Abstract
BACKGROUND:
Patients infected with a parasite often develop opisthorchiasis viverrini, which often progresses into cholangiocarcinoma (CCA) due to the asymptomatic nature of the infection. Currently, there are no effective diagnostic methods for opisthorchiasis or cholangiocarcinoma.
OBJECTIVE:
The aim of this study was to identify the host-responsive protein that can be developed as a diagnostic biomarker of opisthorchiasis and cholangiocarcinoma.
METHODS:
Plasma samples were collected from non-OVCCA, OV, and CCA subjects, and the proteomes were investigated by LC-MS/MS. Venn diagrams and protein network prediction by STITCH were used to identify the potential biomarkers. The level of candidate protein, the plasma checkpoint protein 1 (Chk1), was measured by indirect enzyme-linked immunosorbent assay (ELISA).
RESULTS:
Chk1 was present in the center of the protein network analysis in both the OV and CCA groups. In addition, the plasma Chk1 levels were significantly increased in both groups (
CONCLUSION:
Chk1 was identified by differential plasma proteomes and was increased in O. viverrini-infected and cholangiocarcinoma-derived plasma samples. Higher levels of plasma Chk1 levels may serve as a potential diagnostic biomarker for opisthorchiasis and cholangiocarcinoma.
Background
Opisthorchis viverrini, a food borne-liver fluke, has been identified as a risk factor for cholangiocarcinoma (CCA) in combination with the consumption of nitrosamines. Both diseases have high incidence rates in Southeast Asia, especially in northeastern Thailand [1, 2]. The incidence ratio of CCA to hepatocellular carcinoma (CCA:HCC) in Thailand is the highest worldwide [2], owing to the presence of O. viverrini-endemic areas [1]. Several studies have demonstrated that opisthorchiasis and nitrosamines are the primary risk factors for CCA in Thailand [3, 4, 5, 6], which differs from non-Southeast-Asian countries where the primary risk factor is primary sclerosing cholangitis (PSC) [7]. To prevent the progression of opisthorchiasis to CCA, patients with opisthorchiasis require an early diagnosis and treatment with an anthelmintic drug named praziquantel. Currently, stool examination for O. viverrini eggs in patients is still the diagnostic gold standard, despite its low sensitivity and specificity. False-negative results commonly occur due to bile duct obstruction in heavy infections or decreased eggs in light infections. Several studies have attempted to diagnose patients by detecting O. viverrini antigens, specific antibodies, or nucleic acids; however, the effectiveness of these methods remains unsatisfactory [8, 9, 10, 11, 12, 13, 14, 15, 16]. Due to the asymptomatic nature of both opisthorchiasis and early CCA, patients with CCA typically present in the late stages of the disease. Therefore, early detection of opisthorchiasis, the opisthorchiasis-CCA transition phase, and the early stages of CCA are crucial for the prevention of advanced CCA. Although several candidate biomarkers for non-opisthorchiasis-induced CCA have been identified, biomarkers for opisthorchiasis-induced CCA remain limited. CA19-9 is currently used as a tumor marker for CCA, but with limited success; sensitivity remains at 44%–72% and specificity at 80%–100% [17]. Leelawat et al. determined that the area under the curve of the receiver operating characteristic (ROC) curve in differentiating CCA from benign biliary tract disease using CA19-9 was 0.81 [18]. Other markers, including matrix metalloproteinase 7, matrix metalloproteinase 9 [19], interleukin-6, trypsinogen, mucin-5AC, soluble fragments of cytokeratin 19, and platelet-to-lymphocyte ratios, are still under investigation [20, 21]. Other biomarkers for opisthorchiasis-related CCA, such as proteomic-based markers, have also been proposed [22, 23, 24].
Overall, effective methods for the diagnosis of opisthorchiasis and early stage CCA remain elusive. Alternative methods for diagnosing opisthorchiasis and CCA include the detection of host response proteins. Host proteins that respond to opisthorchiasis and CCA may be useful to indicate opisthorchiasis and CCA. Therefore, this study aimed to identify a novel biomarker in the host response against O. viverrini and CCA, which had the potential for diagnostic biomarker development. We identified a potential diagnostic biomarker using differential plasma proteome analysis. Furthermore, the level of a candidate protein, the checkpoint protein 1 (Chk1), was evaluated from the plasma samples, and its sensitivity, specificity, and accuracy were analyzed.
Methods
Ethics and subjects
This study was approved by the Human Ethics Committee No. 1 of Thammasat University (045/2562) and the Human Ethics Committee of Udonthani Cancer Hospital (UCH-CT 11/2563). The study was conducted in accordance with the code of ethics set by the Declaration of Helsinki and all its amendments. The subjects were divided into three groups: the control group (non-opisthorchiasis and non-cholangiocarcinoma, non-OVCCA), the O. viverrini infected group (OV), and the cholangiocarcinoma group (CCA). Non-OVCCA subjects were defined as patients who had normal alanine transaminase (ALT) plasma levels and were found to have no O. viverrini eggs after stool examination. OV subjects were defined as patients with stool exams positive for O. viverrini eggs and confirmed to not have any signs of CCA on ultrasonographic examination. CCA subjects were defined as patients with a definite diagnosis of CCA via histopathology.
Sample preparation
We collected 50 plasma samples from each group and used them for plasma proteomic analysis. The total protein concentration of plasma was measured using the DC™ Protein Assay Kit (Bio-Rad) according to the manufacturer’s protocol. To prepare a pooled plasma sample for each group, 10
Sample preparation for gel-free based proteomics
Gel-free digestion or an in-solution-based method was used in this study. Total protein pellets (5
Label-free proteomic quantification and data interpretation
LC-MS/MS analysis of the digested peptide mixtures was performed using the Waters SYNAPT HDMS™ system. The 1D-nanoLC was carried out using the Waters nanoACQUITY UPLC system. Four microliters (100 ng) of an equal amount of digested peptide were adjusted for the peptide amount and injected onto the reverse phase (RP) analytical column (20 cm
Bioinformatics
LC-MS/MS datasets for label-free quantification were also evaluated. Relative comparisons of peak intensities of each peptide ion in all datasets of LC-runs were analyzed using the DeCyder MS Differential Analysis software [25]. After the injection (label-free analysis), the expression value was normalized using the R package NormalyzerDE (version 1.1.13) [26], in which quantile normalization was applied to the relative expression data analysis, after adding 1 to all expression values to avoid errors upon log-transformation for visualization. Peak detection (signal/noise ratio
Identification of opisthorchiasis viverrini and cholangiocarcinoma related proteins
The GenInfo Identifier (GI) numbers, a unique number that identifies a specific sequence, of the plasma proteome-derived proteins of the non-OVCCA, OV, and CCA proteomes were used for a Venn diagram analysis using Jvenn (
Evaluation of the checkpoint protein 1 (Chk1) protein in plasma by ELISA
From differential non-OVCCA, OV, and CCA proteomes, Chk1 was remarked and selected as a potential biomarker. Evaluation of the Chk1 as a biomarker was performed using an indirect ELISA. The 1:3200 diluted plasma samples were used, and each sample was performed in duplicate. One hundred microliters of the 28, 32, and 32 diluted plasma samples from the non-OVCCA, OV, and CCA groups, respectively, in coating buffer (100 mM NaHCO
Statistical and diagnostic analyses
Statistical analyses were performed using IBM SPSS Statistics for Windows, version 22.0. (IBM Corp, Armonk, NY, USA). To determine whether there were statistically significant differences between the ELISA values of each group, one-way analysis of variance (ANOVA) and subsequent Scheffe post-hoc test were performed. The data were tested for normal distributions using the Kolmogorov-Smirnov test. The dot plot and the area under the ROC curve (AUC) were analyzed using GraphPad Prism 8. The diagnostic parameters, including sensitivity, specificity, positive likelihood ratio (PLR), negative likelihood ratio (NLR), positive predictive value (PPV), negative predictive value (NPV), and accuracy were calculated using MEDCALC (
The summary of identified protein number of each proteome
The summary of identified protein number of each proteome
PCA analysis of non-OVCCA, OV, CCA proteomes. Each dot represents each replicate. The circles are drawn to cover the area of triplicate of each group.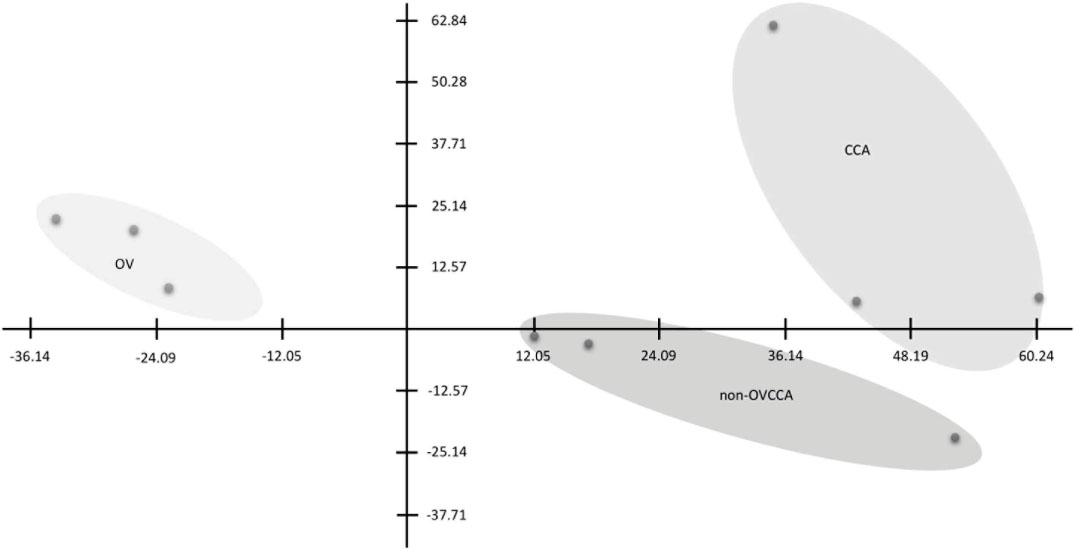
Analyses of plasma proteome
A gel-free LC-MS/MS method was used in this study. A total of 1,822 plasma proteins were identified (Additional File 1). The numbers of retrieved GI numbers, and the number of retrieved UniProt IDs identified from non-OVCCA, OV, and CCA plasma proteomes were 1,758, 1,718, and 1,768 proteins respectively, and are summarized in Table 1. The highest number of identified plasma proteins was found in the CCA group (1,768 proteins), while the lowest number was found in the OV group (1,718 proteins). The heat map and clustering of triplicates of each group were analyzed (shown in Additional File 2) and the triplicates of each group were individually clustered (Additional File 2). The non-OVCCA plasma proteome was more closely related to the CCA than to OV plasma proteomes. The PCA was plotted, and the results were individually separated (Fig. 1) in each group, which agreed with the heat map (Additional File 2). To classify the overall protein task in the cellular machinery, the proteome dataset was subjected to a Gene Ontology analysis by PANTHER according to their biological function. Gene ontology showed similarity in the number and percentage of each molecular function category (Fig. 2).
Gene ontology of molecular function category of non-OVCCA, OV, and CCA proteomes. The numbers indicate protein number tha belongs to each molecular function category.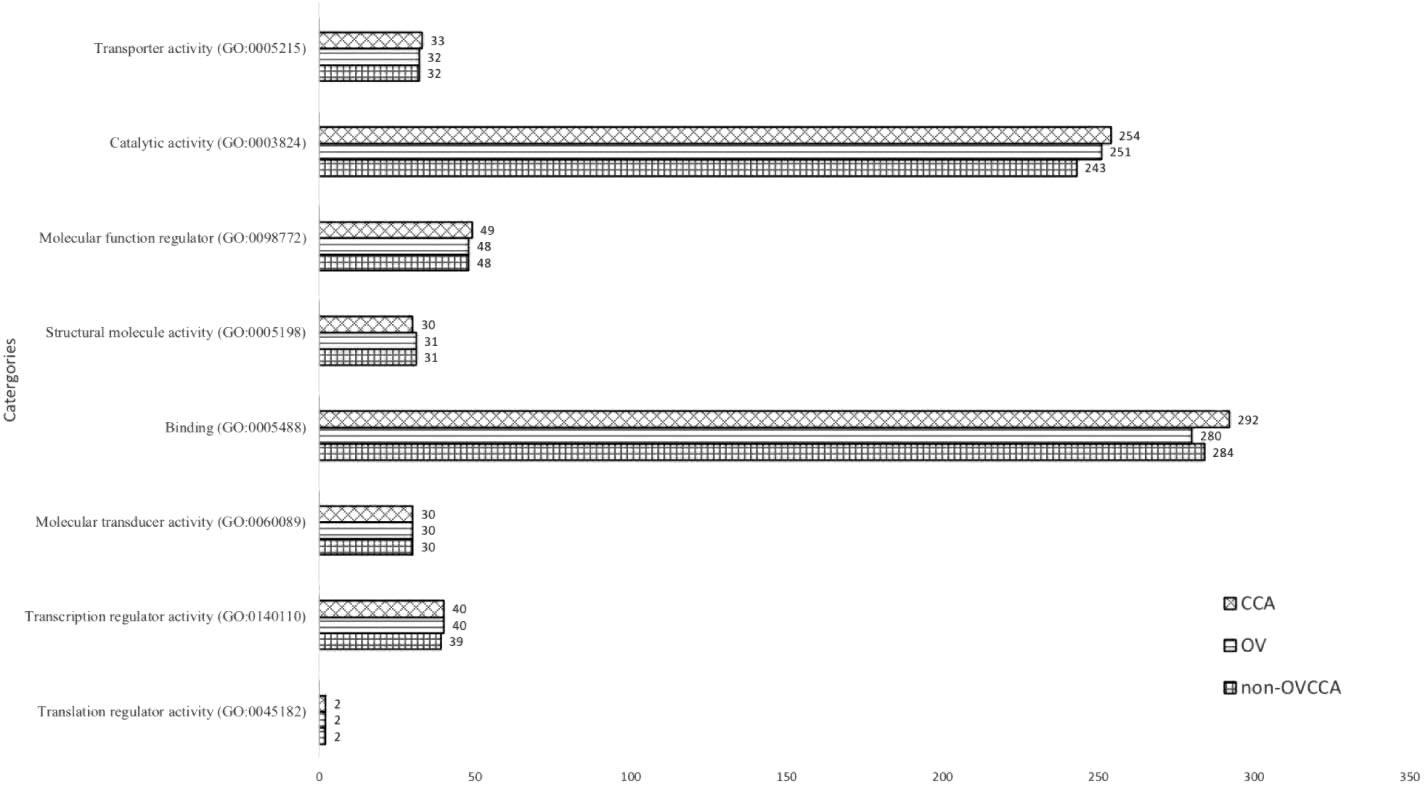
A Venn diagram of the OV against non-OVCCA was performed (Fig. 3). The results showed that 1,663 proteins were shared between the non-OVCCA and OV. The 46 and 86 proteins were group-specific proteins of the OV and non-OVCCA proteomes, respectively (Fig. 3). To investigate the host response proteins against opisthorchiasis, the protein-protein interactions of 46 OV-specific proteins were predicted. The results showed a network of cell checkpoints and proliferation-related proteins which included the checkpoint protein 1 (Chk1), phosphatidylinositol-4-phosphate-3-kinase (PIK3C2B), proto-oncogene with serine/threonine kinase (PIM1), mitogen-activated protein kinase 1 (MAPK1), and vascular cell adhesion molecule 1 (VCAM1). Prediction of the protein and chemical interactions was not performed by STITCH. These results suggested that O. viverrini infection triggered DNA damage-related protein and cell proliferation.
Venn diagram and protein-protein interaction of non-OVCCA and OV proteomes. The protein numbers are indicated in diagram. The protein-protein interaction networks of specific proteins of non-OVCCA and OV are shown in the left and right side, respectively. “X” in the protein ball indicates the predicted functional protein partners. Abbreviations of protein names are available in Additional File 3.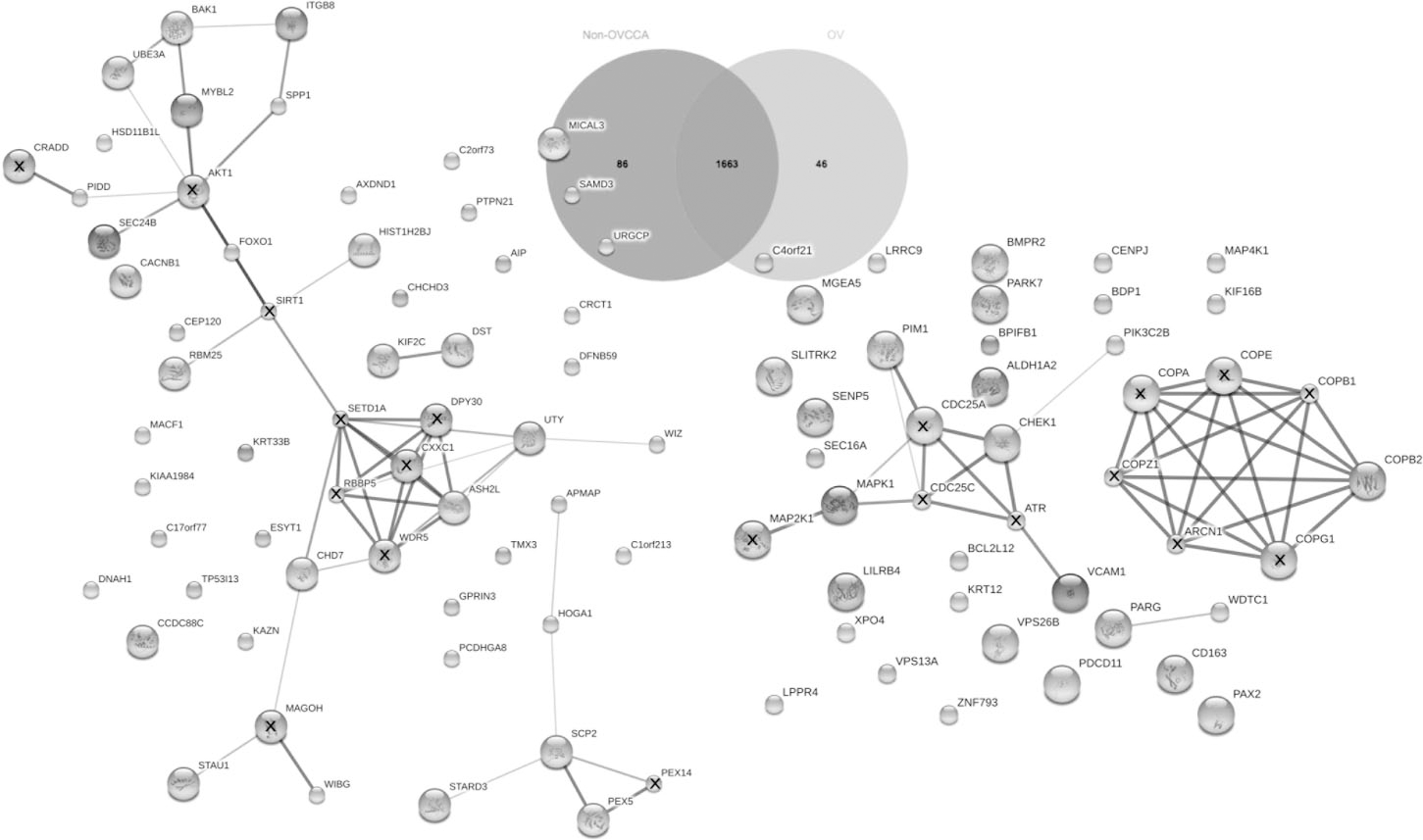
Venn diagram and protein-protein interaction of non-OVCCA and CCA proteomes. The protein numbers are indicated in diagram. The protein-protein interaction networks of specific proteins of non-OVCCA and CCA are shown in the left and right side, respectively. “X” in the protein ball indicates the predicted functional protein partners. Abbreviations of protein names are available in Additional File 4.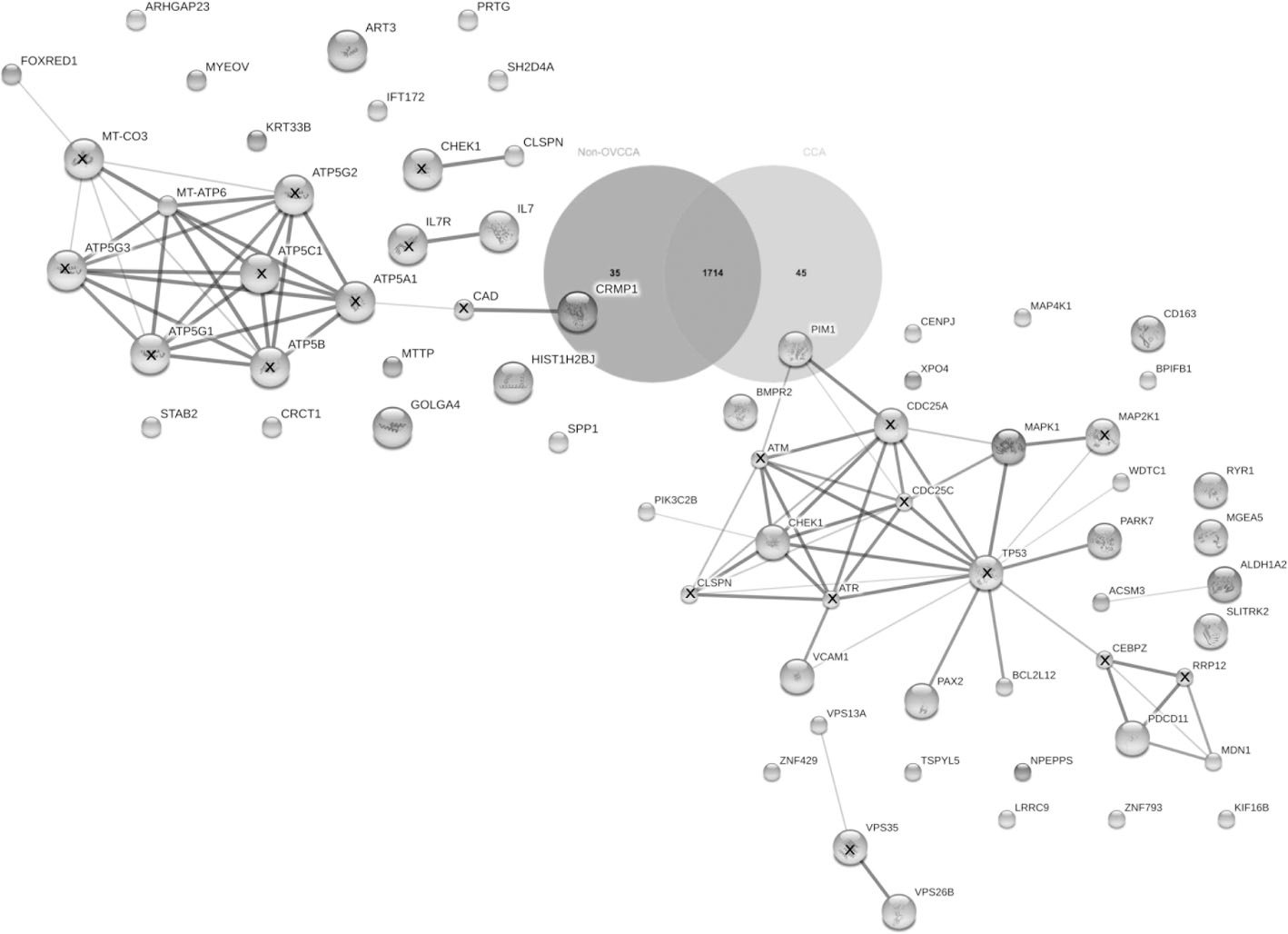
Venn diagram and protein-protein interaction of OV and CCA proteomes. The protein numbers are indicated in diagram. The protein-protein interaction networks of specific proteins of OV and CCA are shown in the left and right side, respectively. “X” in the protein ball indicates the predicted functional protein partners. Abbreviations of protein names are available in Additional File 5.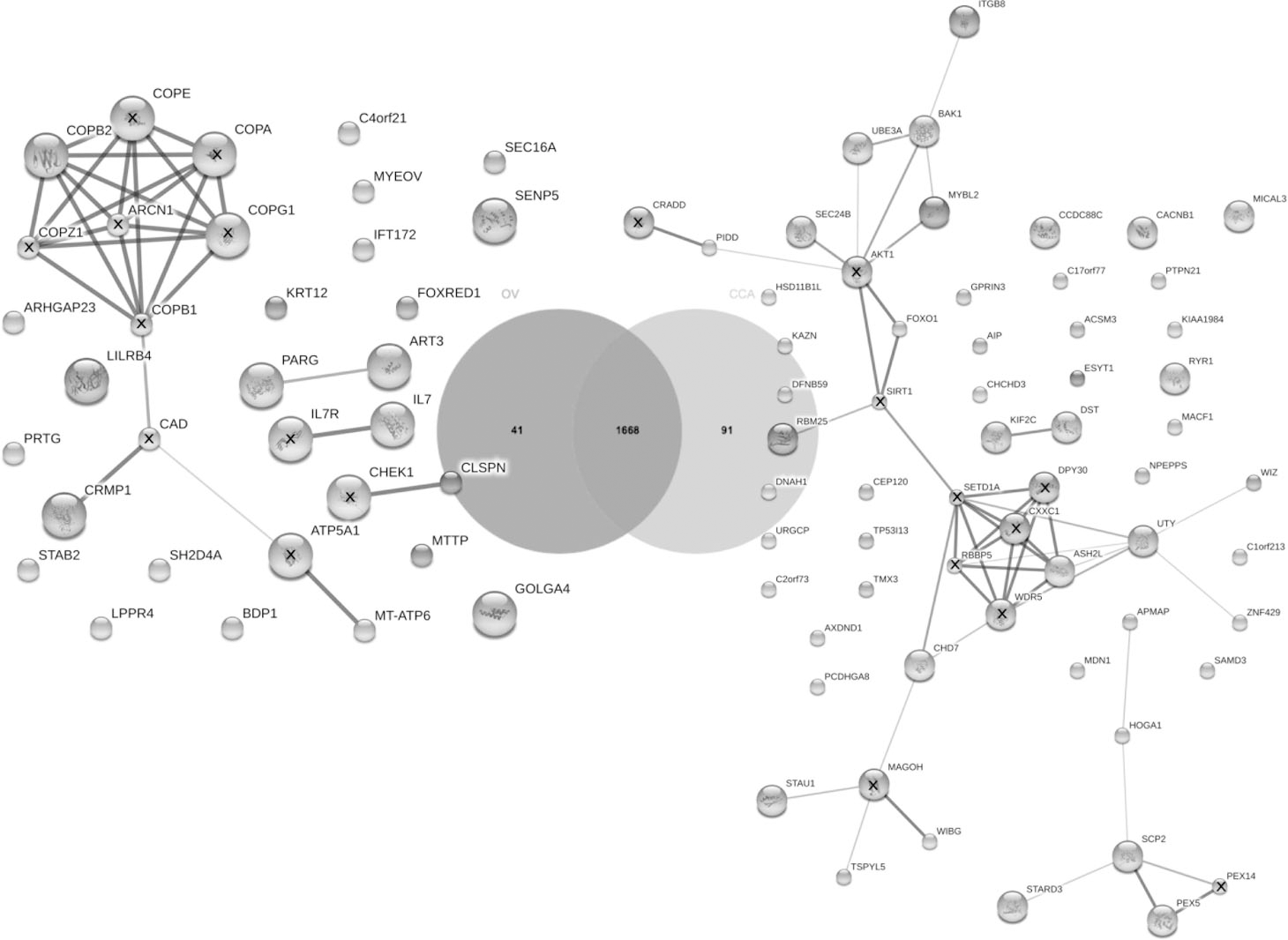
From the Venn diagram, 1,714 proteins were shared between the CCA and non-OVCCA. The 45 proteins were specifically identified in the CCA proteome compared to the non-OVCCA proteome (Fig. 4). Using STITCH, the network of specific proteins identified in the CCA proteome was similar to the OV-specific proteins related to cell signaling of cell proliferation and programmed cell death (Fig. 4). These proteins, which included the mitogen-activated protein kinase 1 (MAPK1), WD, tetratricopeptide repeats 1 (WDTC1), Parkinson protein 7 (PARK7), BCL2-like 12 (BCL2L12), paired box 2 (PAX2), vascular cell adhesion molecule 1 (VCAM1), and checkpoint protein 1 (Chk1), were mainly linked to TP53, the apoptotic pathway protein. Cell proliferation-related proteins, such as phosphatidylinositol-4-phosphate-3-kinase (PIK3C2B) and protooncogene with serine/threonine kinase (PIM1) were also clustered with this network. Furthermore, indirect linkage of midasin homolog (MDN1) and programmed cell death 11 (PDCD11) to TP53 via CCAAT/ enhancer binding protein (CEBPZ) was also predicted. Two vacuolar proteins related to the retrieval of lysosomal enzyme receptors such as IGF2R and M6PR from endosomes to the Golgi were also identified in the CCA plasma proteome. Furthermore, a link between acyl-CoA synthetase medium-chain family member 3 (ASCM3) and aldehyde dehydrogenase 1 family (ALDH1A2) was also observed. Protein and chemical interactions were not recruited using STITCH. These results suggested that changes in the regulation of proteins related to cell signaling of cell proliferation and apoptosis may have accounted for this.
Absorbance at OD
of plasma Chk1 by indirect ELISA
Absorbance at OD
O. viverrini has been identified as a risk factor for cholangiocarcinoma in Thailand. Therefore, the absence and/or presence of proteins in the plasma will be a useful biomarker for differentiating opisthorchiasis from CCA or the transitional phase of O. viverrini infection to CCA. From the Venn diagram, 91 proteins were only found in the CCA plasma proteome, whereas 41 proteins were only found in the OV proteome (Fig. 5). The predicted protein-protein interactions of 91 CCA-specific proteins showed the protein network of apoptotic proteins such as BCL2-antagonist/killer 1 (BAK1), TSPY-like 5 (TSPYL5), and p53-induced death domain protein (PIDD). It has also been linked to several chromatin-related proteins and transcription regulation, which include absent, small, or homeotic-like (ASH2L), widely interspaced zinc finger motifs (WIZ), zinc finger protein 429 (ZNF429), helicase DNA binding protein 7 (CHD7), staufen RNA binding protein homolog 1 (STAU1), within bgcn homolog (WIBG), v-myb myeloblastosis viral oncogene homolog (avian)-like 2 (MYBL2), and RNA binding motif protein 25 (RBM25). Lipid metabolism and transfer-related proteins, such as adipocyte plasma membrane-associated protein (APMAP), StAR-related lipid transfer (START) domain containing 3 (STARDS), and sterol carrier protein 2 (SCP2), were also predicted for their relationship in the protein network. The link between two proteins, cytoskeletal proteins, which include kinesin family member 2C (KIF2C) and dystonin (DST), was also observed. Interestingly, the predicted protein network linked to AKT1, the upregulated proteins in CCA, were related to cell proliferation (Fig. 5). For CCA-absent proteins or OV-specific proteins, COPB2 linked to collapsin response mediator protein 1 (CRMP1) and mitochondrially encoded ATP synthase 6 (MT-ATP6) via predicted functional partners added by STITCH: carbamoyl-phosphate synthetase 2 (CAD) and ATP synthase, H
Evaluation of the checkpoint protein 1 (Chk1) protein in the plasma
Indirect ELISA of plasma Chk1. Each circle is an absorbance of each subjects. The absorbance value is shown in the 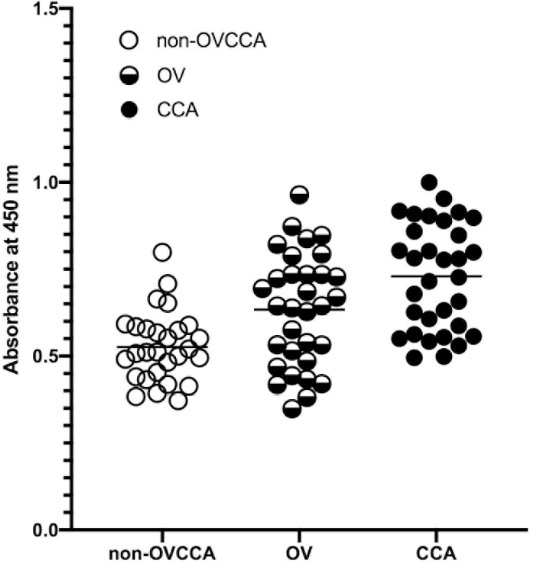
The statistical analysis of non-OVCCA, OV and CCA
The calculated diagnostic parameters of OV and CCA
Prediction of protein-protein interactions by STITCH suggested that the Chk1 was located in the center of the OV and CCA protein networks. These results suggested that the Chk1 played a role in pathogenesis during O. viverrini infection and cholangiocarcinogenesis. Therefore, the plasma Chk1 levels were investigated and evaluated. The average absorbance values of the non-OVCCA, OV, and CCA groups are summarized in Table 2. The ELISA values were normally distributed (
ROC curve of non-OVCCA vs OV. The sensitivity% (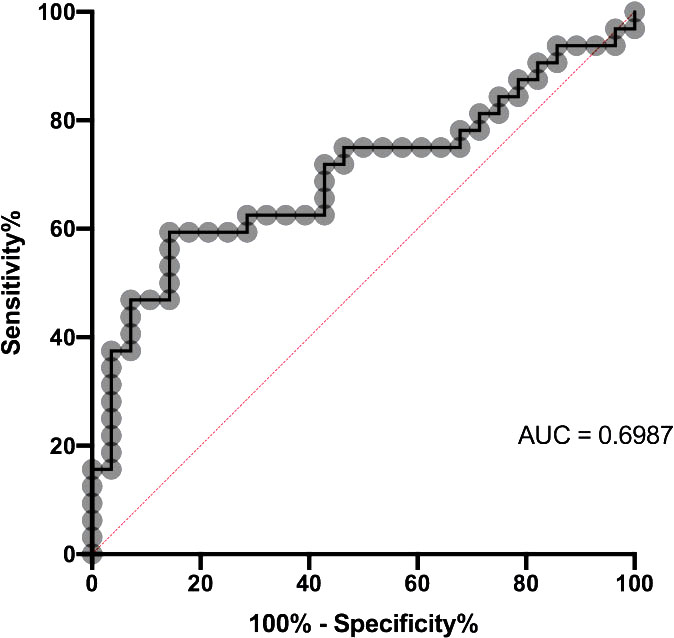
ROC curve of non-OVCCA vs CCA. The sensitivity% (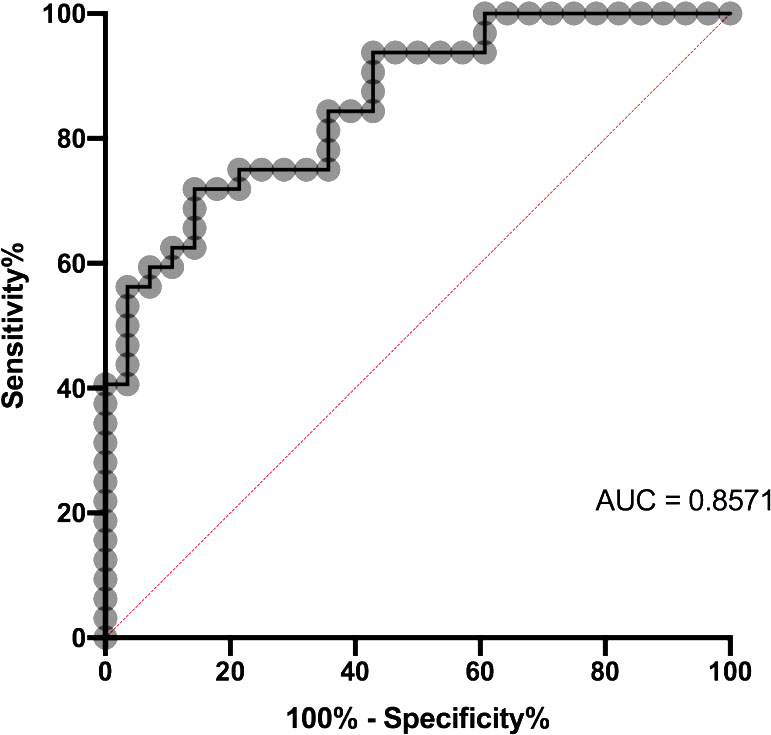
In this study, the differential plasma proteomes, Venn diagram, and protein-protein interactions by STITCH were used to investigate the host response proteins against opisthorchiasis and CCA. Moreover, differential plasma proteomes have also been used as tools in the identification of novel biomarkers for O. viverrini and CCA. From the OV plasma proteome results, the Chk1, VCAM1, PIK3C2B, MAPK1, and PIM1, which have functions related to the cell cycle, cell proliferation, and cell signaling, were observed in the protein network (Fig. 2). This result suggests that increased cell proliferation, cell cycle checkpoints, and DNA damage should occur during opisthorchiasis. During the O. viverrini infection, O. viverrini excretory/secretory proteins (OV-ES) and extracellular vesicles are released from parasites, which can stimulate bile duct epithelial cells [31, 32, 33, 34]. It has been clearly demonstrated that OvGST in OV-ES stimulates AKT and ERK in NIH-3T3 (fibroblast) and MMNK-1 [35]. Stimulation of AKT and ERK may allow either normal or mutated cells to survive and proliferate [36, 37]. Moreover, activation of AKT can be induced by DNA damage, either by single-or double-stranded DNA [38]. Several studies have shown that O. viverrini infection causes DNA damage, which leads to cholangiocarcinoma [39, 40, 41, 42, 43, 44]. The presence of the Chk1, the cell checkpoint protein in predicting the protein network, should play an inhibiting or stalling role on DNA damage cells to proliferate [45] and allow the damaged cells to repair before entering the cell cycle. Overall, an increase in the Chk1 indicates that DNA damage has occurred and involved in the pathogenesis during the O. viverrini infection. An increase in the plasma Chk1 in shotgun plasma proteomes was found not only in the OV proteome but also in the CCA proteome. These results suggest that DNA damage was initiated during the O. viverrini infection period and might have continued during cholangiocarcinogenesis. It is clear that the O. viverrini infection stimulates inflammation and in combination with nitrosamine compounds causing DNA damage [39, 46, 47, 48, 49, 50]. This implies that DNA damage by the carcinogenic liver fluke occurs during an O. viverrini infection, and mutations are accumulated during cholangiocarcinogenesis. This encourages the characteristics of this carcinogenic liver fluke. Recently, a study analyzed and validated the individual serum proteomes of cholangiocarcinoma. At least three potential biomarkers have been proposed, which include S100A9, thioredoxin (TRX), and cadherin-related family member 2 (CDHR2) [51]. In addition, the present study also included a sample of the O. viverrini-infected patients. Interestingly, the Chk1 was present in both the OV and CCA plasma proteomes (Figs 3 and 4). In order to validate the level of plasma Chk1, it was selected for further evaluation. The sensitivity and specificity of cholangiocarcinoma patients were higher than those of patients infected with O. viverrini. Plasma Chk1 may be unsuitable for patient screening due to its low sensitivity. However, its high specificity (85.71%) makes it a potential biomarker in confirming the diagnosis of opisthorchiasis or CCA. However, further investigation should consider a larger sample size to validate the clinical confidence in decision making in the diagnosis of the diseases. In addition, the combination of the plasma Chk1 with other previously identified biomarkers in the diagnosis of CCA may prove to be more effective and should be further investigated [51]. Thus, the application of the plasma Chk1 will shed light on a novel biomarker in the future for diagnostic approaches of O. viverrini-infected and CCA patients. In addition to diagnostic biomarkers, the application of the Chk1 as a prognostic approach is also considered for monitoring treatment regimens, such as the combination of gemcitabine and cisplatin, which is the current recommended treatment for CCA. Gemcitabine targets the Chk1 and induces cancer cell cycle arrest [52, 53]; thus, it is a possible candidate biomarker of prognosis. However, additional studies are required to investigate the prognostic properties of the Chk1. In summary, this study provides novel evidence that the plasma Chk1 is increased in both opisthorchiasis and CCA, and has high specificity and accuracy as a potential diagnostic biomarker of opisthorchiasis and CCA.
Supplementary data
The supplementary files are available to download from
sj-xlsx-1-cbm-10.3233_CBM-210170.xlsx - Supplemental material
Supplemental material, sj-xlsx-1-cbm-10.3233_CBM-210170.xlsx
sj-docx-1-cbm-10.3233_CBM-210170.docx - Supplemental material
Supplemental material, sj-docx-1-cbm-10.3233_CBM-210170.docx
sj-pdf-1-cbm-10.3233_CBM-210170.pdf - Supplemental material
Supplemental material, sj-pdf-1-cbm-10.3233_CBM-210170.pdf
Author contributions
Conception: Teva Phanaksri, Sittiruk Roytrakul, Smarn Tesana, Veerachai Thitapakorn.
Interpretation or analysis of data: Teva Phanaksri, Yodying Yingchutrakul, Sattrachai Prasopdee, Kritiya Butthongkomvong, Thanakrit Sathavornmanee, Veerachai Thitapakorn.
Preparation of the manuscript: Teva Phanaksri, Yodying Yingchutrakul, Sittiruk Roytrakul, Sattrachai Prasopdee, Anthicha Kunjantarachot, Kritiya Butthongkomvong, Thanakrit Sathavornmanee, Veerachai Thitapakorn.
Revision for important intellectual content: Teva Phanaksri, Veerachai Thitapakorn.
Supervision: Smarn Tesana, Veerachai Thitapakorn.
Footnotes
Acknowledgments
This study was financially supported by Thai Government Research Fund through Thammasat University, Fund Contract No.50/2559 and 64/2560 (Dr. Veerachai Thitapakorn) (Year 2016–2017) and Fund Contract No. 66/2560 (Dr. Teva Phanaksri) (Year 2017–2018).
