Abstract
BACKGROUND:
Lung cancer is the leading cause of cancer-related deaths worldwide. With the expectation of improved survival, tremendous efforts and resources have been invested in the discovery of specific biomarkers for early detection of the disease. Several investigators have reported the presence of cancer-associated autoantibodies in the plasma or serum of lung cancer patients. Previously, we used a monoclonal antibody (mAb) proteomics technology platform for the discovery of novel lung cancer-associated proteins.
OBJECTIVE:
The identification of specific protein epitopes associated with various cancers is a promising method in biomarker discovery. Here, in a preliminary study, we aimed to detect autoantibody-leucine-rich alpha-2-glycoprotein 1 (LRG1) immunocomplexes using epitope-specific monoclonal antibodies (mAbs).
METHODS:
We performed sandwich ELISA assays using the LRG1 epitope-specific capture mAbs, Bsi0352 and Bsi0392, and an IgG-specific polyclonal antibody coupled to a reporter system as the detection reagent. We tested the plasma of lung cancer patients and apparently healthy controls.
RESULTS:
Depending on the epitope specificity of the capture mAb, we were either unable to distinguish the control from LC-groups or showed a higher level of LRG1 and IgG autoantibody containing immunocomplexes in the plasma of non-small cell lung cancer and small cell lung cancer subgroups of lung cancer patients than in the plasma of control subjects.
CONCLUSIONS:
Our findings underline the importance of protein epitope-specific antibody targeted approaches in biomarker research, as this may increase the accuracy of previously described tests, which will need further validation in large clinical cohorts.
Introduction
Autoantibodies are generated against endogenous immunogens and, although they are classically associated with autoimmune diseases, they have also proven to be useful tools for the early detection of cancer. Anti-fetal protein New York, esophageal squamous cell carcinoma 1 (NY-ESO-1) [1] or HER2/neu specific antibodies have been detected in the blood of patients with breast and ovarian cancer [2]. Several other antigens that are autoantibody targets have been identified for various cancer types: p53, p62, and HSP70 for hepatocellular carcinoma, p53, HER2/neu, MUC1, and NY-ESO-1 for breast cancer, p53, HSP60, Fas/CD95, and p16 for colon cancer [3], and S100B for lung cancer (LC) patients with early brain metastases [4, 5]. The field is rapidly expanding, as shown by the recent review of de Jonge et al. [6].
Selected characteristics of plasma (sample) donors
Selected characteristics of plasma (sample) donors
LC: lung cancer, ST: subtype, SCLC: small cell lung carcinoma, NSCLC: non-small cell lung carcinoma. *ST: ND indicates that the LC subtype was not determined.
Autoantibodies also show diagnostic value in LC. For example, autoantibodies have been found against p53, SOX1, SOX2, SOX3, and NY-ESO-1 in the plasma or serum of LC patients [3, 7, 8, 9]. In addition, certain autoantibodies have been shown to be good prognostic tools, such as NY-ESO-1, melanoma associated antigen 3 (MAGE3) [10], or MUC1, which negatively correlate with disease progression in non-small cell lung cancer (NSCLC) patients [11]. Human complement factor H (CFH) autoantibody levels have also been found to be significantly higher in the blood of early stage NSCLC patients than late stage NSCLC or control patients [12]. The presence of the ENO1 autoantibody increases the sensitivity of CEA and CYFRA 21-1 in the detection of NSCLC patients [13]. As the measurement of autoantibodies against individual tumor-associated antigens was shown to have low diagnostic performance (at high specificity, the sensitivity is low), a panel approach was necessary to provide additional clues for the early detection of LC [14]. Indeed, the EarlyCDT-Lung blood test is based on autoantibody detection against seven tumor-associated antigens [15, 16, 17, 18, 19] and the autoantibody panels are continuously updated in the hopes of improving the diagnostic performance [20, 21]. Previously, we reported a monoclonal antibody (mAb) proteomics technology for the discovery of novel LC-associated proteins. We found that a panel of plasma proteins, consisting of haptoglobin (HP), complement component 9 (C9), alpha-1-antichymotrypsin (
Autoantibodies are generally detected by ELISA with direct antigen coating (DAC-ELISA; p53, NY-ESO-1, livin, and survivin) or other immunoassays, such as serological analysis of recombinant tumor cDNA expression libraries (SEREX, SOX family proteins) or serological proteome analysis (SERPA; annexin I and annexin II, alpha-enolase, CFH) [7], in which antigens are immobilized on a surface without orientation. Thus, little or no information is obtained concerning the nature of the epitopes recognized by the autoantibodies. Phage display, fluid-phase immunoassays, and antigen microarrays have also been used to investigate autoantibodies with a diagnostic value in various diseases. However, these techniques also suffer from the same shortcomings in terms of the absence of information on the epitopes involved in autoantibody binding [36, 37].
Here, we report our efforts and preliminary results to detect LRG1-specific autoantibody immunocomplexes in plasma samples that contain autoantibodies against known LC-associated autoantigens, such as survivin and/or CFH [12, 38]. Autoantibodies are present in the sera both as free immunoglobulins and as immunocomplexes; measuring either could be of diagnostic value, but the two measurements significantly differ from each other both conceptionally and technically. DAC is the method of choice for autoantibody detection. With this approach, the importance of the epitopes, especially in the context of a neoepitope-specific autoantibody response, has been described [39] and such studies included the use of autoantigen-derived peptides for the broadest possible detection of autoantibodies [40]. Sandwich immunoassays are used for the detection of immunocomplexes, with capture reagents recognizing epitopes that are not blocked by the autoantibodies. The mAb proteomics technology platform that we developed previously resulted in the generation of several antibodies recognizing the same protein via different epitopes. We chose two (Bsi0352 and Bsi0392) LRG1-specific mAbs with non-identical epitope specificity and used them in (epitope) orientation-specific sandwich immunoassays to detect specific autoantibody – LRG1 immunocomplexes in the plasma of LC patients and control subjects.
Materials
LRG1-specific mouse mAbs (Bsi0351, Bsi0352, and Bsi0392) were produced and purified at Biosystems International (BSI) as previously described [41]. The mAbs were biotinylated with a 20-fold excess of EZ-link Sulfo-NHS-Biotin (21217, Thermo Scientific) according to the manufacturer’s protocol. Human recombinant survivin (4160-50) was obtained from Gentaur Europe BVBA (Kampenhout, Belgium). Purified human complement factor H (C5813) and HRP-conjugated anti-human IgG polyclonal antibody (A0170) were purchased from Sigma (Louisiana, MO, USA). SA-HRP (43-4323) was purchased from Invitrogen (Carlsbad, CA, USA) and anti-mouse-HRP (1020-05) from SouthernBiotech, (Birmingham, AL, USA). Plasma samples (20 LC and 22 controls) were obtained from the Department of Pulmonology of the University of Debrecen in Debrecen, Hungary under informed consent and a clinical protocol (RKEB/IKEB: 2422-2005) approved by the Institutional Review Board of the Department of Pulmonology and the Central Regulatory Agency for Clinical Testing in Hungary (the protocol follows the current EU regulations for clinical testing). Selected characteristics of the donors are summarized in Table 1.
Unless otherwise stated, the following tools and reagents were used for the ELISAs: Corning
microplates (CM: 96-well, half-area clear polystyrene, high-binding, non-sterile, plates CLS3690, Sigma); carbonate coating buffer (CCB: 15 mM Na
Purification of recombinant human LRG1
Recombinant human LRG1 was expressed in HEK293T cells transfected with the plasmid, ORIG-SC321789, from OriGene (Rockville, MD), containing the True-Clone (Human NM_052972) for human LRG1 [41]. The recombinant protein was purified from collected supernatants by affinity chromatography and then concentrated and quality tested by SDS-PAGE and ELISA [41].
Phage display
The Ph.D.-12™ Phage Display Peptide Library (New England Biolabs, Ipswich, MA, USA) was used to map the epitope of selected mAbs according to the manufacturer’s suggestions. The peptide sequences (at least 12 per mAb) were deduced via DNA sequencing of the inserts [22]. The individual peptide sequences of selected antibodies were used to manufacture BioTide peptides (JPT Peptide Technologies GmbH, Berlin, Germany), for which the biotin was coupled to the N-terminus of a 12-amino acid peptide with a Ttds linker.
ELISA
Most of the study included ELISA analysis with the methods consisting of several common steps. Coating with autoantigens or streptavidin or immobilization of the capture antibodies were performed in CCB in a volume of 30
Summary of the details for the various ELISAs performed during the study
Summary of the details for the various ELISAs performed during the study
DAC: direct antigen coating; SA-HRP: streptavidin-horseradish peroxidase conjugate; GAM-HRP: HRP-labeled goat polyclonal anti-mouse IgG.
Statistical analysis was performed using the R suite (
Results
The monoclonal antibodies Bsi0352 and Bsi0392 recognize different epitopes of LRG1
We produced several mAbs against the natural human LRG1 protein [22] and tested them in various assays. The three antibodies (Bsi0351, Bsi0352, and Bsi0392) used in this study were characterized in ELISA experiments and their epitope recognition was determined by phage display. Experiments in which recombinant LRG1 (2
Characterization of anti-human LRG1 mAbs. The binding of LRG1-specific biotinylated mAbs to LRG1 was examined by DAC ELISA (A), in which the wells were coated with 1 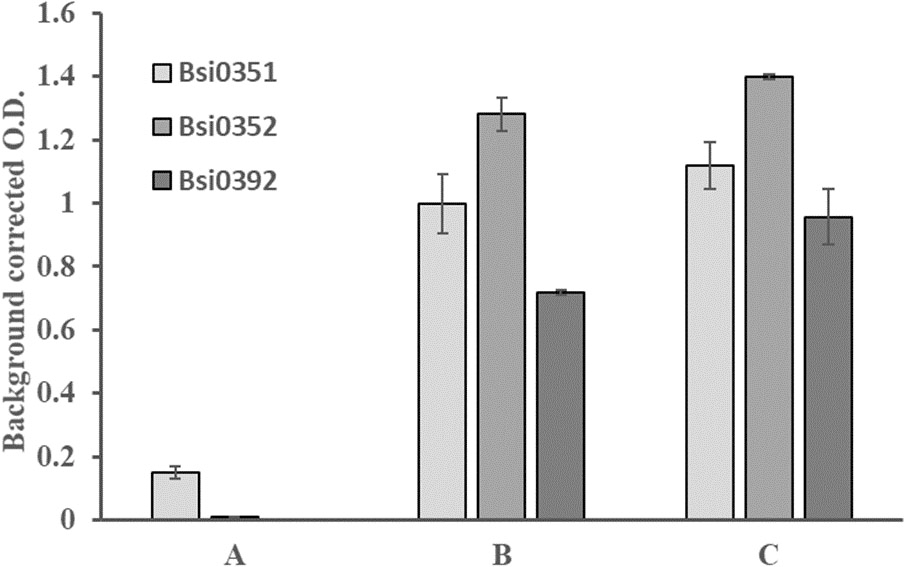
We next used a phage display combinatorial peptide library [37] of random 12-mer peptides fused to the N-terminus of a minor coat protein (pIII) of M13 phage to gain information about the similarity or differences amongst the epitopes recognized by the antibodies. We selected phages for which the displayed peptides were recognized by the antibodies. We were able to select several “binding” phages expressing mimic epitopes (mimotopes) using Bsi0351 and Bsi0352 (Table 3), whereas we were unsuccessful using Bsi0392. Mimotope peptides selected in these phage display experiments were synthesized with an N-terminal biotin and a hydrophilic linker was inserted between the biotin molecule and the peptide to render them flexible. We immobilized the peptides via streptavidin and tested their recognition by the three LRG1-specific antibodies. Peptides selected by Bsi0351 and Bsi0352 were cross-recognized by the two antibodies, mirroring the fact that the peptides showed significant similarity and, for some, even identity (Table 3), suggesting similarity and/or overlap of the epitopes. None of the peptides were recognized by Bsi0392, indicating that the epitope recognized by this antibody is different from those of Bsi0351 and Bsi0352 (Fig. 2).
List of 12 amino acid-long mimotope peptides selected by Bsi0352 (A) and Bsi0351 (B) mAbs in the phage display experiment. Formatted (bold: 0352/7–0351/4, italic
Recognition of phage display-selected biotinylated synthetic peptides. Aliquots of 0.5 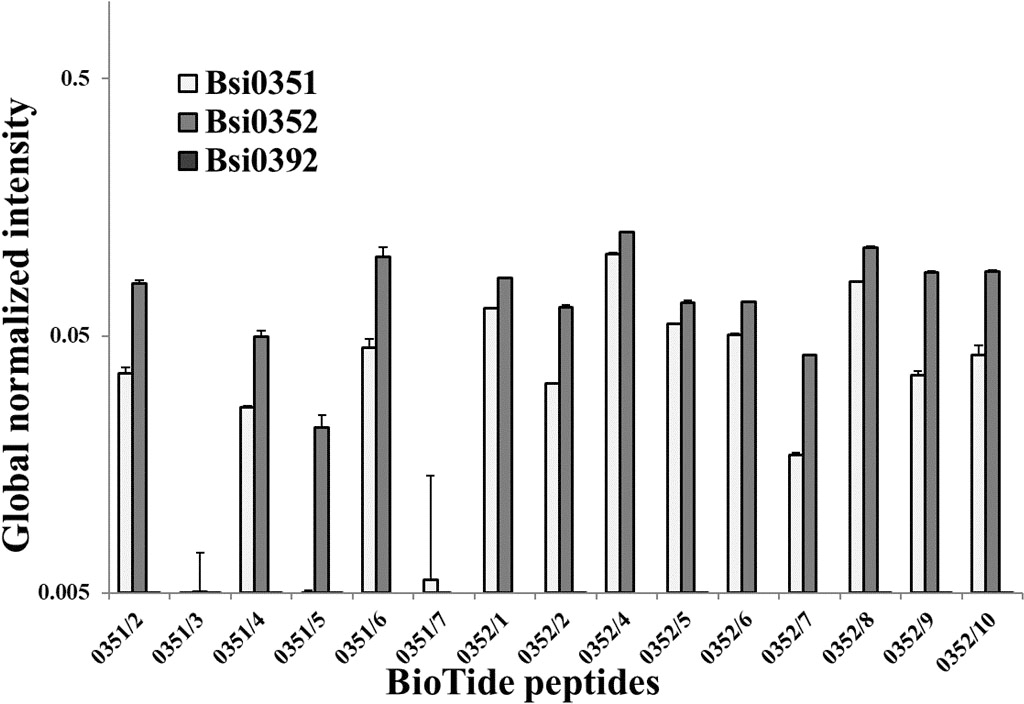
It was previously reported that anti-human BIRC5 (survivin) autoantibody levels in LC patient plasma are significantly higher than those in control samples [42]. CFH autoantibody levels have also been found to be significantly higher in NSCLC patients [12]. We thus assessed our plasma samples for the presence of autoantibodies against survivin and CFH as a positive control.
Detection of autoantibodies with directly immobilized autoantigens. Summary of the direct ELISA results, showing the mean and SD of background-corrected and global normalized intensities of autoantigen-captured antibodies in different sample populations detected by anti-human IgG-HRP. The S–W normality column shows the
-values for the Shapiro–Wilk normality test
Detection of autoantibodies with directly immobilized autoantigens. Summary of the direct ELISA results, showing the mean and SD of background-corrected and global normalized intensities of autoantigen-captured antibodies in different sample populations detected by anti-human IgG-HRP. The S–W normality column shows the
Autoantibody detection using a direct ELISA setup. (A) Recombinant human survivin-coated wells were incubated with 10-fold diluted human plasma samples. The global normalized background-corrected intensities of control and LC samples detected by anti-human IgG-HRP are summarized in boxplots. (B) Purified plasma CFH-coated wells were incubated with 10-fold diluted plasma samples and IgG type autoantibodies were detected by anti-human IgG-HRP. The boxplot shows the distribution of background-corrected and global normalized intensities of control, non-small cell lung cancer (NSCLC), and small cell lung cancer (SCLC) plasma samples. The box of the boxplot shows the data distribution from Q1 to Q3 and the median. The whiskers were shortened to a length of 1.5 times the box length to show possible outliers. All data points beyond the whiskers are plotted as points. Significant differences between sample type are labelled as follows: 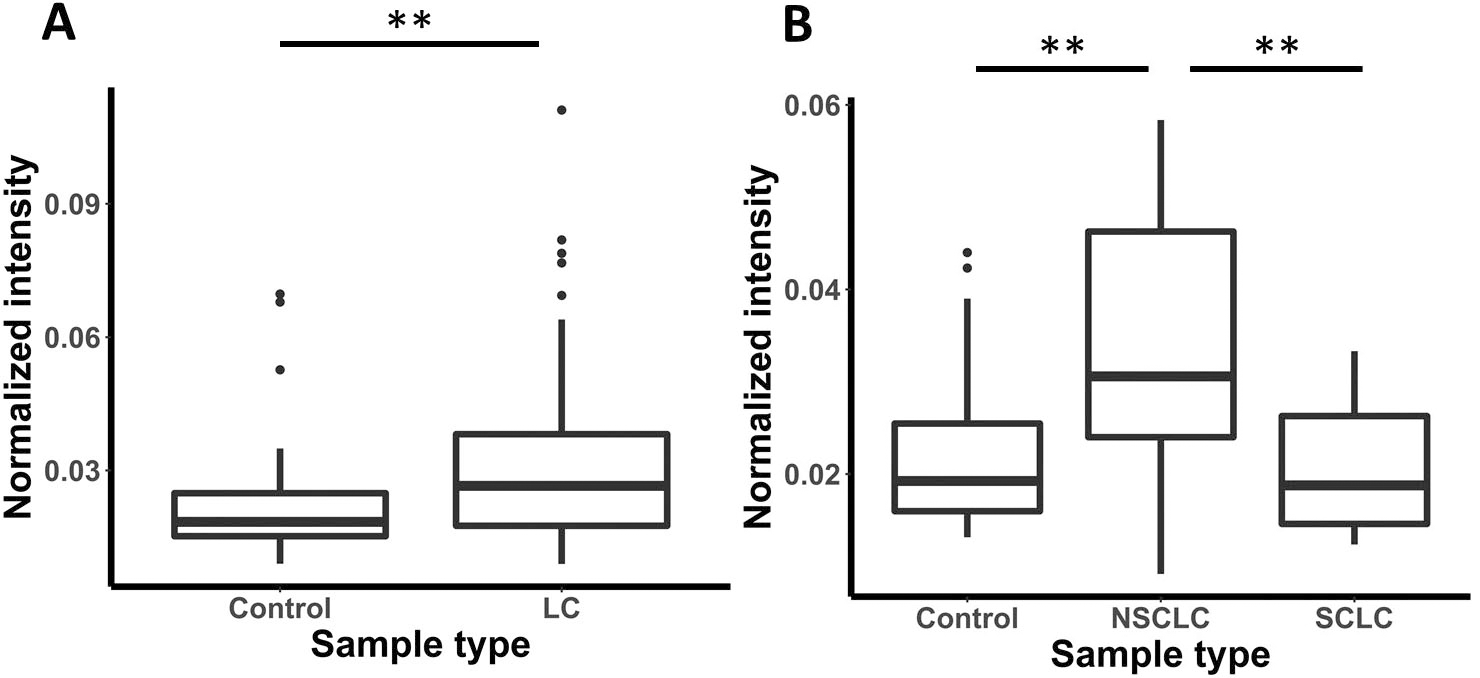
Plates were directly coated with recombinant survivin or purified factor H and incubated with 10-fold diluted human plasma samples of control (
We did not detect LRG1-specific IgG autoantibodies in initial direct antigen coating ELISA experiments in which recombinant LRG1 was immobilized and incubated with human plasma from LC patients or controls (Table 4). We thus used LRG1 captured from plasma samples either by the Bsi0392 or Bsi0352 mAbs to detect LRG1 autoantibody IgG immunocomplexes. In these experiments, LRG1 anti-human LRG1 autoantibody immunocomplexes captured by mAb Bsi0392 resulted in a detectable signal for which the intensities were significantly higher in the LC than control plasma samples (
Detection of LRG1-IgG immunocomplexes by epitope-specific capture. Summary of the LRG1 immunocomplex sandwich ELISA results showing the mean and SD of background corrected and global normalized intensities of alternatively presented (via Bsi0352 or Bsi0392) LRG1 autoantibody immunocomplexes in different subject populations (LC or Control) detected by anti-human IgG-HRP. The S–W normality column shows the
-values for the Shapiro–Wilk normality test
Detection of LRG1-IgG immunocomplexes by epitope-specific capture. Summary of the LRG1 immunocomplex sandwich ELISA results showing the mean and SD of background corrected and global normalized intensities of alternatively presented (via Bsi0352 or Bsi0392) LRG1 autoantibody immunocomplexes in different subject populations (LC or Control) detected by anti-human IgG-HRP. The S–W normality column shows the
Detection of anti-human LRG1 IgG autoantibody immunocomplexes using a sandwich ELISA setup. (A) The indicated antibodies, which recognize LRG1 in human plasma, were used to coat the wells and the coated wells were incubated with plasma samples, as described in Materials and Methods, followed by detection with anti-human IgG-HRP. The boxplot shows the distribution of the background-corrected normalized intensities of the LRG1 autoantibody for the indicated mAb–sample population pairs. (B) Results obtained using Bsi0392 to coat the wells in the sandwich ELISA setup, showing the control and all LC plasma samples divided into the non-small cell lung cancer (NSCLC) and small cell lung cancer (SCLC) plasma populations. Significant differences between sample type are labelled as follows: 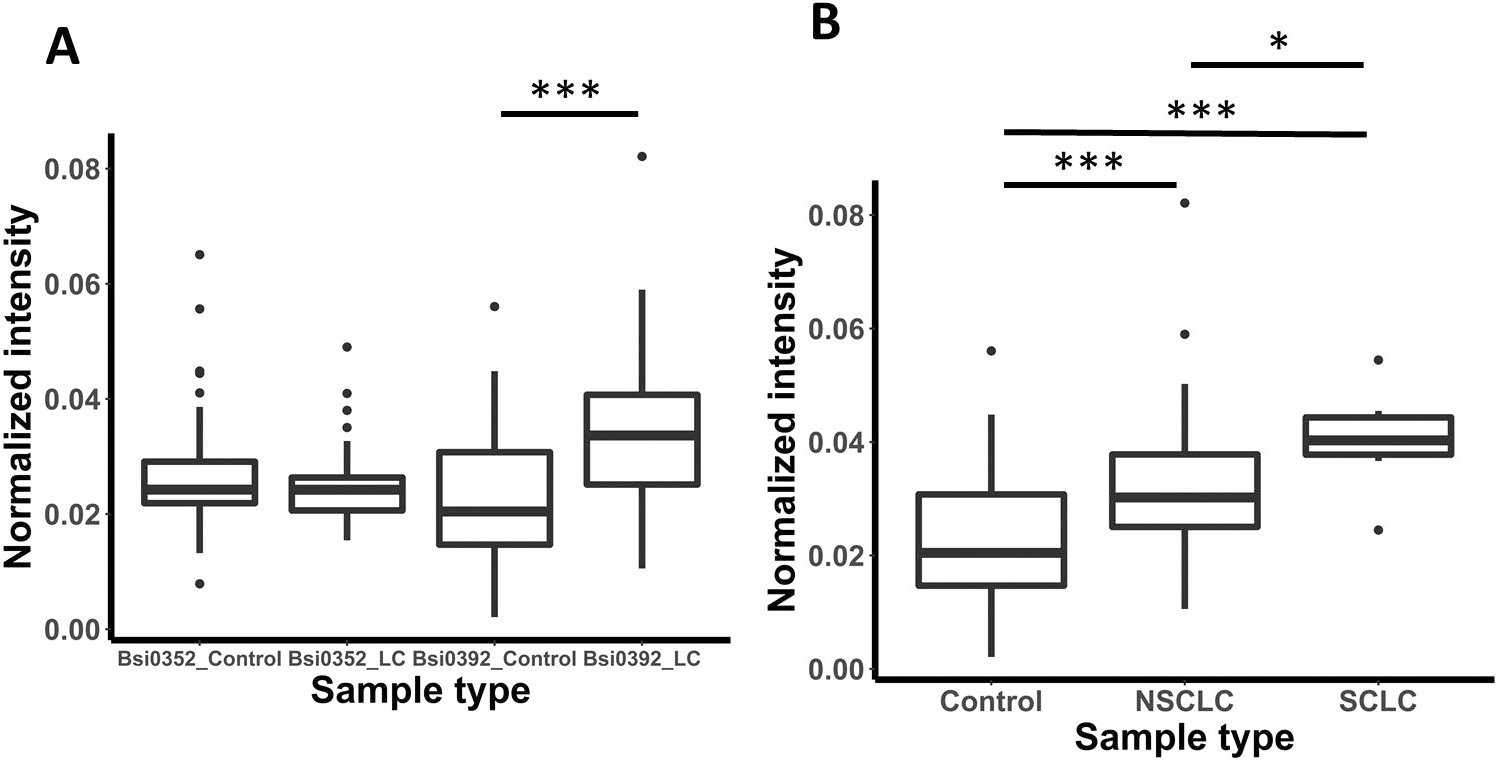
Our long-term goal is to develop widely applicable plasma proteome-specific mAb libraries for the profiling of human plasma. As a part of these larger scale efforts, we previously reported results of LC plasma proteome profiling with nascent mAb libraries. We described the discovery of mAb couples directed against HP, C9,
To our surprise, we did not detect autoantibodies to LRG1 using standard autoantibody measurement technology, in which immobilized antigen is allowed to react with test plasma or serum followed by IgG detection. We then turned to the measurement of circulating LRG1-IgG immunocomplexes by capturing LRG1 using the specific mAbs Bsi0352 and Bsi0392. Indeed, LRG1-IgG immunocomplexes were detectable and in a manner that suggests epitope and/or orientation specificity of the IgG autoantibodies in the complexes. Epitope-specific sandwich ELISA experiments showed significantly (
LGR1 levels have been reported to be elevated in several disease conditions [35, 54, 55, 56, 57]. However, our report is the first to demonstrate LRG1 autoantibodies and LRG1-IgG immunocomplexes. The implications include the appearance of cancer pathology-relevant epitope- and-orientation-specific neoantigens of LRG1 in LC, especially NSCLC. Further molecular studies will be required to determine whether the appearance of such neoantigens actually occurs or the results are due to autoantibodies against epitopes hidden in control subjects. The value of LRG1 immunocomplexes as a biomarker will be tested in the future on larger cohorts to assess its potential to improve LC diagnosis.
Footnotes
Acknowledgments
The authors thank Angéla Berki-Pál, Zsombor Lestár, and János Kádas Ph.D. for their technical assistance in the purification of recombinant LRG1.
This work was supported by the National Office for Research and Technology of Hungary (TECH-09-A1-2009-0113; mAb-CHIC). The sponsor was not involved in the work carried out in the study or the preparation or submission of the manuscript.
Biosystems International in text refers to Biosystems International Kft. (Debrecen, Hungary) and its former mother company Biosystems International SAS (Evry, France) which ceased operations in 2012.
Conflict of interest
All authors declare no conflict of interest.
Author contributions
Conception: JL, LT, and IK.
Interpretation and analysis of the data: JL, LT, and IK.
Preparation of the manuscript: JL, AK, IT, LT, and IK.
LT and IK equally contributed to the manuscript.
