine therapies Fig. 6. On the other hand, patients with high CDK18 expression reacted better to hormonal therapy than patients with low expression. Additionally, CDKs revealed increased sensitivity and specificity as prognostic biomarkers in patients with breast cancer following HER2-directed treatments Fig. 7. Moreover, BC patients receiving HER2-directed therapies with low CDK1 expression had a high pathological complete response with a specificity of 62% and sensitivity of 65 %. CDK11A was found to be less sensitive as a diagnostic marker. CDK18, on the other hand, predicted both pathological complete response and relapse-free survival in BC patients receiving anti-HER2 therapy with high sensitivity and specificity. Besides that, BC patients with high CDK18 expression responded better to anti-HER2 therapy.
Diagnostic analysis of deregulated CDKs in BC receiving anti-HER2 therapy. A ROC plot for the pathological complete response of breast cancer patients receiving anti-HER2 therapy. B. ROC plot for RFS of breast cancer patients receiving anti-HER2 therapy.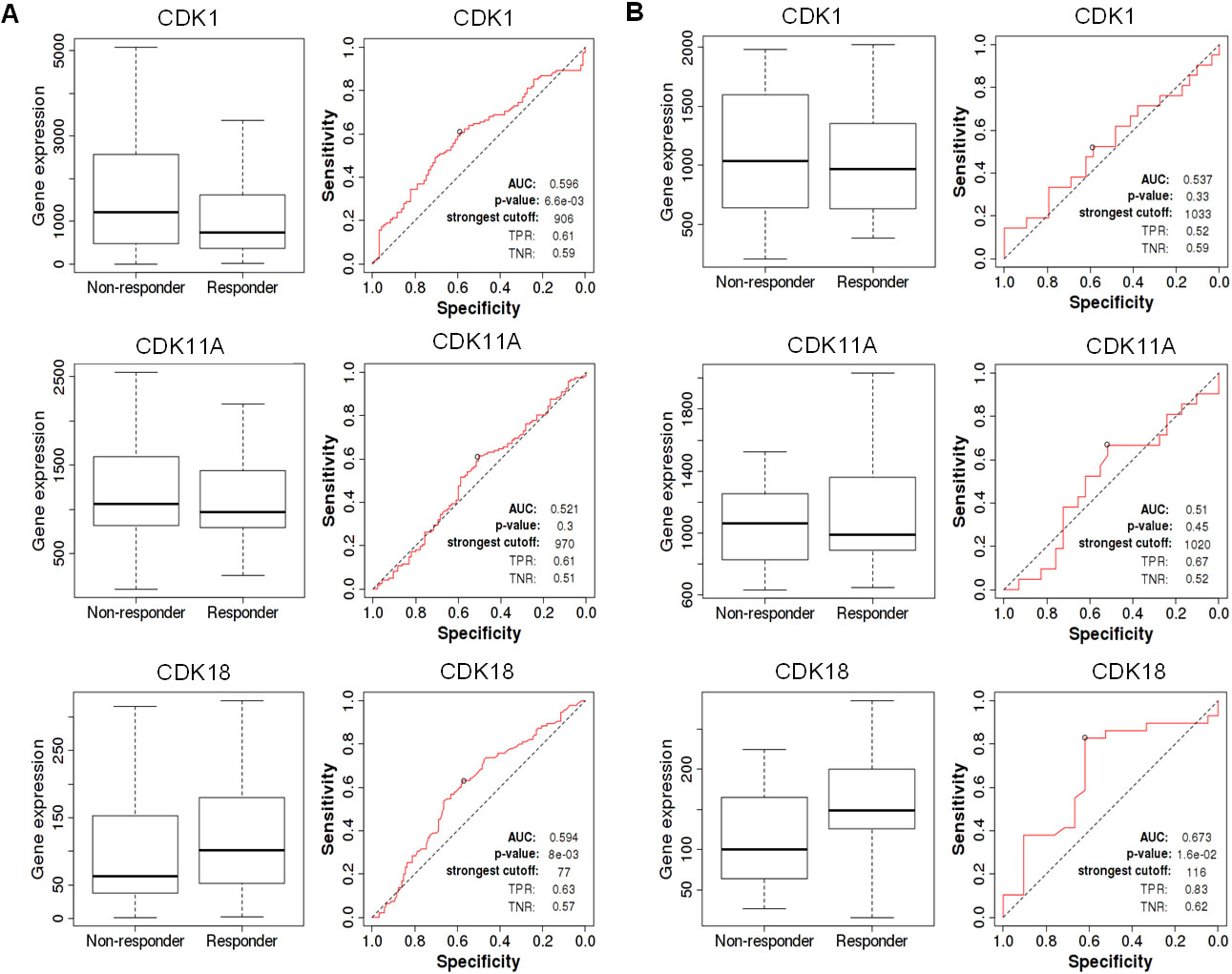
The bc-GenEXMiner database was used to examine the association between severely deregulated CDKs, especially CDK1, CDK11A, and CDK18, and common clinical-pathological characteristics in breast cancer patients. CDK1 expression was shown to be significantly higher in breast cancers lacking hormone receptors (ER and PR) (
Clinicopathological parameters and CDKs in breast cancer. bC-GenEXminer analysis of deregulated CDKs showed a high association with clinicopathological parameters of BC. A. CDK1, B. CDK11A, C. CDK18.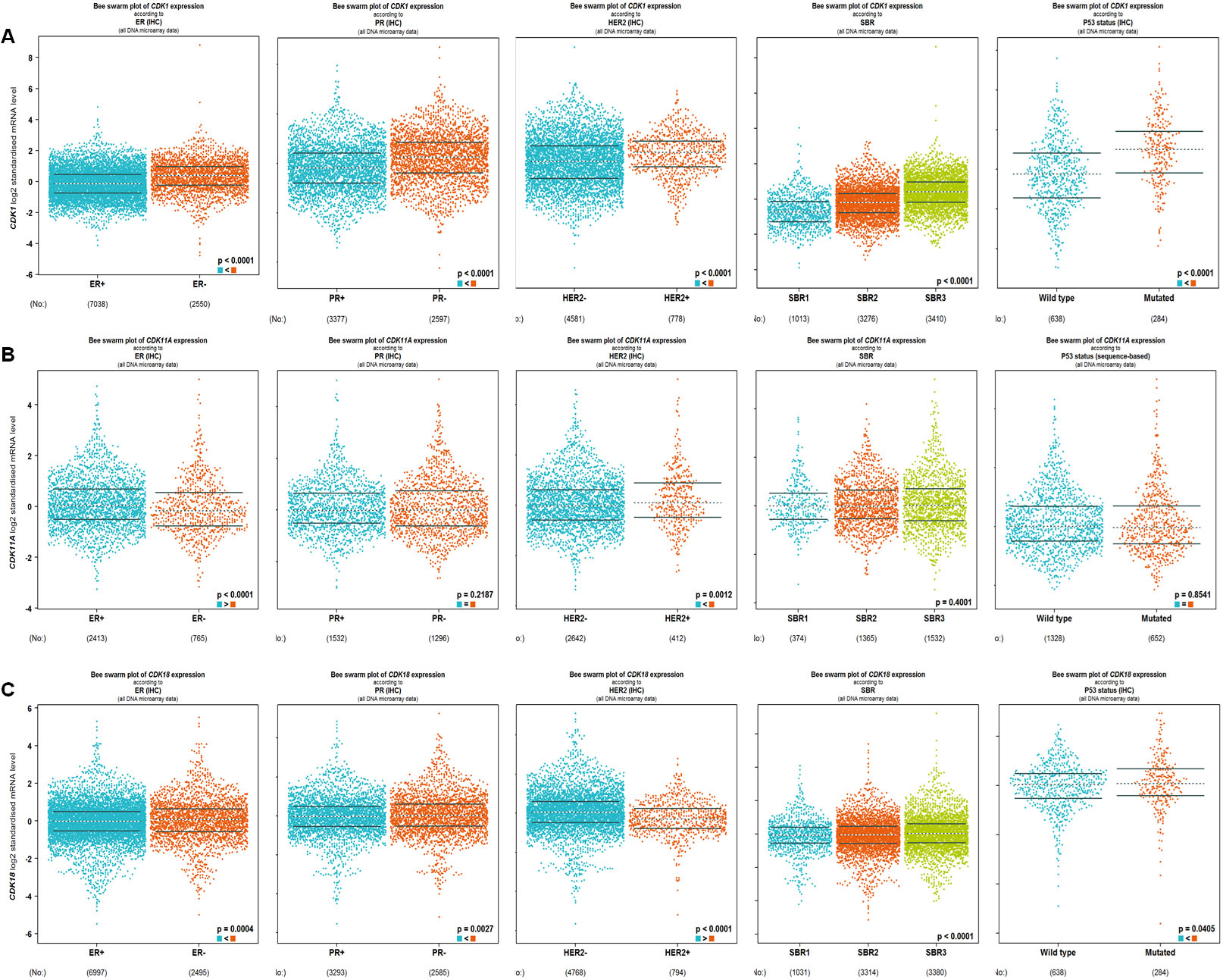
CDK11A expression was considerably lower in patients with ER-negative breast cancer and much elevated in tumors enriched in HER2. CDK11A expression, however, did not vary substantially from PR, SBR grade, or p53 status. CDK18, on the other hand, was shown to be significantly downregulated in patients with breast cancer who expressed hormone receptors (ER, & PR), demonstrating a strong relationship with hormone receptor negativity. CDK18 mRNA levels were discovered to be low in BC patients with HER2 amplification and were observed to correlate with SBR3 grade. In BC patients with wild type 53, CDK18 expression was shown to be substantially linked with p53 status.
The PPI network was constructed using the STRING database and the confidence score of the interactions was set as
Protein-protein interaction and Gene-gene interaction analysis of deregulated CDKs. A. Protein-protein interaction (PPI) network of CDK1 and CDK11A gene constructed with STRING tool had. B. MCODE plugin analysis created a significant module in the constructed network with 31 nodes and 394 edges. C. The top 5 hub genes were identified using the cytoHubba tool in Cytoscape software. D. Gene-gene interaction network of deregulated CDKs.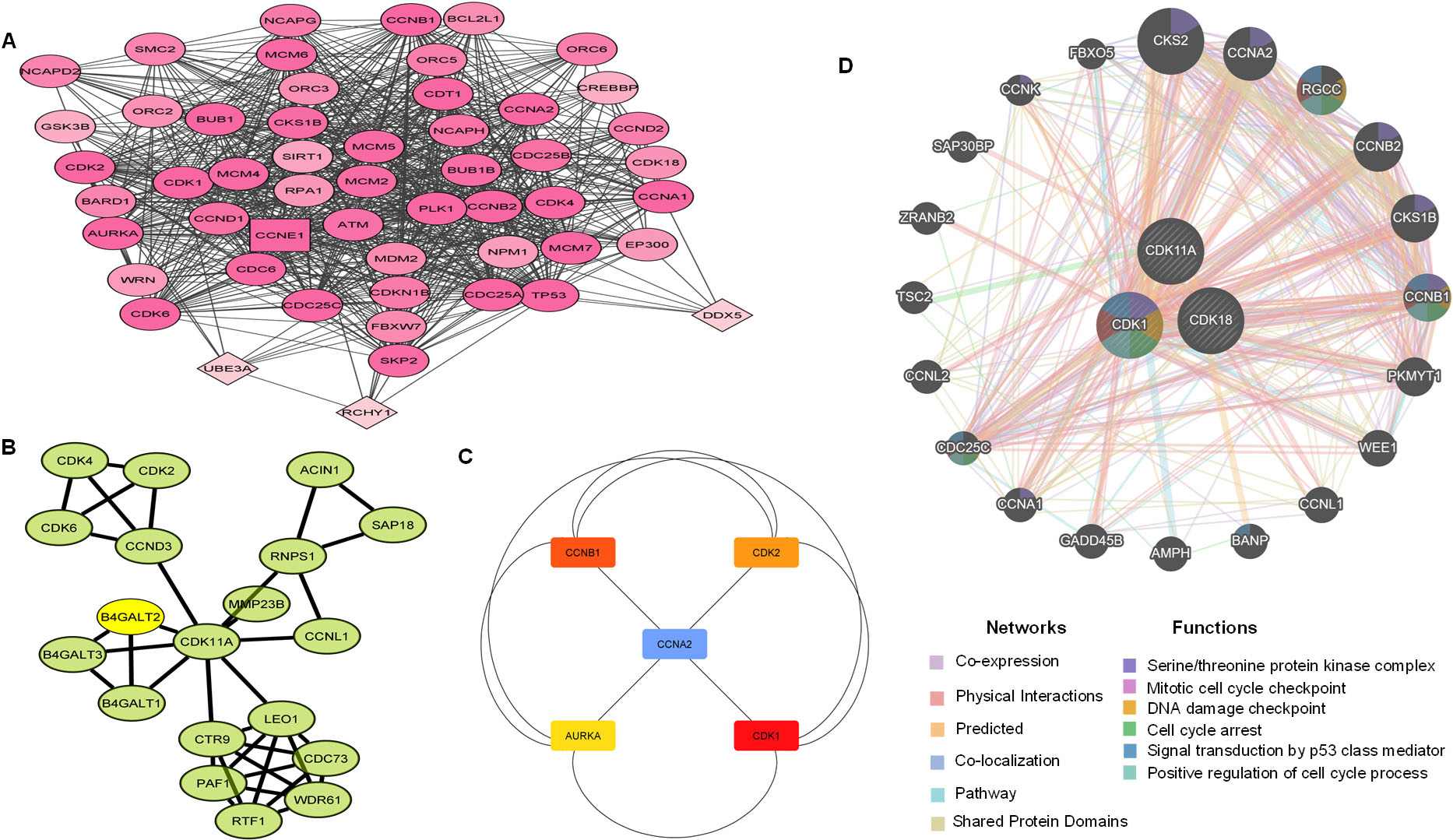
CDK11A formed a significant PPI network with 14 nodes and 30 edges. CDK11A showed direct interactions with LEO1, a component of the PAF1 complex (PAF1C), which has multiple functions during transcription by RNA polymerase II and is implicated in the regulation of development and maintenance of embryonic stem cell pluripotency. CDK11A, also showed linkage with CCNL1 (Cyclin-L1), involved in pre-mRNA splicing and a prime candidate proto-oncogene in head and neck squamous cell carcinomas. CDK11A however didn’t show any interactions with CDK1 or CDK18, indicating that CDK11A acts independently of these two CDKs Fig. 9B.
Furthermore, using the cytohubba plugin, the top five genes regarded as hub genes in the constructed PPI network based on degree were found out to be Aurora kinase A (AURKA), Cyclin A2 (CCNA2), CDK2, CCNB1, and CDK1. The advanced options set as degree cut-off
We further analyzed gene-gene interactions of CDK1, CDK11A, and CDK18, and it was found that deregulated CDKS show high interactions with wee1, CDC25C, CCNB1, and other genes involved in the regulation of the cell cycle. Gene-gene interaction analysis further revealed that deregulated CDKs are central to serine/threonine protein kinase complex, mitotic cell cycle checkpoint, DNA damage checkpoint and signal transduction by p53 class mediator.
Using the Enrichr database, we identified signaling pathways and gene ontology features related to CDK1. GO analysis revealed that CDK1 was highly enriched in biological processes among the three GO terms. The CDK1 gene was found to be enriched in golgi disassembly, positive regulation of the G2/M transition of the mitotic cell cycle, regulation of glial cells, regulation of embryonic development, and other biological processes Fig. 10A. In molecular function (MF) terms, CDK1 showed enrichment in histone kinase activity, protein kinase activity, phosphotransferase activity, RNA Poly II carboxy-terminal domain kinase activity, etc. Fig. 10B. In the cellular compartment group, the suggested GO was primarily enriched in spindle microtubule, mitotic spindle, nuclear chromosome part, centrosome, etc. Fig. 10C.
Gene enrichment and pathway analysis. (A)–(C) Gene Ontology analysis of the BP, CC, and MF and, (D) KEGG pathway enrichment analysis of CDK1, CDK11, and CDK18 genes in breast cancer.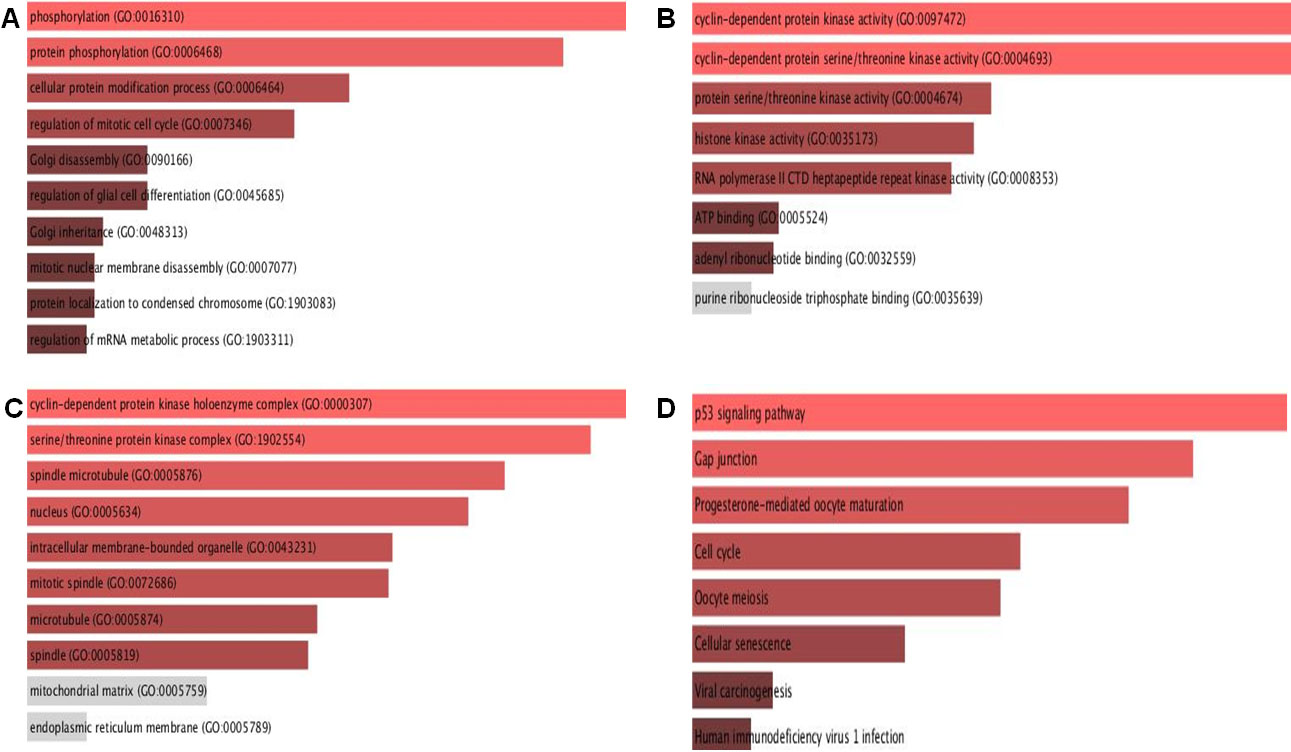
For pathway studies, the KEGG database was evaluated. According to KEGG Human 2019, CDK1 is associated with the cell cycle, P53 signalling pathway, progesterone mediated oocyte maturation, cellular senescence, and cell death pathways Fig. 10D.
Literature-based functional analysis of CDK1 in BC.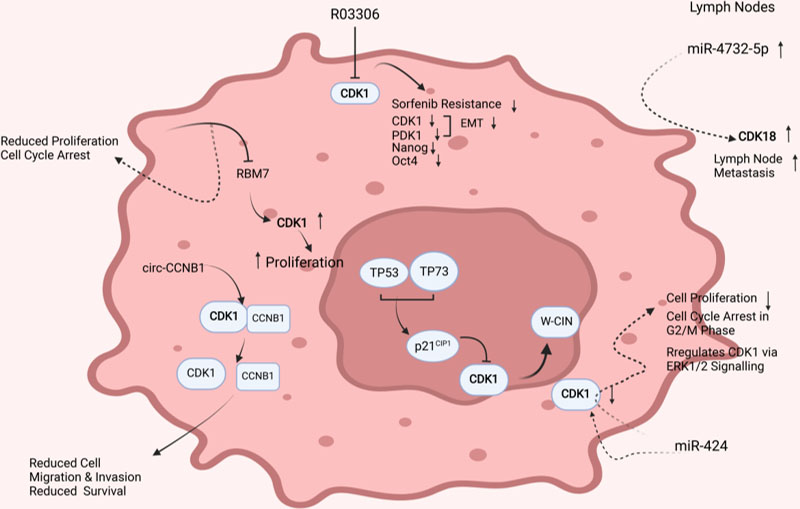
Additionally, a literature-based functional analysis of the deregulated CDKs demonstrated that CDKs regulate key aspects of breast cancer growth and metastasis Fig. 11. CDK1 was found to be central to tumor cell proliferation. Cellular knockdown of RBM7 (RNA binding motif protein 7) reduced CDK1, resulting in decreased cell proliferation and cell cycle arrest in the G1 phase. Besides, circular RNAs have been reported to regulate CDK1 activity in BC. circ-Ccnb1 was found interacting with CCNB1 and CDK1 complex and inducing dissociation of the CCNB1-CDK1 complex, resulting in reduced cell migration and invasion potential of tumor cells. RO3306, a CDK1 inhibitor, significantly reduced CDK1, PDK1, and
Breast cancer is the leading cause of tumor-related deaths in women and one of the most common incident malignancies [2]. Though advances in early detection and advances in therapies have improved the OS and RFS of breast tumor patients, the development of metastatic and resistant tumors requires urgent focus [24, 25]. Also, tumors are complex and don’t merely comprise the highly proliferating tumor cells, but also the surrounding cellular and non-cellular components of the tumor defined as the tumor microenvironment (TME) [26, 27]. Tumor cells and the surrounding stroma influence each other through secreted factors and physical interaction and enhance tumor progression, metastasis, and therapeutic resistance [27]. The stromal cells have been found to induce epigenetic changes in the tumor cells and influence gene expression, aiding in tumor progression and affecting prognosis [28].
The cell cycle is an evolutionarily conserved process, a requisite and regulated phenomenon in mammalian cell growth and development. Abnormalities in the cell-cycle control system are one of the hallmarks of human cancers, including BC [29]. Several studies have analyzed the role of CDKs in controlling the growth, development, and survival outcomes of tumor patients. However, a deep understanding of the role of CDKs in tumor biology is needed to develop rationale therapies for the betterment of cancer patients [30]. In the present study, we employed a bioinformatic approach to unravel the expression pattern, prognostic and diagnostic significance, and functional role of the deregulated CDKs in breast tumor patients. Our analysis revealed that CDKs are highly deregulated in BC, with CDK1, CDK11A, and CDK18 deregulated with a log2 FC of
The CDK1 gene was found to be substantially expressed in individuals with BC, and prognostic analysis indicated that increased CDK1 expression was associated with a worse OS and RFS. These data suggest that CDK1 plays a critical role in tumor development promotion. Interestingly, patients with reduced CDK1 expression had a favorable response to endocrine and anti-HER2 treatments, indicating that CDK1 has a high diagnostic potential. Further investigation established a link between CDK1 and the HER2 receptor, since patients with elevated levels of HER2 expressed a high level of CDK1. Additionally, HER2
In contrary to CDK1, CDK11A and CDK18 were considerably downregulated in patients with breast cancer, with both being highly downregulated in HER2
Furthermore, a literature-based survey demonstrated a key role for CDK1 in breast tumorigenicity. For example, miR-424 downregulated in BC was found to be central to CDK1 regulation. Upregulation of miR-424 induces cell cycle arrest and regulates CDK1 expression via ERK1/2 signaling. The expression of CDK1 was found downregulated in cells overexpressing miR-424 with reduced cell survival, demonstrating that targeting CDK1 is a promising candidate in breast tumorigenicity [37]. Also, inhibiting CDK1 with R03306 enhanced sensitivity to sorafenib in resistant models with reduced EMT potential [38].
Additionally, TP53 and TP73 act as tumor suppressors and induce expression of the CDK1 inhibitor, p21
In terms of clinical characteristics, CDK11A showed association with ER and HER2 receptor with ER
BC patients with high expression of CDK18 receiving anti-HER2 therapy showed a high response and had better pCR and RFS and the results were highly significant with
Previously, CDK18 was found to interact with RAD9, a part of the 9-1-1 replication stress signaling complex [3]. This is a critical axis for establishing an appropriate cellular response to replication stress and, by extension, to chemotherapy [44]. CDK18 expression may impact tumor cell biological responses to chemotherapy. The authors used CRISPR activation to enhance endogenous CDK18 to better understand the function of CDK18 amplification in breast cancer. Cells were more prone to accumulating DNA damage after CDK18 amplification, as seen by staining with a biomarker of double-strand breaks, -H2AX. Importantly, -H2AX activation was pan-nuclear, indicating a widespread problem with DNA replication. Furthermore, the response to replication stress caused by restricting nucleotides was reduced in cells expressing high amounts of CDK18. The scientists discovered a defect in ATR signaling activation, as well as an increase in susceptibility to DNA damaging compounds that impede nucleotide synthesis, such as methotrexate or 5-FU [43, 45]. A literature-based survey further revealed that CDK18 is regulated by miR-4732-5p with enhanced expression at metastatic sites such as lymph nodes, indicating that CDK18 at metastatic sites may regulate the migratory potential of breast tumor cells [46].
Together, these findings provide a picture of CDKs in breast cancer, but elucidating the molecular mechanisms involved in CDK regulation will aid in establishing the rationale for CDK modulation in breast cancer research.
Conclusion
In conclusion, our study found that CDK1, CDK11A and CDK18 may serve as possible predictors of OS and RFS in patients with breast cancer following hormonal or anti-HER2 therapy. Additionally, although modulating deregulated CDKs may be a potential strategy for treating BC patients, well-designed prospective pre-clinical and clinical studies are required to elucidate the molecular mechanism behind CDK differential expression and its function in BC.
Author contributions
Conception: MAM.
Interpretation or analysis of data: UM and SS.
Preparation of the manuscript: UM.
Revision for important intellectual content: MAM and BA.
Supervision: MAM.
Funding
This work was funded by the JK Science Technology & Innovation council DST, Govt. of J&K, India with grant No. JKST&IC/SRE/885-87 sanctioned to Dr. Manzoor Ahmad Mir.
Footnotes
Acknowledgments
Mr. Umar Mehraj is a recipient of a Senior research fellowship (SRF) from UGC-CSIR, Govt. of India.
Conflict of interest
The authors declare no conflicts of interest.
