Abstract
BACKGROUND:
Blood-based protein biomarkers for detection of colorectal cancer (CRC) have been submitted to intense research to improve the full potential in screening for CRC.
OBJECTIVE:
The aim was to explore the diagnostic performance of 92 proteins related to inflammation and carcinogenesis in detection of CRC or precancerous lesions.
METHODS:
Blood-samples were collected from 4,698 individuals undergoing colonoscopy. An explorative unmatched case-control study was designed with 294 cases (individuals with CRC or high-risk colorectal adenoma) and 490 controls (individuals with low-risk colorectal adenoma, non-malignant findings or clean colorectum at colonoscopy). Protein profiling was performed by multiplex proximity extension assay. Statistical analyses were performed as univariate and multivariate logistic regression analyses.
RESULTS:
Univariably, CSF-1, MMP12 and IL8 demonstrated superior performance in discrimination of individuals with CRC. Recurrently, IL8 was included as contributor in majority of multivariate models discriminating individuals with CRC. The multivariate evaluation in discrimination of individuals with CRC demonstrated AUC
CONCLUSIONS:
A subset of biomarker candidates, specifically IL8, investigated in the present study suggest a potential as blood-based biomarkers in screening of CRC.
Keywords
Abbreviations
Introduction
Early detection of colorectal cancer (CRC) and precancerous bowel lesions (high-risk adenoma (HRA)) by screening has been shown to reduce mortality [37], which has led to the implementation of national screening programs in some countries, while others are still considering to initiate such programs [36]. Bowel examinations including colonoscopy and sigmoidoscopy are well-established modalities for diagnostic approaches. However, bowel endoscopy as a screening concept is challenged by disadvantages, such as bowel preparation, inconvenience for the individual, invasiveness and risk of severe complications [35], in addition to costs and limited healthcare budgets. Current screening programs for CRC are mostly based on non-invasive stool-based test for occult blood (fecal occult blood test (FOBT)) [36]; fecal immunochemical test (FIT) is the preferred choice due to superior performance, cost and convenience compared to other available FOBTs. However, the required self-sampling and handling of stool for testing may challenge the adherence to feces based screening [34]. In Denmark for instance, the compliance to FIT is 58% [28] and the applied FIT test (OC-Sensor, Eiken Pharmaceuticals, Japan) has a sensitivity 76% at 94% specificity [22] and thereby population screening does only identify 44% of the individuals with a CRC (compliance 58% X sensitivity 76%
Consequently, major research efforts have been put into the development of blood-based, cancer-associated biomarkers or panels of combined biomarkers for the detection of colorectal neoplasia. Recent promising results suggest a variety of blood-based biomarkers for early detection of colorectal neoplasia including proteins [41, 6, 2], circulating tumor DNA methylations/ mutations/fragmentations [31, 14, 8], microRNA [33], nucleosomes and histone modifications [32] and meta- bolomes [4]. However, a blood-based biomarker test as an alternative to FIT is still anticipated. Presently, the test for methylated Septin 9 (mSEPT9) (Epi proColon, Epigenomics Inc.) remains the only FDA approved blood-based biomarker test in CRC screening, but reported diagnostic performance is not competitive to FIT [7, 16]. The combination of biomarkers and even combination of different modalities in screening test could potentially provide the required performance.
The development of CRC occurs predominantly from accumulated genetic and epigenetic DNA alterations of the epithelial cells in the colorectum, which eventually undergo malignant transformation into adenocarcinoma [24]. The solid tumor-tissue of CRC is inhomogeneous and consists of a variety of different cells including malignant, immune and inflammatory cells [38, 25]. Solitary or in combination, these cells contribute to the surrounding inflammatory and immune response, growth of tumor cells, metastasis and angiogenesis by cell-to-cell contact and/or the production of inflammatory mediators such as cytokines. Consequently, the expression and secretion of various proteins from tumor cells and the surroundings are altered and possibly released into the blood stream with a potential influence on protein levels in the circulation.
The aim of the present study was to explore the diagnostic performance of 92 proteins related to inflammation and carcinogenesis in detection of early CRC or precancerous lesions (HRA) in a population of individuals with symptoms attributable to CRC. In addition, the diagnostic performance of a combined biomarker panel consisting of the best performing biomarkers was identified and subsequently evaluated.
Methods
Study design and population
The present study was conducted in the context of the Endoscopy II protocol as previously described [41, 32] and was approved by the Ethics Committee of the Capital Region of Denmark and the Danish Data Protection Agency and performed according to the Helsinki II Declaration. The Endoscopy II protocol has founded a biobank comprising blood samples from individuals undergoing colonoscopy due to symptoms attributable to CRC from May 1, 2010 to November 30, 2012 at seven hospitals in Denmark (Aarhus, Bispebjerg, Herning, Hillerød, Horsens, Hvidovre and Randers). In addition to symptoms, the inclusion criteria included age
The proposed design of the present study was an explorative unmatched case-control study with cases defined as individuals with either CRC or HRA and controls defined as individuals with low-risk adenoma (LRA), non-malignant findings or no findings (clean colorectum) at colonoscopy. Cases and controls were randomly selected from the Endoscopy II population including 196 individuals with CRC (49 individuals with stage I CRC, 49 individuals with stage II CRC, 49 individuals with stage III CRC and 49 individuals with stage IV CRC), 98 individuals with HRA, 98 individuals with LRA, 196 individuals with non-malignant findings and 196 individuals with clean colorectum at colonoscopy. The total number of included individuals was 784. Additional data regarding age, gender, localization of CRC or colorectal adenoma (colon or rectum) were recorded along with the presence of comorbidity (diabetes I/II, cardiovascular, lung or rheumatic disease or previous history of malignant disease). Figure 1 provides a flowchart which demonstrate descriptive data and random selection of the study participants.
Flowchart illustration the inclusion and descriptive data of the present study. CRC: colorectal cancer, CC: colon cancer, RC: rectal cancer, HRA: high-risk adenoma, LRA: low-risk adenoma.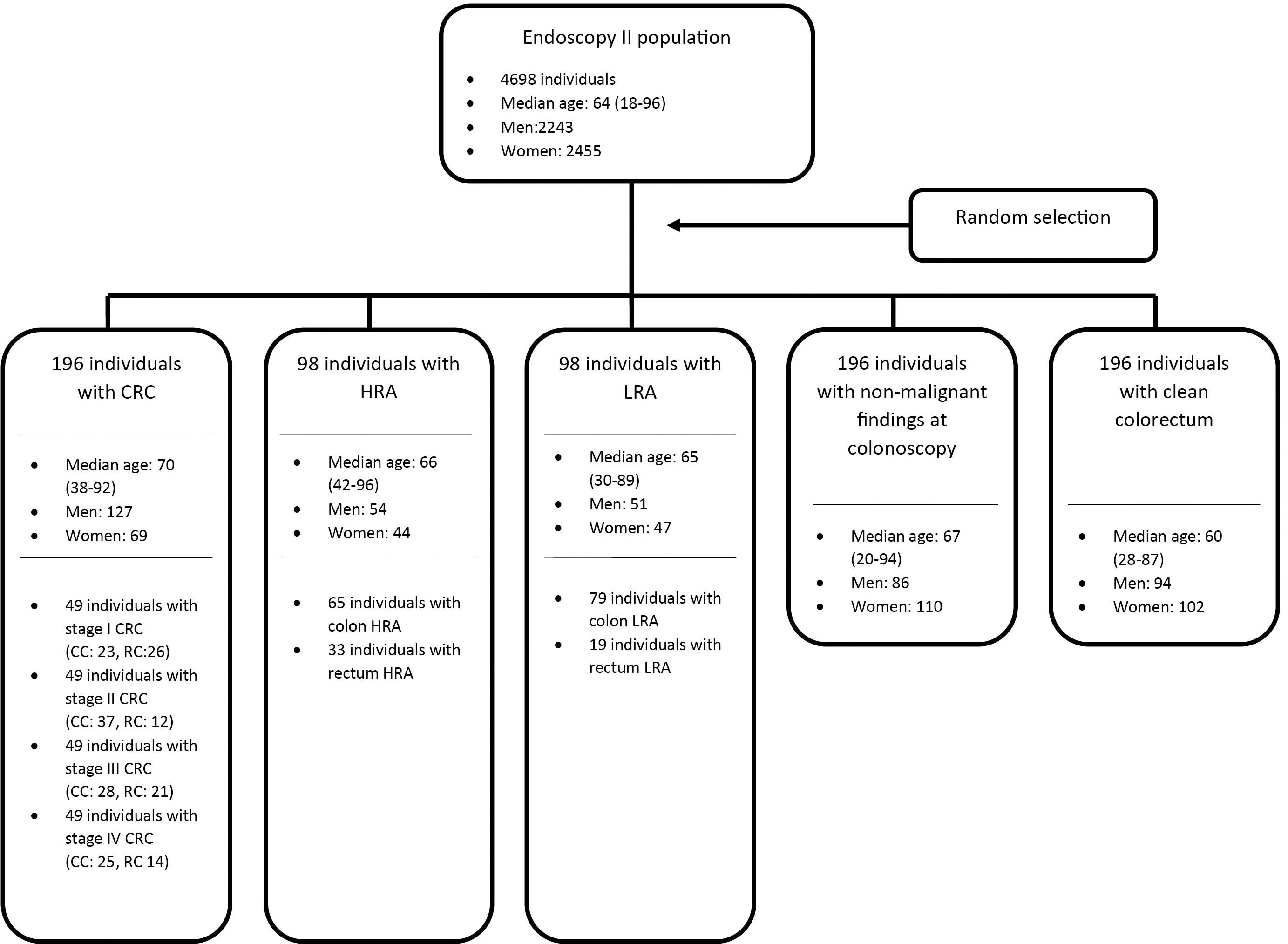
Three primary Endpoints and five secondary Endpoints were defined based on outcome at colonoscopy:
Primary Endpoints: Endpoint 1: The discrimination of individuals with CRC or HRA from all other individuals. Endpoint 2: The discrimination of individuals with CRC from all other individuals. Endpoint 3: The discrimination of individuals with HRA from all other individuals but excluding individuals with CRC. Secondary Endpoints: Endpoint 4: The discrimination of individuals with CRC stage I or II from all other individuals but excluding individuals with CRC stage III or IV. Endpoint 5: The discrimination of individuals with CRC stage III or IV from all other individuals but excluding individuals with CRC stage I or II. Endpoint 6: The discrimination of individuals with CRC stage I or II from individuals with clean colorectum. Endpoint 7: The discrimination of individuals with CRC stage III or IV from individuals with clean colorectum. Endpoint 8: The discrimination of individuals with HRA from individuals with clean colorectum.
The blood samples for EDTA plasma were collected by dedicated research nurses on the day of planned colonoscopy prior to the procedure and after bowel preparation. The blood samples were handled according to a predefined, validated standard operating procedure including centrifugation at 3,000 g for 10 minutes at 21
Laboratory procedures
The protein profiling was performed by using the Olink AB (Stockholm, Sweden) custom multiplex proximity extension assay (PEA) which allows simultaneous quantification of 92 human immunological and tumor-associated proteins (IMMUNO-ONCOLOGY panel). A full list of the 92 protein biomarkers is specified in the Supplementary (Table S1). The IMMUNO-ONCOLOGY panel was chosen as it reflects various cellular mechanisms in carcinogenesis and inflammation associated with early CRC or its precancerous lesions. The laboratory analyses were performed according to manufactures instructions at BioXpedia A/S (Aarhus, Denmark). In brief, 92 oligonucleotide-labeled antibodies in pairs were allowed binding to their respective target protein in the plasma sample thereby bringing the paired oligonucleotides into proximity to provide proximity-dependent DNA polymerization. The subsequent detection and quantification were performed by quantitative real-time PCR (qPCR).
Standardization and normalization of raw data were performed according to manufactures instructions [29]. For each sample analyzed, internal controls (incubation, extension and detection controls) were added. In addition, external controls (inter-plate controls, negative controls and sample controls) were added to each 96-well plate. For each protein-assay, controls were used to calculate limit of detection and to normalize raw data as Normal Protein Expression (NPX); an arbitrary, relative quantification on the log2 scale, where a higher NPX value represents a higher level of the target protein in plasma with a background level at approximately zero.
Statistics
Data management and statistical analyses were done using SAS (v9.4, Cary, N.C., USA), and further statistical analyses were performed with R version 3.6.3 (2020-02-29) using the package “rms” (Frank E Harrell Jr (2019). rms: Regression Modeling Strategies. R package version 5.1-4). The significance level was 5%.
The primary as well as secondary Endpoints are described above. The explanatory variables were age, gender and the 92 markers (expressed as NPX). The statistical analysis was done using logistic regression analysis modelling the probability of the respective endpoints. All NPX values were log transformed base 2, with results presented by the p-values. In addition, sensitivities for 70, 80 and 90% specificities were presented for multivariate models. The area under the receiver operating characteristic curves (AUC
Initial analysis for the selection of relevant biomarkers was done using univariate logistic regression for each of the 92 biomarkers, choosing those biomarkers with
Comorbidity was assessed by including each comorbidity in the multivariate model with an interaction between the biomarkers and the comorbidity.
Results of the multivariate analyses for Endpoint 1–8 are presented with regulation (
/
),
-value and AUC
with sensitivities at specificities 70%, 80% and 90%
Results of the multivariate analyses for Endpoint 1–8 are presented with regulation (
Characteristics of the population
The study design is shown in Fig. 1 including descriptive statistics.
Individuals protein assessment
Results of the univariate analyses of all 92 proteins for all 8 Endpoints are presented by the
In the overall identification of individuals with CRC (Endpoint 1, 2 and 4–7), three proteins (colony stimulating factor 1 (CSF-1), matrix metalloproteinase 12 (MMP12) and interleukin 8 (IL8)) consistently demonstrated significant superior performance in the univariate evaluation. Restricting the discrimination to individuals with late stage CRC (stage III–IV) demonstrated improved performance, and further improvements were shown by restricting the control group to individuals with clean colorectum. In details, 50 of the 92 proteins demonstrated significance in discrimination of individuals with CRC or HRA (Endpoint 1) with superior performance by CSF-1, MMP12 and IL8 demonstrating AUC
The overall identification of individuals with HRA (Endpoint 3 and 8) demonstrated superior discrimination by two proteins (hepatocyte growth factor (HGF) and MCP-3). In specific, nine proteins demonstrated significance in discrimination of individuals with HRA from all other individuals excluding individuals with CRC (Endpoint 3). MCP-3, Chemokine ligand 20 (CCL20), heme oxygenase 1 (HO-1) and TNF receptor superfamily member 12A (TNFRSF12A) all demonstrated AUC
Assessment of the multivariate models
Results of the cross validated multivariate analyses for Endpoint 1–8 are presented with p-values and AUC
An overall improved discrimination of individuals with CRC (Endpoint 1,2 and 4–7) was observed in the multivariate evaluation compared to univariate evaluation. A subset of proteins from the 92 protein-panel was included in the multivariate models with IL8 recurrently identified as contributor. In specific, the discrimination of individuals with CRC or HRA (Endpoint 1) included Inducible T-cell co-stimulator ligand (ICOSLG) and IL8 in the model and demonstrated AUC
The overall discrimination in the multivariate evaluation of individuals with HRA (Endpoint 3 and 8) was inferior compared to the discrimination of individuals with CRC. A single protein (T-cell surface glycoprotein 28 (CD28)) was included in the model in discrimination of individuals with HRA (Endpoint 3); AUC
Assessment of influence of co-morbidity
The potential influence of co-morbidity (diabetes I/II, cardiovascular, lung or rheumatic disease or previous history of malignant disease) in the multivariate analyses of Endpoints 1–8 was evaluated, and significance was not demonstrated for the interaction between levels of proteins and the presence of co-morbidity for any of the eight Endpoints as demonstrated in Table 2.
The influence of comorbidity in the multivariate analyses of Endpoint 1–8 presented with
-value for the comorbidity and the interaction between comorbidity and the protein biomarkers included in the multivariate model
The influence of comorbidity in the multivariate analyses of Endpoint 1–8 presented with
The aim of the present study was to explore and evaluate the diagnostic potential of 92 proteins associated with inflammation and carcinogenesis, individually or in combination, in detection of individuals with CRC or HRA.
Evaluated individually, majority of the proteins demonstrated significance in discrimination of individuals with CRC (Endpoint 1, 2, 4–7). Three proteins (CSF-1, MMP-12 and IL8) demonstrated consistent superior performance with IL8 achieving the highest performance in discrimination of individuals with CRC. Superior results for CSF-1, MMP-12 and IL8 were obtained when the analysis was restricted to discriminate individuals with late stage CRC (Endpoint 5 and 7 with AUC
In the combined multivariate evaluation in prediction of individuals with CRC (Endpoint 1, 2, 4–7), improved diagnostic performance was observed. Noteworthy, IL8 was recurrently included as contributor in majority of the multivariate models. The overall superior performance was demonstrated in identification of individuals with late stage CRC from individuals with clean colorectum (Endpoint 7) with inclusion of ARG1, ICOSLG and IL8 in the multivariate model (AUC
The results of the present study indicate a potential of CSF-1, MMP12 and, specifically, IL8 as biomarker candidates in CRC screening due to the persistent selection as superior discriminators of CRC in the univariate evaluation and the recurrent selection of IL8 as contributor in the multivariate evaluation. In the literature, these three proteins are involved in diverse biological processes and classified as modulators of the inflammatory and carcinogenic response, and changes in levels in the circulation as well as changes in expression in tumor tissue have been associated with CRC. In specific, CSF-1 is a cytokine primarily involved in the migration, activation and proliferation of macrophages in inflammation and carcinogenesis [13, 18, 10]. In relations to CRC, significantly altered expression in tumor tissue have been described [39]. MMP12 is an extracellular enzyme involved in breakdown and re-modeling of the extracellular matrix and are associated with cell proliferation, differentiation, inflammation, tissue invasion as well as angiogenesis [9]. In CRC, upregulation of MMP12 in tumor-tissue of CRC have been described as well as an indication of increased levels in the circulation of individuals with CRC [20, 43]. IL8 is a chemotactic cytokine inducing migration and activation of neutrophils as well as being associated with tumor growth, angiogenesis and metastasis [40, 1, 27, 5, 17, 19]. Upregulation in CRC tissue as well as in the circulation of individuals with CRC have been described [17, 26], and a potential as prognostic marker for CRC has been previously suggested [42].
An essential requirement to a screening test for CRC is the identification of individuals with early stage CRC prior to dissemination and with a potential of surgical resection. In the present study, discrimination of individuals with early stage CRC from individuals with clean colon (Endpoint 6) demonstrated an AUC
In the individual as well as combined evaluation, improved diagnostic performance was observed when discrimination was restricted to individuals with late stage CRC (Endpoint 5 and 7). Progression of CRC with increased tumor burden, increased inflammatory response in the surrounding tissue as well as dissemination may have influence on levels of proteins associated with carcinogenesis and inflammation which provides an explanation for the observed superior discrimination of individuals with late stage CRC compared to early stage CRC.
The discrimination of individuals with HRA (Endpoint 3 and 8) was inferior to that of individuals with CRC. In the individual evaluation, a smaller subset of the investigated proteins reached significance with inferior performance. HGF and MCP-3 demonstrated superior performance for Endpoint 3 and 8; AUC
Superior discrimination of individuals with CRC or HRA was obtained when the control group was restricted to individuals without findings at colonoscopy (clean colorectum). The study design includes a potential risk of underestimation of performance as the study cohort include individuals undergoing colonoscopy due to symptoms attributable to CRC which, consequently, lead to a symptomatic control group. The purpose of restricting the control group to individuals with clean colorectum was to indicate a potential true diagnostic performance. However, the design of the present study as an explorative unmatched case-control study specifies limitations for the interpretation of results and conclusions. A trend and a possible potential of the investigated proteins as biomarkers in identification of individuals with CRC or HRA can be insinuated, but the results are not referable to the study population of Endoscopy II or to the Danish population in general due to a selected and unrepresentative cohort. To refer the results to the background population, the results and conclusion of the present study must be validated in a non-selected, representative and asymptomatic CRC-screening population.
The analysis technique of PEA applied in the present study enables simultaneous quantification of 92 proteins in a single sample with high reproducibility and modest requirements to sample size (1
The various protein-candidates for biomarkers in CRC identified by PEA in present and previous studies could possibly reflect the diversity of biological pathways and mechanisms involved in the carcinogenic and inflammatory response affecting levels of various proteins and mediators. Furthermore, the proteins of interest are not specific in development and progression of CRC but are associated to inflammation and carcinogenesis in general.
The promising potential of IL8 demonstrated by PEA technique in the present study is recognized in previous research by different techniques such as biochip array technology [5], enzyme-linked immunosorbent assay [17] and multiplex immunoassay technology [19]. Diversity in study designs hinders comparison of the different techniques. To determine the optimal technique for IL8 measurement, future studies comparing the different techniques are needed.
The potential of specific proteins such as IL8 as biomarker for CRC indicated in the present study contribute to the foundation of knowledge to form hypotheses for future validation studies and could lead to improvement of the existing screening possibilities. The individual performance of IL8 indicated by the present study is insufficient to qualify as a biomarker for CRC. Nevertheless, possible combination with other promising or validated blood-based biomarkers could improve the diagnostic potential and provide a screening concept superior or competitive to FIT. It could even provide the opportunity to choose between modalities of screening (blood or fecal sampling) and possibly improve the adherence to a screening program by offering an alternative to fecal-based testing leading to improved efficiency and cost/benefit of existing screening programs.
The advantage of the applied technique of PEA is the simultaneous determination of 92 proteins in a single sample providing the opportunity to perform large scale studies in the search for blood-based protein biomarkers in various cancer types. The design of the protein panel of the present study was predefined as a commercial panel by the manufacturer (Olink AB, Stockholm, Sweden). The panels offered by Olink AB are designed in consideration of various biological processes (panels including proteins associated with cell-regulation, metabolism, organ damage, neurological disease and cardiovascular disease in addition to the IMMUNO-ONCOLOGY panel included in the present study). The future perspective of PEA in the context of cancer screening test could be the design of a protein panel composed of subunits of validated organ-specific cancer-associated protein biomarkers. The panel could identify “at risk” individuals with increased incidence of the various cancer types, and the organ-specific subunit protein profile could then be used as guidance for the choice of further diagnostic tests or examinations.
Conclusion
In summary, a subset of the proteins associated with inflammation and carcinogenesis from the panel of 92 proteins investigated in the present study with a systematic, prospectively collected cohort suggest a potential as blood-based biomarkers in screening of CRC. Of specific interest is CSF-1, MMP12 and particularly IL8 with superior performance in discrimination of individuals with CRC. The future perspectives could include the combination of IL8 with other biomarkers or screening modalities for CRC (established as well as experimental); a combination which could lead to further improvement in the screening of CRC.
Author contributions
Conception: LR, HJN, IJC.
Interpretation or analysis of data: LR, IJC.
Preparation of the manuscript: LR.
Revision for important intellectual content: LR, HJN, IJC.
Supervision: HJN, IJC.
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-203211.
sj-pdf-1-cbm-10.3233_CBM-203211.pdf - Supplemental material
Supplemental material, sj-pdf-1-cbm-10.3233_CBM-203211.pdf
Footnotes
Acknowledgments
The research nurses, secretaries and technicians at the participating hospital departments and laboratories are thanked for their skillful work.
The study received financial support from The Andersen Isted Fund, The Augustinus Foundation, The Beckett Fund, The Inger Bonnén Fund, The Hans & Nora Buchard Fund, CEO Jens Bærentsen (private donation), The Walter Christensen Family Fund, The P.M. Christiansen Family Fund, The Aase & Ejnar Danielsen Fund, The Erichsen Family Fund, The Knud & Edith Eriksen Fund, The Svend Espersen Fund, The Elna and Jørgen Fagerholt Fund, The Sofus Carl Emil Friis Fund, The Torben & Alice Frimodt Fund, The Eva & Henry Frænkel Fund, The Gangsted Fund, The Thora & Viggo Grove Fund, The H Foundation, The Erna Hamilton Fund, The Sven & Ina Hansen Fund, The Søren & Helene Hempel Fund, The Henrik Henriksen Fund, The Jørgen Holm Family Fund, Foundation Jochum, The KID Fund, The Kornerup Fund, The Linex Fund, The Dagmar Marshall Fund, The “Midtjyske Bladfund”, The Axel Muusfeldt Fund, The Børge Nielsen Family Fund, The Michael Hermann Nielsen Fund, The Arvid Nilsson Fund, The Obel Family Fund, The Krista & Viggo Petersen Fund, The Willy & Ingeborg Reinhard Fund, The Kathrine & Vigo Skovgaard Fund, The Toyota Fund, The Vissing Fund, The Wedell-Wedellsborg Fund and Hvidovre University Hospital (The Capital Region of Denmark).
