Abstract
BACKGROUND:
Metastatic castration-resistant prostate cancer (mCRPC) is the lethal stage of prostate cancer and the main cause of morbidity and mortality, which is also a potential target for immunotherapy.
METHOD:
In this study, using the Approximate Relative Subset of RNA Transcripts (CIBERSORT) online method, we analysed the immune cell abundance ratio of each sample in the mCRPC dataset. The EdgeR (an R package) was used to classify differentially expressed genes (DEGs). Using the Database for annotation, visualisation and interactive exploration (DAVID) online method, we performed functional enrichment analyses. STRING online database and Cytoscape tools have been used to analyse protein-protein interaction (PPI) and classify hub genes.
RESULTS:
The profiles of immune infiltration in mCRPC showed that Macrophages M2, Macrophages M0, T cells CD4 memory resting, T cells CD8 and Plasma cells were the main infiltration cell types in mCRPC samples. Macrophage M0 and T cell CD4 memory resting abundance ratios were correlated with clinical outcomes. We identified 1102 differentially expressed genes (DEGs) associated with the above two immune cells to further explore the underlying mechanisms. Enrichment analysis found that DEGs were substantially enriched in immune response, cell metastasis, and metabolism related categories. We identified 20 hub genes by the protein-protein interaction network analysis. Further analysis showed that three critical hub genes, CCR5, COL1A1 and CXCR3, were significantly associated with prostate cancer prognosis.
CONCLUSION:
Our findings revealed the pattern of immune cell infiltration in mCRPC, and identified the types and genes of immune cells correlated with clinical outcomes. A new theoretical basis for immunotherapy may be given by our results.
Keywords
Introduction
Prostate cancer is one of the most common malignant tumors of the urinary system [1]. In 2018, 1,276,106 new prostate cancer cases were reported, which caused 358,989 deaths (3.8% of all cancer deaths in men) [2]. Men with localized prostate cancer (defined as no identifiable regional lymph nodes or distant metastases) often have 3 primary treatment options: expectant management, surgery, and radiation. As for men with metastatic prostate cancer, androgen deprivation therapy (ADT) is the first-line treatment [3]. While after ADT treatment, the majority of these patients will develop to CRPC within 1–3 years [4, 5, 6, 7]. CRPC and metastatic castration resistant prostate cancer (mCRPC) were defined as the advanced prostate cancer, which are the lethal stages of prostate cancer and the leading cause of patient death [8, 9].
Several mechanisms of how CRPC developed have been proposed, many of which are driven by androgen receptor (AR). Previous studies provide evidence that some patients with CRPC still rely on AR signaling, which are always defined as AR-dependent CRPC [10, 11]. Such type of CRPC would develop through multiple AR-related mechanisms: increased AR expression, the emergence of AR splice variants, AR mutation, elevated intratumoral steroid hormone synthesis or modulation of co-factor activity [11]. For some other patients (AR-independent CRPC), the development of CRPC does not rely on AR. In these cases, tumor cells always origin from several epithelial cell populations (including basal cells, stem cells, neuroendocrine cells, and some luminal cells) that lack AR expression [12, 13]. Drugs targeting AR could attack and inflict therapeutic pressures on cells with targetable AR protein. As a consequence, treatment (such as ADT) will stimulate the development of these populations of AR negative cells and give rise to a tumor that is inherently resistant to AR antagonism, resulting in CRPC [11, 14, 15]. In addition, the metastatic burden is also crucial for development of castration resistance. Chemohormonal Therapy Versus Androgen Ablation Randomized Trial for Extensive Disease in Prostate Cancer (CHAARTED) trial has shown that patients with high-volume (presence of visceral metastases and/or more than four bone metastases with at least one outside of the vertebral column and pelvis) are more prone to develop to CRPC with shorter time in clinical progression [16].
In recent years, the treatment of CRPC has developed rapidly [17, 18]. For patients with mCRPC, sipuleucel-T, abiraterone acetate, enzalutamide, docetaxel, cabazitaxel and radium-223 have been proven to have improved clinical therapeutic effects [18, 19, 20, 21]. Recent clinical trials revealed that docetaxel, with or without ADT, could prolong the time to biochemical recurrence and progression, and improve overall survival in patients with locally advanced or metastatic prostate cancer [16, 22, 23]. PARP (poly ADP-ribose polymerase) inhibitors such as olaparib and DNA damaging agents such as platinum chemotherapy have been shown to have greater benefit in specific patients with mutations in homologous recombination repair and DNA repair pathways (such as BRCA1/2) [24]. In addition, some new generation anti-androgen drugs (such as apalutamide and dalolutamide) are currently under investigation in clinic [25, 26]. At present, there are many investigations of the significance of novel drugs in the treatment of prostate cancer, among which immunotherapy has become a potentially promising treatment [27, 28].
The immune system is critical to fight against pathogens and to maintain the normal physiological functions of our body. Previous studies on tumor pathogenesis have indicated that immune escape is a critical contributor to malignancy [29]. Therefore, it is possible that tumor cells could be effectively eliminated if any treatment could mobilize the immune system by braking the immune escape. Followed studies showed the ability of immune system to fight with cancer, which inspired a strong focus on novel immunotherapy of cancer [30].
In the past decades, immunotherapy has gradually shown better therapeutic effect on advanced tumors, including urogenital tumors [31, 32, 33]. Especially in high-risk non-muscle-invasive bladder cancer (NMIBC), intravesical Bacillus Calmette-Guérin (BCG) is one of the most successful immunotherapies as the standard of treatment [33]. PD-1 inhibitor (such as pembrolizumab) and PD-L1 inhibitor (such as atezolizumab) have also been shown to have clinical activity in patients with advanced bladder cancer [33, 34]. In renal cell carcinoma, checkpoint inhibitors such as nivolumab (a PD-1 inhibitor) and ipililumab (a CTLA-4 inhibitor) demonstrated improved clinical therapeutic effects on metastatic renal cell carcinoma (mRCC), in addition to the early application of interleukin-2 (IL-2) and interferon alpha (IFNa) in the 1990s [32, 35, 36, 37, 38]. Currently, some new drugs (such as VEGF inhibitors) are under research in mRCC to understand the clinical activity [39].
Compared with other urogenital tumors above, prostate cancer is not an immunologic solid tumor and the tumor microenvironment of prostate cancer is immunosuppressive. Recruitment of regulatory T cells and Th17, higher transforming growth factor-beta levels, and low cytolytic activity of natural killer (NK) cells could inhibit NK and T cell function, resulting in the downregulation of tumor immunity [40, 41]. These characteristics of prostate cancer lead to the limitation of immunotherapy in the treatment. Sipuleucel-T, an autologous vaccine, is a personalized immunotherapeutic agent that is collected by white blood cell separation and then processed by peripheral dendritic cells (DC). Sipuleucel-T is a therapeutic cancer vaccine for mCRPC patients with low-volume diseases and asymptomatic or mildly symptomatic patients with a slower disease course, and is the first of its kind approved by the FDA [19]. Previous studies indicated that prostate cancer is less reactive to immune checkpoint inhibitors, resulting in the resistance to immune checkpoint blockade [9, 42]. One research showed that lethal prostate cancers with defective mismatch repair (dMMR) defects constitute a subtype with PD-L1 IHC staining and decreased survival time, which may be the only genomic subgroup shown to respond well to checkpoint blockade [43]. However, recent results showed that enhancing the immune response against cancer cells by modulating the tumor microenvironment using immune checkpoint inhibitors has also shown encouraging results in the treatment of mCRPC [36]. Pembrolizumab was shown to have activity in metastatic prostate cancer [44]. In the phase 1b KEYNOTE-028 trial and phase 2 KEYNOTE 199 trial, Pembrolizumab showed anti-tumor activity in advanced prostate cancer (including mCRPC) patients regardless of PD-L1 status [44, 45]. Further research and application of immunotherapy in prostate cancer is of significance and promising.
To further study the role of immunotherapy in prostate cancer, we should understand its immune cell infiltration pattern and immune status, which remains to be studied in depth. We focused on mCRPC in this study, and used the latest data [46]. to study the immune cell infiltration pattern of mCRPC, hoping to find out central immune cells and related genes in the tumor microenvironment. Our findings may help to better understand mCRPC ’s immune status and could include some new theoretical foundations for better immunotherapy implementation.
Materials and methods
Data information
We downloaded the gene expression data with corresponding clinical profiles of a published mCRPC dataset of the SU2C/PCF Prostate Cancer International Dream Team (accession number: prad su2c 2019) [46]. from online datasets cBioPortal for Cancer Genomics website and GitHub website (Websties:
Identifying infiltrated immune cells
CIBERSORT online analysis tool is mainly used to estimate the abundance ratio of member cell types in mixed cell populations based on gene expression data [49]. We uploaded the mCRPC RNA-Seq data to the online platform to obtain the abundance ratio matrix of 22 immune cells. Finally, 112 samples with
Identification of outcome-related immune cells
To find out the immune cells related to clinical outcome, we used the data of time on treatment with a first-line androgen receptor signaling inhibitor (ARSI; abiraterone or enzalutamide) in the dataset to analyze. The immune cells we identified were defined as outcome-related immune cells. These patients received an ARSI in the first-line setting without another agent. We dealt with the analysis by Graphpad using the Kaplan-Meier with log rank test.
Analysis of outcome-related immune cell abundance ratio and treatment history
Data on patient exposure status to ARSI and taxanes were used to analyze the relationship between outcome-related immune cell abundance ratio and treatment history. This could help us determine whether the infiltration of outcome-related immune cells changed after receiving the drugs above. The analysis was undertaken using the
Identification of differentially expressed genes (DEGs) in patients with different outcome-related immune cell infiltration pattern
DEGs related to the different infiltration patterns of the two outcome-related immune cells were identified. We identified the DEGs between individuals with high and low infiltration levels of Macrophage M0, and DEGs between individuals with high and low infiltration levels of T cell CD4 memory resting, respectively. EdgeR, an R package for examining differential expression of RNA-Seq count data, was used according to the user’s guide [50, 51]. We used the following criteria to determine DEGs: fold change (FC)
Functional annotation and pathway enrichment analysis of DEGs
We did enrichment analysis of the DEGs above using gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) to annotate the functions and pathways. The analysis was undertaken using the Database for annotation, visualization and integrated discovery (DAVID) website by uploading the DEGs identified to the online tool [52].
Protein-protein interaction (PPI) network and module analysis
We used the STRING online database for PPI evaluation and integration [53]. The cutoff was a combined score 0.4. We then imported the PPI network into Cytoscape to complete module screening by Molecular Complex Detection (MCODE) and to identify the hub genes [54].
Statistical analysis
The
Summary of the clinical characteristics of patients with mCRPC
Summary of the clinical characteristics of patients with mCRPC
The relationship between the abundance ratios of immune cells and time on treatment with a first-line ARSI. A. Histogram of the proportion of 22 immune cells in all samples. B. The stacked histogram shows the distribution of 22 immune cell infiltration in mCRPC samples. C. The relationship between the abundance ratios of various immune cells. Red represents a positive correlation, and the blue represents a negative correlation. D. The time on treatment with a first-line ARSI in patients with different abundance ratios of the two immune cells. Time on treatment analysis was evaluated for patients who received an ARSI in the first-line setting (abiraterone or enzalutamide) without another agent.
Data source and related information
We obtained the gene expression data with follow-up profiles of 212 patients with mCRPC from cBioPortal and GitHub websites. Tumor metastasis sites were summarized as follows: 2 (0.95%) in adrenal; 83 (39.15%) in bone; 27 (12.74%) in liver; 81 (38.2%) in lymph nodes; 14 (6.6%) in other soft tissues and 5 (2.36%) in prostate. In the following analyses, we excluded the 5 patients whose metastatic site was in the prostate. These data were obtained directly from the online website. We summarized all the available clinical characteristics in Table 1.
Relationship between immune cell abundance ratio and clinical characteristics. A. Violin plots of immune cells in bone metastasis tumors and tumors in other locations. B. C. The relationship between the abundance ratios of the two immune cells and Taxane or abiraterone (ABI) and enzalutamide (ENZA) exposure. 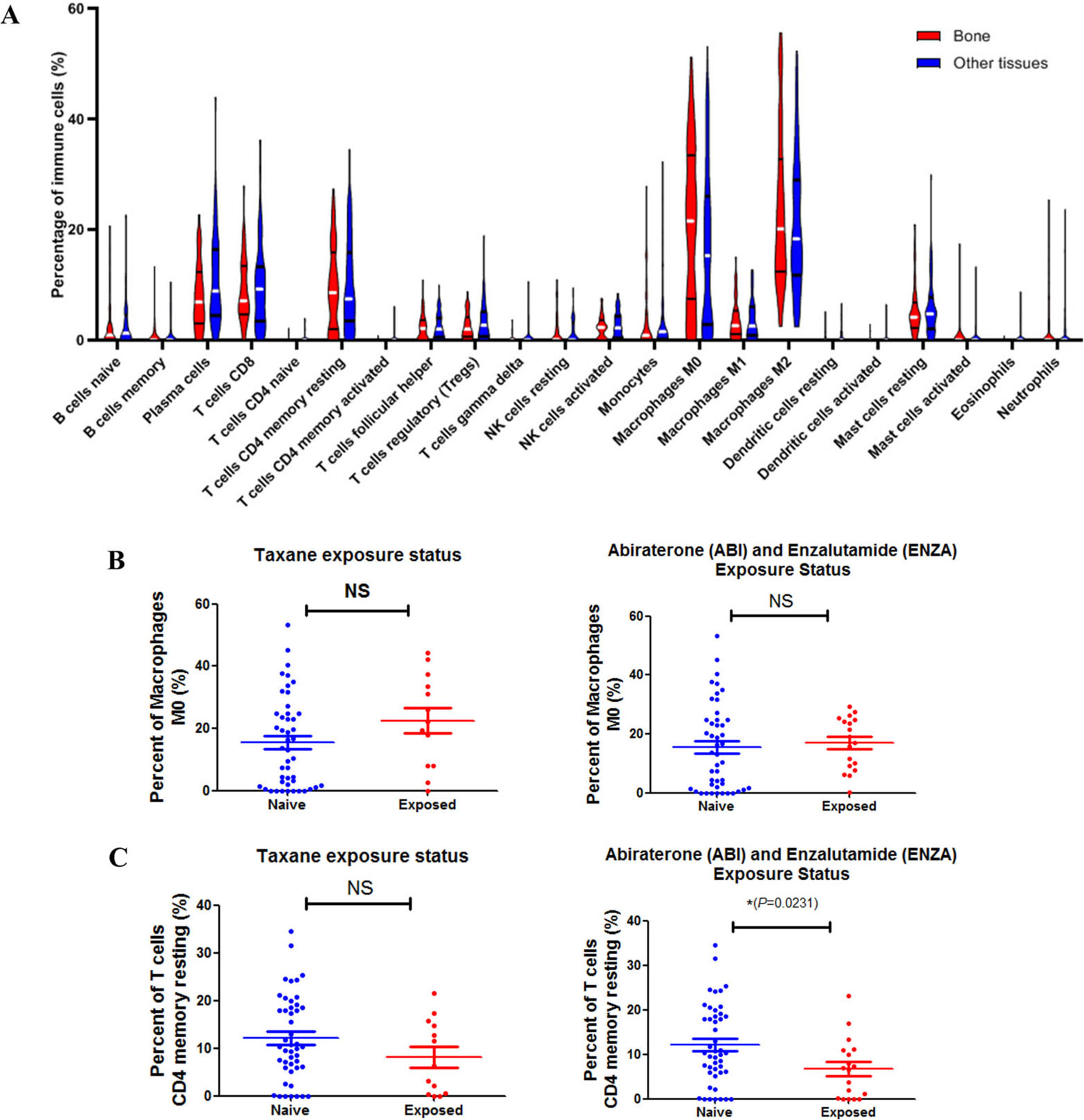
To identify the immune cell infiltration pattern in mCRPC, we uploaded the gene expression data to CIBERSORT website. The abundance ratios of 22 immune cells and their correlations were analyzed in 207 samples (Figs 1A–C). Macrophages M2, Macrophages M0, T cells CD4 memory resting, T cells CD8 and Plasma cells were the main infiltration cell types in mCRPC samples. Macrophage M1 and T cell CD4 memory resting (
Identification of genes related to immune cell infiltration and DAVID enrichment results of the differentially expressed genes. A. B. Volcanic diagram of prostate cancer oncogene expression profile grouped by the memory of macrophages M0 and T cells CD4. Red / blue symbols classify genes that are up / down-regulated. C. The Venn calculations yielded genes involved in the infiltration of two immune cells. D. The GO enrichment terms of differentially expressed genes. E. The KEGG pathway analysis of differentially expressed genes.
Prognosis analyses of the hub genes. A–C. Three hub genes were significantly related to prognosis by analyzing the data from GSE21032. A total of 140 non-metastatic prostate cancer tumor samples with prognostic information were obtained from patients treated by radical prostatectomy at Memorial Sloan-Kettering Cancer Center.
We first compared whether the distribution of these 22 immune cells in metastasis tumors was different in different locations. As shown in Fig. 2A, violin plots of immune cells in different locations indicated no significant differences. Then we analyzed other clinical relationships with the two outcome-related immune cells. We analyzed whether the abundance ratio of the two immune cells correlated with Gleason score or prostate-specific antigen (PSA). However, we found that there was no correlation between the content of the two outcome-related immune cells above and these clinical features (Sup Fig. s1). Then we analyzed whether the abundance ratio of the two immune cells correlated with Taxane or abiraterone/enzalutamide exposure. Results showed that Taxane or abiraterone/enzalutamide exposure had no influence on Macrophage M0 infiltration in patients (Fig. 2B), while the abundance of T cell CD4 memory resting in patients who had abiraterone and enzalutamide exposure was lower than other patients (Fig. 2C). Our results indicate that drug intaking might affect the composition of immune cells in the tumor microenvironment.
To confirm our results, we then used a dataset named prad-su2c-2015 from the SU2C/PCF Prostate Cancer International Dream Team for analysis. Due to the limited clinical data of this database, we only analyzed the immune cell infiltration pattern, and explored the relationship between the abundance of the two immune cells above and Taxane or abiraterone and enzalutamide exposure. The results also confirmed that only T cell CD4 memory resting in patients who had abiraterone and enzalutamide exposure was lower than other patients (Sup Fig. s2 A–D).
Identifying immune-related genes
To further explore the underlying mechanisms and to find out some potential therapeutic targets, we analyzed DEGs related to the levels of two outcome-related immune cell infiltration. We identified the DEGs between individuals with high and low infiltration levels of Macrophage M0, and DEGs between individuals with high and low infiltration level of T cell CD4 memory resting, respectively. We found that 310 DEGs were associated with Macrophage M0 and 911 were associated with T cell CD4 memory resting (Sup Table s1). Figure 3A and B showed the volcano plots. The Venn map analysis shown in Fig. 3C revealed all the 1102 genes related to immune cell infiltration, which we defined as immune-related genes in the following studies.
GO and KEGG analyses of DEGs
To analyze biological classifications of the DEGs, we submitted all the 1102 DEGs online for further analyses with DAVID. The GO analysis of DEGs (Figure 3D) suggested significant enrichment of the immune-related genes in transmembrane receptor protein tyrosine kinase signaling pathway, type I interferon signaling pathway, cell-cell signaling, positive regulation of cytosolic calcium ion concentration, proteolysis, defense response to virus, negative regulation of viral genome replication, T cell co-stimulation, oxygen transport, collagen fibril organization, immune response, cell adhesion, collagen catabolic process, ossification, extracellular matrix organization and skeletal system development.
Furthermore, in KEGG pathway analysis, the immune-related genes were significantly enriched in chemokine signaling pathway, T cell receptor signaling pathway, allograft rejection, rheumatoid arthritis, osteoclast differentiation, staphylococcus aureus infection, cell adhesion molecules (CAMs), ribosome, systemic lupus erythematosus, focal adhesion, PI3K-Akt signaling pathway, primary immunodeficiency, protein digestion and absorption, hematopoietic cell lineage, and ECM-receptor interaction (Fig. 3E). We also did GO and KEGG analyses on the 119 DEGs shared in both Macrophage M0 and T cell CD4 memory resting groups (Sup Fig. s3).
Top three modules from the PPI network. A. PPI network of module 1. B. GO and KEGG analyses of module 1. C. PPI network of module 2. D. GO and KEGG analyses of module 2. E. PPI network of module 3. F. GO and KEGG analyses of module 3.
To further explore the interrelationships of immune-related genes and to gain hub genes, we did analysis in the STRING database. Ranked by degree, we identified the top 20 genes as hub genes. These genes were TNF, IL2, CXCL10, CCR5, CCR7, COL1A1, CXCL9, CXCR3, MMP9, GNB3, RUNX2, COL1A2, CD28, STAT1, CCR2, ISG15, PPBP, SELL, LCK and SPP1. TNF was identified as the top hub gene with a degree of 118.
In order to find out whether the above 20 hub genes correlated to prostate cancer prognosis, we then did the Kaplan-Meier prognostic analysis. Since not much prognostic information is contained in the mCRPC database we used, we used another GSE21032 authoritative prostate cancer database for research, not limited to mCRPC. The results showed that six of the 20 hub genes strongly associated with clinical outcomes, including CCR5, COL1A1, STAT1, COL1A2, CXCR3 and PPBP. To confirm the hub genes associated with clinical outcomes, we then used another dataset from TCGA and found that only CCR5, COL1A1 and CXCR3 correlate to prostate cancer prognosis (Fig. 4 and Sup Figs s2 E–G).
Finally, we imported the above network into Cytoscape software to classify gene modules via the MCODE plate. We did enrichment research on the top three main modules in order to analyse the modules at the significant level (Fig. 5, respectively). Results showed that functional annotation in module 1 genes was primarily enriched in G-protein coupled receptor signaling pathway, chemokine-mediated signaling pathway, inflammatory response, taste transduction and neuroactive ligand-receptor interaction. In Module 2, the analysis showed that the enrichment was mainly related to the SRP-dependent co-translational protein-targeting membrane, viral transcription, nuclear catabolic mRNA catabolic processes, nonsense-mediated decay, ribosomes, protein digestion and absorption, and amoeba Disease related. While in module 3, genes were enriched in type I interferon signaling pathway, defense response to the virus, activation of adenylate cyclase activity, neuroactive ligand-receptor interaction, measles, and osteoclast differentiation.
Discussion
Prostate cancer, especially the metastatic prostate cancer, is still one of the most common malignancies that cause patients to death in men [1]. Such patients have lost the chance of surgery, and only endocrine therapy and chemotherapy could delay their disease progression. In recent years, with the development of drug research, some new drugs targeting androgens and AR have been put on the market (such as abiraterone and enzalutamide). Some of the treatment might benefit overall survival, while some could alleviate the disease and improve the quality-of-life of a large number of patients with CRPC and mCRPC [55, 56]. Nowadays, immunotherapy has become a promising therapeutic method, whose effectiveness could be affected by the immune cell infiltration pattern [57]. The composition of prostate cancer immune microenvironment is complicated. Previous studies showed that monocytes and macrophages were the most common inferred immune cell populations in mCRPC tumor microenvironment [43]. Tumor infiltrating lymphocyte was shown to be a positive prognostic indicator and a predictor of immune checkpoint blockade response [58]. The low immunotherapy response rate in prostate cancers may also be due to the scarce infiltration of T cells in prostate cancer [59]. In-depth research is necessary for the application of immunotherapy in prostate cancer [58].
Previous study evaluated the immune infiltration in non-metastatic prostate cancer tissues and established a novel nomogram to assess the recurrence risk [60]. Here in this study, we analyzed an RNA-Seq dataset of mCRPC downloaded from GitHub and cBioportal to describe the immune cell infiltration pattern in mCRPC using bioinformatics analysis approaches. We found that Macrophages M2, Macrophages M0, T cells CD4 memory resting, T cells CD8 and Plasma cells were the main infiltration cell types in mCRPC samples. We then analyzed the time on treatment with a first-line ARSI in patients with different composition of immune cells. As a result, we found an association of higher Macrophage M0 or lower T cell CD4 memory resting with a shorter time on treatment with a first-line ARSI. The correlation with clinical outcomes made the two outcome-related cells to be most likely to play a critical role in disease progression. In addition, we found that the abundance of T cell CD4 memory resting in patients who had abiraterone and enzalutamide exposure was lower than other patients. This result might indicate that patients who had abiraterone and enzalutamide exposure may not be suitable for potential immunotherapy against this cell type.
The enrichment analysis showed that the PI3K-Akt, chemokine, and Toll-like receptor signaling pathways and other metabolism related progressions are mainly associated with outcome-related cell infiltration. Excessive metabolism is one of the important characteristics of tumors. In order to meet the energy and biosynthesis needs for rapid diffusion, tumor cells rapidly provide energy through aerobic glycolysis [61]. In addition, different subsets of immune cells also use different nutrients to provide energy. For example, effector T (Teff) cells, activated T cells, activated DCs and activated M1 macrophages all use aerobic glycolysis. Subpopulations of immunosuppressive cells, such as regulatory T (Treg) cells, myeloid suppressor cells, DC resting and naive T cells, using fatty acid oxidation to provide energy [62, 63]. Therefore, for a long time, energy competition will occur between tumour cells and immune cells and will then affect tumour cell survival. A large number of previous studies have confirmed that the way immune cells function can be affected by changes in nutritional factors in the tumour microenvironment. For example, severe nutritional deficiencies will make Treg cells use lactic acid as an energy substrate, and inhibiting lactic acid metabolism can significantly reduce the Treg cell’s content [64].
The PI3K-Akt signaling pathway is the most important pathway in regulating the immune environment, which can significantly affect immune cell effector functions and regulate intrinsic characteristics of the immune system [65]. In the tumor microenvironment, multiple immune cells coexist and interact, and the activation of most immune cells is affected by the PI3K-AKT signaling pathway [66, 67]. AKT could regulate the balance of terminal differentiation and memory CD8
In PPI network analysis, we identified 20 hub genes with the highest degree of interaction. TNF (tumor necrosis factor), which is mainly secreted by macrophages, can bind to its receptors TNFRSF1A/ TNFR1 and TNFRSF1B/TNFBR to exert its function. TNF could involve in the development of tumors by regulating inflammatory and immune responses. The role of these 20 hub genes in prostate cancer progression was further analysed. In the classic and authoritative prostate cancer database GSE21032, we used the disease prognosis information and found that the expression levels of three genes are associated with the outcome of the disease, including CCR5, COL1A1 and CXCR3. Chemokine receptor CCR5 play important roles in inflammatory processes and in immune system regulation and has been shown to contribute to proliferation and metastasis of prostate cancer [70, 71]. CCR5 antagonists, such as anibamine, could block specific metastasis of prostate cancer cells to bone and could be potential therapeutic agents targeting prostate cancer in the future [72, 73]. COL1A1 is involved in the development of mutiple cancer types, such as breast cancer cells [74], gastric cancer [75], and pancreatic cancer [76]. CXCR3 could inhibit cell proliferation and invasion ability through the PCL
Moreover, mutations of critical genes could also influence the effect of immunotherapy in prostate cancer. BRCA1 and BRCA2 are two tumor suppressor genes first described in hereditary breast cancer [78]. BRCA1/2 are involved in homologous recombination (HR)-mediated repair of DNA double-strand breaks, and mutations in BRCA1/2 often lead to an increase of the somatic mutation rate [79, 80, 81]. Prostate cancer patients with BRCA1/2-deficient not only show increased sensitivity to platinum-based chemotherapies [82], but also show sensitivity to PARP inhibitors [24, 83]. Moreover, previous results have shown that BRCA2-mutated prostate cancer could influence intratumoral immune infiltration: such patients have an enhanced intratumoral immune infiltration, in particular T lymphocytes expressing CD4, CD8, and FOXP3, compared to BRCA1/2 wild-type tumors [84]. These results indicate the significance of BRCA1/2 mutations in future research.
Our study contained several limitations. We utilized a retrospective RNA-seq data to deconvolute immune cell population by bioinformatics analysis. The exact immune populations will need to be validated using an enriched stromal population or at the protein levels. Patients’ different systemic immune status and different tissue sites might result in different immune, cytological and histological backgrounds, which could affect the infiltration of immune cells and cause bias when compared. More samples are needed for further validation in future. Moreover, the current clinical data in related databases are relatively limited. In the following research, we will use the updated database and our own tissue samples for further research to dig out the correlations between the identified types and genes of immune cells and clinical data. Furthermore, association between immune infiltrating cells and the response to chemotherapy or immunotherapy will need to be explored. The present study supports further functional studies to identify the roles of the hub genes in clinical mCRPC progression.
Conclusions
In conclusion, this study illustrated the immune cell infilitration pattern in mCRPC, and identified that a high level of M0 macrophages and a low level of CD4 T cells predicted poor prognosis. Of the 20 hub genes, high expression levels of CCR5, COL1A1, and CXCR3 were shown to predict shorter biochemical recurrence. These results can help us better understand the immune cell infiltration pattern in mCRPC lesions and provide a certain theoretical basis for future immunotherapy. However, the evidence of this study remains indirect and more clinical and basic researches are needed for further research.
Data availability statement
The gene expression data (accession number: prad su2c 2019 and prad su2c 2015) were downloaded from online datasets cBioPortal for Cancer Genomics website and GitHub website (Websites:
Author contributions
Conception: JW and GD.
Interpretation or analysis of data: CF, FW, CZ and WL.
Preparation of the manuscript: CF, KL and WL.
Revision for important intellectual content: CF, WL and JW.
Supervision: JW and GD.
Supplementary data
The supplementary files are available to download from
sj-xlsx-1-cbm-10.3233_CBM-203222.xlsx - Supplemental material
Supplemental material, sj-xlsx-1-cbm-10.3233_CBM-203222.xlsx
sj-pdf-1-cbm-10.3233_CBM-203222.pdf - Supplemental material
Supplemental material, sj-pdf-1-cbm-10.3233_CBM-203222.pdf
Footnotes
Acknowledgments
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the National Natural Science Foundation of China (grant no. 81802565); Natural Science Foundation of Jiangsu Province (grant no. BK20180216); Key Project of the Scientific Research Project of Nanjing Medical University Affiliated Suzhou Hospital (grant no. szslyy2017005).
Conflict of interest
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
