Abstract
BACKGROUND:
Triple-negative breast cancer (TNBC) is a highly malignant breast cancer subtype with a poor prognosis. The cell cycle regulator cyclin A2 (CCNA2) plays a role in tumor development. Herein, we explored the role of CCNA2 in TNBC.
METHODS:
We analyzed CCNA2 expression in 15 pairs of TNBC and adjacent tissues and assessed the relationship between CCNA2 expression using the tissue microarray cohort. Furthermore, we used two TNBC cohort datasets to analyze the correlation between CCNA2 and E2F transcription factor 1 (E2F1) and a luciferase reporter to explore their association. Through rescue experiments, we analyzed the effects of E2F1 knockdown on CCNA2 expression and cellular behavior.
RESULTS:
We found that CCNA2 expression in TNBC was significantly higher than that in adjacent tissues with similar observations in MDA-MB-231 and MDA-MB-468 cells. E2F1 was highly correlated with CCNA2 as observed through bioinformatics analysis (
CONCLUSION:
Our data indicate that E2F1 promotes TNBC proliferation and invasion by upregulating CCNA2 expression. E2F1 and CCNA2 are potential candidates that may be targeted for effective TNBC treatment.
Keywords
Introduction
The latest data show that breast cancer is the most common cancer worldwide, and it is the primary cause of cancer-related deaths among women [1, 2]. Breast cancer incidence and mortality rank first among all cancers, respectively, with 2.3 million new cases and 0.68 million deaths [1]. Triple-negative breast cancer (TNBC) is characterized by a lack of expression of the estrogen receptor (ER), progesterone receptor (PR), and human epidermal growth factor receptor 2 (HER2). This breast cancer subtype is highly proliferative and associated with a poor prognosis, accounting for approximately 15–20% of all breast cancers [3]. Although many molecular biomarkers have been found [4], TNBC lacks effective therapeutic targets, and its treatment is still primarily based on radiotherapy and chemotherapy [5]. Therefore, there is an urgent need to find potential and effective molecular therapeutic targets for TNBC.
The cell cycle controls proliferation and plays an important role in the early stage of DNA synthesis (G1 stage), the DNA synthesis stage (S stage), the late DNA synthesis stage (G2 stage), and the division stage (M stage); notably, misregulation of this process can contribute to cancer occurrence and development [6, 7]. Cyclin A2 (CCNA2) is a core component of the cell cycle and is abnormally expressed in cervical, colorectal, and breast cancers, thereby affecting cell proliferation [6, 8, 9]. Previously, we found that CCNA2 is highly expressed in TNBC tissues based on bioinformatics analysis [10]. Some studies have found that the transcription factors E2F4 and GATA3 have direct binding sites for CCNA2 [11, 12].
Transcriptional regulation is very important in cancer progression. The E2F transcription factor family proteins, including E2F1, play an important role in controlling cell cycle progression as well as in promoting proliferation and tumorigenesis [13]. Lu et al. [14] found that the LINC00511/miR-185-3p/E2F1/Nanog axis can promote breast cancer tumorigenesis. Recent findings from Xiong et al. [3] confirmed that ABP32E induces TNBC, a process regulated by the expression of E2F1. Furthermore, higher E2F1 expression levels are often observed in breast cancer tissues than in normal tissues [15]. Therefore, E2F1 plays a key role in the occurrence and development of TNBC. Interestingly, the expression levels of E2F1 and cyclin A2 were downregulated in MSK2-depleted cervical cancer cells [16]. In hepatocellular carcinoma cells, E2F1 reduces the hyperphosphorylation level of cyclin A2 mRNA pRb, leading to a decrease in the number of cells in the S-phase and promoting the occurrence and development of liver cancer [17]. However, the relationship between E2F1 and CCNA2 and whether E2F1 mediates the transcriptional regulation of CCNA2 to promote TNBC progression have not been fully elucidated.
How CCNA2 promotes the progression of TNBC is our core concern. Here, we found that CCNA2 expression promoted the proliferation of TNBC cells through a series of in vivo and in vitro experiments. Our results showed that CCNA2 expression was highly correlated with E2F1. Notably, we revealed that E2F1 could promote CCNA2 expression through an interaction with the CCNA2 promoter at
Materials and methods
Cell culture
MCF10A cells, a normal human mammary epithelial cell line, were obtained from Fuheng Company (Shanghai, China) and cultured in a special medium. Breast cancer cell lines, including HS578T, MCF-7, MDA-MB-231, and MDA-MB-468 cells, were obtained from Fuheng Company (Shanghai, China) and were cultured in DMEM supplemented with 10% fetal bovine serum (Gibco, Thermo Fisher Scientific, Waltham, MA, USA) at 37
Tissue microarray (TMA) cohorts and immunohistochemical staining
TMA cohorts containing 15 TNBC tissues and 15 adjacent tissues were obtained from the First Hospital of Lanzhou University. The study was performed with the approval of the Ethics Committee of the First Hospital of Lanzhou University. All participants agreed to participate in this study and signed an informed consent form. TMA sections were deparaffinized, hydrated, and blocked for endogenous peroxidases and antigen retrieval. After sealing for 1 h at room temperature, anti-cyclin A2 (1:50) was added, followed by incubation at 37
RNA extraction and quantitative PCR (qPCR)
TRIzol reagent (Invitrogen, Carlsbad, CA, USA) was used to extract total RNA according to the manufacturer’s protocol. A NanoDrop spectrophotometer (Thermo, Rockford, IL, USA) was used to quantify the RNA. Reverse transcription was then used to synthesize cDNA. Using cDNA as the template and GAPDH as the internal reference, the expression levels of CCNA2 and E2F1 were calculated using the 2
Western blotting
Cellular proteins were extracted using lysis buffer. An equal amount of cell lysate was separated via sodium dodecyl sulfate polyacrylamide gel electrophoresis using a 10% gel and then transferred onto a polyvinylidene fluoride membrane. The membrane was blocked, incubated with the indicated primary antibody (1:200), and then incubated with a secondary antibody (1:1,000). The cyclin A2 and E2F1 antibodies were purchased from Abcam (Cambridge, MA, USA).
Cell transfection
For siRNA-mediated knockdown, cells were transiently transfected using Lipofectamine 2000 (GenePharma, Shanghai, China) according to the manufacturer’s instructions. The siRNA sequences (sense strand) are shown in Table S2. The lentiviral vector used to overexpressession and interferes with E2F1 was constructed by GenePharma.
Cell proliferation assay
The Cell Counting Kit-8 (CCK-8) assay was used to detect cell proliferation. Cell suspensions (100
Colony formation assay
MDA-MB-231 and MDA-MB-468 cells infected with CCNA2-si and E2F1-si for 72 h were seeded in a 6-well plate with 300 cells per well. Three parallel replicate wells were set up for each group, incubated at 37
Transwell assay
Cells that were serum-starved were resuspended in serum-free medium and seeded into the upper chamber of a Transwell insert (Shanghai) with 8
Wound healing assay
The transfected cells were seeded in a 24-well plate and cultured for 24 h. A linear wound was generated, and the cells were washed three times with PBS. Then, complete medium was added, and the cells were cultured for 36 h. Images were taken at 0 and 36 h, and the scratched area was recorded.
Cell apoptosis analysis
We used the terminal deoxynucleotidyl transferase dUTP nick end labeling (TUNEL) kit to detect apoptosis in TNBC tumor cells. The transfected cells were seeded on glass sheets in a 6-well plate (5
Luciferase reporter gene assay
The CCNA2 promoter region was amplified and then subcloned into the pGL3-Basic vector (GenePharma) for the luciferase assay. The cells were co-transfected with the pGL3 vector and cultured for 48 h, and then the luciferase reporter gene detection system (Promega) was used. To further clarify the transcriptional regulation of CCNA2 via E2F1, a wild-type or mutant binding site of CCNA2 was synthesized for luciferase reporter gene analysis.
In vivo tumorigenesis in nude mice
BALB/c female nude mice (Charles River, Beijing, China) were randomly divided into two groups, with five in each group. All mice were kept in a specific pathogen-free environment. MDA-MB-468 cells in which endogenous CCNA2 was knocked down and then stably overexpressed were regarded as the CCNA2-si group. The control group was designated as the negative control (NC) group. After disinfecting the skin with alcohol, the cells were implanted into the upper abdomen of the nude mice (8
Statistical analysis
All data are expressed as the mean
Results
CCNA2 is overexpressed in breast cancer based on The Cancer Genome Atlas (TCGA) database
CCNA2 mRNA level was found to be upregulated in breast cancer tissues compared with the levels in adjacent tissues based on TCGA database (Fig. 1A). Further subgroup analysis showed that the expression of CCNA2 mRNA was upregulated in different pathological stages compared to that in stage I (Fig. 1B). Multivariate Cox regression analysis showed that the prognosis of breast cancer patients with high expression of CCNA2 is poor. It was also found to be an independent risk factor for the prognosis of breast cancer patients with an HR (95% CI) of 1.20 (1.02–1.40) (Fig. 1C).
Cyclin A2 (CCNA2) is overexpressed in breast cancer based on TCGA database. (A) CCNA2 mRNA is upregulated in breast cancer tissues compared with adjacent tissues. (B) The expression of CCNA2 in different pathological stages. (C) Multivariate Cox regression analysis of breast cancer prognosis.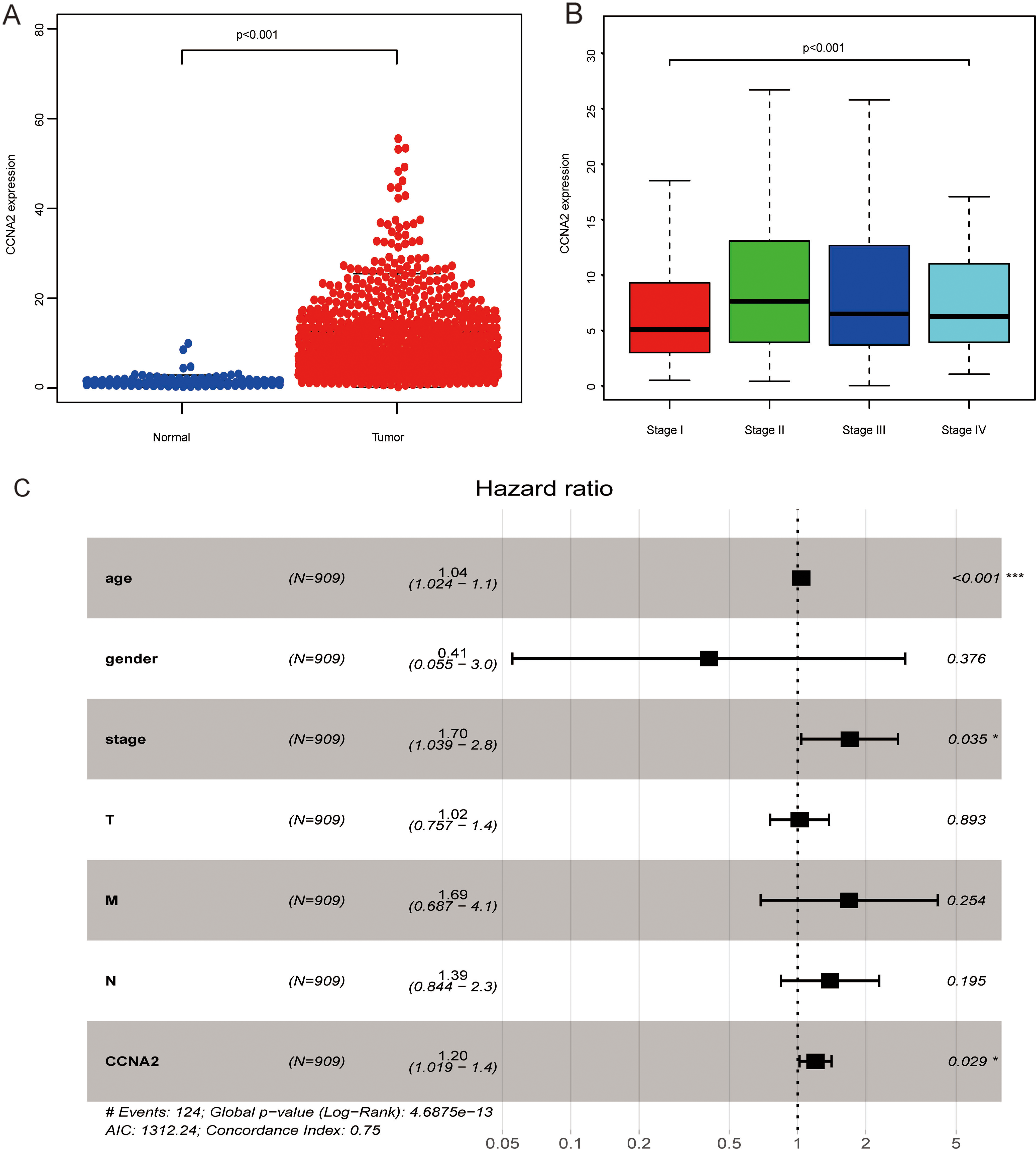
Cyclin A2 mRNA and protein expression levels were upregulated in 15 pairs of TNBC tissues compared to those in adjacent tissues (Fig. 2A and B). Immunohistochemistry results showed that cyclin A2 is primarily located in the nucleus; notably, the positive expression rate of cyclin A2 in TNBC tissues was significantly higher than that in adjacent tissues (Fig. 2C). Moreover, the expression levels of cyclin A2 in multiple breast cancer cell lines, including BT-20, HS578T, MDA-MB-231, and MDA-MB-468 cells, were higher than those in MCF10A cells (Fig. 2D). Interestingly, cyclin A2 was the most abundantly expressed protein in MDA-MB-231 and MDA-MB-468 cells. Therefore, these two cell lines were used for the subsequent in vitro studies.
CCNA2 is overexpressed in triple negative breast cancer (TNBC). (A) CCNA2 mRNA is upregulated in TNBC tissues compared with adjacent tissues, analyzed by quantitative PCR (qPCR). (B) CCNA2 expression is higher in TNBC tissues than in adjacent tissues, analyzed by western blot (WB). (C) Correlation between immunohistochemical grading and CCNA2 expression in TNBC. (D) mRNA and protein levels in a normal cell line (MCF10A cells) and four breast cancer cell lines (HS578T, MCF-7, MDA-MB-231, and MDA-MB-468 cells) via qPCR and WB, respectively. Data were compared using a Student’s 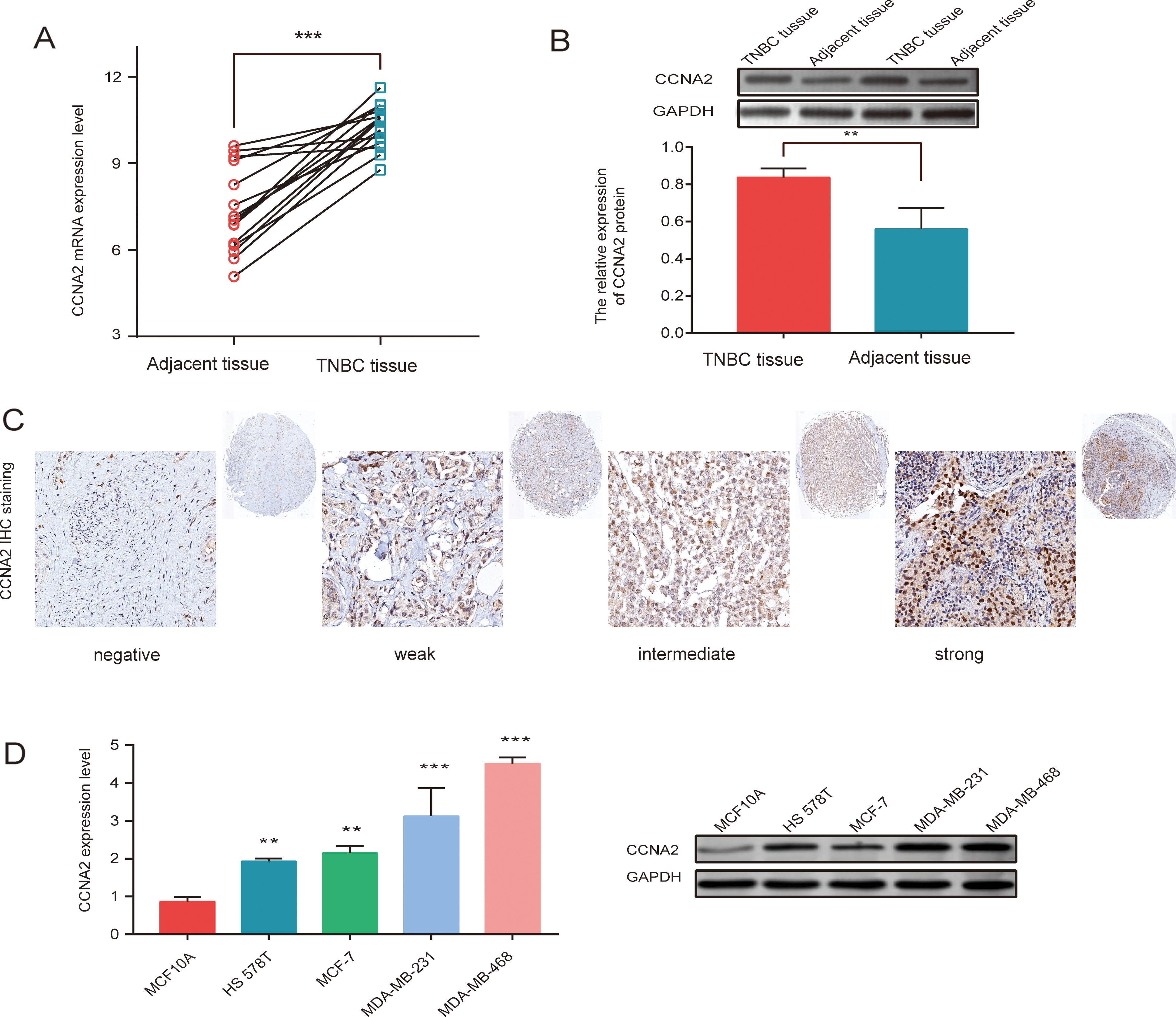
To study the biological functions of CCNA2, we generated a stable CCNA2 knockdown cell line using both MDA-MB-231 and MDA-MB-468 cells (
CCNA2 promotes the proliferation, invasion, and migration but inhibits apoptosis of MDA-MB-231 and MDA-MB-468 cells in vitro. (A) The CCNA2 knockdown effects were confirmed via WB. Knockdown of CCNA2 inhibited (B) cell proliferation, determined using CCK-8 kit and (C) colony formation assays, in MDA-MB-231 and MDA-MB-468 cells. (D) Knockdown of CCNA2 inhibited invasion, analyzed via a transwell assay in MDA-MB-231 and MDA-MB-468 cells. (E) Knockdown of CCNA2 suppressed migration as shown via a wound-healing assay in MDA-MB-231 and MDA-MB-468 cells. (F) Knockdown of CCNA2 promoted cell apoptosis as shown with TUNEL detection in MDA-MB-231 and MDA-MB-468 cells. (G) Knockdown of CCNA2 inhibited cell cycle-related genes expression in MDA-MB-231 and MDA-MB-468 cell lines. Data were compared using a Student’s 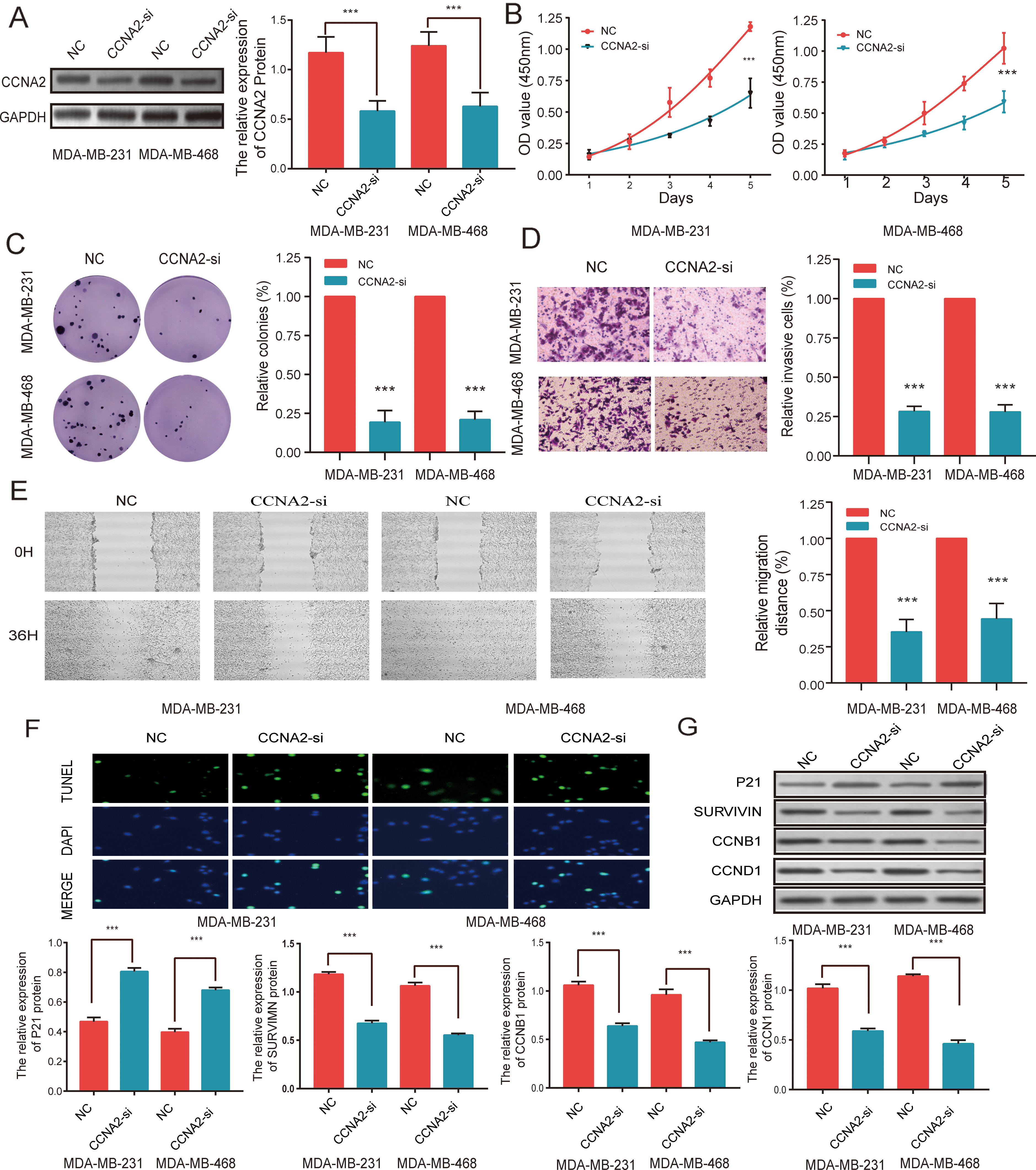
We further investigated the metastatic ability of TNBC cells with different CCNA2 expression levels. In the transwell assay, cells treated with CCNA2-si were significantly less invasive compared to the cells in the NC group (
Using the TUNEL assay, we observed that CCNA2-si promoted apoptosis in both MDA-MB-231 and MDA-MB-468 cells (Fig. 3F); moreover, the expression of cell cycle-related proteins, including SURVIVIN, CCNB1, and CCND1, was significantly reduced in the CCNA2-si group, compared to that in the NC group (Fig. 3G).
Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis indicated that upregulated differentially expressed genes (DEGs) in TNBC included those genes that are involved in the spliceosome formation, pyrimidine metabolism, and cell cycle pathways (Fig. 4A). Functional annotation was performed using representative Gene Set Enrichment Analysis (GSEA) (Fig. 4B). Among the gene sets, G2 M_CHECKPOINT, CLEE_CYCLE, and MTTOTIC_SPINDLE pathways were well established in TNBC (Fig. 4C–E).
Pathway enrichment analysis in TNBC tissues. (A) KEGG analysis of TNBC. (B) Enrichment analysis of genes that target CCNA2. (C–E) Representative Gene Set Enrichment Analysis (GSEA) plots enriched in TCGA database of a TNBC group with high levels of CCNA2 expression. Data were compared using a Student’s 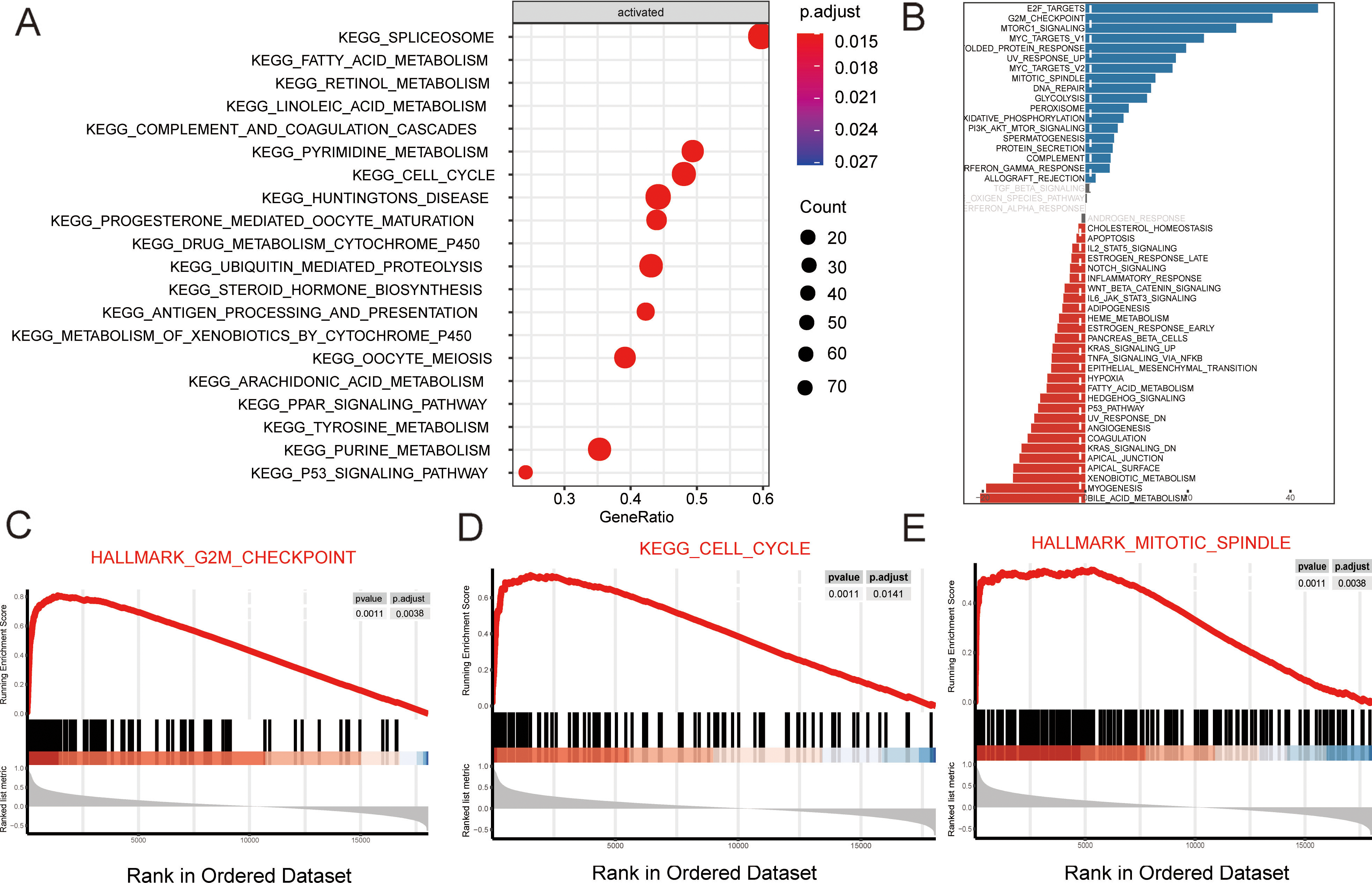
To investigate the effect of CCNA2 on TNBC cell proliferation in vivo, we generated MDA-MB-468 cells stably overexpressing CCNA2, which were injected subcutaneously into the upper abdomens of nude mice. The volumes and weights of tumors from the MDA-MB-468/NC cells were significantly smaller than those from the MDA-MB-468/CCNA2-si cells (
CCNA2 promotes cell proliferation in vivo. (A, B) Knockdown of CCNA2 reduces tumor weight and volume in mice. (C) CCNA2, and Ki-67 staining of subcutaneous xenograft tumors from the NC and CCNA2-si groups. Data were compared using a Student’s 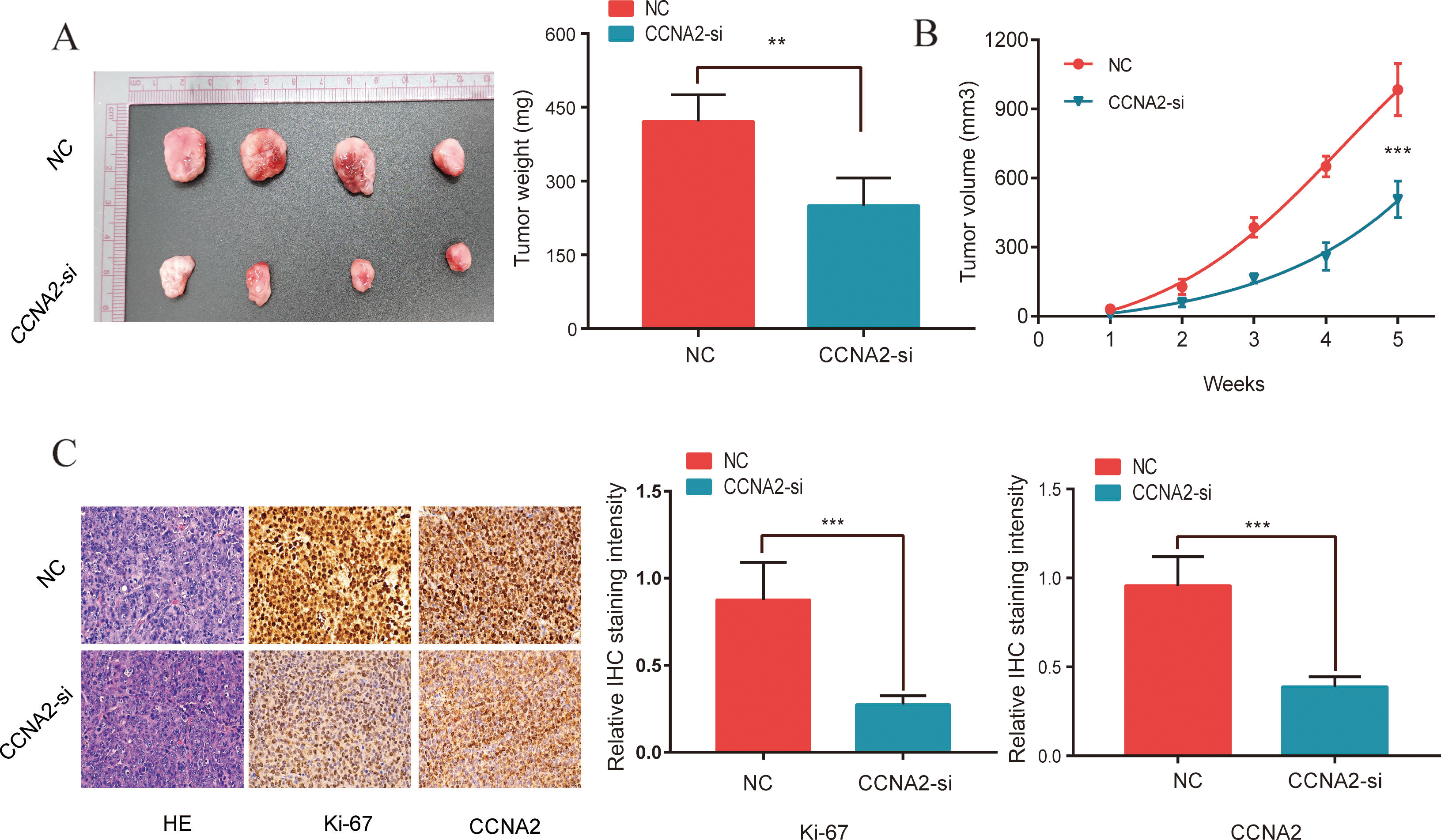
GSEA plots enriched in TCGA showed that CCNA2 is a target gene of E2F1 (Fig. 6A). Results from 25 cancer databases suggested that there is a notable correlation between CCNA2 and E2F1 (Fig. 6B). We further analyzed its correlation through TCGA BRCA cohort database and found that there was a significant correlation between CCNA2 and E2F1, and the correlation coefficient was as high as 0.80 (Fig. 6C). Similarly, using our TNBC cohort, we observed a similar trend and found that CCNA2 expression significantly correlated with E2F1 (
CCNA2 is a transcriptional target of E2F1. (A) Representative Gene Set Enrichment Analysis (GSEA) plots enriched in TCGA database showed that CCNA2 is the target gene of E2F1. (B) Correlation between CCNA2 and E2F1 in 25 cancer databases; the red mark represents invasive breast cancer. (C) Correlation between CCNA2 and E2F1 mRNA levels in TCGA BRCA cohort. (D) Correlation between CCNA2 and E2F1 mRNA levels in TNBC tissues. (E) Predicted E2F1 binding sequence, GGCGCC, in the CCNA2 promoter 5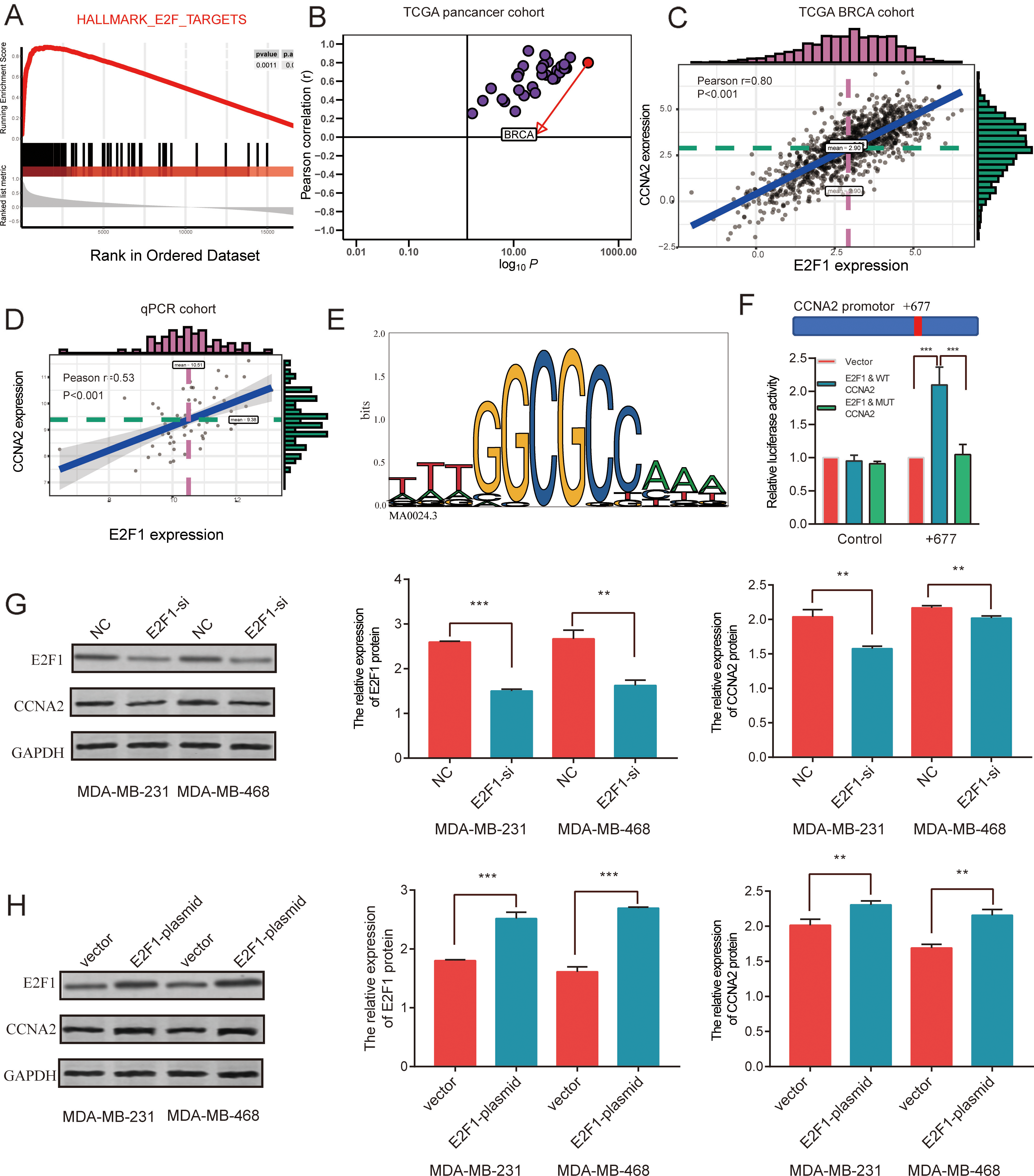
Overexpression CCNA2 increases cell proliferation, invasion, and migration but inhibits apoptosis via E2F1-mediated regulation in TNBC cells. (A) E2F1 silencing downregulates the expression of CCNA2 in MDA-MB-231 and MDA-MB-468 cells, analyzed via WB. (B, C) E2F1 silencing inhibits cell proliferation by downregulating the expression of CCNA2 in MDA-MB-231 and MDA-MB-468 cells, as shown by CCK-8 and colony formation assays. (D) E2F1 silencing inhibits cell invasion by downregulating the expression of CCNA2 in MDA-MB-231 and MDA-MB-468 cells, as shown by transwell assays. (E) E2F1 silencing inhibits cell migration by downregulating the expression of CCNA2 in MDA-MB-231 and MDA-MB-468 cells, as shown by wound healing assays. (F) Silencing E2F1 promotes cell apoptosis by downregulating the expression of CCNA2 in MDA-MB-231 and MDA-MB-468 cells, as detected by TUNEL staining. Data were compared using a Student’s 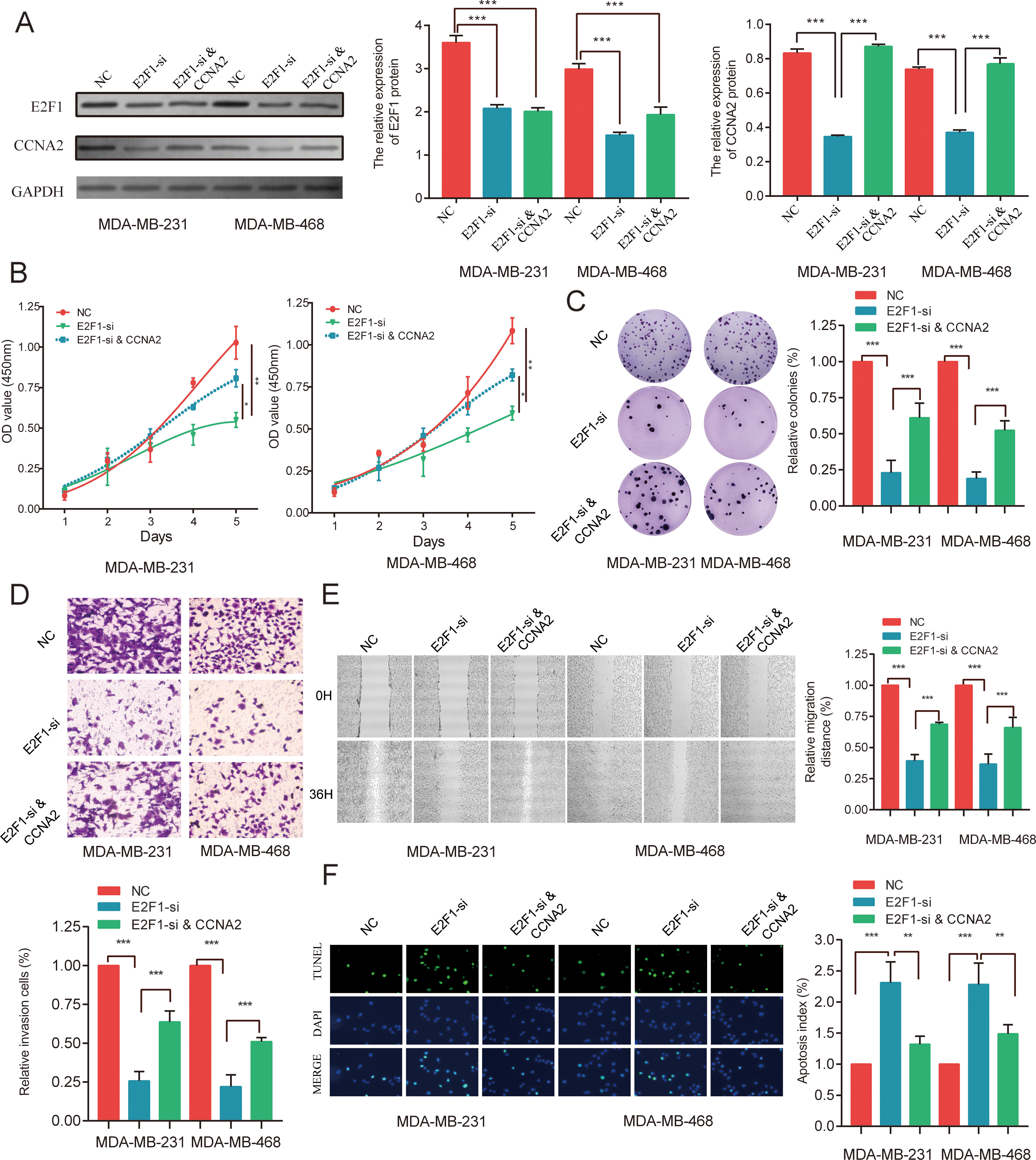
The scientific hypothesis of E2F1 mediates the transcriptional regulation of CCNA2 expression to promote TNBC cell proliferation.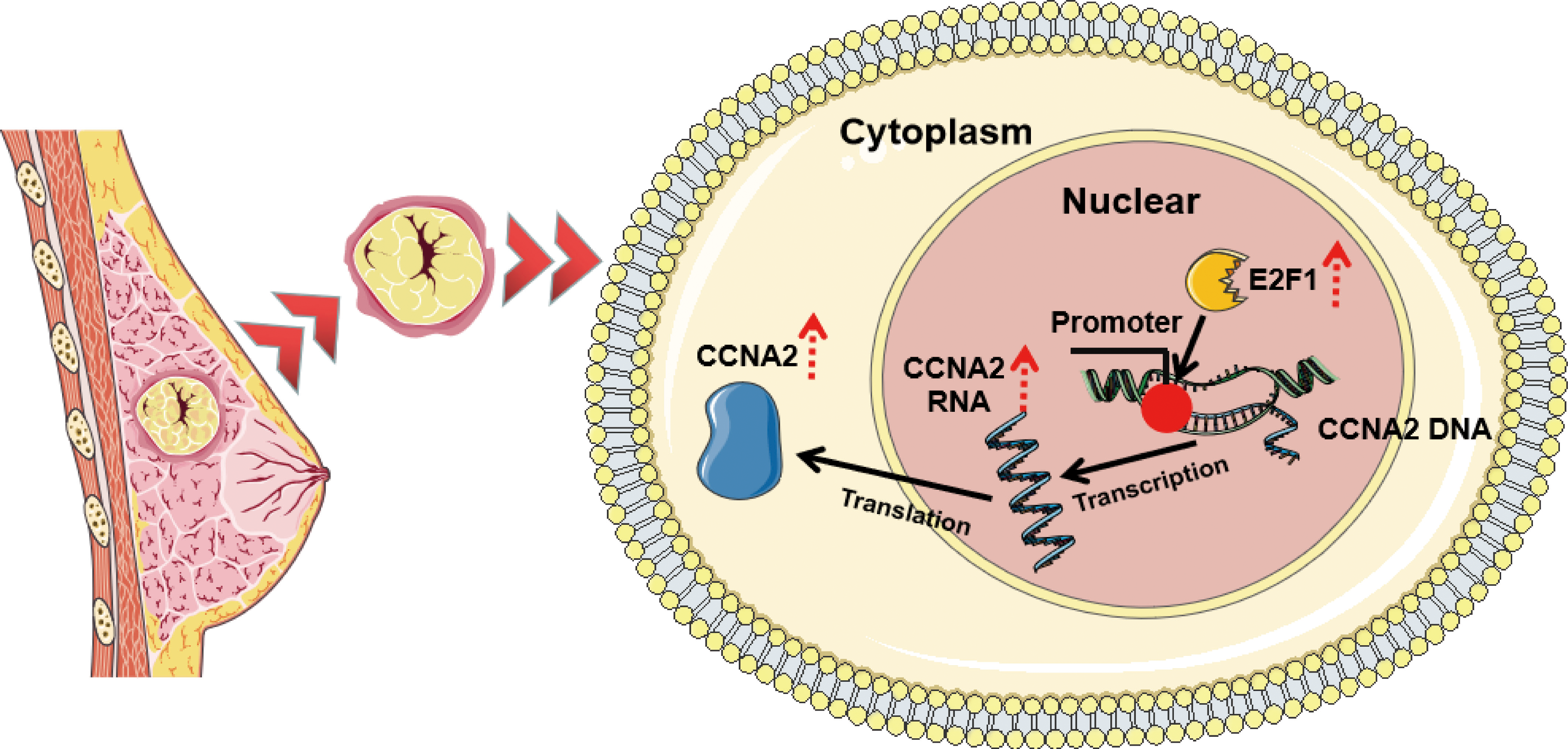
Cell rescue experiments revealed that the expression of cyclin A2 was significantly downregulated between the E2F1-si and E2F1-si plus CCNA2 overexpression groups using both MDA-MB-231 and MDA-MB-468 cells (
Scientific hypothesis diagram
In summary, upregulation of the transcription factor E2F1 mediates the transcriptional regulation of CCNA2, which promotes an abundance of CCNA2 by increasing its transcription level leading to its high expression, which ultimately promotes the proliferation of TNBC cells (Fig. 8).
Discussion
Our study revealed that E2F1 and CCNA2 were overexpressed in TNBC cells; moreover, they showed a strong positive correlation. Using a luciferase reporter assay, we examined the association between E2F1 and the promoter sequence for CCNA2 and found that E2F1 binds to the CCNA2 promoter at the
Transcriptional regulation plays an important role in promoting cancer cell proliferation. We used UCSC (
In conclusion, for the first time, we demonstrated that E2F1 actively regulates the expression of CCNA2 by binding to a specific region in its promoter. We revealed that overexpressed E2F1 promotes tumor cell proliferation by regulating the transcription of CCNA2 both in vitro and in vivo. Therefore, E2F1 and CCNA2 could be attractive therapeutic targets for effectively inhibiting tumor growth in TNBC.
Abbreviations
CCK-8: cell counting kit-8; DAPI: 4
Author contributions
Interpretation or analysis of data: Yongbin Lu,Tao Zhang and Yi Xiao.
Preparation of the manuscript: Yongbin Lu, Fei Su and Hui Yang.
Revison for important intellectual content: Xiaoling Ling and Yana Bai.
Supervision: Xiaobin Zhang, Hongxin Su and Taozhang.
Funding
This work was founded by the Gansu Provincial Innovation team Funding project (No. 20JR5RA352 and 20JR10FA686), the Lanzhou Guidance Plan Project (No. 2019-ZD-42 and 2020-ZD-75), the First Hospital of Lanzhou University Hospital Found (NO. ldyyyn2019-82 and ldyyyn2018-66).
Competing interests
The authors have declared that no competing interest exists.
Supplementary data
The supplementary files are available to download from
sj-pdf-1-cbm-10.3233_CBM-210149.pdf - Supplemental material
Supplemental material, sj-pdf-1-cbm-10.3233_CBM-210149.pdf
