Abstract
BACKGROUND:
Immunomodulatory genes play significant roles in the regulation of immunological properties of gastric cancer, but the effect of epigenetic regulation of these genes on the immune properties is unknown.
METHOD:
I analyzed the methylation-expression correlation among all immunomodulators and compared with the non-immunomodulators. The association between epigenetically regulated immunomodulators (ERI) and tumor microenvironment is evaluated. A key immunomodulator TIGIT is further selected to investigate the potential value in the regulation of immunologic properties. Furthermore, the prognostic value and the immunotherapeutic potential of TIGIT are also explored.
RESULT:
Four genes are identified as ERIs based on the negative correlation between expression and methylation. Association analysis shows that three ERIs participate in the regulation of the immune microenvironment of gastric cancer. Among these ERIs, TIGIT is identified as a key immunomodulator. TIGIT is found to be significantly associated with immune properties. The high TIGIT expression group tends to display an active immune landscape. TIGIT expression is also found to be associated with survival and immunotherapeutic sensitivity. High TIGIT expression group has a favorable prognosis and is more likely to respond to immunotherapy than the low expression group.
CONCLUSION:
TIGIT is an epigenetically regulated immunomodulator of gastric cancer which can modify the immune activity and affect immunotherapeutic sensitivity. These findings can promote the research of epigenetic therapies and improve the survival of cancer patients by sensitizing tumors to immune therapies.
Introduction
Gastric cancer is the third leading cause of cancer-related death and the fifth most common cancer in the world [1]. Most of the patients are diagnosed at an advanced stage, leading to poor survival. Immunotherapy has revolutionized the traditional treatment of gastric cancer in the past few years. But only a small proportion of patients respond to the treatment. Furthermore, some patients will fail to respond to the secondary treatment and develop resistance properties to the immunotherapy [2]. Thus, it is of great significance to identify the potential patients that will respond to the immune therapies. Extensive studies have concentrated on the identification of signature responding to immunotherapy. Mutational burden and neoantigen load are found to be associated with immunotherapy [3]. Expression of immune checkpoints and interferon-gamma is reported to affect the response to immune therapies [4, 5]. Composition of tumor microenvironment is also supported to be a significant factor affecting the sensitivity to immunotherapy [6, 7]. Although combinations of immune drugs have proven to be effective protesting against resistance, they often lead to severe immune-related toxicities [8]. Thus, combination of immune and non-immune drugs that epigenetically modify the immune properties emerges as a promising option to sensitize the tumors to immune therapies.
In recent years, epigenetic regulation is found to be associated with immunological properties [9, 10]. DNA methylation loss is reported to promote immune evasion of tumors [11]. Furthermore, DNA methylation is reported to affect the innate immunity through the regulation of a variety of immune cells including NK cells [12], DCs [13], and T cells [14]. DNA methylation inhibitors have become the mainstay for treatment of certain haematological malignancies [15]. But the therapeutic potential in solid tumors is unclear. Currently, an increasing number of immunomodulatory genes have been discovered to play significant roles in the regulation of immune properties. But the effect of epigenetic regulation of these immunomodulators to the immunological properties is unknown. Deeply investigating the interaction between methylation of immunomodulators and the immunological properties can improve my understanding of the epigenetic regulation to the immunity and advance the research of epigenetic therapies.
In this study, I aim to characterize the interactions between methylation and immune properties of gastric cancer. To determine the interactions, I will start my work from the following three ways. First, I will evaluate the association between methylation and expression to identify ERIs, as well as comparison with non-ERI to check whether immunomodulators are more likely to be methylated. Next, I will investigate the association between these ERIs with tumor microenvironment to determine the significant value of these genes in regulating immune activity. Finally, I will evaluate the association of these ERIs in regulating immunotherapeutic sensitivity. In brief, these analyses can provide insights of ERIs in modification of immune properties and promote the development of epigenetic therapies, optimizing the treatment options in the clinical management of cancer.
Result
Characterization of ERI
A total of 78 immunomodulatory genes (Supplementary Table 1) are retrieved from the published literature [16]. In all of these immunomodulators, 49 genes have both methylation and expression data and are selected for the further analysis(Supplementary Table 2). ERIs are identified based on the significant negative association between expression and methylation intensity which is implemented in MethylMix package [17]. As a result, four immunomodulators are identified as epigenetically regulated genes. Association analysis shows that there is a negative correlation between expression and methylation of these genes (Fig. 1a–d), suggesting that these immunomodulators are epigenetically regulated by methylation. Comparing with non-immunomodulators, I found that the methylation-expression correlation of immunomodulators is smaller than the non-immunomodulators (Fig. 1e), suggesting that the immunomodulators are more likely to be epigenetically regulated by methylation than the non-immunomodulators.
Characterization of ERI. (a-d) Four immunomodulatory genes are identified as ERI. Point diagram shows the negative association between the expression and the methylation. (e) Density curve shows the distribution of expression-methylation correlation among immunomodulators and non-immunomodulators. They are colored into red and green separately. Dashed line is the median of the correlation. The four ERI are annotated on the graph. (f, g) Boxplot and violinplot show the methylation and expression of these four ERI separately. (h, i) Heatmap shows the methylation and expression of these ERI. 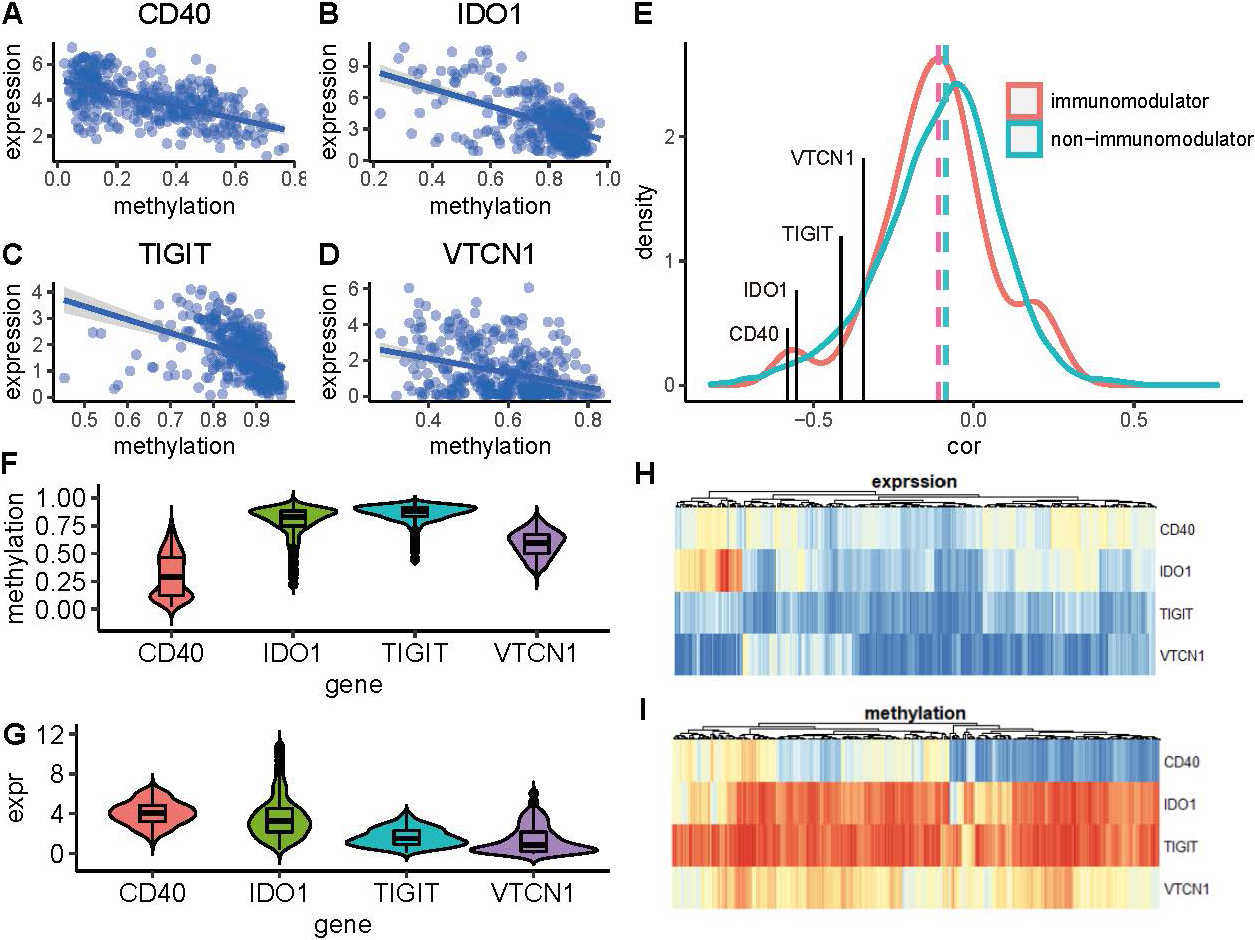
In all of these ERIs, TIGIT, which encodes a member of the poliovirus receptor family of immunoglobin proteins, tends to have the highest methylation than other genes and a relatively low expression (Fig. 1f–i). CD40, which is a member of the TNF-receptor superfamily has the highest expression and the lowest methylation than other ERIs (Fig. 1f–i). Comparing with these two genes, the other two ERIs IDO1 and VTCN1 have moderate expression and methylation.
To identify the interactions between these ERIs and immune microenvironment of gastric cancer, sixty-four kinds of immune cells and stromal cells are quantified using xCell [18]. Immune score of each sample is also evaluated based on these immune cells. A cluster consisted of 10 kinds of immune cells is found to be positively regulated by ERIs based on the association analysis (Fig. 2b). This cluster is positively associated with the expression of TIGIT, CD40, and IDO1 and negatively associated with the methylation of these immunomodulators (Fig. 2c), suggesting that this cluster is epigenetically regulated by these ERIs. Among these four immunomodulators, both the expression and methylation of VTCN1 are negatively associated with the immune cells, which is distinct from other ERIs and is removed in the further analysis.
ERI in the regulation of immune microenvironment. (a) Point diagram shows the positive association between the expression of ERI and immune score. (b) Heatmap shows the correlation between the expression and methylation of ERI and the 64 immune cells and stromal cells. Red indicates positive association and blue indicates negative association. Ten kinds of infiltrating immune cells that are significantly associated with these ERI are marked in the red rectangle. (c) Correlation matrix shows the association of the ten kinds of immune cell and the expression and methylation of ERI in TCGA cohort. (d, e) Validation of the association of ERI and the immune cells in two external datasets GSE15459 and GSE62254. (f) ERI-immune cells interaction network constructed using the correlation threshold 0.6. Blue node represents the ERI and red node represents the immune cells. (g) Two kinds of immune cells are most positively associated with the ERI.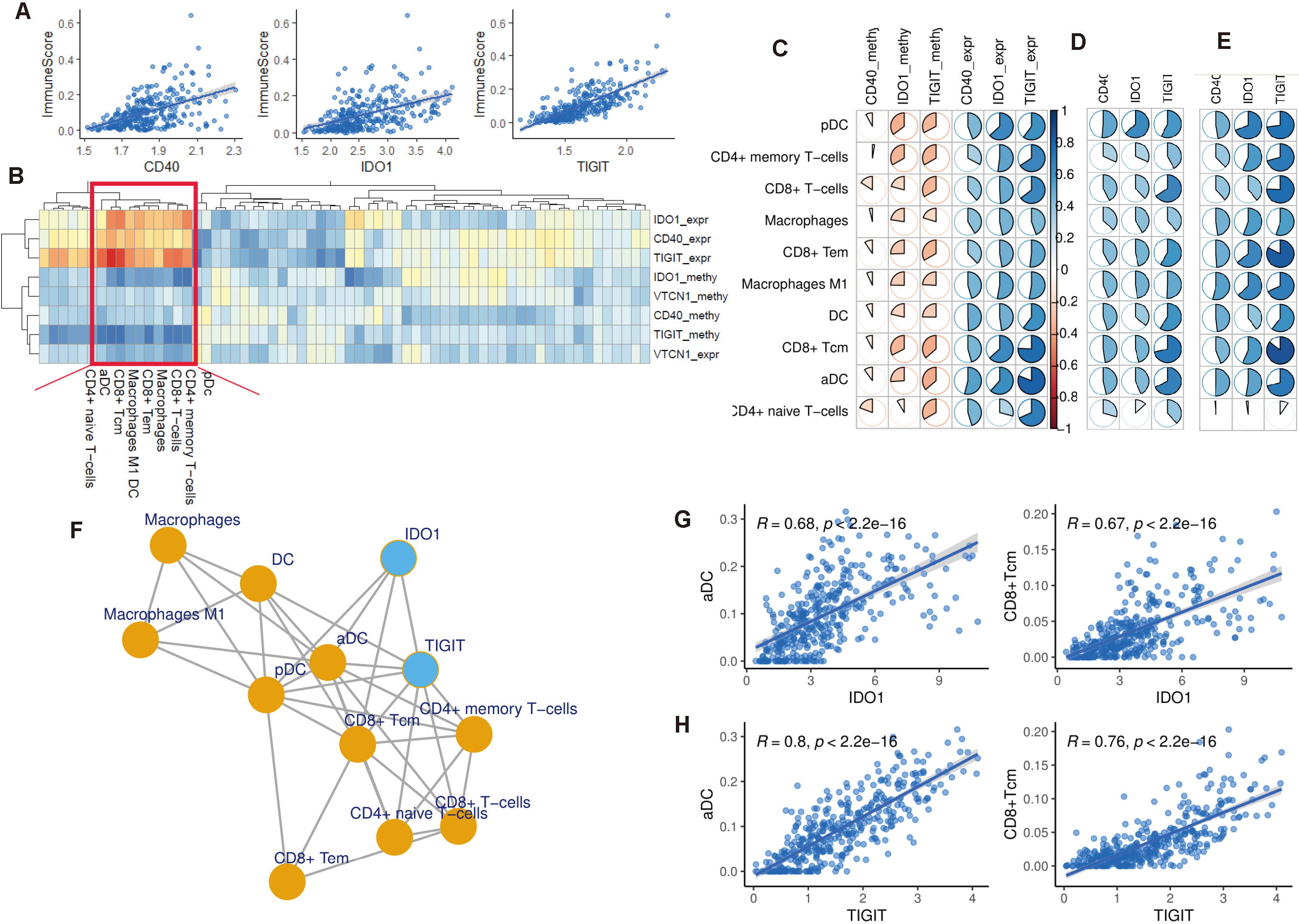
Next, I analyzed the regulation of these ERIs in the immune activity of gastric cancer. I found that the expression of these genes is positively associated with the immune score (Fig. 2a), suggesting that these immunomodulators participate in the regulation of immune-associated pathway and activate the immune landscape of gastric cancer.
To validate the regulation of these ERIs in the immune-related cluster, two external datasets are collected for the association analysis. I found that almost all of these immune cells in the cluster are positively associated with the expression of these ERIs. Among the ten immune cells in the cluster, positive regulation of pDC, CD4+memory T-cells, CD8+T-cells, macrophages, CD8+Tem, macrophages M1, DC, CD8+Tcm, and aDC, are observed in GSE15459 and GSE62254 (Fig. 2d and e), suggesting the significant role of these three ERIs in the activation of immune microenvironment. However, CD4+ naïve T-cells is not found to be associated with the expression of these ERIs in the GSE62254 cohort.
I constructed an interaction network for these ERIs and immune cells based on correlation threshold 0.6. Network analysis of these immunomodulators and immune cells shows that pDC, aDC, and CD8+Tcm are the hub cells in the network which connect with other nodes tightly (Fig. 2f). Furthermore, I found that TIGIT has more interactions than IDO1, suggesting that TIGIT plays a more significant role in the regulation of immune microenvironment than IDO1. Among these immune cells, aDC and CD8+Tcm have the most positive correlation compared with other cells. The positive association of these two immune cells and the ERIs is showed in Fig. 2g and h.
Immunomodulators play significant roles in the regulation of individual immune activity. They can be classified as epigenetically regulated and non-epigenetically regulated in my study. But the effect of epigenetic regulation on these immunomodulators is unknown and the difference between these ERIs and non-ERIs is unclear. Thus, I want to identify the difference between the immunomodulators that are epigenetically regulated and those that are not methylation derived.
I calculated the methylation-expression correlation for all immunomodulators and rank them by the correlation coefficient. I found that the immunomodulators that are epigenetically regulated have the most negative correlation compared with the non-ERIs (Fig. 3a). Most of the immunomodulators are found to be positively associated with immune score, aDC, and CD8+Tcm (Fig. 3b–d), suggesting their positive regulation in the immune activity of gastric cancer. However, VTCN1 is found to be negatively associated with these immune signatures, suggesting suppressed mechanism of immune regulation. Association analysis shows that TIGIT has a supreme positive correlation than almost all of other immunomodulators (Fig. 3b–d). These results show that TIGIT, which is epigenetically regulated, is a key immunomodulator of gastric cancer. Thus, it is of great significance to elucidate the underlying interaction between TIGIT and immune properties.
Comparison between ERI and non-ERI. (a) Bar diagram shows the methylation-expression correlation among all immunomodulators. ERI are colored in blue and located on the left part of the diagram. (b) Bar diagram shows the expression-immune correlation among all immunomodulators. (c, d) Bar diagram shows the expression-immune cell association among all immunomodulators.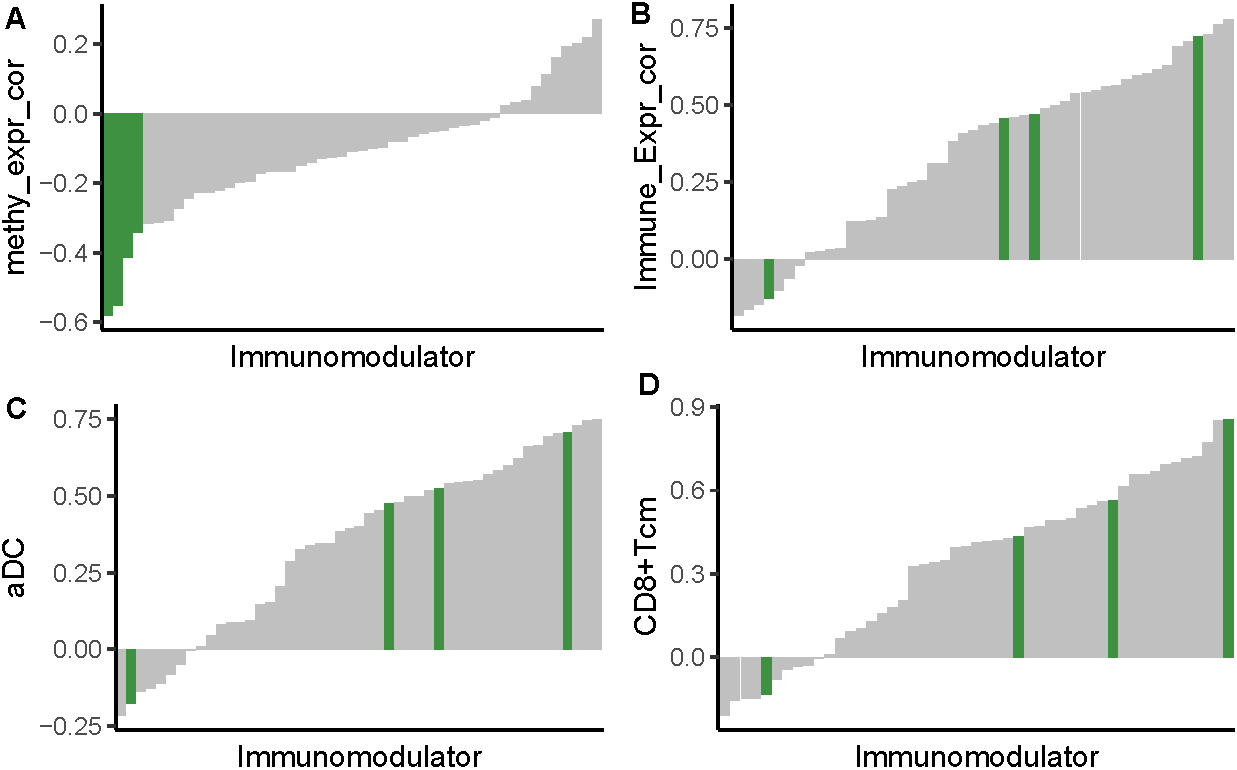
To identify the potential value of TIGIT in the regulation of immune activity of gastric cancer, I divided gastric cancer patients into two groups based on TIGIT expression. Additionally, I collected 17 immunity-related gene sets covering both innate and adaptive responses from published literature. The enrichment score (ES) of these gene sets for each sample is evaluated using single sample gene set enrichment analysis (ssGSEA). Clustering analysis shows that gastric cancer samples are clustered based on different TIGIT expression. Samples within the same TIGIT expression group tend to be clustered together and samples in different TIGIT groups are clustered separately (Fig. 4a), suggesting that TIGIT is a key immunomodulators in distinguishing gastric cancer with distinct immune landscape. Furthermore, TIGIT high expression group is found to have a higher ES than the low expression group in all of these 17 immunity-related gene sets (Fig. 4b), suggesting that TIGIT participate in the regulation of a variety of immune-related pathways.
TIGIT expression perturbation shapes the immune landscape of gastric cancer. (a) Heatmap shows the immune activity of gastric cancer patients. A total of 17 immunity-related gene sets annotated in the rows represents both innate and adaptive responses. Samples are annotated into different TIGIT groups. (b) Boxplot shows the enrichment score of 17 immunity-related gene sets for different TIGIT expression groups. (c-f) Comparison between TIGIT expression and previous gastric cancer subtypes. (g) Survival analysis between different TIGIT groups.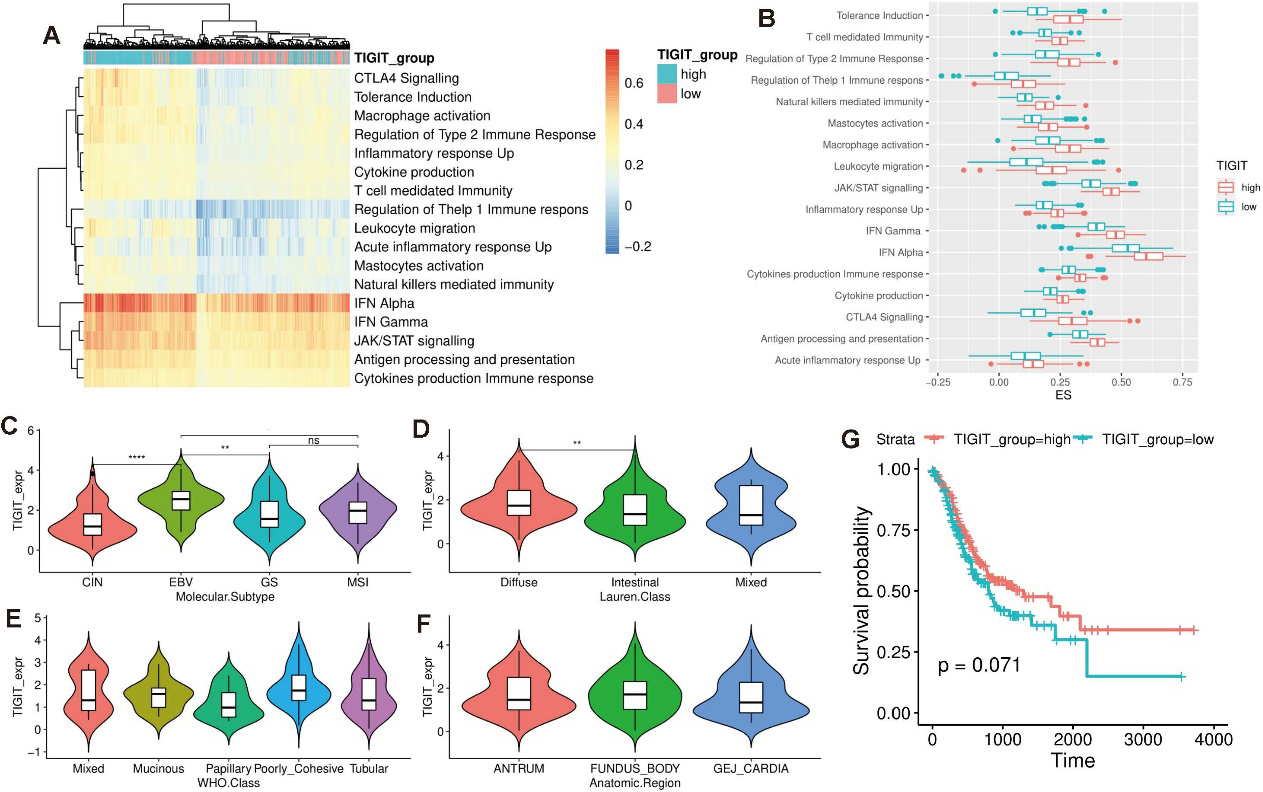
To explore the clinical utility of TIGIT, I connected my TIGIT group with current gastric cancer subtypes. Molecular classification which divides gastric cancer into tumors positive for Epstein–Barr virus (EBV), microsatellite unstable tumors (MSI), genomically stable tumors (GS), and tumors with chromosomal instability (CIN) is first investigated [19]. EBV subtype is found to have the highest TIGIT expression than other subtypes and CIN group has the lowest TIGIT expression compared with other groups (Fig. 4c). Lauren classification, WHO classification, and anatomic region are also evaluated in my analysis. Diffuse type gastric cancer tends to have a higher TIGIT expression than the intestinal group (Fig. 4d). Papillary type gastric cancer is found to have the lowest TIGIT expression than other WHO subtypes (Fig. 4e). But no difference of TIGIT expression is observed across anatomic region (Fig. 4f).
Immune activity is found to affect the prognosis in a variety of cancers [20, 21]. In this analysis, TIGIT is found to activate a series of immune-related pathways. Based on these reasons, I hypothesize that TIGIT expression is associated with the prognosis with gastric cancer and patients with high TIGIT expression have an activated immune landscape and thus favorable prognosis. Survival analysis between different TIGIT groups shows that TIGIT high expression group has a favorable prognosis than the low expression group (Fig. 4g). Additionally, I validate the favorable survival of TIGIT high expression group in IMvigor210 cohort and found the same result (Fig. 5a), suggesting that TIGIT is a robust factor in the assessment of prognosis of gastric cancer.
Evaluation of TIGIT in the immunotherapeutic sensitivity. (a) Survival analysis of different TIGIT expression in IMvigor210 cohort. (b) Bar graph shows the TIGIT expression in different immunotherapeutic response groups (complete response (CR), partial response (PR), stable disease (SD), and progressive disease (PD)) in IMvigor210 cohort. (c) Boxplot shows the TIGIT expression in responder and non-responder groups in GSE35640 cohort. (d) Bar graph shows the proportion of responder and non-responder in different TIGIT expression groups. (e) Bar graph shows the TIGIT expression for all samples. Responders are colored in blue. 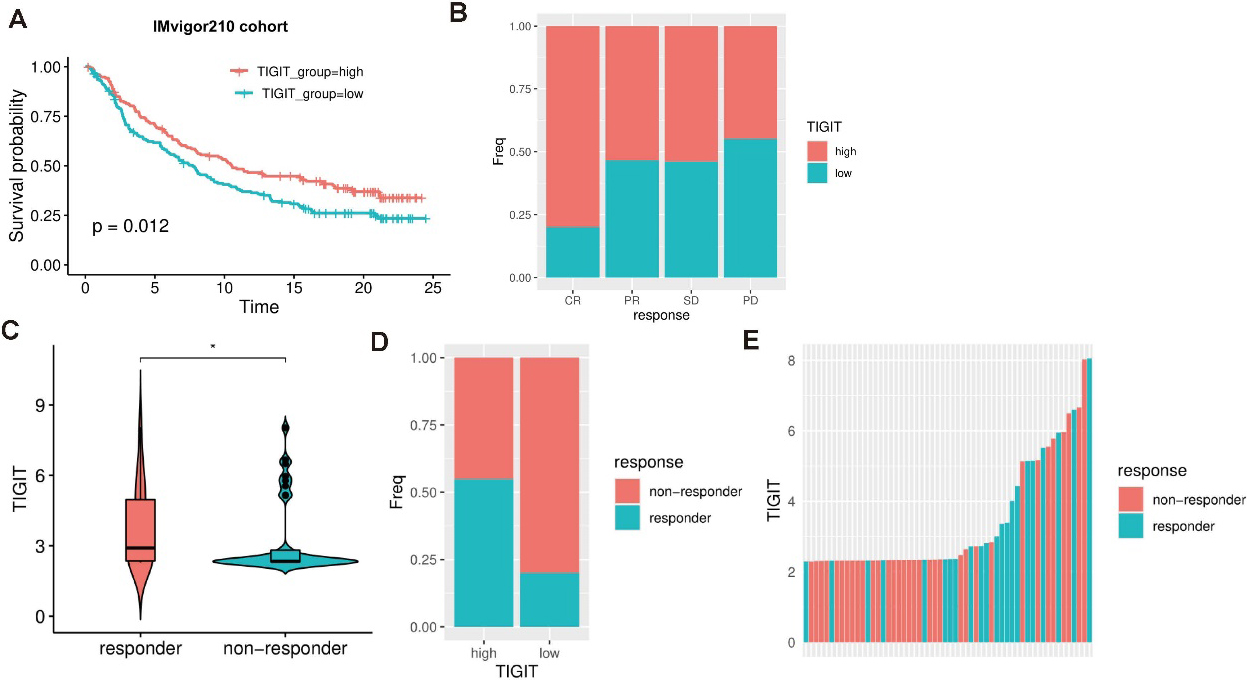
Tumor immune microenvironment and individual immune landscape are found to be significant determinant affecting the prognosis and immunotherapeutic response of gastric cancer [22]. Patients with different immune signature tend to respond distinctly to the same treatment even though they are in the same or similar clinical traits [23]. In this study, I find that TIGIT is a significant immunomodulator regulating a large number of immune-related pathways. Therefore, it is of great significance to identify the association between TIGIT expression and the immunotherapeutic sensitivity.
I examine the immunotherapeutic sensitivity of TIGIT groups in two datasets IMvigor210 cohort and GSE35640 cohort. I found that the proportion of TIGIT high expression samples in response group is higher than the non-response group in IMvigor210 cohort. Especially in the complete response (CR) group, the proportion of TIGIT high expression samples is the highest than other groups (Fig. 5b). In the GSE35640 cohort, I found that the responder group has higher TIGIT expression than the non-responder group (Fig. 5c). The proportion of responder in TIGIT high expression group is higher than in the low expression group (Fig. 5d). Furthermore, I rank all samples by TIGIT expression and find that responder samples tend to have higher TIGIT expression than the non-responder samples (Fig. 5e). In conclusion, all these results suggest that TIGIT expression is significantly associated with the immunotherapeutic response.
Materials and methods
Data collection
Multi omics data containing methylation and expression data of gastric cancer is downloaded from TCGA-STAD cohort. A total of 375 gastric adenocarcinoma samples are collected in this study to identify ERI. Clinical information including overall survival, histological subtypes, WHO classification, and anatomic region is also downloaded for survival analysis and evaluation of clinical features. The methylation data in the TCGA cohort is produced from different platforms. To maximize the usage of these methylation data, I combine the data from different platform and eliminate the platform-based bias using combat algorithm [24]. For multiple probes matching to the same gene, I select the probe that has the most negative correlation with the expression of this gene. The 79 immunomodulators and 17 immunity-related gene sets covering both innate and adaptive responses are retrieved from published literature [25]. Gastric cancer subtype information of TCGA cohort that has been previously classified is downloaded using the TCGAbiolink package [26].
Additionally, two external datasets GSE15459 and GSE62254 are collected to analyze the association between ERIs and immune microenvironment. GSE15459 cohort contains mRNA expression profiles of 200 primary gastric tumors and GSE62254 contains microarray profiles of 300 gastric cancer samples. Expression matrix of these samples is downloaded to characterize the tumor microenvironment of gastric cancer, as well as to investigate the association of ERI with immune microenvironment.
For the analysis of immunotherapeutic sensitivity, patients with metastatic urothelial cancer treated with anti-PD-L1 agents are obtained from the R package IMvigor210CoreBiologies [27]. The melanoma GSE35640 cohort treated with MAGE-3 antigen-based immunotherapy is also collected in this analysis. Clinical records of immunotherapeutic response containing complete response, partial response, stable disease, and progressive disease of these patients are downloaded to check whether TIGIT can affect immunotherapeutic sensitivity.
Identification of ERI
ERI is identified based on the negative correlation between methylation and expression data which is implemented using MethylMix package. MethylMix identifies DNA methylation driven genes by modeling DNA methylation data in cancer vs. normal and looking for homogeneous subpopulations and evaluate the negative association between methylation and expression data of a specific gene.
Immune microenvironment quantification
Tumor microenvironment is quantified using xcell algorithm which is a single sample gene set enrichment analysis-based methodology. It evaluates the expression of a pre-defined gene set relative to the expression of genes in the whole cohort to check whether they are enriched in the upper or bottom of the ranked gene list. It contains 64 kinds of immune cells and stromal cells and gives each sample enrichment score representing the proportion of a specific cell involved in tumor microenvironment.
Gene set enrichment analysis
Enrichment score for these 17 immunity-related gene sets is evaluated using ssGSEA [28]. This algorithm calculates an enrichment score for each specific gene set based on the expression of pre-defined gene list relative to the expression of genes in the whole cohort to check whether they are enriched in the upper or bottom of the ranked gene list. It is conducted using R package gsva.
Survival analysis and statistical analysis
To analyze the effect of TIGIT expression for patient survival, gastric cancer patients are divided into high expression and low expression groups based on the median expression of TIGIT. KM curve is plotted to describe the survival trend between different gastric cancer groups. Log-rank test is used to evaluate the difference between the survival curves. To evaluate the association of TIGIT with immunotherapeutic sensitivity, patients are divided into responder and non-responder groups based on the clinical records of sensitivity. Wilcoxon test is used to evaluate the difference of TIGIT expression between different groups.
Discussion
In this study, I identified several epigenetically regulated immunomodulators. IDO1, CD40, and TIGIT are all found to be positively associated with a variety of immune cells in the tumor microenvironment. TIGIT is found to be a key regulator of the immune activity of gastric cancer. The expression of TIGIT is observed to be positively associated with a variety of immune-related pathways. The expression perturbation of TIGIT is found to affect the prognosis and immunotherapeutic sensitivity. These findings can provide insights into the interactions between methylation and immune regulation, advancing the research of epigenetic therapies for precision medicine in the future.
Traditional tumor staging system has been widely used in the clinical assessment of prognosis in gastric cancer [29]. But it does not perform well due to the molecular heterogeneity among the cancer patients. In this study, TIGIT is found to be associated with the prognosis of gastric cancer, demonstrating that TIGIT is a potential factor in evaluating survival of gastric cancer. The prognostic value of TIGIT should be validated widely in large scale cohort and clinical trial in the future. Once the prognostic value of TIGIT is confirmed, it can facilitate the clinical utility and precisely evaluate the prognosis of gastric cancer in combination with current criterion.
A main strength of this study is the discovery of epigenetic regulation of methylation in immunomodulators, which builds a bridge between methylation and immune properties. Immunomodulators, defined as significant regulators in modification of immune properties and microenvironment, have potential to stimulate or inhibit the immune activity. Due to the unique attribute of immunomodulator in regulating immune activity, they are promising and candidate biomarkers in identification of potential responders of immunotherapy. In this study, three immunomodulators are identified as epigenetically regulated, suggesting that epigenetically targeting these genes can change and improve the immune properties and microenvironment. Thus, it is significant and promising to take these genes into consideration for the design of epigenetic therapies to sensitize tumors for immunotherapy.
This study is limited by the absence of gastric cancer dataset for the evaluation of TIGIT in immunotherapeutic sensitivity which may cause biased result due to the heterogeneity of different tumor. To best of my knowledge, there is no immunotherapy-related gastric cancer dataset available in the online database. To maximize the usage of immunotherapeutic datasets for the assessment of TIGIT in predicting therapeutic sensitivity, I utilized urothelial cancer and melanoma to evaluate the interaction between TIGIT and immunotherapeutic sensitivity which will leads to an incorrect result. With the development and construction of database in the future, predictive value of TIGIT will be determined in multiple cancers.
The effect of methylation on the immunomodulators remains unknown in most cancers. my study suggests that the immune landscape of gastric cancer can be epigenetically regulated by the methylation of immunomodulators. The positive regulation of ERIs involved in my study can be examined in other cancer types to check whether they have the same regulation pattern for the immune activity. Furthermore, the method used in this study can be applied in other cancer types to identify the ERIs. Whether the ERIs identified in different cancer types are common or not is unknown. Thus, further analysis should concentrate on the identification of ERIs and investigate the functional regulation of these ERIs that are shared across different cancer types.
Epigenetic modulation of immune cells is reported to modify the immunological properties and sensitize tumors to immunotherapy [30, 31]. A variety of proteins involved in the histone modifications are identified as targets of epigenetic therapy [32]. In this study, TIGIT is found to be epigenetically regulated immunomodulator. The immunological property of gastric cancer is regulated by the methylation of TIGIT, suggesting that TIGIT can be a potential target of epigenetic therapy. It seems feasible that I can modify the immunological properties of gastric cancer indirectly by epigenetically targeting TIGIT. Thus, it is of great significance to investigate the immunotherapeutic utility of TIGIT as a potential epigenetic target in the future.
In conclusion, TIGIT is identified as an epigenetically regulated immunomodulator which can affect the prognosis and immunotherapeutic sensitivity of gastric cancer. The prognostic value of TIGIT can be utilized to precisely evaluate the survival of gastric cancer and has potential to identify responder of immunotherapy. The immunomodulatory attribute of TIGIT suggests that it can be used as potential target for the design of epigenetic drugs to sensitize tumors to immune therapies.
Author contributions
Conception: Jianfei Ma.
Interpretation or analysis of data: Jianfei Ma.
Preparation of the manuscript: Jianfei Ma.
Revision for important intellectual content: Jianfei Ma.
Supervision: Jianfei Ma.
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-210159.
sj-xlsx-1-cbm-10.3233_CBM-210159.xlsx - Supplemental material
Supplemental material, sj-xlsx-1-cbm-10.3233_CBM-210159.xlsx
Footnotes
Acknowledgments
I acknowledge TCGA and GEO database for providing their platforms and contributors for uploading their meaningful datasets.
