Abstract
BACKGROUND:
FAS can serve as both an oncogene and a suppresser in different malignancies, and the prognostic value of FAS remains controversial.
METHODS:
The Oncomine database, KM-Plotter and bc-GenExMiner platform were adopted to analyze the prognostic value of FAS in breast cancer. Breast cancer tissue microarrays were further used to verify these data. The Cell Miner Tool was used to predict the value of FAS mRNA expression in predicting the efficacies of clinical drugs.
RESULTS:
We found that both FAS mRNA and protein expression level significantly reduced in breast carcinoma. In addition, high FAS expression indicates a better metastatic relapse-free survival. Interestingly, FAS was associated with a better prognosis in different subtypes of breast cancer patients, namely, only in grade II and III, lymph nodal positive or p53 wild-type patients. The data from the Cell Miner Tool revealed that FAS mRNA expression was correlated with the efficacy of the first-line chemotherapeutic taxane agents and target drugs including olaparib and everolimus.
CONCLUSIONS:
FAS expression correlates with a better prognosis in breast cancer and may provide an effective clinical strategy to predict the sensitivity of taxanes and targeted drugs.
Introduction
Breast cancer is the most common and lethal cancer in women worldwide. The estimated new cases caused by breast cancer were up to 252,710 in the USA in 2017, which places it first among all types of cancers [1]. Due to the improvement of therapy, especially the targeted drugs, the survival rate of breast cancer patients has improved significantly in the past 3 decades. However, breast cancer is still the second leading cause of cancer-related death, causing 40,610 deaths in the USA in 2017. Drug resistance is the main cause of cancer-associated deaths.
Breast cancer drug therapy often consists of chemotherapy, targeted therapies and endocrine therapy. Anthracyclines and taxanes are the first-line chemotherapy cytotoxic agents, but only 50% of patients benefit from these drugs because of the development of resistance [2]. In addition, more than 30% of metastatic breast cancer patients do not respond to these drugs, and their overall survival is typically less than 1 year [3]. Moreover, approximately 50% of luminal patients receiving endocrine therapy experience hormonal resistance, and it has also been reported that approximately 60% of HER2
FAS had been reported to reverse the cisplatin resistance of human small cell lung cancer [5]. FAS enhances the sensitivity of human soft tissue sarcoma to anticancer drugs through activation of p38 [6], and FAS is deficient in drug-resistant leukemia cells [7]. These findings suggest that FAS is a potential marker to predict the sensitivity of anticancer agents. FAS was first identified as a death receptor on the surface of cells. It can interact with FAS ligand (FasL), recruit the downstream caspases via the death inducing signaling complex (DISC) and induce apoptotic cell death. Thus, FAS had been recognized as a tumor suppressor gene in cancer cells since it can interact with FasL of immune cells and induce apoptosis of cancer cells. However, FAS also serves as an oncogene that enhances tumor growth and invasion by non-apoptotic signaling pathways, including NF-kB, MAPK and Src [8]. It has been reported that FAS protein expression is negatively associated with lymph nodes metastasis and the number of recurrences [9]. The loss of FAS expression in primary lesions means an increased risk of bone metastasis [10]. In addition, low expression of FAS in tumors is associated with longer overall survival of patients with human glioblastoma [11]. Therefore, the prognostic role of FAS in malignancies remains controversial.
In this retrospective analysis, we compared the expression of FAS mRNA between breast cancer and normal tissue using the Oncomine database. Through KM-Plotter and the bc-GenExMiner platforms, we analyzed the relationship of FAS mRNA expression and the prognosis of breast cancer. Breast cancer tissue microarrays were used to verify the prognostic value of FAS in breast cancer. In addition, we investigated the value of FAS mRNA expression in predicting the efficacies of chemotherapeutic drugs, targeted agents and endocrine drugs in breast cancer by applying the Cell Miner Tool.
Materials and methods
Oncomine analysis
We compared the FAS mRNA expression both in breast cancer and normal tissue in Oncomine 4.5. The fold-change threshold was set as 1.5 and the gene rank of the top was set as 10%. The significant difference of expression level was analyzed by Student’s
Breast cancer gene-expression miner
The prognostic value of FAS mRNA in all breast cancer patients was analyzed by the breast cancer gene-expression miner (bc-GenExMiner v4.1). The cases were divided into two groups based on the median expression level of FAS mRNA. The metastatic relapse (MR) free survival and any events free (AE) survival were analyzed. The pooled HR with 95% CIs and log-rank
Kaplan-Meier-plotter analysis
The prognostic value of FAS mRNA in different subtypes of breast cancer and the clinicopathological parameters were evaluated by KM-Plotter. Affymetrix probe ID: 204781_s_at was selected for further analysis. The patients were divided into two groups based on the median expression of FAS mRNA, and the overall survival (OS) and relapse-free survival (RFS) were analyzed. The significant differences of OS and RFS were estimated by the Kaplan-Meier method. The hazard ratios (HRs) and 95% intervals between two groups were calculated by Cox regression analysis and the
The FAS mRNA expression in breast cancer and normal tissue. (A) Fas mRNA expression in normal breast tissue and invasive ductal carcinoma (IDC) from the TCGA database, (B) Fas mRNA expression in normal breast tissue and IDC and breast invasive lobular carcinoma from the TCGA database, (C) Fas mRNA expression in normal breast tissue and IDC from GSE3971, (D) Fas mRNA expression in normal breast tissue and invasive breast cancer (IBC) from GSE22358, (E) Fas mRNA expression in normal breast tissue and breast cancer from human fresh tissue, (F) Fas protein expression in normal breast tissue and breast cancer from tissue microarrays.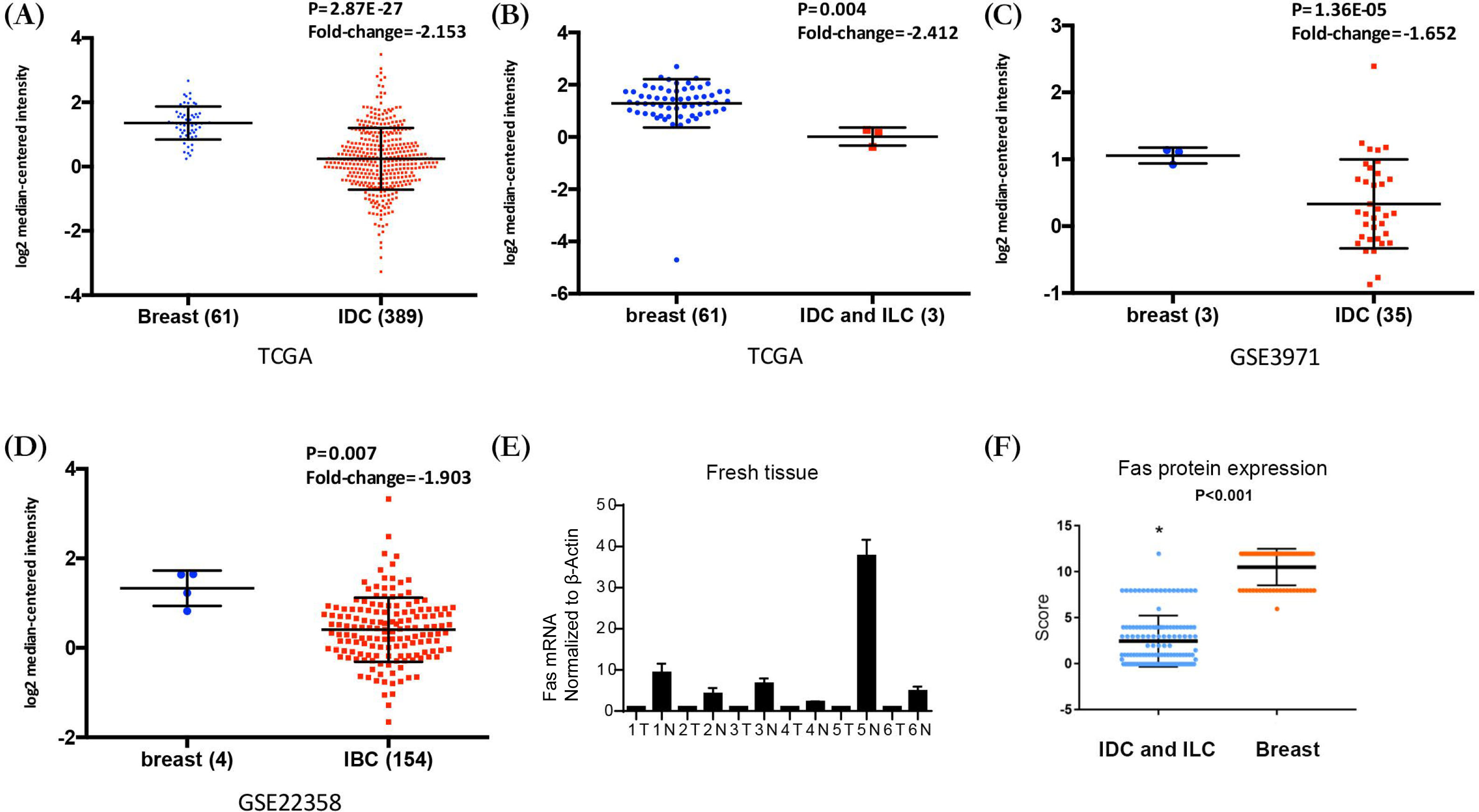
The data of FAS mRNA expression and sensitivity to compounds were converted to Z-scores. The association between FAS and the efficacy of different compounds were tested by linear regression analysis. The correlation coefficients (R squares) and
Tissue microarrays and immunohistochemistry
Human breast tumor (HBreD140Su07) and normal tissue arrays (HBreD077Su01) were obtained from Shanghai Outdo Biotech. The EnVision™ FLEX detection system (Dako) was used following the manufacturer’s instructions. FAS antibody (Abcam, ab133619) was added at a dilution of 1:300 and incubated at 4
Ethical statement
This retrospective study analyzed the FAS mRNA expression, clinical parameters and drug resistance from publicly available datasets. All methods were carried out based on relevant guidelines and approved by the ethics committee of the Second Affiliated Hospital of Zhejiang University.
The pooled analysis of the correlation between FAS mRNA expression and OS of patients with breast cancer.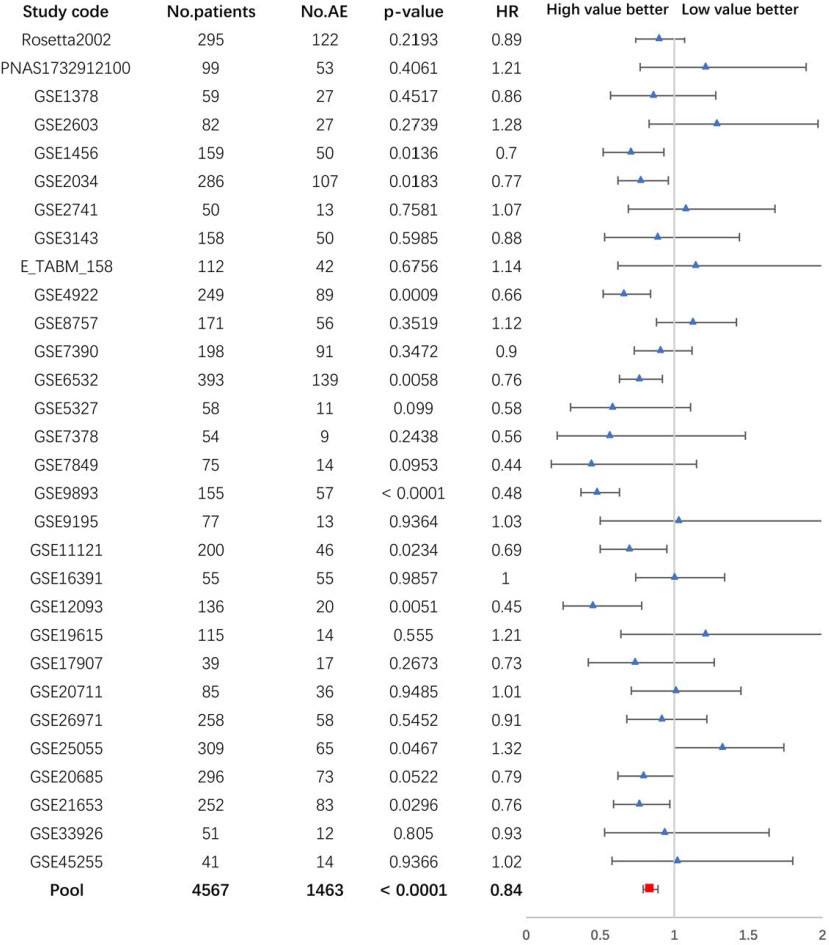
Correlation of FAS expression with relapse-free survival in breast cancer. (A) Pooled analysis of the relationship between Fas mRNA expression and relapse-free survival of breast cancer from the online database; (B) the relationship between Fas protein expression and relapse-free survival of breast cancer from tissue microarrays.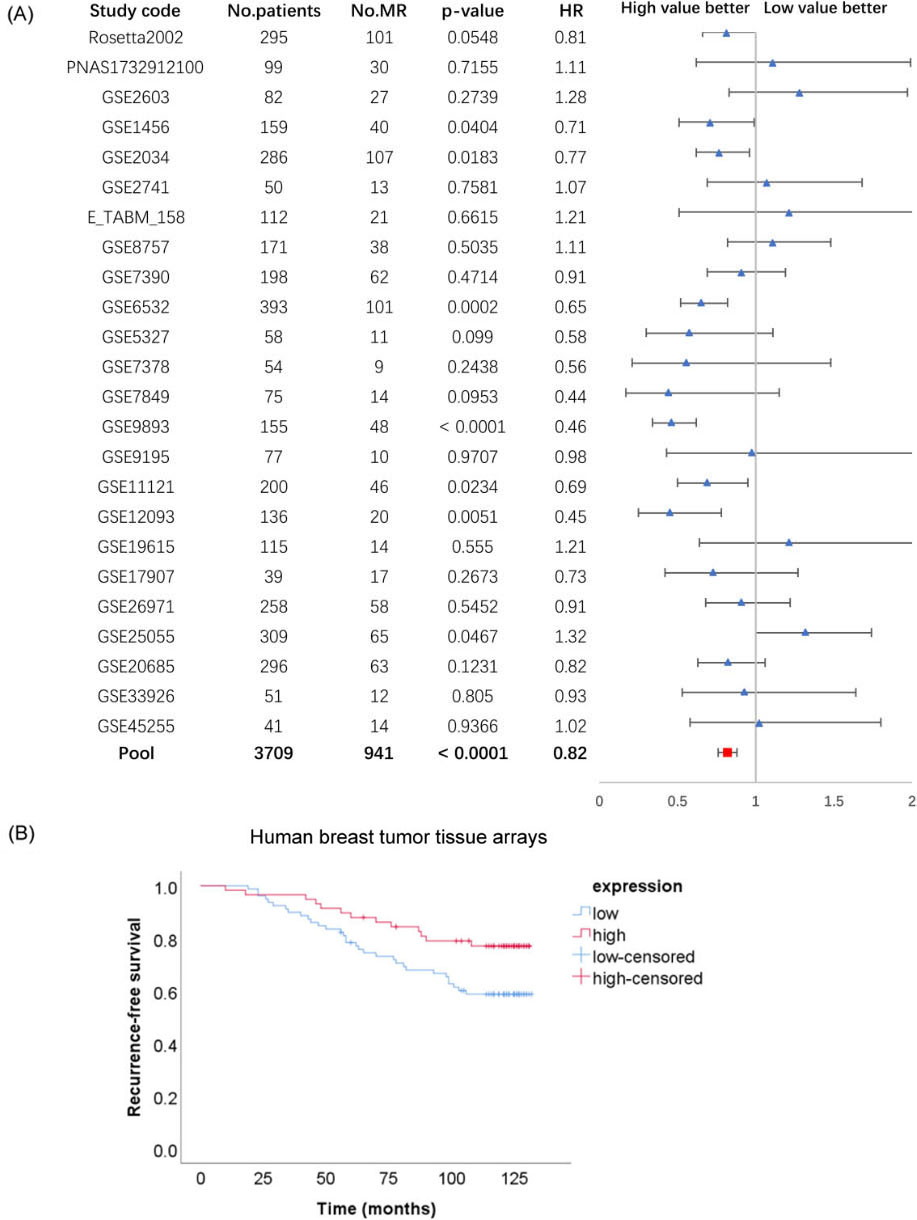
Evaluation of the correlation between FAS mRNA expression and the overall survival of different types of breast cancer patients. Survival curves by FAS mRNA expression were plotted for luminal A patients (A), luminal B patients (B), basal-like patients (C), and Her-2 amplified patients (D).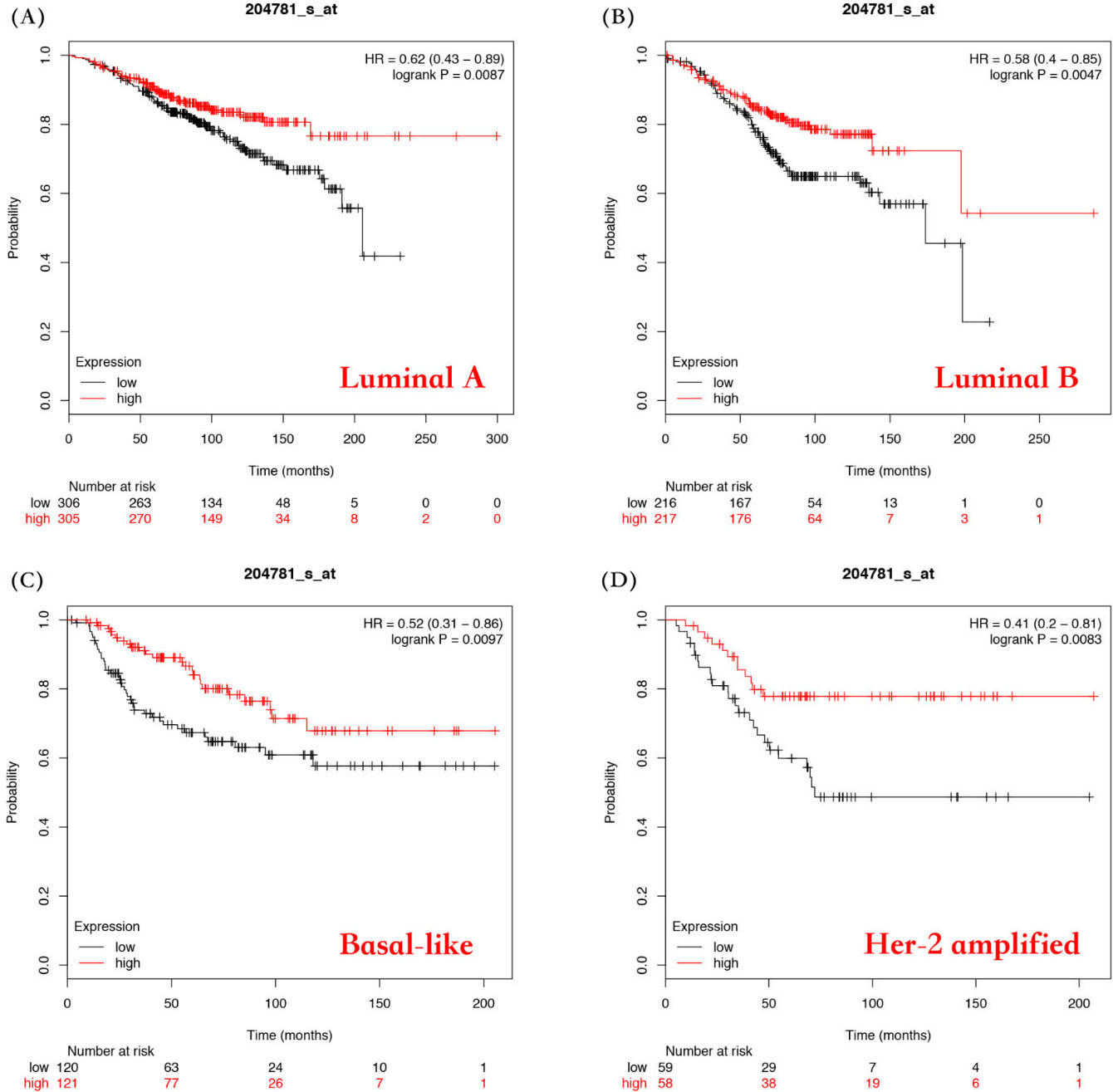
Evaluation of the correlation between FAS mRNA expression and relapse-free survival of different types of breast cancer patients. Survival curves by FAS mRNA expression were plotted for luminal A patients (A), luminal B patients (B), basal-like patients (C), and Her-2 amplified patients (D).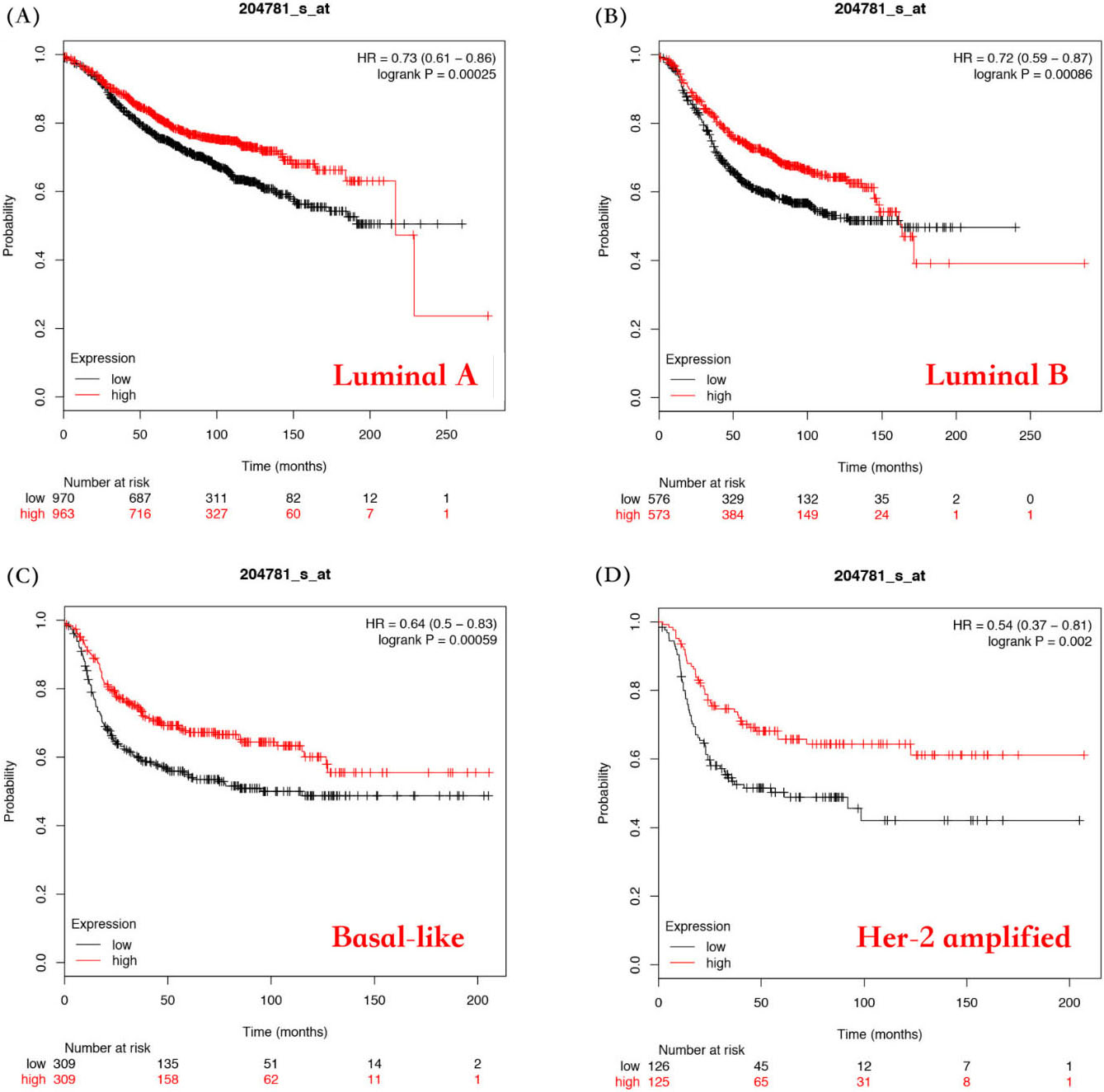
The FAS expression in breast cancer is lower than that in normal tissue
We searched the Oncomine database and downloaded the data from different datasets. We compared the expression of FAS mRNA between breast cancer and normal breast tissue (Fig. 1). The data from TCGA indicated that the FAS mRNA expression level was significantly lower in breast invasive ductal carcinoma (IDC) (Fold-change
High FAS expression indicates a better prognosis in breast cancer patients
The prognostic value of FAS mRNA expression was analyzed with all breast cancer patients by bc-GenExMiner. The results showed that overexpression of FAS mRNA was correlated with a better OS (HR
The prognostic value of FAS mRNA expression in different subtypes of breast cancer patients
Next, we investigated the prognostic value of FAS in different subtypes of breast cancer including luminal A, luminal B, basal and Her-2 amplified subtype. As shown in Fig. 4 and Table S1, the expression of FAS mRNA was significantly related to a better OS in luminal A (HR
The prognostic value of FAS mRNA expression in breast cancer patients with different clinico-pathological features
We also performed subgroup analysis based on different clinicopathological features such as grade, nodal status, and p53 status (Table S1). First, we divided the patients into 3 groups based on the grade from I to III. The results showed that high FAS mRNA expression was significantly correlated with a better OS in grade II patients (HR
Evaluation of the correlation between FAS mRNA expression and prognostic value of different grades of breast cancer patients. Overall survival curves by FAS mRNA expression were plotted for grade 1 patients (A), grade 2 patients (B), and grade 3 patients (C). Relapse-free survival curves of FAS mRNA expression were plotted for grade 1 patients (D), grade 2 patients (E), and grade 3 patients (F).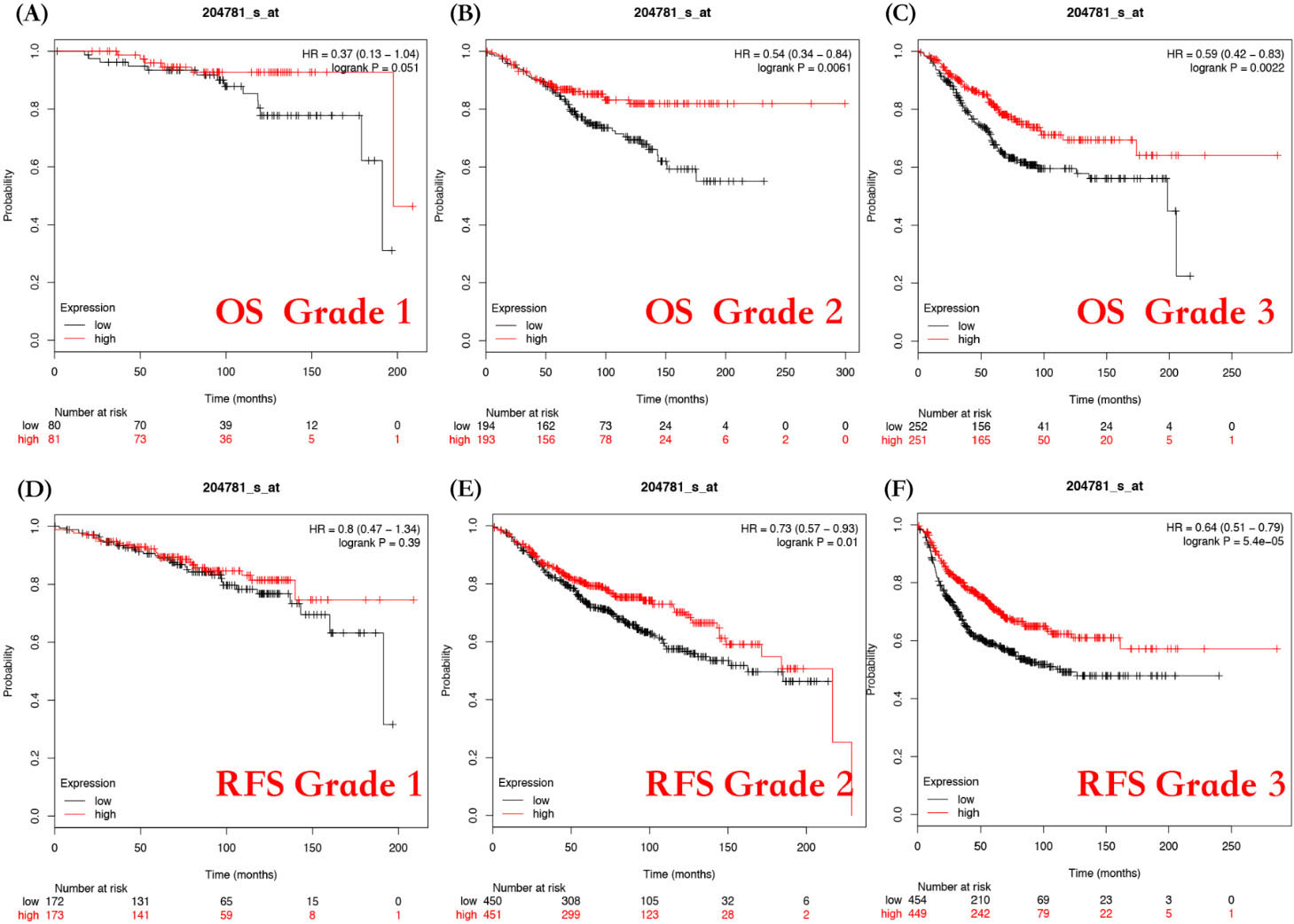
Second, we tested the association of FAS mRNA expression with the lymph node status. As is shown in Figure, high FAS mRNA expression was significantly correlated with better OS and RFS in lymph node positive patients (HR
Evaluation of the correlation between FAS mRNA expression and the prognostic value of lymph node status of breast cancer patients. Overall survival curves by FAS mRNA expression were plotted for lymph node positive patients (A) and lymph node negative patients (B). Relapse-free survival curves by FAS mRNA expression were plotted for lymph node positive patients (C) and lymph node negative patients (D).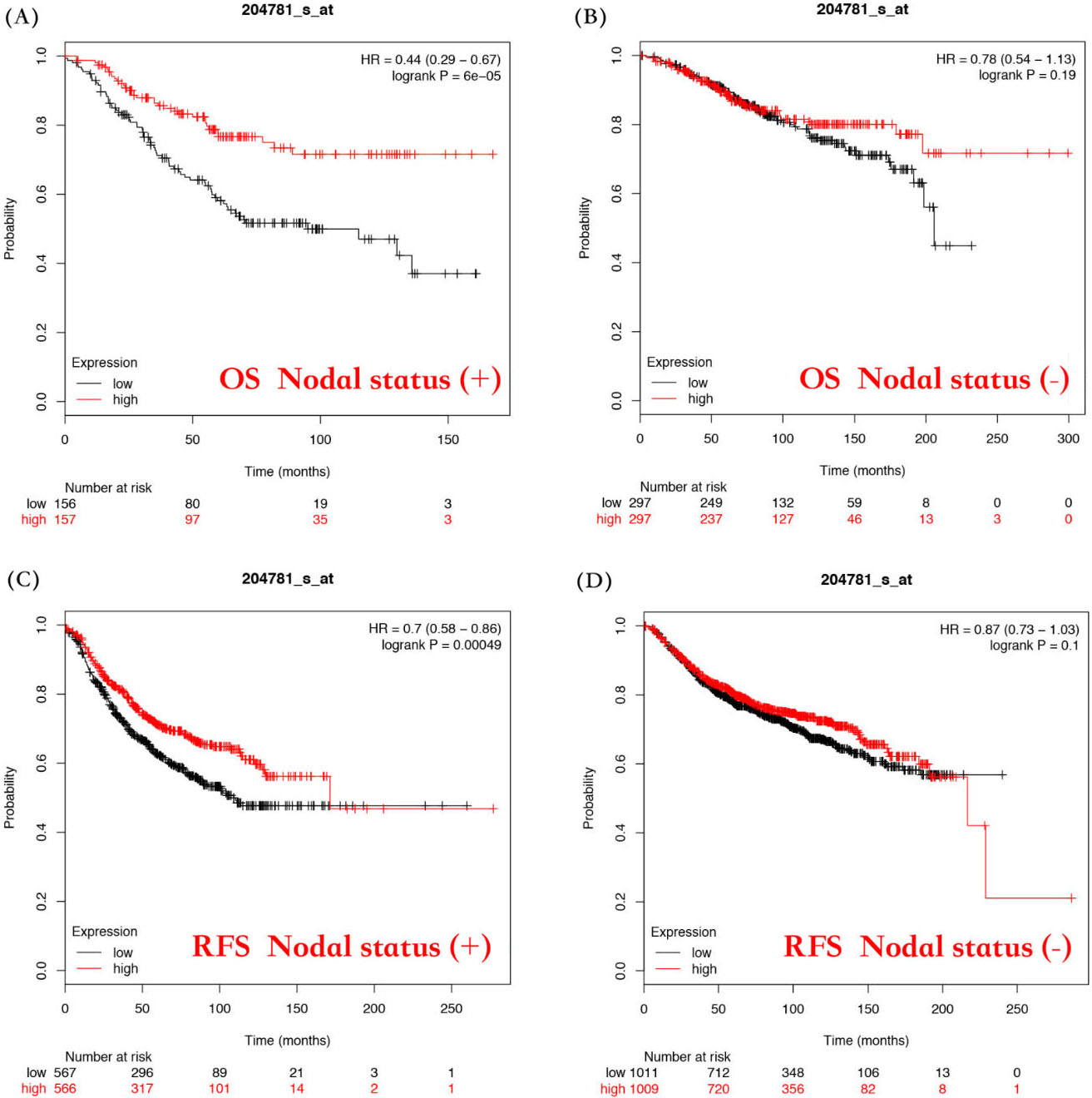
Third, we performed an analysis of FAS mRNA expression with OS based on the p53 status. The results showed that longer OS and RFS were correlated with p53 wild-type patients (OS: HR
Evaluation of the correlation between FAS mRNA expression and the prognostic value of P53 genotype of breast cancer patients. Overall survival curves by FAS mRNA expression were plotted for P53 wild type patients (A) and P53 mutant patients (B). Relapse-free survival curves by FAS mRNA expression were plotted for P53 wild type patients (C) and P53 mutant patients (D).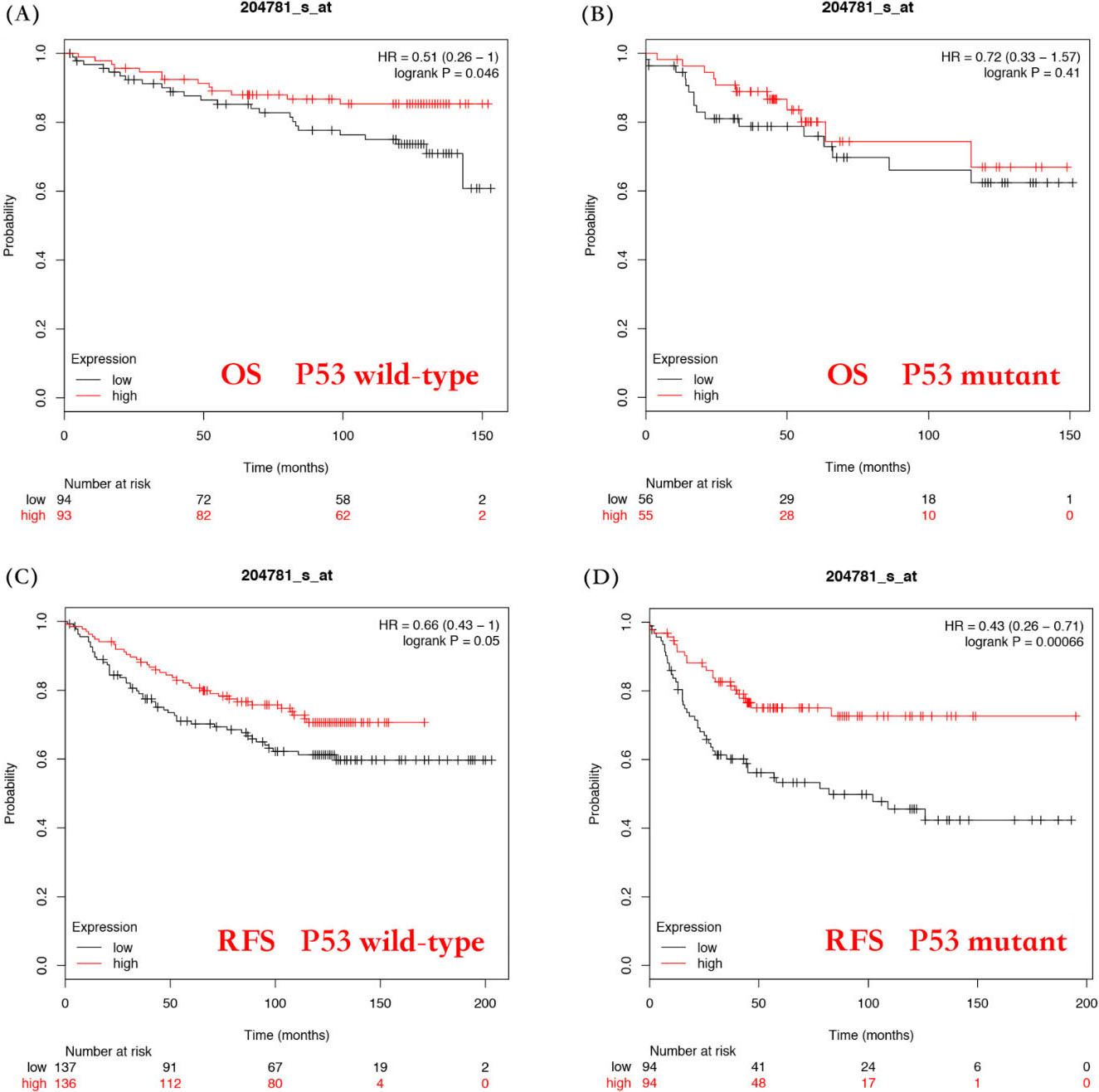
To determine the predictive value of FAS mRNA expression for the efficacy of therapeutic agents including chemotherapeutic and target agents, we analyzed the data from the Cell Miner Tool database. The results showed that FAS mRNA expression was negatively associated with the efficacies of the chemotherapeutic drugs paclitaxel (
Evaluation of the association between FAS mRNA expression and the sensitivity of different cell lines to clinical drugs. The regression figures of docetaxel (A), paclitaxel (B), everolimus (C), and olaparib (D). The data were transformed to Z-Scores and were downloaded from Cell Miner Tools. 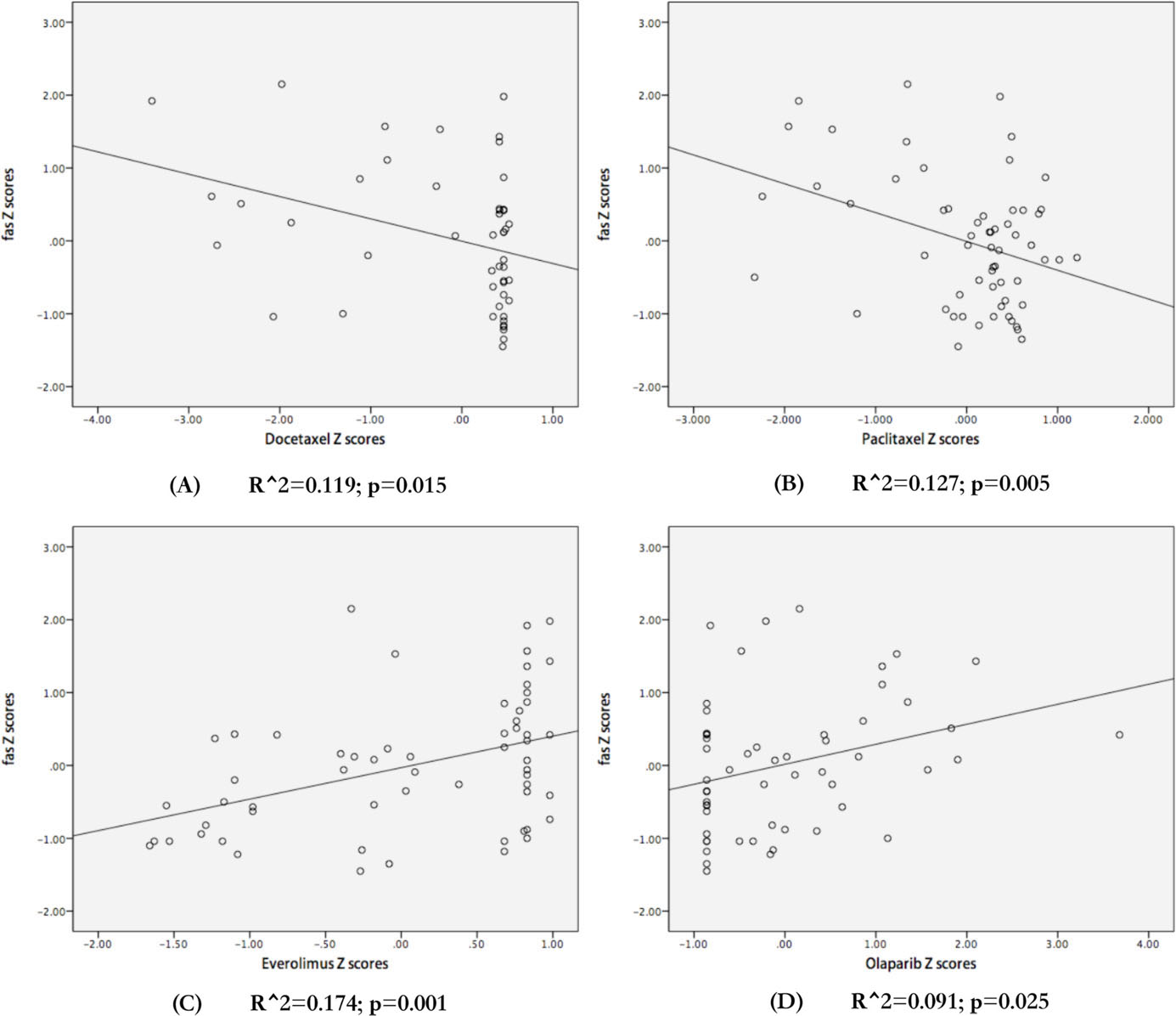
Furthermore, we tested the efficacies of the targeted agents including lapatinib, olaparib and everolimus. As shown in Fig. 9, FAS mRNA expression was positively associated with the efficacies of olaparib (
We also studied the correlation between FAS mRNA expression and the efficacy of four endocrine drugs, including tamoxifen, fulvestrant, megestrol acetate and fluoxymesterone. The results showed that FAS mRNA expression was not related to the sensitivity to tamoxifen (
FAS can serve as both oncogene and suppresser in different types of tumors [12, 13, 14, 15, 16, 17, 18]. Therefore, there is controversy about the relationship between FAS mRNA expression and the prognosis of breast cancer. In this retrospective analysis, we performed a meta-analysis of online databases and included 4,567 patients with breast cancer. The results showed that FAS mRNA expression in breast cancer is significantly lower than in normal breast tissue. Then, we compared the association of FAS mRNA expression with prognosis and found that high FAS mRNA expression is significantly associated with a better prognosis in breast cancer. When we performed subgroup analysis based on the subtype of breast cancer, we found that high FAS mRNA expression was significantly associated with a better prognosis in all subtypes of breast cancer, including luminal A, luminal B, basal and Her-2 amplified subtype.
Interestingly, we also found that high FAS mRNA expression was correlated with a better OS and RFS in Grade II or III patients, lymph node positive patients and p53 wild-type cases when we performed subgroup analysis based on grade, nodal status, p53 status and therapy history. It has been reported that the FAS can be activated by p53 in tumor cells during chemotherapy treatment [19]. In p53 wild type tumors, p53 can interact with the promoter region of the FAS gene and activate transcription, while FAS mRNA transcription cannot be activated in mutant p53 tumors [20]. This may explain why high FAS mRNA expression is associated with a better prognosis in the p53 wild-type patients, but not the p53 mutant cases. In line with our results, previous studies had reported that high FAS expression was associated with less node metastasis, a smaller tumor size, less recurrence and longer disease-free survival [9, 21]. However, the mechanisms still need to be studied further.
In addition, we investigated the value of FAS expression in the efficacy of clinical agents used in breast cancer, and found that FAS may have potential to predict the efficacy of chemotherapeutic agents (paclitaxel and docetaxel) and targeted agents (everolimus and olaparib). Currently, anthracyclines (doxorubicin and epirubicin) and taxanes (paclitaxel and docetaxel) are widely used in adjuvant or neoadjuvant chemotherapy of breast cancer [22, 23]. However, only 50% of patients benefit because of resistance to these cytotoxic drugs. Chemo-resistant cancer cells can survive, develop high metastasis potential and promote disease progression [3, 24, 25]. Cancer cells can be intrinsically chemo-resistant or acquire resistance during treatment [26, 27].
In this retrospective analysis, we found that high FAS mRNA expression was negatively associated with the efficacies of chemotherapeutic taxanes (paclitaxel (
Trastuzumab, a humanized mAb and the first anti-HER2 targeted agent for breast cancer treatment, has been reported to enhance OS and RFS of patients with breast cancer [28]. Lapatinib is a dual inhibitor of the tyrosine kinase activity of HER2 and EGFR [29]. It has been reported that lapatinib combined with trastuzumab can be used to treat advanced or metastatic HER2 positive breast cancer [30]. Everolimus is an oral mechanistic target of rapamycin (mTOR) inhibitor. Olaparib is an oral poly-adenosine diphosphate (ADP)-ribose-polymerase inhibitor (PARPi). In this retrospective analysis, we found that FAS mRNA expression was positively associated with the efficacies of olaparib (
Endocrine therapy is an important treatment for hormone receptor-positive breast cancer patients. It reduces the risk of breast cancer recurrences and mortality. In this retrospective analysis, we also studied the correlation between FAS mRNA expression and the efficacy of four endocrine drugs, including tamoxifen, fulvestrant, megestrol acetate and fluoxymesterone. The results showed that FAS mRNA expression was not related to the sensitivity to tamoxifen (
In conclusion, higher expression of FAS is correlated with a better prognosis of patients with breast cancer. Furthermore, FAS mRNA expression can serve as a potential sensitivity marker for taxanes (paclitaxel and docetaxel) and targeted regents (olaparib and everolimus) for breast cancer. However, certain limitations exist in this study. First, the sample size is relative small. We performed immunohistochemistry assays to evaluate the expression and prognosis of FAS protein in 138 human samples with breast cancer in tissue microarrays (HBreD140Su07). Second, we found that FAS mRNA expression can serve as a potential biomarker for taxanes, olaparib and everolimus, but the mechanisms involved in this process are still unknown. Our data provides a starting point. Deeper studies are warranted to rigorously investigate the mechanisms involved in the drug sensitivity. We expect that more preclinical and clinical studies will be designed to study the prognostic value of Fas in the future studies.
Authors’ Contributions
Conception: Zhigang Chen and Jun Wu
Interpretation or analysis of data: Yi Zhang and Xuan Shao
Preparation of the manuscript: Yi Zhang and Jun Wu
Revision for important intellectual content: Danying Xu, Chenyi Gao and Xuan Zhu
Supervision: Zhigang Chen
Competing financial interests
The authors declared that they have no conflicts of interest to this work.
Supplementary data
The supplementary files are available to download from
Footnotes
Acknowledgments
This study is supported by the National Natural Science Foundation of China (NO. 81502564, 8170 3498, 81802750, 81802633, 81972598) and the Natural Science Foundation of Zhejiang province (NO. LY16H160018). This work is also supported in part by the scholarship from China Scholarship Council (CSC) under the Grant CSC NO. 201706325003.
