Abstract
BACKGROUND:
Few trials have evaluated the utility of liquid biopsies to detect epidermal growth factor receptor mutations (EGFRm) at the time of response evaluation and its association with the clinical characteristics and outcomes of non-small-cell lung cancer (NSCLC) patients.
OBJECTIVE:
This study aimed to evaluate, in a real-world clinical setting, the prevalence of plasma EGFRm and its association with the clinical characteristics, response and survival outcomes of NSCLC patients under treatment with EGFR-tyrosine kinase inhibitors (EGFR-TKIs).
METHODS:
This observational study enrolled advanced or metastatic NSCLC patients, with confirmed tumor EGFRm, receiving treatment with first- or second-generation EGFR-TKIs. Blood samples for the detection of plasma EGFRm were collected at the time of response evaluation and processed using the Target Selector™ assay. The main outcomes were the detection rate of plasma EGFRm, median Progression-Free Survival (PFS) and Overall Survival (OS) according to plasma EGFR mutational status.
RESULTS:
Of 84 patients, 50 (59.5%) had an EGFRm detected in plasma. After a median follow-up of 21.1 months, 63 patients (75%) had disease progression. The detection rate of plasma EGFRm was significantly higher in patients with disease progression than in patients with partial response or stable disease (68.3% versus 33.3%;
CONCLUSIONS:
In NSCLC patients receiving EGFR-TKIs, the detection of plasma EGFRm at the time of tumor response evaluation is associated with poor clinical outcomes.
List of abbreviations
Introduction
Lung cancer is the leading cause of cancer-related deaths worldwide, with non-small-cell lung cancer (NSCLC) accounting for approximately 85% of all newly diagnosed cases [1]. Approximately 70% of NSCLC patients are diagnosed with advanced-stage disease, which is associated with a poor prognosis. Improved outcomes have been achieved using targeted drugs to treat NSCLC patients with oncogenic aberrations [2]. Therefore, histological and molecular analysis from tissue biopsies or cytology specimens, collected at the time of diagnosis, is now routine in metastatic NSCLC to identify subsets of patients who are more likely to benefit from specific therapies. The most common oncogenic driver mutations in NSCLC are epidermal growth factor receptor (EGFR), in 14%–32% of the cases [3]. These include short in-frame deletions in exon 19 (Ex19del) and substitution of leucine-858 with arginine (L858R) in exon 21, together accounting for approximately 90% of all EGFR mutations (EGFRm) [4, 5]. Moreover, the frequency of EGFRm variates depending on race, ethnicity [5], with a particularly high prevalence in Asian and Latin American countries and among younger, non-smoker women [6].
In patients with these genotypes, treatment with first- or second-generation EGFR-tyrosine kinase inhibitors (TKIs), such as erlotinib, gefitinib, and afatinib, has resulted in better objective response rates (ORR), PFS, and quality of life (QOL), as compared to platinum-based chemotherapy [7, 8, 9]. However, the overwhelming majority of patients who initially respond to treatment with first- or second-generation EGFR-TKIs, eventually develop acquired resistance within 9–12 months [10], which, in approximately 50–60% of patients, is caused by a secondary threonine-to-methionine mutation at position 790 (T790M) of exon 20 [11, 12].
Although tissue biopsies remain the “gold standard” to identify specific EGFRm, they are subject to several limitations, including the risk of complications, tumor inaccessibility, and lack of sufficient and high-quality tissue for genetic testing [13]. Given these limitations, which are even more pronounced in patients with disease progression, liquid biopsy (LB) has emerged as a clinically validated alternative for molecular testing. Several prior studies demonstrated that different tumor biomarkers detected in blood samples, like EGFR, are upregulated in metastatic tissues and circulating tumor cells (CTC) as in primary tumor [14]. For molecular purposes, the concordance between plasma and tissue samples has been estimated at 89% in the ASSESS trial, a real-world data analysis, showing that the assay has high specificity but low sensitivity (46%) [15]. The concordance rate between tissue and plasma biopsies has been evaluated in several studies using different platforms and ranges between 74% to 97% [16, 17, 18].
In recent years, there has been an increase in the number of studies evaluating the clinical utility of LB in different settings [19]. LBs provide valuable information without the need of an invasive procedure or the bias from sampling a localized cell population, as is the case with standard biopsying procedures [20]. However, most studies to date have been performed in the context of controlled clinical trials, which include newly diagnosed patients for whom tissue and LB are available at baseline and at different time points after treatment initiation. In clinical practice, LBs are not routinely performed throughout the entire follow-up of patients. Thus, the correlation between the detection of plasma EGFRm and the outcomes of patients in a real-world clinical setting remains unclear. This study aimed to assess through an ultrasensitive LB the impact of plasma EGFRm detection on PFS and OS, in a cohort of advanced NSCLC patients treated with first- or second-generation EGFR-TKIs, and for whom LB were obtained after treatment initiation.
Materials and methods
Study design and oversight
In this single-center observational study, a total of 84 NSCLC patients harboring an EGFRm (EGFRm; Ex19del and/or L858R) and treated with first- or second-generation TKIs were recruited from February 2016 to October 2017 at the Thoracic Oncology Unit of the “Instituto Nacional de Cancerología” of Mexico (INCan) and followed during this time with liquid biopsies. The study was conducted in accordance with The Code of Ethics of the World Medical Association (Declaration of Helsinki), printed in the British Medical Journal (18 July 1964), following local laws on observational studies, and all applicable regulatory requirements. The present research was performed under a protocol approved by the Institutional Review Board and Ethics Committee of the Instituto Nacional de Cancerología of Mexico, [INCAN (018/022/ICI-CEI/1207/17)]. All patients provided a written informed consent form before enrollment and before any study-specific procedures were performed.
Patients
Eligible patients were adults (
Study end points and assessments
Main outcomes were progression-free survival (PFS) and overall survival (OS). PFS was defined as the time from EGFR-TKI therapy initiation to the time of confirmed disease progression, death, or loss of follow-up. PFS was analyzed according to the LB result (mutation detected vs. mutation non-detected). OS was defined as the time from diagnosis to death, or last contact at a cutoff date July 6
Tumor biopsies and EGFR mutational status
Tumor tissue was obtained from all patients at an initial diagnostic procedure and was paraffin embedded. Histopathological confirmation of NSCLC and determination of tumor EGFR mutational status was performed by an independent and centralized pathology unit using validated techniques and a clinically approved, allele-specific, quantitative real-time PCR test kit (Therascreen EGFR 29; Qiagen, Manchester, UK).
Tumor response
All patients were evaluated by chest, abdomen, and pelvic computed tomography (CT) or positron emission tomography-CT at baseline, then every 8–9 weeks during the first year of targeted therapy and subsequently, every 12 weeks or as clinically indicated. Brain MRIs were performed at baseline, and then every six months or as clinically indicated. Tumor responses were evaluated by an independent central imaging group, consisting of trained radiologists and oncologists who specialize in thoracic malignancies, and classified in accordance with RECIST 1.1 criteria.
Patient enrollment and testing flowchart. Flowchart summarizes patient enrollment, tests conducted, and mutations detected.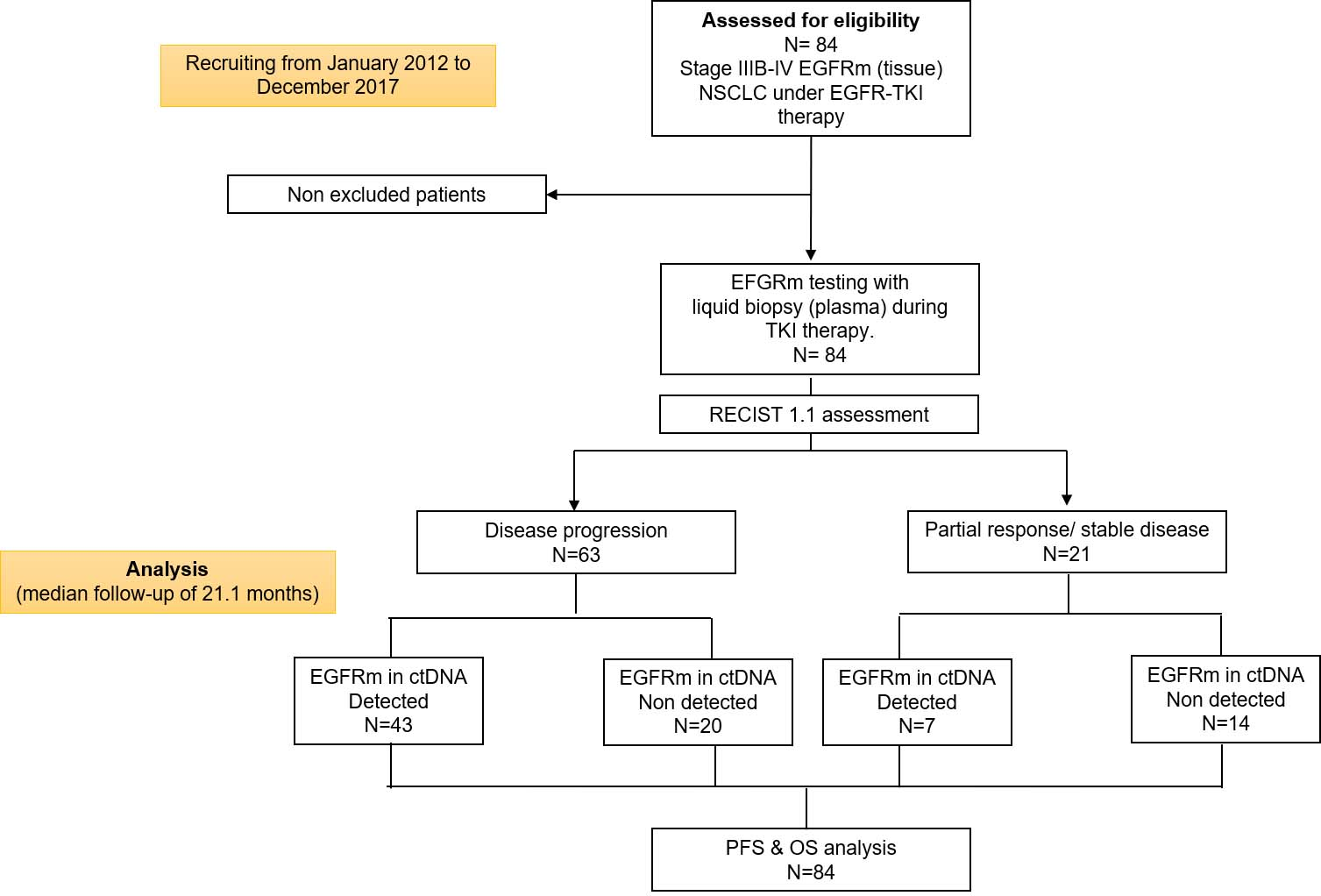
Liquid biopsies were taken at the time of response evaluation and at disease progression. Blood samples were collected in 10 mL CEE-Sure™ tubes (Biocept, Inc.). Samples were shipped to Biocept Laboratories (San Diego, CA) for processing. Circulating nucleic acid was isolated from blood plasma using standard procedures (QIAmp Circulating Nucleic Acid Kit, Cat. No. 55114) as of current recommendations in guidelines [21]. Then isolated DNA was used in Target Selector™ assays for the specific amplification of EGFRm: (1) deletion 746–750 in exon 19; (2) the L858R point mutation within exon 21; and (3) T790M within exon 20. Proprietary Target Selector™ switch blocker methodology uses forward and reverses primers, as well as a wild-type PCR blocker probe, to enrich the sample with mutant target sequences while specifically blocking PCR amplification of the wild-type allele. Subsequent Sanger sequencing of the Target Selector™ amplicons is used to confirm the presence of the specific mutations.
Carcinoembryonic antigen quantification
Carcinoembryonic antigen (CEA) quantification was performed at the Clinical Pathology Laboratory of the INCan using a solid-phase, two-site, sequential chemiluminescent immunometric assay (IMMULITE-2000-CEA Analyzer, Siemens, Inc., LA, CA). Serum samples (15
Statistical analysis
Continuous variables were summarized as medians with ranges, or as means with standard deviations (SDs), depending on data distribution. The distribution was assessed using the Shapiro-Wilk test with a p-value greater than 0.05 considered as normally distributed. Two group comparisons were tested using Student’s t-test or Mann-Whitney U depending on data distribution, while nominal data were analyzed using the chi-square (X
Results
Patient population
A total of 84 patients with advanced EGFRm NSCLC patients were included in the study. At a median follow-up of 21.1 months, 63 (75%) patients had DP, and 21 (25%) had a partial response or stable disease. The patient enrollment and testing flowchart is summarized in Fig. 1. Population baseline characteristics are shown in Table 1. The median age was 61 years (range, 31–88), 59 (70.2%) women, 48 (58.3%) current or former smokers, 28 (33.3%) had a history of wood-smoke exposure, and 68 (81%) had stage IV disease. Most of the patients had an ECOG-PS
Patients characteristics at diagnosis
Patients characteristics at diagnosis
Abbreviations: ECOG-PS, Eastern Cooperative Oncology Group Performance Statu; EGFRm, epidermal growth factor receptor mutations; CEAi, Carcinoembryonic antigen at diagnosis; ctDNA, circulating tumor DN; EGF-TKI, epidermal growth factor receptor-tyrosine kinase inhibitor.
Of 84 patients, 50 (59.5%) had an EGFRm detected in plasma regardless of the state of the disease. Among these patients, T790M was the most common genetic aberration detected in 21 (42%) patients (alone or combined with other mutations). T790M mutation was detected alone in 3 (6%) patients, in combination with Ex19del in 12 (24%) patients, and with L858R in 6 (12%) patients. The second most frequent genetic aberration was Ex19del, which was detected in 17 (34%) patients, followed by the L858R mutation in 12 (24%) patients. (Fig. 1).
The detection rate of plasma EGFRm was 68.3% (43 of 63) in patients with DP and in patients with partial response or stable disease 33.3% (7 of 91). A chi-square test of independence was performed and confirmed that EGFRm were more likely to be detected in patients with DP than in patients with partial response or stable disease
Clinical factors associated with a positive EGFRm in ctDNA
Clinical factors associated with a positive EGFRm in ctDNA
Abbreviations: ECOG PS, Eastern Cooperative Oncology Group Performance Statu; CNS, Central nervous system; CEA
Frequency of EGFRm. Histograms showing the proportion of EGFRm in diagnostic tissue and liquid biopsies after initiation of EGFR-TKI treatment in all patients or classified according to progression status. Abbreviations: PD, Progressive disease; SD, stable disease; PR, partial response.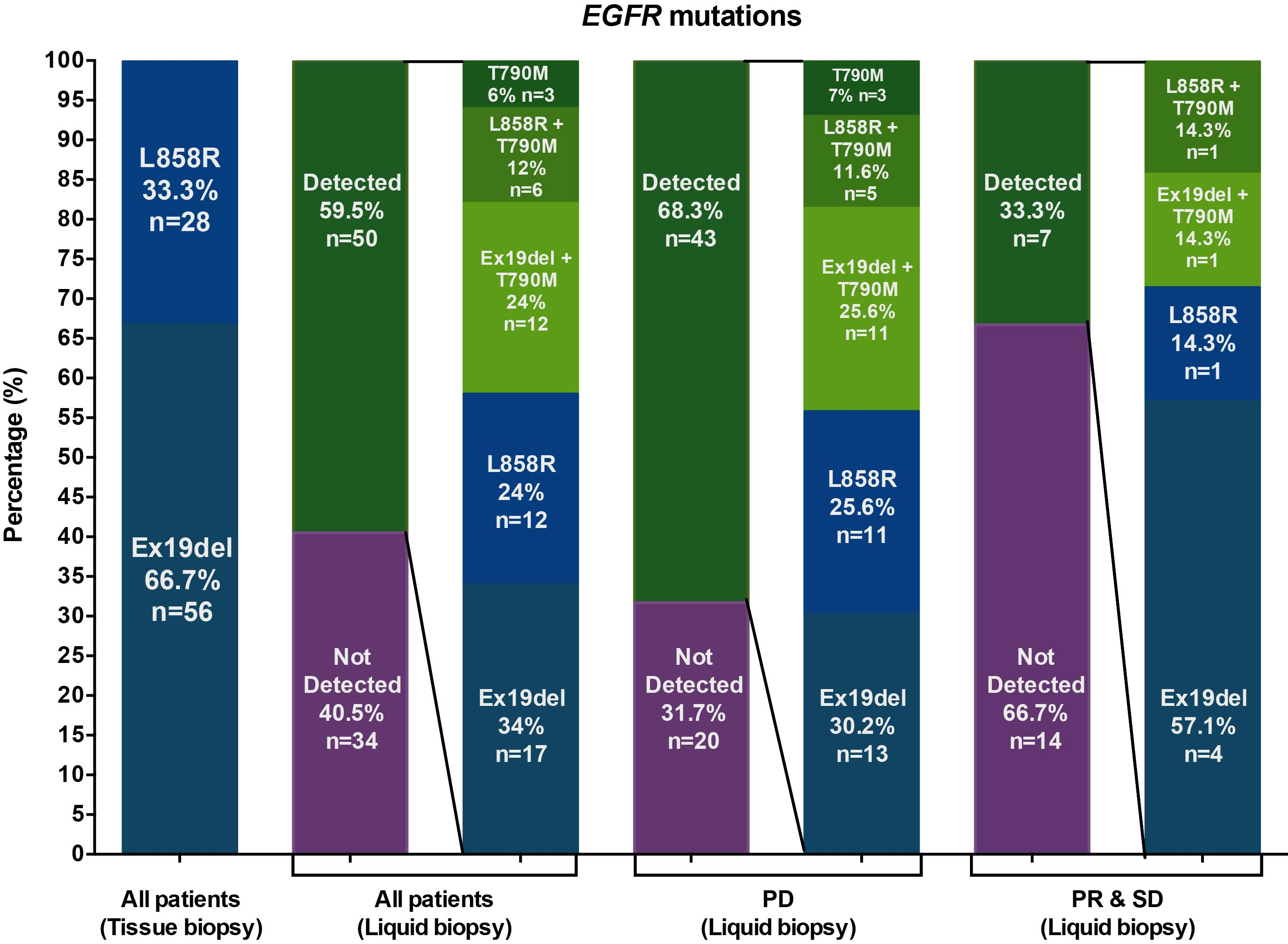
In patients with partial response or stable disease, the most prevalent genetic aberrations were sensitizing mutations: Ex19del was detected in 4 (57.1%) patients, or as a commutation with T790M in 1 (14.3%) patient. The L858R aberration was found as a single or in combination with T790M in 2 (28.6%) patients (Fig. 2). Conversely, among patients with DP, the most frequently detected mutation was T790M in 19 (44.2%) of 43 patients, either alone or in combination with other alterations. In this subset of patients, T790M was detected as a single mutation in 3 (7%). A commutation was presented in 11 (25.6%) with Ex19del, and in 5 (11.6%) patients with L858R mutation. The second mutation in frequency was Ex19del, which was detected alone in 13 (30.2%) patients, followed by L858R in 11 (25.6%) patients (Fig. 2).
The clinical characteristics associated with a higher probability of detecting a positive EGFRm in ctDNA were the presence of
Survival outcomes according to EGFR status in the LB
After a median follow-up time of 21.1 months (95% CI, 16.31–25.93), the median PFS since EGFR-TKI initiation therapy was 11.0 months (95% CI 9.25–19.39) in the subgroup of patients with a positive plasma EGFRm vs. 14.3 months (95% CI 8.61–13.46) in those without detected mutations. (
Kaplan-Meier curves. Progression-free survival according to the A) detection status of an EGFRm in plasma; B) presence of sensitizing or resistance EGFRm in plasma. Overall survival from diagnosis according to the C) detection status of an EGFRm in plasma; D) presence of sensitizing or resistance EGFRm in plasma.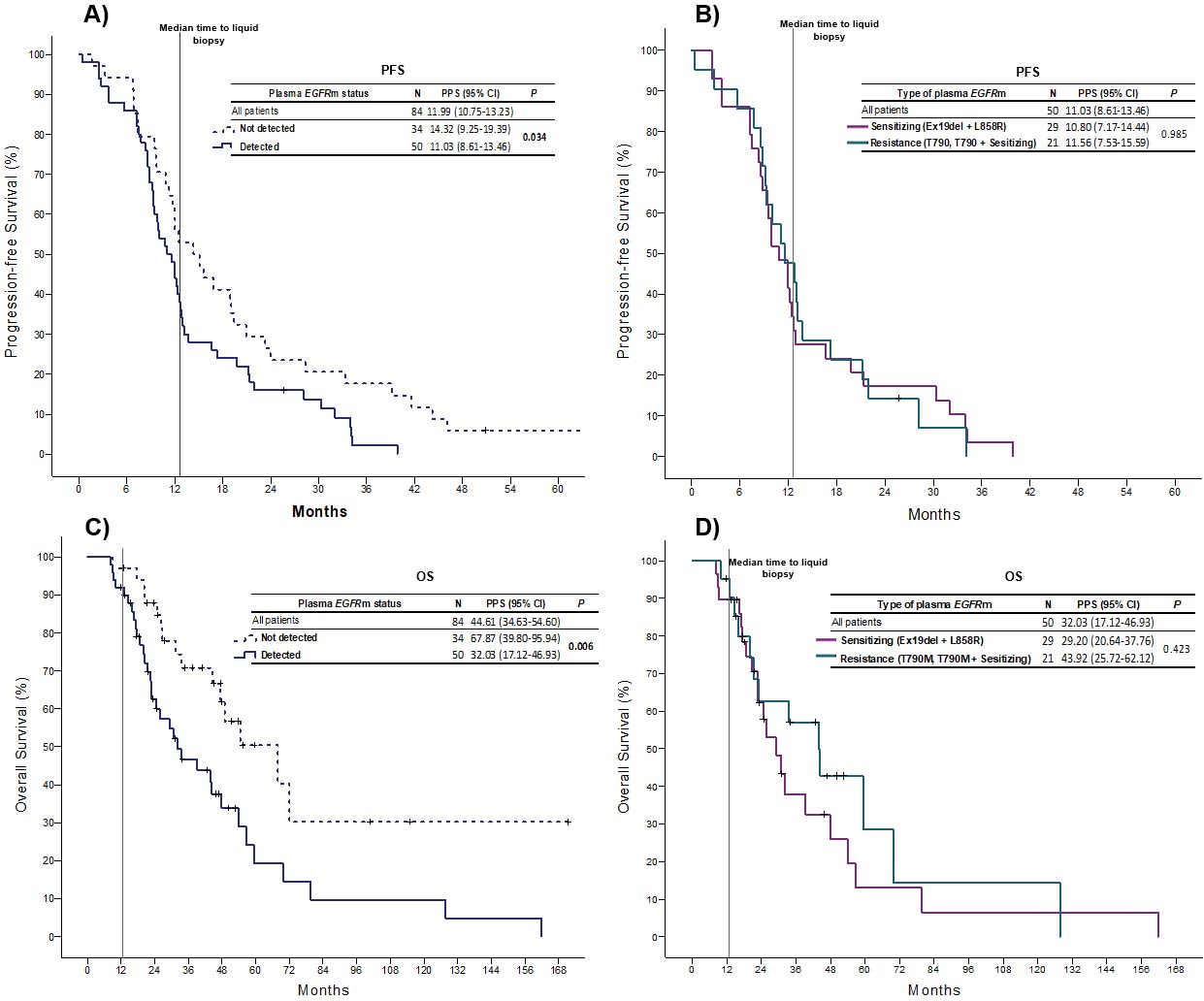
The median OS for the total cohort from diagnosis was 44.6 months (95% CI 34.6–54.6). A significant difference in the median OS was observed between patients with detected and non-detected EGFRm in plasma, 32.0 months (17.1–46.9) vs 67.8 months (95% CI 39.8–95.9), respectively. (Fig. 3C) In patients with an acquired resistant EGFRm (T790M) compared with sensitizing mutations, the median OS was 43.9 months (95% CI, 25.72–62.12) vs. 29.20 months (95% CI, 20.64–37.76). However, this difference was not statistically significant (
Multivariate Cox regression analysis of factors correlated OS after liquid biopsy
Abbreviations: PPS, post-progression surviva; EGFRm, epidermal growth factor receptor mutations; DP disease progressio; PR partial response; SD stable disease.
Kaplan-Meier curves. Overall survival from liquid biopsy according to A) detection of any EGFRm in plasma; B) detection of sensitizing or resistance EGFRm in plasma.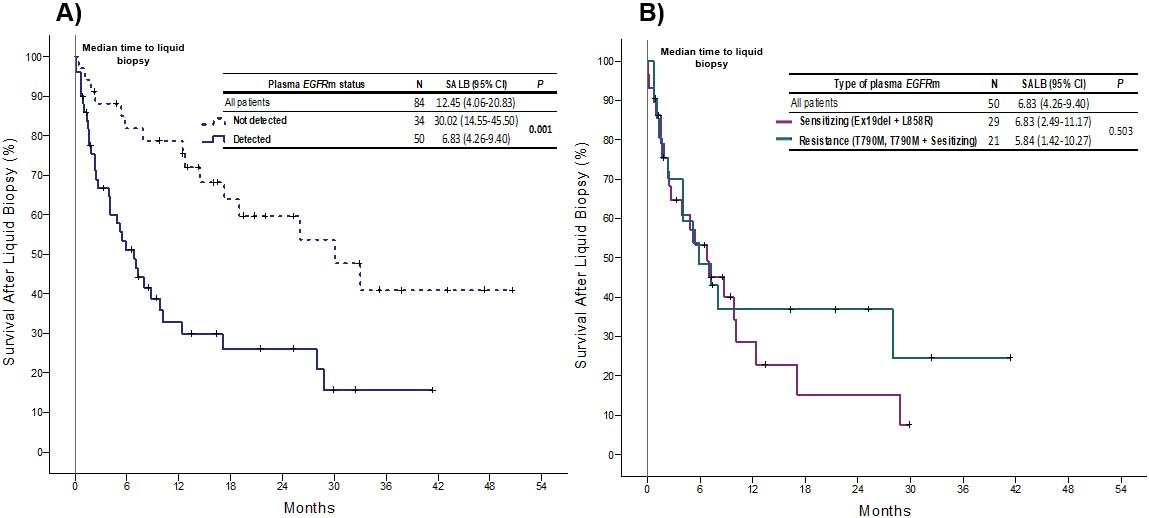
The median OS after LB testing was 12.45 months (95% CI, 4.06–20.83); irrespective of the EGFR status; the median OS from LB testing was 30.0 months (95% CI, 14.55–45.50) versus 6.83 months (95% CI, 4.26–9.40) in patients with non-detected and detected EGFRms,
Two multivariate Cox proportional hazard models were performed. In the first model, negative plasma EGFRm was independently associated with a lower risk of death (hazard ratio [HR]: 0.706 [95% CI, 0.535–0.931;
Until recently, the use of liquid biopsies was restricted to research settings, with the “Cobas
With very few exceptions [15, 24, 25, 26], information on the clinical application and utility of plasma EGFRm comes from controlled clinical trials, which generally include patients for whom tissue and liquid biopsies are collected at baseline and at different time points after treatment initiation. Therefore, much uncertainty exists about the validity of using liquid biopsies to detect EGFRm in patients that are already under EGFR-TKI treatment, and for whom longitudinally collected and matched biopsies (tissue and liquid) are not available. This study evaluated, in a real-world clinical setting, the utility of plasma EGFRm testing, at the time of response evaluation, in patients with advanced and metastatic NSCLC receiving EGFR-TKI treatment.
Controlled trials evaluating whether the outcomes of patients would change, if mutational status and treatment allocation were based on the results from liquid biopsies, rather than from tumor tissue biopsies, are limited. However, there is evidence indicating that the clinical outcomes of patients would not change if treatment allocation were based on the results from liquid biopsies [27].
A strength of the current study is that, in contrast to other studies where plasma EGFRm testing has been performed using different platforms (with varying analytical detection limits), all samples were tested using the Target Selector™ assay, an ultrasensitive LB test processed in a single CLIA-certified and CAP-accredited BIOCEPT laboratory, which minimized variation due to sample processing. A recent study of the analytical validation of Target Selector™ showed that the assay: a) has an analytical sensitivity and specificity higher than 99%; b) can detect a single mutant copy for EGFR, BRAF, and KRAS mutations in plasma ctDNA (the lowest possible limit for any DNA-based diagnostic test); c) has a clinical specificity higher than 99%. Finally, the rate of detection of EGFRm using this platform highly correlates with the overall frequency of EGFRm in NSCLC patients in North America [28].
According to our results, the overall detection rate of plasma EGFRm was 59.5% but the detection rate was significantly higher in patients with DP (68.3%) than in patients with partial response or stable disease (33%). These results are agreement with previous studies showing that disease burden positively correlates with a higher probability of detecting genetic aberrations in plasma [29, 30].
In the current study we report a detection rate of plasma T790M of 44.2% (19/43) in patients with DP, and of 9.5% (2/21) in patients without progression. These results are broadly consistent with the findings of other studies [31, 32, 33], showing that the detection rate of plasma T790M mutations ranges between 28.6% to 36.5% in patients with progression after first- or second-generation EGFR-TKIs. Of note, in patients with radiological progression, plasma T790M was more frequently detected in combination with the original activating EGFRm (Ex19del or L858R), detected in diagnostic tissue biopsies. [34, 35, 36].
The prognostic utility of detecting EGFRm in plasma from NSCLC patients treated with EGFR-TKIs throughout the course of the disease has been extensively explored, with available data showing that the timing of the liquid biopsy affects whether EGFRm are detected or not. At baseline, some studies have shown that the presence of plasma EGFRm at is associated with longer PFS and OS [37, 38, 39, 40], while others have shown that patients with high baseline levels of EGFRm, or in whom EGFRm persist , have a shorter PFS and OS [41, 42, 43].
Our results indicate that detection of plasma EGFRm after treatment initiation with a first- or second-generation EGFR-TKI is associated with a significantly worse PFS (14.3 vs. 11 months,
Although in the current study data was not collected to explain the markedly different survival outcomes between patients with plasma EGFRm and those without plasma EGFRm, it has been suggested that the quantity of ctDNA in circulation is associated with a more disseminated disease, extrathoracic site involvement, and a higher disease burden [20], which correlate with poor clinical and radiological outcomes [30]. Other studies have shown that radiological measurements of progression by RECIST 1.1 may be inadequate in patients with pleural effusions, diffuse nodules, or tumors with poorly defined margins. Under these circumstances the detection of EGFRm in plasma might be a complementary tool to obtain additional information for monitoring purposes [48, 49, 50]. Further, there are numerous reports showing that detection of plasma EGFRm in ctDNA often precedes radiological progression by 12 to 344 days [36, 41, 51, 52], which stands in stark contrast with the low detection rates of the T790M mutation in tissue biopsies obtained before confirmed radiological progression, as opposed to biopsies collected after radiological progression in which T790M can be detected in approximately 50–60% of cases [11, 12, 53].
In our study, no significant differences were found in the survival outcomes of patients according to the specific plasma EGFRm detected (Ex19del, L858R, T790M, or combinations). This finding is in agreement with previous studies showing that patients with EGFRm detected in plasma have worse survival outcomes regardless of the specific mutations detected in plasma [9, 54]. Although we report no differences in OS according to specific plasma EGFRm, a favorable trend was observed in patients with the T790M mutation. Indeed, previous studies showing that the sustained presence of plasma EGFRm or the appearance of T790M in plasma is a prognostic factor for poor PFS and OS in NSCLC patients treated with EGFR-TKIs [31, 41, 52, 55, 56, 57, 58], with only a few studies reporting that the presence of plasma T790M is associated with a better PFS and OS [59].
Of note, a longer PFS and OS has been more commonly reported in studies where the detection of T790M is performed on tissue biopsies collected after initiation of EGFR-TKI treatment [12, 60, 61], which has been generally ascribed to a more indolent tumor behavior. However, an important bias in both retrospective and prospective studies using serial tissue biopsies is the unavoidable exclusion of patients who cannot undergo a repeat biopsy because of poor health, or whose disease progressed rapidly and died before a second tissue biopsy could be collected.
A retrospective analysis of the AURA phase 1 trial did not find significant differences in the outcomes between patients with a positive T790M mutation detected in plasma, compared with a tissue-based assay [34]. Similarly, the AURA phase 2 trial found an ORR of 64% regardless of the detecting method for the T790M mutation, using plasma or tissue [62]. The first prospective study evaluating the efficacy of osimertinib in advanced NSCLC after DP to first-line therapy based on a positive LB test result for T790M mutation demonstrated comparable ORR and PFS, consistently with previous reports based on tumor tissue biopsies [63].
Insufficient information exists regarding other clinical characteristics associated with the detection of plasma EGFRm and its implications. Some studies have found that the type of EGFRm, such Ex19del and the local pattern of progression could be associated with an increased rate of T790M detection in plasma [33]. However, these results have not been consistent across studies. In our study there were no clinical characteristics significantly associated with the detection of T790M mutation in plasma. On the other hand, our results indicate that the probability of detecting a plasma EGFRm is associated with the presence of
The use of osimertinib for the second-line treatment of NSCLC patients with the T790M mutation, has led to longer PFS and OS [66]. It might be expected that the previously described association between the presence of T790M and a lower PFS and OS would be reversed. However, it has recently been reported that NSCLC patients with plasma EGFRm (including T790M) have shorter PFS and OS, even if treated with osimertinib as a second-line therapy [67]. Similarly, a retrospective analysis of ctDNA samples from the AURA3 trial found that in patients with tissue T790M, the presence of plasma T790M at baseline is associated with a shorter PFS, regardless of whether treatment with platinum-pemetrexed or osimertinib is administered [68]. This might indicate that the presence of T790M is associated with the burden of disease and hence patients who have the mutation detected in plasma derive less benefit, but it might also indicate that the acquisition of EGFR-independent mechanisms is an early event and thus, clones with EGFRm continue to expand. Some proposed EGFR-independent mechanisms include MET amplification, HER2 mutation, and SCLC transformation. Further, off-target resistant mechanisms currently have fewer effective alternatives for treatment.
Some of the potential limitations of our study include the lack of longitudinally collected, and matched, tissue and liquid biopsies, which precludes the interpretation of our results in terms of “mutation clearance”, “negativization”, “mutation reappearance”, or the classification of patients as “shedders” and “non-shedders”, as has been done in other studies. Second, the relatively small number of patients warrants caution when interpreting the results from the association of clinical characteristics in subgroups of patients stratified by progression and plasma EGFRm status. Thus, subgroup analyses must be taken as hypothesis-generating and exploratory information that should be confirmed in future studies.
Conclusions
To the best of our knowledge, this is the first study in Latin America showing that detection of plasma EGFRm, during first- or second-generation EGFR-TKIs, is associated with worse survival outcomes in advanced and metastatic NSCLC patients, regardless of progression status.
This study addressed the question of whether there is any use for liquid biopsies in NSCLC patients that have already started treatment, and for whom longitudinally collected and matched samples are not available. The hesitancy to request a liquid biopsy after treatment initiation can be largely attributed to the belief that a baseline liquid biopsy is essential to interpret the results of subsequent liquid biopsies. Our results, from a real-world clinical setting, show that liquid biopsies can be used, alongside radiological evaluation, to monitor NSCLC patients treated with EGFR-TKIs. Importantly, our results indicate that the prognostic value of plasma EGFRm detection is preserved beyond the first-line treatment and that liquid biopsies can provide useful information even in the absence of serially collected and matched samples. Therefore, a lack of a baseline liquid biopsy should not be a deterrent to obtain one after treatment initiation. The present study adds to existing evidence encouraging health institutions to adopt liquid biopsies in order to obtain useful clinical information that can be used to evaluate the mutational status of patients, to guide treatment selection and to assess prognosis, which could potentially improve patient care.
Authors disclosures
Dr. Oscar Arrieta has received honoraria as an advisor, participated in speakers’ bureau, and given expert opinions to Pfizer, AstraZeneca, Boehringer-Ingelheim, Roche, Lilly, and Bristol-Myers Squibb. Dr. Cardona reports grants from Merck Sharp and Dohme, grants from Boehringer Ingelheim, grants from Roche, grants from Bristol-Myers Squibb, grants from The Foundation for Clinical and Applied Cancer Research – FICMAC, personal fees from Merck Sharp and Dohme, personal fees from Boehringer Ingelheim, personal fees from Roche, personal fees from Bristol-Myers Squibb, personal fees from Pfizer, personal fees from Novartis, personal fees from Celldex Therapeutics, personal fees from Foundation Medicine, personal fees from Eli Lilly, personal fees from Foundation for Clinical and Applied Cancer Research – FICMAC, outside the submitted work; and Other from Pfizer, Boehringer Ingelheim, Astra Zeneca, MSD, BMS, Celldex, Roche. The other authors declare no competing interests.
Author contribution
Conception: Oscar Arrieta, Juan-Manuel Hernán dez-Martínez, Edgar Montes-Servín, Andrés F. Cardona, Camilo Molina-Romero.
Interpretation or analysis of data: Oscar Arrieta, Juan-Manuel Hernández-Martínez, Edgar Montes-Servín, Camilo Molina-Romero, Diego Diaz-García, Dolores L. Mendoza Oliva, Andrés F. Cardona.
Preparation of the manuscript: Oscar Arrieta, Juan-Manuel Hernández-Martínez, Edgar Montes-Servín, Camilo Molina-Romero, Luis Lara-Mejía, David Heredia, Antonio Bahena-González.
Revision for important intellectual content: Oscar Arrieta, Andrés F. Cardona, Luis Lara-Mejía, David Heredia, Juan-Manuel Hernández-Martínez, Edgar Montes-Servín, Camilo Molina-Romero.
Supervision: Oscar Arrieta, Juan-Manuel Hernán dez-Martínez, Edgar Montes-Servín, Camilo Molina-Romero, Dolores L. Mendoza Oliva, Antonio Bahena-González.
Final approval of the manuscript: All authors; Oscar Arrieta, Juan-Manuel Hernández-Martínez, Edgar Montes-Servín, David Heredia, Andrés F. Cardona, Camilo Molina-Romero, Luis Lara-Mejía, Diego Diaz-Garcia1, Antonio Bahena-Gonzalez1 and Dolores L. Mendoza-Oliva.
Supplementary data
The supplementary files are available to download from
Footnotes
Acknowledgments
This work was financially supported by AstraZeneca. However, the sponsor had no role in the design, conduct or oversight of the study. Data collection, management, analysis, and interpretation was carried out independently, as was the preparation, review, and final approval of the manuscript.
