Abstract
In this paper, we have studied the role of chromosomal abnormalities in the expression of small nucleolar RNAs (snoRNAs) of radioresistant (K562) and radiosensitive (HL-60) leukemia cell line. Cells were exposed to an X-ray dose of 4 Gy. SnoRNA expression was investigated using NGS sequencing.
The distribution of expressed snoRNAs on chromosomes has been found to be different for two cell lines. The most significant differences in the expression of snoRNAs were found in the K562 cell line based on the analysis of the dynamics of log2fc values. The type of clustering, the number and type of snoRNAs slightly differed in the chromosomes with trisomy and monosomy and had a pronounced difference in pairs with marker chromosomes in both cell lines. In this study, we have demonstrated that chromosomal abnormalities alter the expression of snoRNA after irradiation. Trisomies and monosomies do not have such a noticeable effect on the expression of snoRNAs as the presence of marker chromosomes.
Introduction
Cancer treatment faces a serious challenge of tumor radioresistance [1, 2] and chemoresistance [3]. Progress in treatment is possible only if there is an adequate protocol developed in accordance with the sensitivity of the tumor to the therapy used. To determine the cancer tumor radiosensitivity, multiple markers, in particular, small non-coding RNAs, have been studied [4, 5, 6, 7].
To date, numerous data on the role of long non-coding RNAs and micro RNAs in the mechanisms of chemo- and radioresistance of cancer tumors have been reported [8, 9, 10, 11, 12]. However, the contribution of small nucleolar RNA to the radioresistance of cancer tumors is still unclear [13, 14].
Small nucleolar RNAs (snoRNAs) are a class of RNAs that regulate biogenesis of ribosomal and transport RNAs in eukaryotic nucleolus [15]. SnoRNAs are divided into two groups depending on their structure and main function: H/ACA and C/D box snoRNAs. The C/D box snoRNAs are responsible for the 2’O-methylation of ribose, and the H/ACA box snoRNAs provide pseudouridylation [16].
In the treatment of leukemia, one of the key factors in detection is genetic changes, such as a deletion of chromosome 13q14 in chronic lymphocytic leukemia [17]. The level of expression in this case of small non-coding RNA plays an important prognostic value. Expression values are correlated in both circulating siRNAs and tumor cells, and can serve as an indicator of a therapeutic response. Elevated levels of snord112–114 snoRNAs have been found in acute promyelocytic leukemia, which also indicates their role in the development of cancer [18, 19]. SnoRNAs could be involved in oncogenesis. For example, SNORD50A and SNORD50B directly bind and block K-Ras (proto-oncogene) and are not expressed during cancer progression in humans [20]. So, in non-small cell lung cancer, snoRNA78 contributes to oncogenesis. snoRNAU50 has been detected in breast cancer [21]. In prostate cancer progression, the expression of SNORA42 is significantly increased compared to normal tissue [22].
Warner et al. and Valleron et al. have demonstrated a decrease in the snoRNA expression in acute myeloid leukemia [23] and acute lymphoblastic leukemia compared with normal cells [24]. In [16], it is shown that the C/D snoRNA SNORD114–1 expression increases in acute myeloid leukemia and correlates with the proliferation of the K562 cell line, but the target suppression of the C/D gene expression induces cell death. A number of other studies have shown that there is no general pattern in the expression change for all snoRNAs in acute promyelocytic leukemia, chronic lymphocytic leukemia and multiple myeloma because different types of snoRNAs change their expression profile in a positive or negative direction [25, 26, 27, 28, 29].
The above studies reveal contradictions in the role of snoRNAs in cancer progression. The dependence of small non-coding RNA expression, in particular snoRNA, on the number of chromosomes and chromosomal abnormalities could be responsible for inconsistencies in these data [30]. For example, in cancer cells the number of deletions and copies of chromosomes contributes to changes in the expression of suppressor genes for tumors and oncogenes [31]. The miRNA expression profiles in the K562 and HL-60 cancer cell lines [32], in chronic lymphocytic leukemia [33] and in neurons [34] depend on the presence of chromosomal aberrations and genomic mutations. A change in expression is observed in miR-486-5p [35], let-7c, miRNA-99a, miRNA-125b, miRNA-155 [36] in patients with trisomy of chromosome 21. The expression level of miR-92b, miR-199-5p and miR-223 miRs increases in patients with monosomy of chromosome 3 [37].
These results suggest that the expression of small non-coding RNAs is dependent on chromosomal abnormalities. Here we present a study of the effect of chromosomal abnormalities on snoRNA expression after the irradiation of K562 and HL-60 cell lines with different levels of chromosomal mutations.
Materials and methods
A radiosensitive cell line of human promyelocytic leukemia HL-60 and a radioresistant cell line of chronic myeloid human leukemia K562 were the objects of research. The cell lines were obtained from the Russian collection of cell cultures (Institute of Cytology RAS, St. Petersburg) [38, 39]. Cancer cells were cultivated at a temperature of 37
The cell lines were exposed to X-ray irradiation for 55 seconds at room temperature and the dose of 4 Gy (photon energy 10 MEV) using the Elekta Synergy linear accelerator (Elekta, Stockholm, Sweden). Prior to irradiation, the cells were placed to 6 well plates transported to and from the irradiation site in a thermostatically controlled container at a temperature of 37
Cell culture survival was evaluated using propidium iodide and acridine orange dye mixtures. A fluorescence microscope was used for the subsequent analysis [40].
The total RNA containing snoRNAs was isolated from the cells using the Absolutely RNA miRNA Kit (Agilent Technologies, USA) 1, 4, and 24 hours after irradiation. Using the Agilent 2100 Bioanalyzer capillary electrophoresis device (Agilent Technologies, USA), the quality of the isolated RNA was evaluated by the 18S/28S RNA ratio. Samples with RIN
The sequencing data were recorded in the FASTQ file format and processed using the GenXProomiRas service. The tables summarizing the data on the snoRNA names and the number of snoRNAs normalized by 10
The localization of snoRNA in chromosomes was determined using the National Center for Biotechnology Information Search database (
Each experiment was repeated tree times and the data were given as
Results
Cell viability
Percentage of the HL-60 and K562 cells with signs of necrosis registered after irradiation at the dose of 4 Gy is shown in Fig. 1. In the control group of the HL-60 cell line, the percentage of dead cells was 5.4%, and, after exposure, it increased up to 21% that is about 4 times higher than that in the control. In the control group of radioresistant K562 cell line, the percentage of dead cells was 7.5% and, after irradiation, this value increased up to 15% (* – Statistically significant difference in relation to the control,
The percentage of cells with signs of necrosis in the control and after radiation exposure at a dose of 4 Gy in cell lines HL-60 and K562.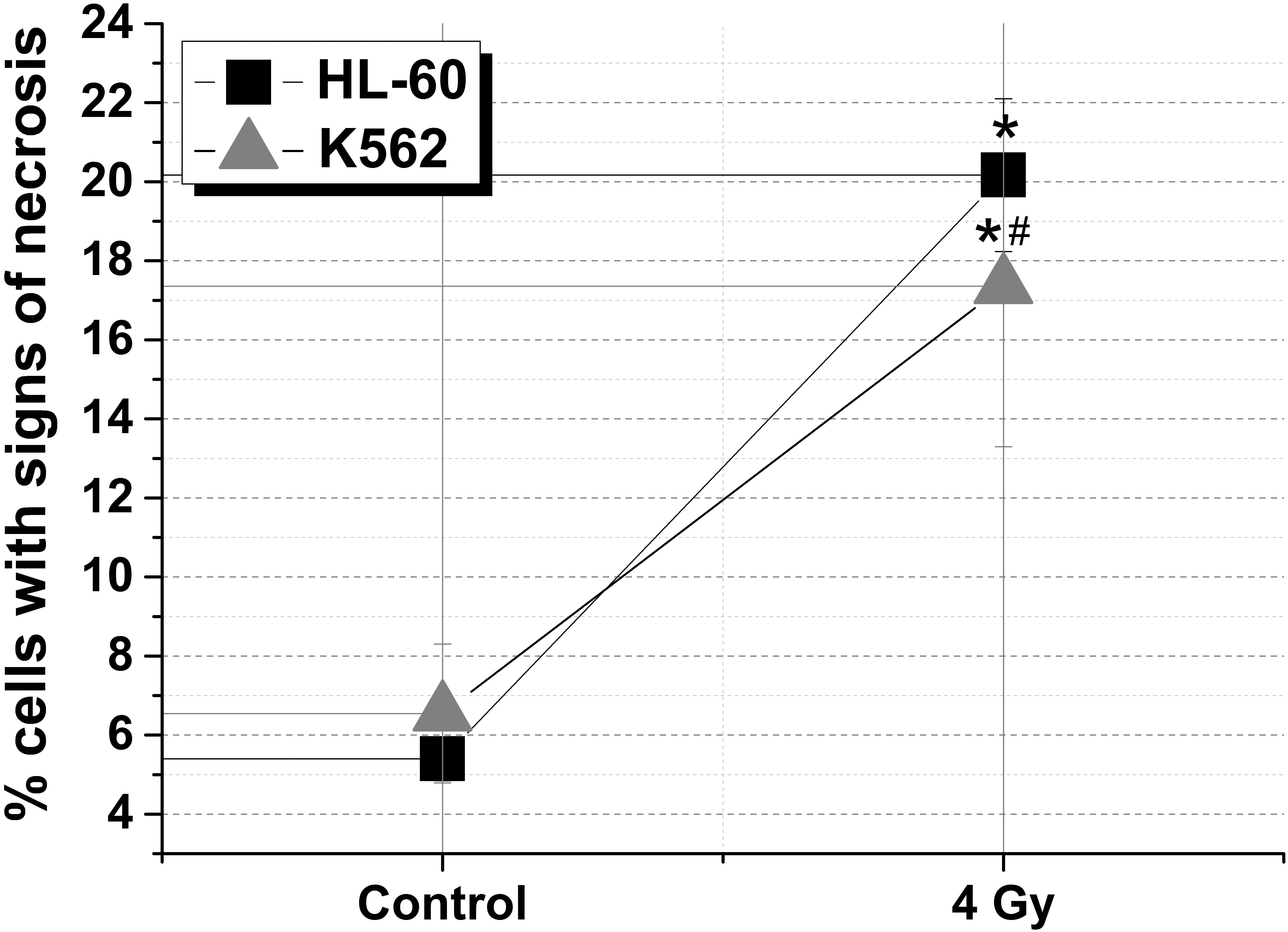
The number of differentially expressed snoRNAs in the K562 and HL-60 cell lines.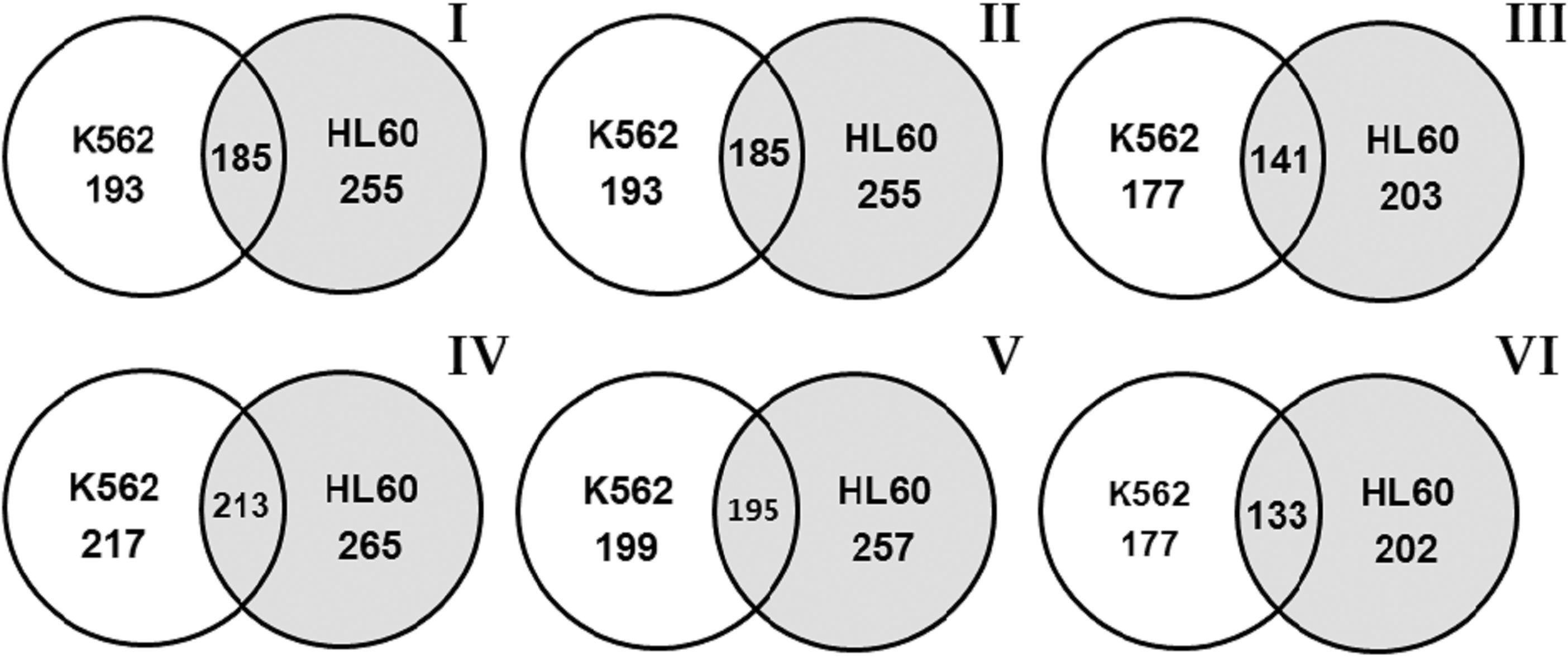
In Fig. 2, the Venn diagrams summarize the number of differentially expressed snoRNAs in the control in both cell lines and after irradiation (the number is given under the cell line name) the number of differentially expressed snoRNAs in the K562 and HL-60 cell lines (the number is given in intersection of circles) (I – the number of differentially expressed snoRNAs in the control in 1 hour; II – the number of differentially expressed snoRNAs in the control after 4 hours; III – the number of differentially expressed snoRNAs in the control after 24 hours; IV – the number of differentially expressed snoRNAs 1 hour after irradiation; V is the number of differentially expressed snoRNAs 4 hours after irradiation; VI – the number of differentially expressed snoRNAs 24 hours after irradiation.). The number of snoRNAs in both control groups was the same 1 h and 4 h. In the experimental groups, 1 h after irradiation of the cells, 213 differentially expressed snoRNAs inherent of the two studied cell lines were detected (Fig. 2, IV). In the intersections, 4 and 24 hours after irradiation there were 195 (Fig. 2, V) and 133snoRNAs (Fig. 2, VI) detected, respectively. Thus, the analysis of snoRNA dynamics showed that, in more than a half of all snoRNAs inherent to both radiosensitive and radioresistant cell lines, the expression changed after irradiation.
The number of expressed snoRNAs depending on the chromosomal localization in the K562 and HL-60 cell lines at 1, 4 and 24 hours after irradiation.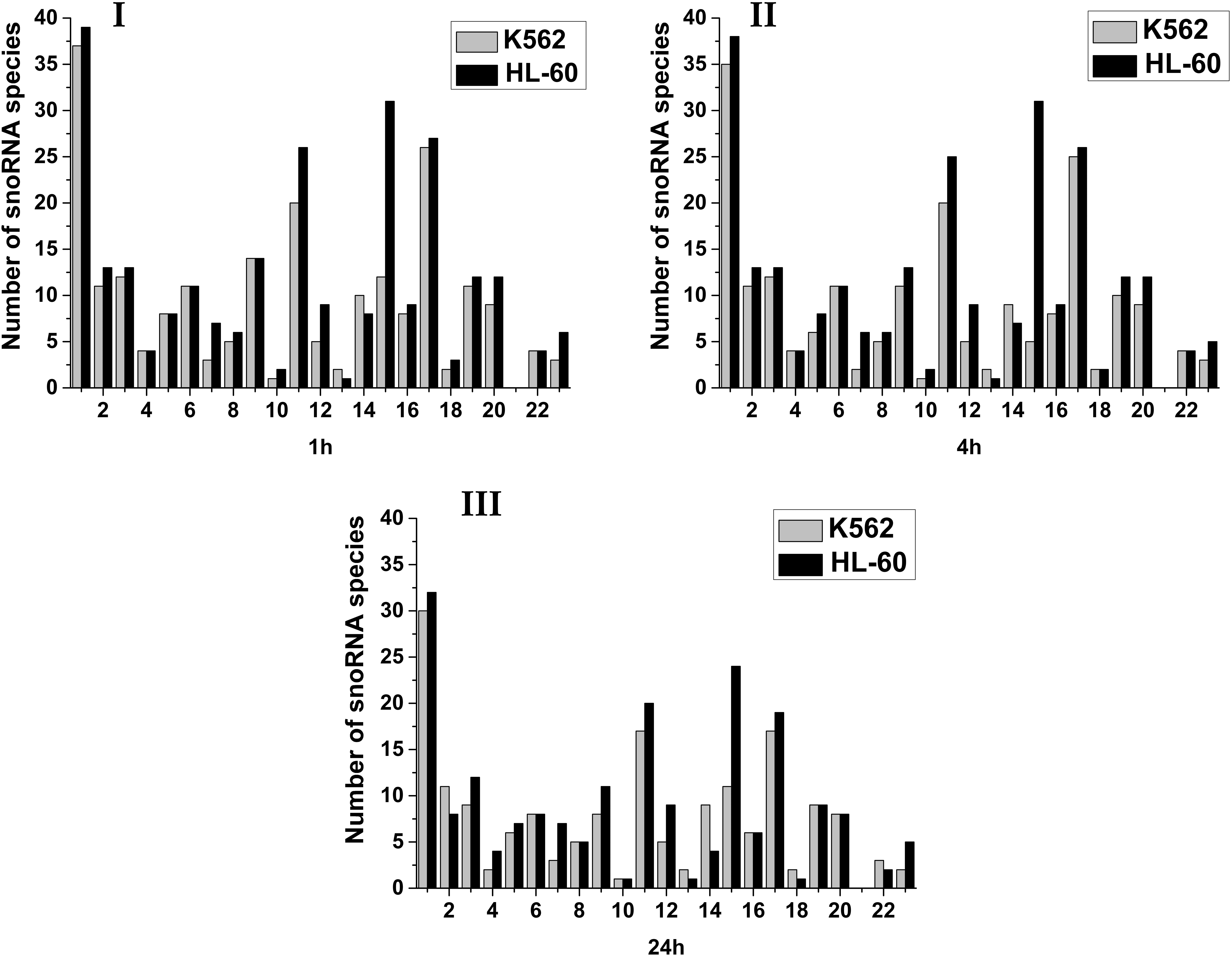
Figure 3 illustrates the data on the number of expressed snoRNAs localized in either chromosome at each stage of the experiment (I is the amount of expressed snoRNA 1 hour after irradiation; II – the number of expressed snoRNAs 4 hours after irradiation; III is the number of expressed snoRNAs 24 hours after irradiation). It was shown that in the K562 and HL-60 cell lines, the largest number of expressed snoRNAs was localized in chromosomes 1, 11, 17, and 1, 11, 15, 17, respectively. The smallest number of snoRNAs was localized in chromosomes 10, 13, 18 in both cell lines. It can be seen that the distribution of the expressed snoRNAsin chromosomes is different for the two cell lines. The number of the expressed snoRNAs changed during the experiment in both cell lines.
Analysis of the dynamics of snoRNA expression depending on the chromosomal localization after radiation exposure at the dose of 4 Gy in comparison with the control.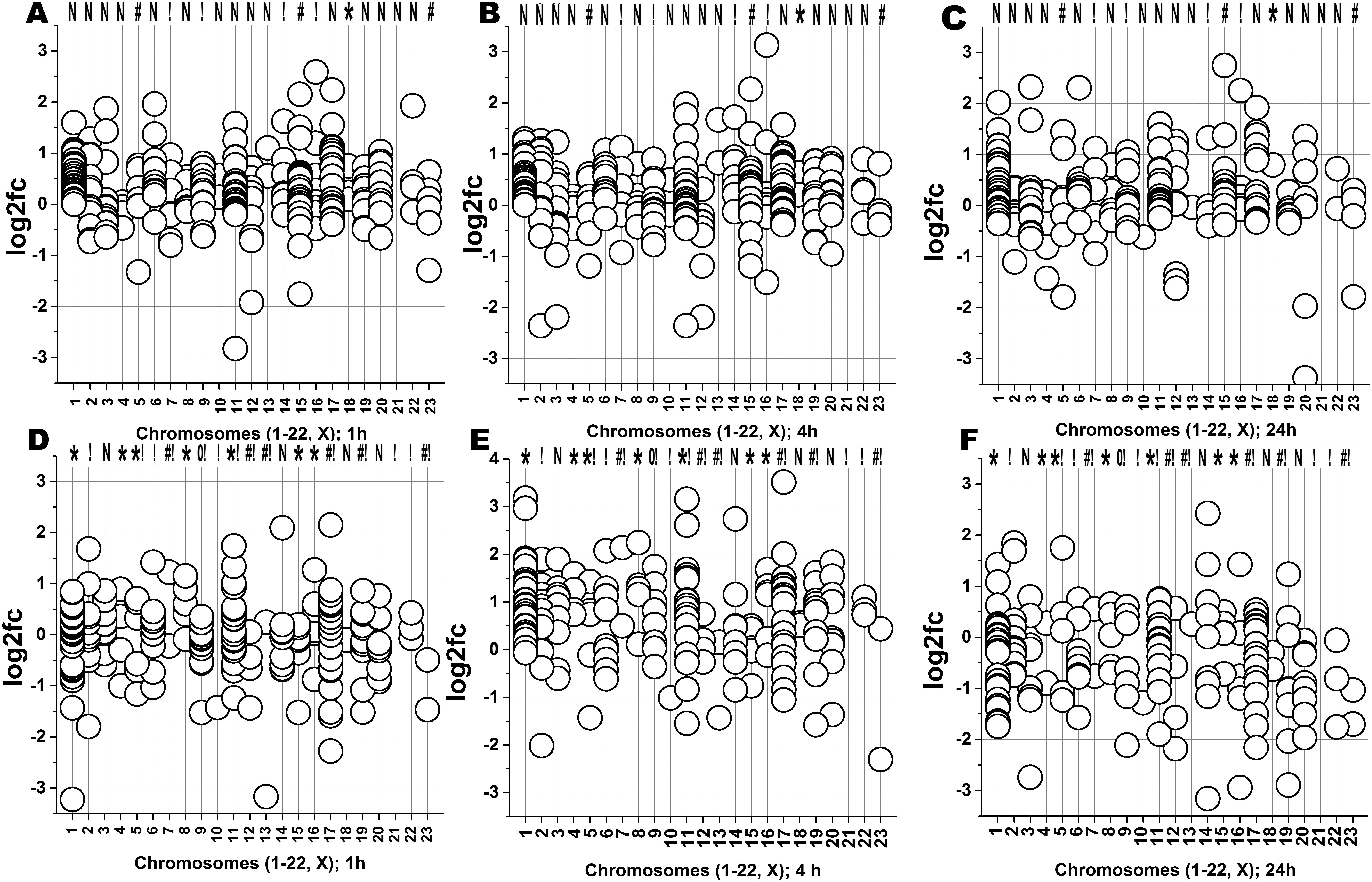
Figure 4 shows the log2fc value scattering for all expressed snoRNAs in the HL-60 and K562 cell lines distributed according to their localization in chromosomes (A – dynamics of snoRNA expression in the HL-60 cell line 1 hour after irradiation; B – dynamics of snoRNA expression in the HL-60 cell line 4 hours after irradiation; C – dynamics of snoRNA expression in the HL-60 cell line 24 hours after irradiation; D – dynamics of snoRNA expression in the K562 cell line 1 hour after irradiation; E – dynamics of expression of snoRNA in the K562 cell line 4 hours after irradiation; F – dynamics of expression of snoRNA in the K562 cell line 24 hours after irradiation; 0 – nulisomy, N – normal chromosome, * – trisomy, # – monosomy, ! – marker chromosome). In the K562 cell line at all points in the experiment (Fig. 4, D, E, F), snoRNAs have a wider log2fc value scattering (
The expressed snoRNAs localized in the chromosome 3 are shown in dendrograms (Fig. 5 (I), (II) for the K562 and HL-60 cell lines, respectively). In both cell lines, chromosome 3 comprises 2 copies of normal chromosomes. One can see that the type and number snoRNAs are the same but there is a slight difference in clustering: 12 and 13 snoRNAs for the K562 and HL-60 cell lines, respectively. Clustering of snoRNAs localized in chromosome 5 is shown in Fig. 5 (III, IV). In the HL-60 and K562 cell lines, monosomy and trisomy of chromosome 5 is observed, respectively. The hsa-sno-U63 snoRNA is isolated into a separate cluster in both cell lines due to its expression, which is ten times higher compared to that in other snoRNAs. In general, there is no significant difference in the number and name of the expressed snoRNAs in the considered cell lines. In both cell lines, snoRNAs are combined into two clusters. The first is small and consists of two snoRNAs and the second, which includes the rest. In the second cluster snoRNAs are interconnected into subclusters.
Cluster analysis of expressed snoRNAs located on chromosomes 3, 5, and 11 in cell lines K562 and HL-60.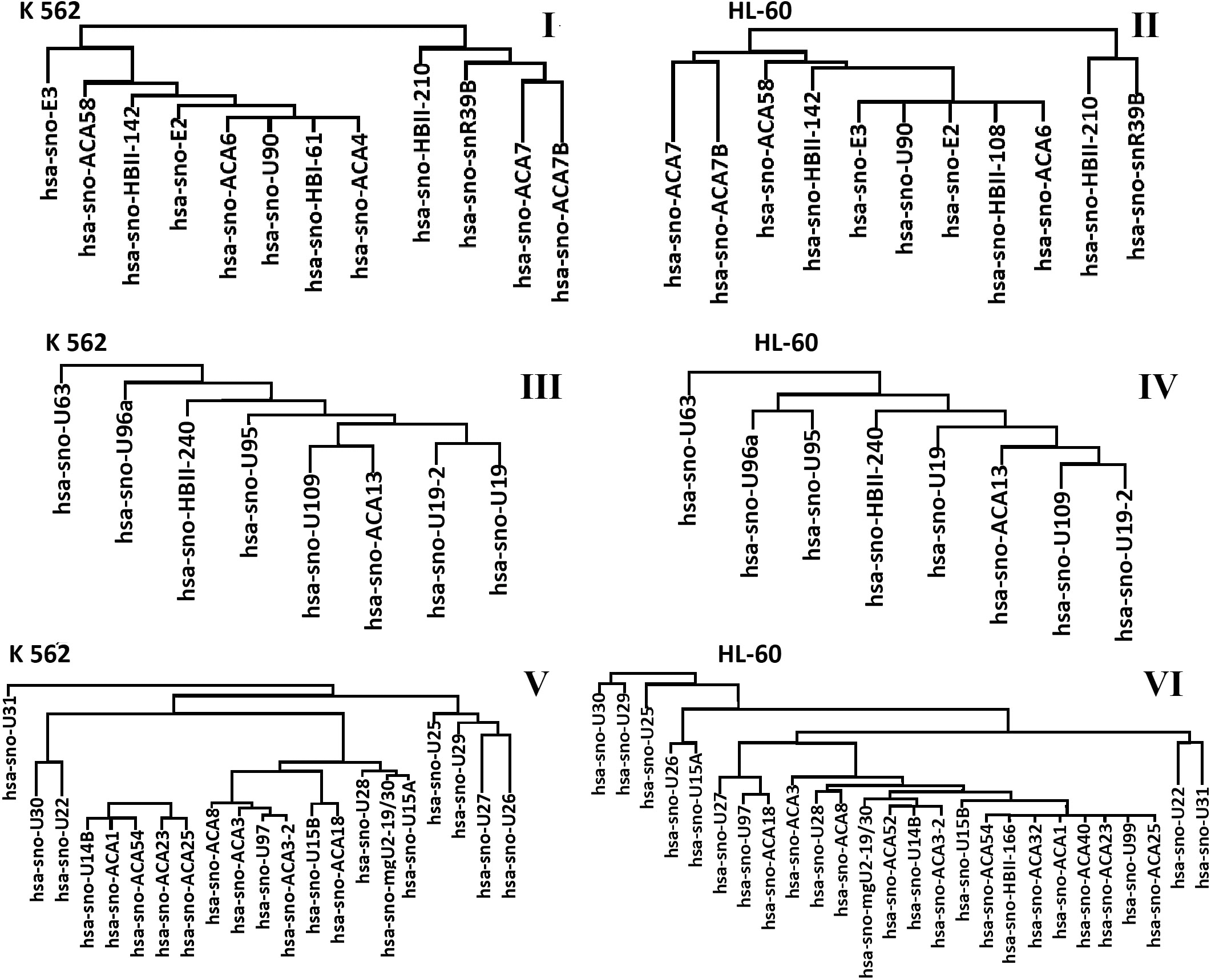
Figure 5 (V, VI) shows the dendrograms of the expressed snoRNAs localized in chromosome 11 of the HL-60 and K562 cell lines (HL-60 – two copies of normal chromosome 11, K562 – trisomy of chromosome 11 and marker chromosome). A number of the expressed snoRNAs differs in two cell lines. 20 and 25 snoRNAs are expressed in the K562and HL-60 cell lines, respectively. In the HL-60 cell line, 5 additional snoRNAs are expressed: hsa-sno-ACA32, hsa-sno-ACA40, hsa-sno-ACA52, hsa-sno-HBII-166, hsa-sno-U99. They form the first type clusters with the snoRNA expressed in the K562 cell line.
In both cell lines, snoRNA expression and clustering are approximately the same in chromosome 3, which has two normal chromosomes from each pair. In chromosome 5 with the detected trisomy (the K562 cell line) and monosomy (the HL-60 cell line), snoRNA names and the way of clustering are similar and demonstrate slight differences. The way of clustering and types of the expressed snoRNAs in chromosome 11 in both cell lines are quite different. These differences are associated with the effect of chromosomal mutations on gene expression.
In this work, we have used two cancer cell lines of leukemia, which have different sensitivity to radiation exposure and different levels of chromosomal abnormalities. The HL-60 cell line has a small number of genomic abnormalities (monosomy of chromosomes 5, 15 and X, trisomy of chromosome 18) and moderate radiosensitivity. The K562 cell line has 15 genomic mutations (monosomy of chromosomes 7, 12, 13, 17, 19, X, trisomy of chromosomes 1, 4, 5, 8, 11, 15, 16, nulisomy of chromosome 9) and 21 marker chromosomes and is less radiosensitive. Such strong differences in the genome affect the expression of all genes. According to our experiments, about half of all snoRNAs in K562 and HL 60 cells change their expression after an exposure to a dose of 4 Gy (Fig. 2), which is confirmed by previous studies that have demonstrated changes in snoRNA expression in response to ionizing radiation [41]. An analysis of the change in the amount of expressed snoRNA taking into account the chromosome localization after radiation exposure shows that the changes in the number of snoRNA are characteristic of all chromosomes and are more pronounced in chromosomes 11, 15 and X (Fig. 3). The greatest differences in the profile of snoRNA expression were observed in the K562 radioresistant cell line (Fig. 4).
Figure 5 (I and II) shows expressed snoRNAs located in the chromosome 3 of the K562 and HL-60 cell lines. Both cell lines have two normal copies in chromosome 3. Clustering of snoRNAs in both cell lines is similar, and identical snoRNAs are expressed Studies of circadian (diurnal) rhythms of snoRNA expression normally have shown a similar dynamics [44, 45]. We have obtained similar results for other normal chromosomes of the cell lines K562 and HL-60 (chromosomes 14 and 20). These results suggest that the expression of snoRNAs localized in the normal chromosomes of related cell lines may have slight differences.
The K562 cell line has three normal copies of chromosome 5 in the karyotype (trisomy) [36, 37, 38]. One normal chromosome 5 (monosomy) has been detected in the karyotype of the HL-60 cell line [39] (Fig. 5 III and IV). The number, clustering, and types of expressed snoRNAs located on chromosome 5 are the same in the two studied cell lines. It has been shown that, with trisomy, an increase in gene expression in blastocysts is observed, while reduced expression is observed in monosomal blastocysts [46]. However, our experiment did not confirm the previous research, as snoRNA expression in the presence of trisomy and monosomy not significantly different in the two cell lines (Fig. 4). In our opinion, this is due to the fact that this type of chromosomal abnormality does not carry functional rearrangements within the chromosome itself, and it can be assumed that the regulation of gene expression should be the same as that of normal chromosomes.
For further analysis, chromosome 11 was selected, which was represented in the HL-60 cell line by two normal copies, and in K562 by three normal copies of chromosome 11 and a marker chromosome. In our experiments, it was demonstrated that, in the presence of an additional marker chromosome, snoRNA expression and clustering are significantly different compared to normal chromosome 11 (Fig. 5 V and VI). A similar picture was observed when comparing the expression of snoRNAs of other normal chromosomes of the HL-60 cell line with abnormal chromosomes of the K562 cell line (chromosomes 2, 6, 17, 19, 22). There is still little information about marker chromosomes in humans and their effect in gene expression. Studies of sperm with marker chromosomes have shown that this violation leads to a change in gene expression and infertility [47, 48, 49], developmental defects and various pathologies are observed in patients with marker chromosomes [50, 51]. The above data indicate that marker chromosomes have a significant effect on snoRNA expression and gene expression in general. But it’s difficult to correlate a certain marker chromosome with a specific disorder or disease due to the wide variety of marker chromosomes.
The mechanisms of the effect of radiation on snoRNA have not yet been studied, but it can be assumed that snoRNAs affect radioresistance indirectly, through interaction with the processes occurring in eukaryotic nucleolus. Recently, data has begun to appear confirming the role of the processes occurring in the nucleolus in response to cellular stressors, maintaining the stability of the genome and repairing DNA damage, as well as the regulation of gene expression and biogenesis of ribonucleoproteins [52]. Previously, it has been demonstrated that radiation exposure causes significant changes in the nucleolus after radiation exposure [53]. Also, it should be borne in mind that RNA is more radiosensitive than DNA or proteins [54] and the presence of an additional amount of snoRNA due to the presence of additional marker chromosomes can mechanistically reduce radiosensitivity. In this case, according to the target theory [55], there will be more intact snoRNAs, which will allow the processes in the nucleolus to change less fatally after radiation exposure.
After analyzing the dynamics of snoRNAs expression in the K562 and HL-60 cancer cell lines, we can conclude that the nature and change in expression of snoRNA in response to ionizing radiation in our experiments may depend on the presence of chromosomal abnormalities. Trisomies and monosomies do not have such a significant effect on the expression of snoRNAs as the presence of marker chromosomes. Marker chromosomes lead to impaired expression of snoRNAs, so it should be done with caution to use snoRNAs located in abnormal chromosomes as markers of radioresistance.
Footnotes
Acknowledgments
This research was funded by the Ministry of Higher Education and Science of the Russian Federation, grant number 0830-2020-0008.
Author contributions
Conception: Yury Saenko, Daria Liamina, Eugenia Rastorgueva.
Interpretation or analysis of data: Ivan Panchenko, Dmitrii Sugak and Yury Saenko, Eugenia Rastorgueva, Daria Liamina.
Preparation of the manuscript: Elena Iurova, Daria Liamina, Eugenia Rastorgueva, Evgeniya Pogodina.
Revision for important intellectual content: Evgenii Beloborodov, Sergei Slesarev, Yury Saenko.
Supervision: Yury Saenko.
