Abstract
BACKGROUND:
Secreted phosphoprotein 1 (SPP1), also known as osteopontin (OPN), is a multifunctional protein expressed in diverse normal tissues, and functionally is involved in cellular matrix and signaling processes. Many studies have linked SPP1 to pathophysiological conditions including cancer.
OBJECTIVE:
The aim of this study is to evaluate the 3’UTR length of SPP1 gene in glioblastoma cell line.
METHODS:
3’ Rapid Amplification of cDNA End (3’-RACE) was used to determine the 3’ end of SPP1 gene. APAatlas data base, GEPIA web server, and miRcode were also used to extract related information and bioinformatic analysis part.
RESULTS:
In this study we show that SPP1 gene undergoes Alternative cleavage and Polyadenylation (APA) mechanism, by which it generates two 3’ termini, longer isoform and shorter isoform, in glioblastoma derived cell line, U87-MG. Further bioinformatic analysis reveals that SPP1 alternative 3’UTR (aUTR), which is absent in shorter isoform, is targeted by two families of microRNAs-miR-181abcd/4262 and miR-154/872. These miRNAs also target and perhaps negatively regulate NAP1L1 and ENAH genes that are involved in cell proliferation and cell polarity, respectively. Relative expression difference (RED), obtained from RNA-seq data of diverse normal tissues, representing APA usage appears to be negatively correlated with expression of NAP1L1 and ENAH, emphasizing co-expression of SPP1 longer isoform with these two genes, indicating miRNA sponge function of aUTR (longer 3’UTR). Bioinformatic analysis also shows that in normal brain tissue longer APA isoform of SPP1 is expressed; however shorter isoform appears to be expressed in cancer condition.
CONCLUSION:
Together, this study reveals that SPP1 APA isoforms have different pattern in normal and cancerous conditions, which can be considered as a diagnostic and prognostic marker in cancers.
Abbreviations
Introduction
Secreted phosphoprotein 1 (SPP1), also known as osteopontin (OPN), belongs to Intrinsically Disordered Proteins (IDPs) that is a class of proteins lacking any known secondary or tertiary structure [1]. SPP1 is a multifunctional secreted protein that is expressed in a variety of cells such as macrophages, endothelial cells and osteoclasts, and serves as lymphokine, integrin binding protein, and hydroxyapatite binding protein [2, 3, 4]. It is also associated with various pathophysiological conditions ranging from autoimmune diseases [5] to osteoarthritis [6], inflammatory diseases [7], and cancers [3].
Overexpression of SPP1 gene has been reported in a variety of cancers such as lung cancer [8], head and neck cancer [9], colorectal cancers [10, 11], breast cancer [12, 13, 14], liver cancer [15], prostate cancer [16, 17] and brain cancers [18, 19, 20]. Increased level of SPP1 in blood and tumor tissues is associated with poor prognosis [21, 22]. SPP1 contributes to cancer progression in a variety of ways including enhancing angiogenesis, increasing cell proliferation, migration and invasion, as well as cancer cell self-renewal [18].
Alternative cleavage and polyadenylation (APA) is a mechanism of post transcriptional gene regulation, by which multiple distinct 3’ termini on the same messenger RNA (mRNA) and long non-coding RNA (lncRNA) are generated [23], that may differ in functions. 3’UTR of mRNAs (and lncRNAs) harbors key nucleotide sequences or signals, called cis-regulatory elements which are very crucial in defining the fate of those mRNA (and lncRNAs). These regulatory elements affect translation, stability (or half-life), and localization of mRNAs [24]; thus any changes to the pattern of polyadenylation mechanism may lead to changes in the fate of affected mRNAs and lncRNAs, which may subsequently lead to pathophysiological conditions.
Even though overexpression of SPP1 gene has been repeatedly reported in a variety of cancers, the mechanisms underlying this overexpression have not been addressed yet. In this study, we aimed to investigate the polyadenylation status of SPP1 gene in glioblastoma. Experimental and bioinformatics analyses have been conducted to delineate the alternative cleavage and polyadenylation status of SPP1 gene. Here we explore how SPP1 APA might contribute to the elevated expression level of SPP1 gene in glioblastoma.
Materials and methods
Cell culture and RNA extraction
U87-MG cell line was used in this study. Cells were cultured in DMEM medium supplemented with 1% pen/strep and 10% FBS and maintained at humidified 37
3’ Rapid Amplification of cDNA End (3’RACE)
Total RNA (1
GEPIA web server
For comparing the expression of SPP1 gene between tumor and normal tissues, GEPIA web server (
APAatlas database analysis
APAatlas data base (
RNA-seq (SAAP-RS) algorithm was applied to calculate polyadenylation sites on GTEx RNA-seq data.
miRNA prediction
UCSC genome browser was employed to visualize any potential miRNA seeding site on APA-3’UTR. Using Track Hubs, miRcode which predicts microRNA target sites in GENECODE transcripts (hg19) was imported to UCSC Genome Browser to predict and visualize potential miRNAs targeting aUTR.
Results
SPP1 is highly expressed in glioblastoma
To evaluate the expression of SPP1 in glioblastoma we used Gene Expression Profiling Interactive Analysis (GEPIA), which provides gene expression data (RNA-seq) of 9,736 tumors and 8,587 normal samples from the TCGA and the GTEx projects. Analyzing the expression of SPP1 in 163 glioblastoma and 207 matched normal samples, showed a significant upregulation of this gene in tumor tissues (Fig. 1).
SPP1 gene expression in glioblastoma vs normal matched tissues (obtained from GEPIA).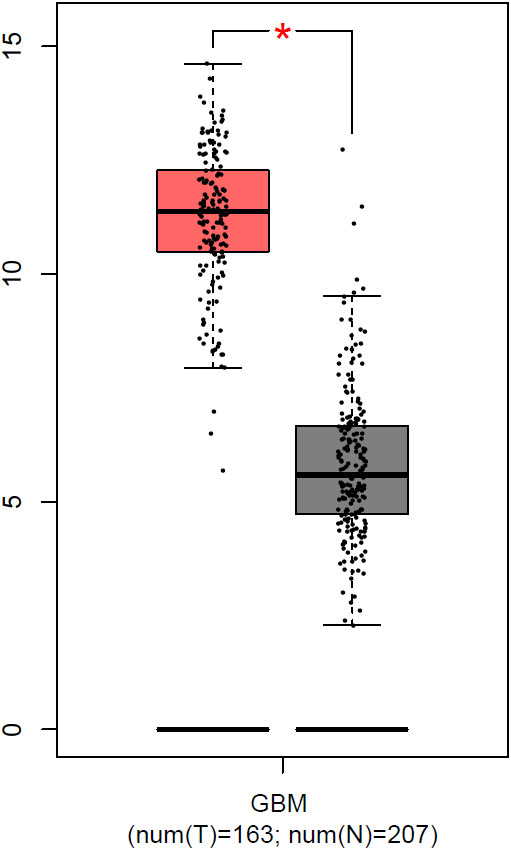
In human genome, more than half of genes utilize alternative polyadenylation mechanism of gene regulation to generate different length of 3’-UTR [26]. In an effort to study the 3’UTR of SPP1 gene we have taken advantage of 3’RACE method (Fig. 2a) to figure out whether this gene has single 3’UTR or multiple ones in glioblastoma derived U87-MG cell line. Designing specific forward primers against beginning of SPP1 last exon enabled us to capture all possible different 3’UTRs of this gene. According to 3’RACE approach, two rounds of PCR were performed in a semi-nested manner (round 1 with forward 1 and universal reverse primers, and round 2 with forward 2 and universal reverse primers) to amplify the 3’terminus of the SPP1 transcripts (Fig. 2a). By separation of PCR amplified fragments with agarose gel-electrophoresis we noticed that two clear bands (shorter and longer bands) have been amplified (Fig. 2b), indicating SPP1 may have at least two cleavage and polyadenylation sites at 3’UTR. Both shorter and longer isoforms appear to be expressed at same level (Fig. 2c). For further validation, the corresponding bands were sequenced, and aligned to the human genome, which confirmed that the amplified bands belong to SPP1 gene 3’end. To depict the deference between two isoforms, next the sequenced reads were aligned with the sequence of SPP1 RefSeq retrieved from GeneBank (NCBI) using Multiple Sequence Alignment by CLUSTALW (
3’RACE reveals alternative polyadenylation isoforms of SPP1. a) Schematic illustration of 3’RACE work flow. cDNA synthesis using anchored oligo d(T) followed by semi-nested PCR reveals 3’ terminus status of the gene. b) Experimental validation of APA in SPP1 using 3’RACE. c) quantification of SPP1 alternative isoforms expression.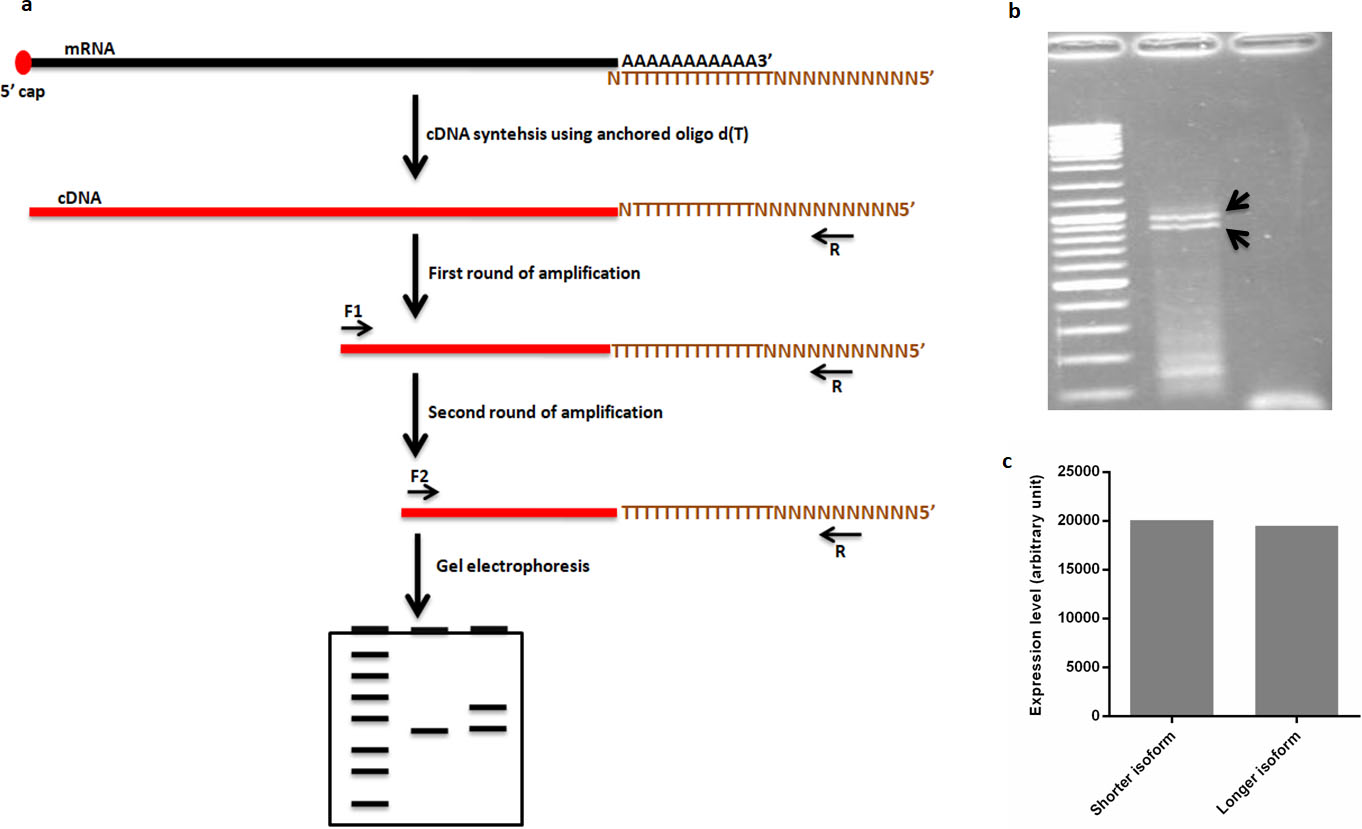
Alignment of SPP1 RefSeq 3’UTR with nucleotide sequence of PCR amplified SPP1 APA isoforms. Spp1Ref: 3’UTR of SPP1 RefSeq. Shorter: shorter amplified isoform; Longer: longer amplified isoform.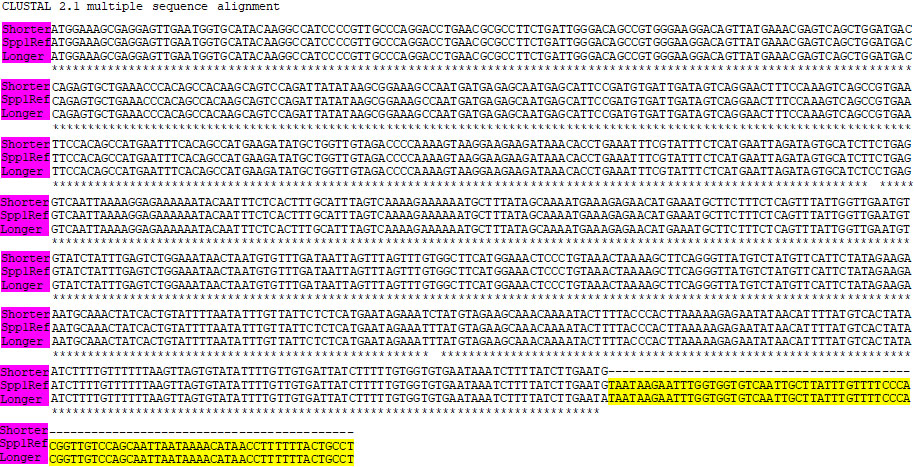
aUTR of SPP1 harbors two seeding sites for two families of microRNAs (miR-181abcd/4262 on the 5’ side of aUTR and miR-154/872 on the 3’ side of a UTR). a) The figure is an screen shot of UCSC genome browser showing the aUTR of SPP1 has seeding sites for abovementioned. The prediction is performed by using miRcode. b) Schematic representation of SPP1 gene structure showing start codon, stop codon, and polyadenylation sites at the 3’ end where habors miRNA seding sites.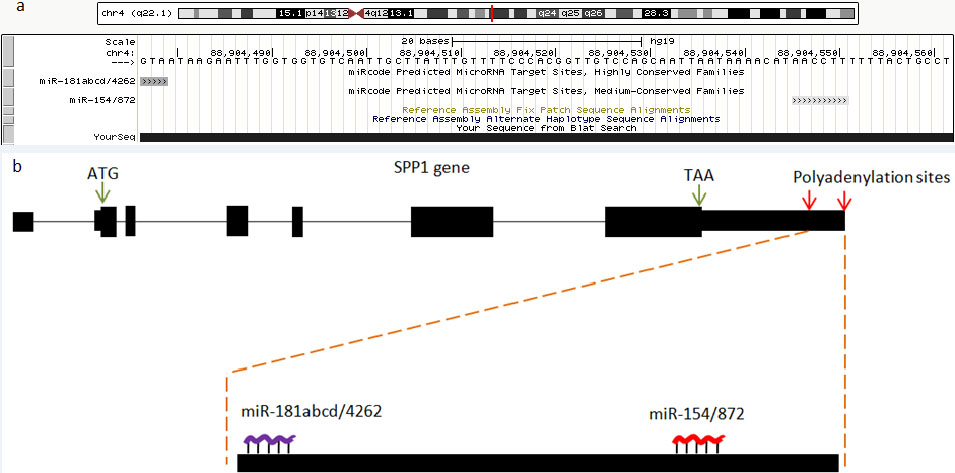
Given that miRNA target sites are mostly located in 3’UTR of mRNAs [23], we sought to figure out whether alternative 3’UTR (aUTR) of SPP1 is targeted by already known miRNAs. Transcriptome-wide microRNA target prediction tool [27] implemented on UCSC genome browser (
SPP1 uses different cleavage and polyadenylation sites in different tissues
To determine whether APA of SPP1 exhibits tissue specificity, we used APAatlas database (
Polyadenylation site position on SPP1 3’UTR. a) There are no significance changes in the position of poly A site in different parts of brain according to RNA-seq data of GTEx project. b) Poly A site position significantly differs across various normal tissues of human body.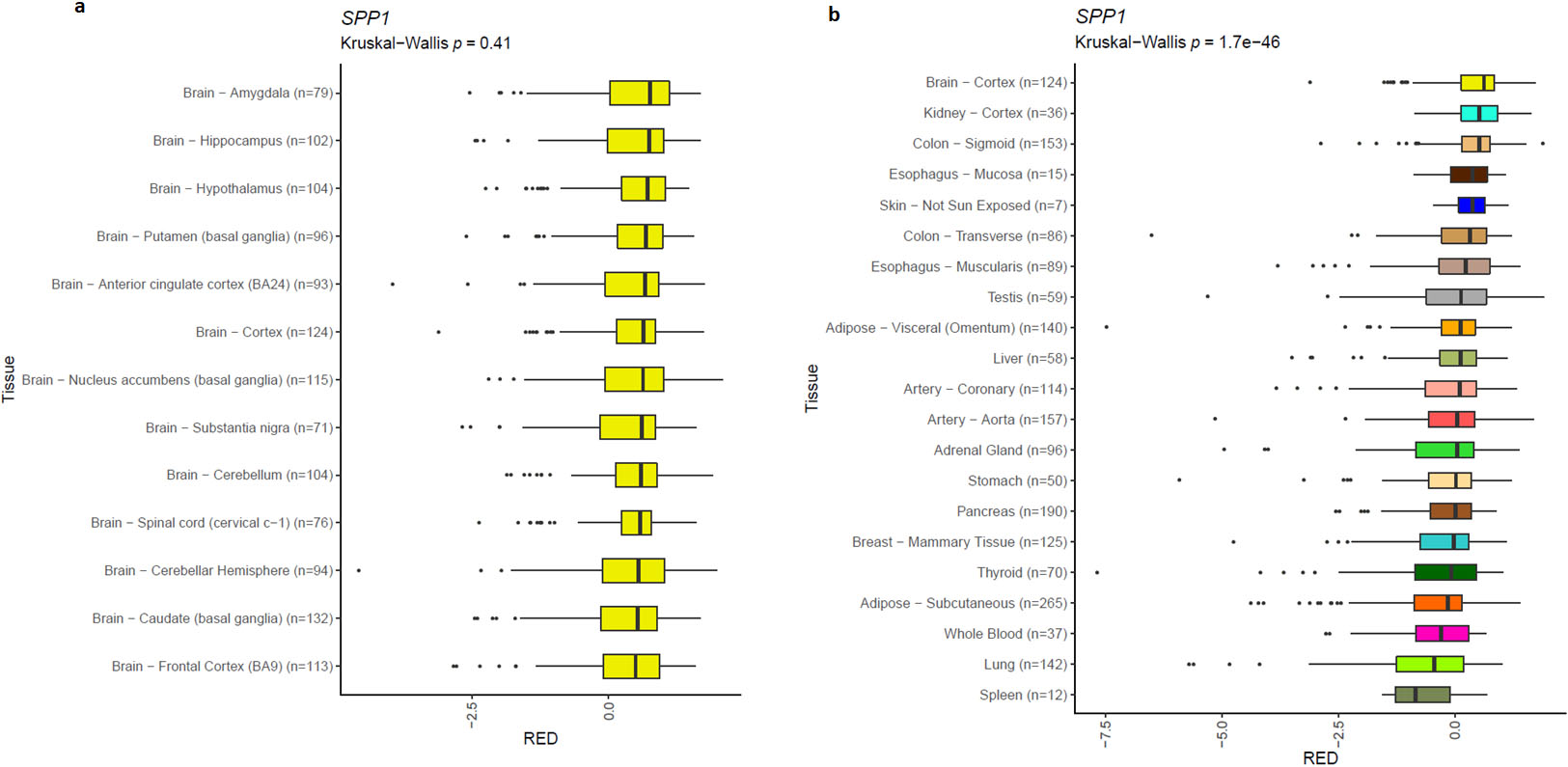
Given that aUTR of SPP1 is targeted by two families of microRNAs (Fig. 4), we sought to figure out if these miRNAs have other potential targets. Using miRcode prediction we found that miR-181abc/4262 targets NAP1L1 and ENAH genes, beside SPP1. NAP1L1 is a member of nucleosome assembly protein family involved in DNA replication and cell proliferation, and ENAH gene is associated with actin and cell polarity. To check how loss of aUTR in SPP1 affects NAP1L1 and ENAH through miR-181abc/4262, we checked the correlation between SPP1 aUTR presence and expression of NAP1L1 and ENAH genes. For this analysis, relative expression difference (RED) representing APA usage [28], from APAatlas data base was used, which showed that SPP1 APA is negatively correlated with the expression of NAP1L1 and ENAH (Fig. 6a and b) indicating SPP1 aUTR is likely functions as miRNA sponge to reduce miRNA suppression effect on NAP1L1 and ENAH. miRNA-154/872 also targets both SPP1 and NAP1L1 which further implies that aUTR of SPP1 may function as miRNA sponge.
NAP1L1 and ENAH gene expression is negatively correlated with SPP1 relative expression difference (RED) representing APA usage. RED is calculated using log2 (DN/UP); DN: number of reads mapped into the downstream regions of polyA site; UP: number of reads mapped into the upstream regions of polyA site [28].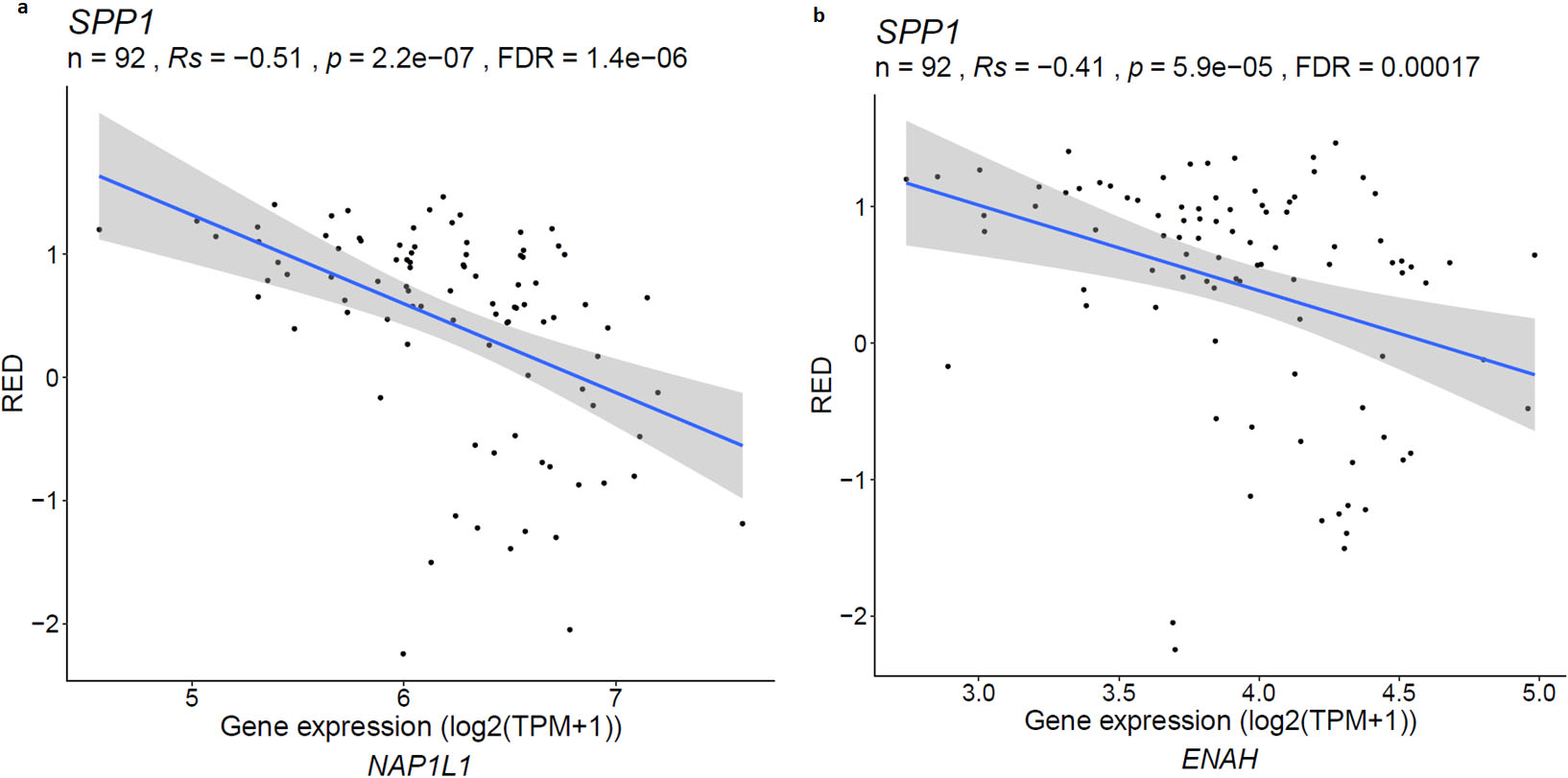
In this study we have identified two alternative polyadenylation (APA) isoforms of SPP1 gene in glioblastoma derived U87-MG cell line. Both isoforms are expressed at same level. The shorter isoform loses a 85-nucelotide fragment at the end of 3’UTR. Cleavage and polyadenylation process in these two isoforms appears to be mediated through canonical AAUAAA consensus sequence, which resides 17 and 21 nucleotides upstream of cleavage site of shorter and longer isoforms, respectively. Further bioinformatic analysis revealed that longer isoform of SPP1 is expressed in normal brain samples; however, in other tissues the shorter form is mostly predominant. Longer isoform harbors seeding sites for two miRNA families -miR-181abcd/4262 (a highly conserved family) and miR-154/872 (medium conserved family). Besides SPP1, miR181abcd/4262 and miR-154/872 target other genes as well. For example, miR-181abcd/4262 targets NAP1L1 and ENAH, and miR-154/872 targets NAP1L1. Interestingly, we noticed that there is a negative correlation between usage of shorter isoform and expression of these genes, emphasizing miRNA sponging role of SPP1 aUTR.
SPP1 has been shown to be deregulated in pathophysiological conditions including cancers. Due to the nature of this protein, it can affect in autocrine and paracrine manner on its targets; thus can mediate cellular cross talk and can be an influential factor in tumor microenvironment [3]. SPP1 longer 3’UTR isoform is predominant in normal brain tissues. However, we found that glioblastoma cell line expresses shorter isoform as well. This finding is in line with another study which has showed that most of APA genes have shorter APA isoforms in cancers that can avoid miRNA mediated repression [29]. We also showed two miRNAs can target aUTR of SPP1. These two miRNAs likely repress SPP1 expression if longer isoform is used. However, shorter SPP1 isoform may avoid miRNA-mediated repression and can be more stable, by which influences the transformation process of cancer. These findings are in line with the result of another group that showed SPP1 gene is upregulated in liver cancer [15]. This group also found that SPP1 gene is targeted by miR-181c and this interaction can contribute to posttranscriptional regulation of this gene [15] which leads to upregulation of SPP1 gene.
In human tissues, it has been demonstrated that APA sites usage is biased [30], leading to the tissue-specificity of APA. Generally brain uses distal polyadenylation sites, while in testis, for example, proximal poly A site predominantly is used [23], which further supports our data and analysis. Thus, it is deduced that shorter isoform found in U87-MG cell line is a deregulated form of SPP1 isoform in glioblastoma, which warrants further investigation to delineate its role in cancer initiation and progression.
Kejevaska et al., 2017 [18] also found overexpression of SPP1 gene in both glioblastoma and U87-MG cell line as well as other glioblastoma derived cell lines. Authors have demonstrated in this study that interaction of SPP1-CD44 is very important in self-renewal and pluripotency of glioma initiating cells [18]. Thus this study emphasizes the importance of SPP1 gene in initiaing the cancer progression in glioblastoma. Our finding that SPP1 generates two 3’UTR termini (short and long) in U87-MG cell line also further emphasizes the importance of APA mechanism in generating more stable variants that can contribute to the initiation and progression of glioblastoma.
Although the best known consequence of APA is its effects on miRNA functions, other mechanisms also may be affected due to alternative usage of UTR. Two most important of such processes are RNA binding proteins (RBPs) and long non coding RNAs (lncRNAs) binding to 3’UTR [23]. How changes of poly A site in SPP1 influence these processes, and how these changes are linked to cancer also need further investigation.
In sum, cleavage and polyadenylation site appears to be tissue specific. Its deregulation leads to pathophysiological conditions and can act as a biomarker for diagnostic and prognostic forecast. We found that SPP1 has at least two APA isoforms. Even though, longer isoform supposedly should be expressed in brain tissues, we found the expression of shorter isoform in brain cancer derived cell line as well, indicating deregulation of poly A site in SPP1 gene in brain cancer. aUTR of SPP1 harbors seeding site for at least two group of miRNAs. Shorter isoform usage may affect SPP1 gene expression by avoiding from miRNA mediated-suppression and also can abolish sponge effect that longer isoform may apply on shared miRNAs, by which it can lead to suppressing other common targets of miRNAs. Further functional study is needed to evaluate sponge activity of SPP1 gene regarding abovementioned miRNAs.
Authors contributions
Conception: Majid Mehravar, Fatemeh Ghaemimanesh, Ensieh M. Poursani.
Interpretation or analysis of data: Majid Mehravar.
Preparation of the manuscript: Majid Mehravar.
Revision for important intellectual content: Fatemeh Ghaemimanesh, Ensieh M. Poursani.
Supervision: Fatemeh Ghaemimanesh, Ensieh M. Poursani.
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-210135.
Footnotes
Acknowledgments
Authors would like to thank the GEPIA and APAatlas for their free availability.
The results shown here are in part based upon data generated by the TCGA Research Network: https:// www.cancer.gov/tcga.
