Abstract
BACKGROUND:
Stemness factors associated with tumorigenesis in different types of cancers have not been specifically studied in oral tongue SCC (OTSSC). Here, we aimed to quantify expression levels and distribution of KLF4 and SOX2, two relevant stemness factors, in oral SCC including OTSCC samples from different subsites.
METHODS AND RESULTS:
We determined KLF4 and SOX2 expression levels by immunostaining 35 biopsies of OSCC. Stained wholeslide images were digitized and subjected to automatic cell detection and unbiased quantification using Qupath software. We found statistically significant reduction in KLF4 positive cells density (
CONCLUSIONS:
KLF4 and SOX2 exhibit opposite expression profiles regarding subsite localization and differentiation level in OSCC. Our study prompts future OTSCC prospective studies looking for clinical prognosis to incorporate detailed subsite information in the analysis.
Keywords
Introduction
Head and neck cancers (HNC), including cancers of the oral cavity, are highly prevalent worldwide, characterized by high morbidity and limited therapeutic options [1, 2]. Oral cancers show variations in aetiology, incidence rates, clinical progression and prognosis based on the anatomical subsite affected [3, 4, 5]. The most common histologic type is squamous cell carcinoma (SCC) comprising over 90% of oral cancers [6, 7]. In this regard, oral (mobile) tongue squamous cell carcinoma (OTSCC) is one of the most common malignant tumors of the oral cavity [8, 9]. Moreover, current epidemiological records show that oral tongue cancers are becoming more common amongst the younger patients than in previous decades, although the exact reason of this trend is yet to be elucidated [8, 9]. OTSCC is generally associated with poor prognosis [9, 10] and behaves aggressively with higher rates of occult metastasis and neck nodal metastasis than any other cancer of the oral cavity [10, 11]. Patients with OTSCC have a significantly worse prognosis than those with similar lesions of the oropharynx, larynx, hypopharynx and others oral cavity sites [11]. These clinical and epidemiological information highlight the importance of developing more effective diagnostic modalities associated with different oral cancer subsites that allow for targeted therapeutic approaches.
In the last years, new research on molecular mechanism underlying HNC development has revolutionized the treatment for patients with metastatic disease. In particular, immune modulation with checkpoint inhibitors has been established and applied to HNC patients [12, 13]. However, the overall survival of treated patients after development of metastatic disease has barely improved [14], in part due to treatment resistance. The latter is thought to be related to intratumoral heterogeneity as well as the presence of specific population of resistant cancer stem cells (CSC) [15, 16]. In this regard, previous studies have shown that the expression of the Yamanaka factors (Oct3/4, Sox2, Klf4, c-Myc) is sufficient to reprogram differentiated cells to pluripotent stem cells [17]. One of these factors, the transcription factor Krppel-like factor 4 (KLF4), in addition to its role in cell stemness, suppresses cell proliferation, promotes differentiation, and thus helps to maintain homeostasis of adult epithelial cells [18, 19]. Furthermore, KLF4 has a tumor suppressor function in many human malignancies including gastric, colorectal and lung cancers [19, 20]. Nevertheless KLF4 was also reported to have an oncogenic role in breast and other cancers in a context dependent-manner [18, 20]. However, the role of KLF4 in head and neck cancer remains controversial. Increased KLF4 expression has been reported in head and neck SCC by Shyh-Kuan Tai et al. [21]. Conversely, KLF4 expression was shown decreased in oral SCC (OSCC) [22]. Furthermore, sex-determining region Y-box 2 (SOX2) is other Yamanaka factor that has been suggested as a marker of CSC [23, 24]. SOX2 is a transcription factor expressed in a wide range of stem and progenitor cells in diverse adult tissues, including squamous epithelia of the tongue and genetically amplified in different human cancers [24, 25]. However, to date, the expression pattern of KLF4 and SOX2 has not been specifically studied in OTSSC. We hypostatize that OTSCC may show a differential expression of these Yamanaka factors connected with tumor localization and histologic grade. In the present study we quantified KLF4 and SOX2 expression levels by immunohistochemical staining in OSCC biopsies including OTSCC samples from different oral tongue subsites.
Materials and methods
Thirty five excisional specimens with histological diagnosis of squamous cell carcinoma were identified retrospectively at the Surgical Pathology Laboratory, Oral Pathology Department of the School of Dentistry of the University of Buenos Aires, Argentina. Clinicopathological information, including age, gender and location was obtained from patient records (Table 1). Tumors were graded using World Health Organization criteria [26]. The protocol for this study was approved by the Ethics Committee of the School of Dentistry of the University of Buenos Aires.
Clinicopathologic characteristic for 35 selected cases of OSCC
Clinicopathologic characteristic for 35 selected cases of OSCC
B, lateral border of the tongue; F, female; M, male; WD, well differentiated; MD, moderately differentiated; PD, poorly differentiated.
Before immunohistochemical studies, H&E-stained section from each specimen was evaluated and the suitability of inclusion for the study was determined. Five micrometer serial sections were used for all subsequent analysis. The tissues slides were dewaxed and hydrated through graded alcohols and incubated in 3% hydrogen peroxide in PBS for 30 min to quench the endogenous peroxidase. Antigen retrieval was done using 10 mmol/L citric acid in a microwave and the following steps were performed as described previously [27]. Primary antibodies used were: KLF4 (H180) at dilution 1:400 and Sox-2 (E4) at dilution 1:50 from Santa Cruz Biotechnology. The slides were washed and developed in 3, 3-diaminobenzidine (DAB) (FASTDAB tablet; Sigma) under microscopic control and after counterstained with Mayer’s hematoxylin.
Qualitative staining evaluation
Initially, tissue sections immunohistochemically stained for KLF4 and SOX2 were independently classified by two pathologists blinded to the clinicopathological information. A case was marked negative when both observers considered that nuclear DAB labeling was absent or only present within a few limited tumor cells (less than
Image acquisition and analysis
Stained histological slides were scanned and digitized for unbiased cell detection and automatic quantification using Qupath software (
Automatic cell detection with mean DAB nuclear intensity measurement in ROIs using Qupath software. A. Low magnification whole slide composite image depicting the ROIs drawn by a trained pathologist in a tongue case immunostained for KLF4. B. Same image with the superimposed color-coded cell classification obtained after Qupath analysis (red positive, blue negative). C. Digital zoom within a ROI of the whole slide composite image depicted in A showing cells with different degrees of nuclear DAB staining intensity. D. Frequency histogram of DAB nuclear intensity values obtained for all detected cells in the ROIs of the case presented in A. The arrow indicates positive cell threshold in this case. Cells with DAB nuclear signal above threshold are considered positive (black). Inset: schematic representation of threshold value determination for case A, frequency histogram of DAB nuclear signal of case A lymphocytes. Red bar indicates mean 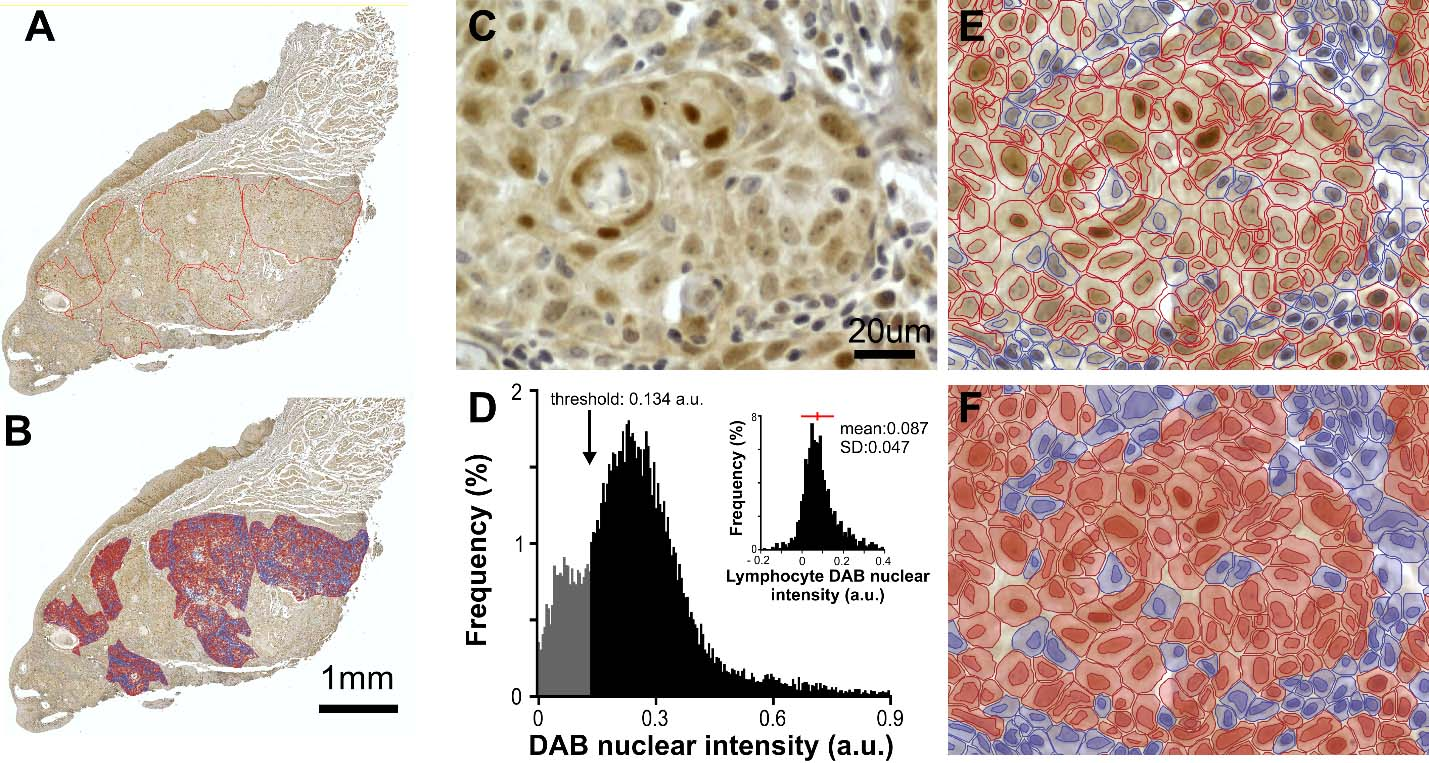
Stain separation (Hematoxylin/DAB) was achieved with Qupath color deconvolution tool using each image RGB modal values for automatic classification. Next, automatic cell detection was achieved using optical density sum. Detection parameters initially optimized by visual inspection and kept constant across images with few exceptions. Qupath output parameters were collected for all individual cells included in the ROIs. No filtering or exclusion criteria were adopted (Fig. 1B).
To establish an observer-independent threshold for positive cell detection (allowing automatic cell classification as positive/negative) we quantified DAB nuclear staining in a subset of 50–300 lymphocytes per slide. Nuclear lymphocytes DAB intensity followed a normal/quasi-normal distribution (Fig. 1D) reflecting a homogenous population (Fig. 1D). Threshold was independently set for each case as the mean nuclear DAB signal
Positive cells fraction was defined as the number of positive tumor cells divided by total number of detected cells, positive cells density was calculated as the number of positive tumor cells divided by ROIs combined area (positive cells/mm
The Cancer Genome Atlas (TCGA) data
TCGA mRNA expression analysis was conducted using a publicly available dataset. Patient data from TCGA (Head and Neck Squamous Cell Carcinoma. Source data from GDAC Firehose. Previously known as TCGA Provisional), including site location, were retrieved using cBioportal web site (
Statistical analysis
Data are presented as mean
Assessment of KLF4 immunohistochemical expression in oral squamous cell carcinomas. A. Normal representative oral epithelium with positive cells with KLF4 expression in the intermediate layer and strongest staining at upper layers. B. Representative image of SCC developed in the cheek with clear nuclear staining for KLF4 throughout the malignant epithelial cells. C. A representative carcinoma from gingiva immunostained for KLF4 detection displays positive cells with a wide range of intensity labeling. D. OTSCC developed in the dorsal tongue shows KLF4 positive staining with variable intensity. E. OTSCC from the lateral border of the tongue positive for KLF4 staining. F, G. Detail at higher magnification of the boxed areas in E. Images F and G depict two different areas with positive KLF4 expression exemplifying regional differences in fraction and density of KLF4 positive cells.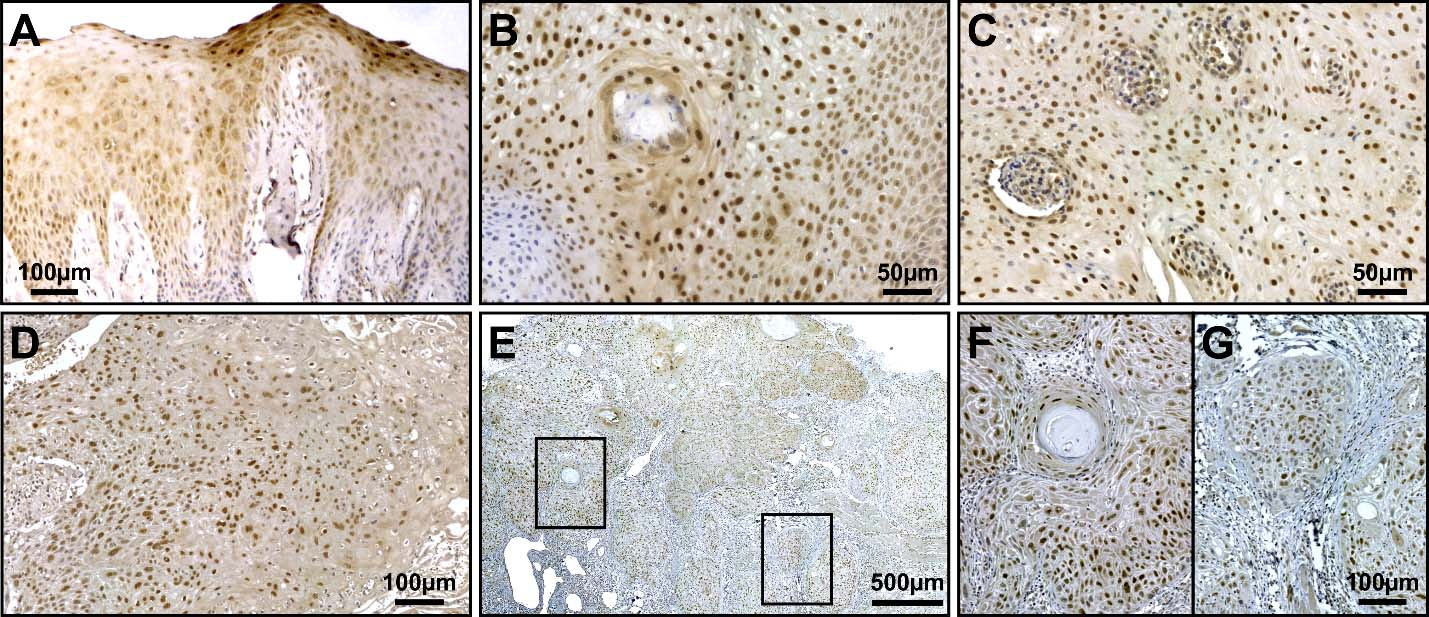
Quantitative analysis of KLF4 expression in OTSCC versus OSCC. A. Fraction, density and nuclear labeling intensity of KLF4 positive cells for OTSCC (T) and the rest of the SCC developed in the oral cavity (NT, non-tongue). Bars represent mean 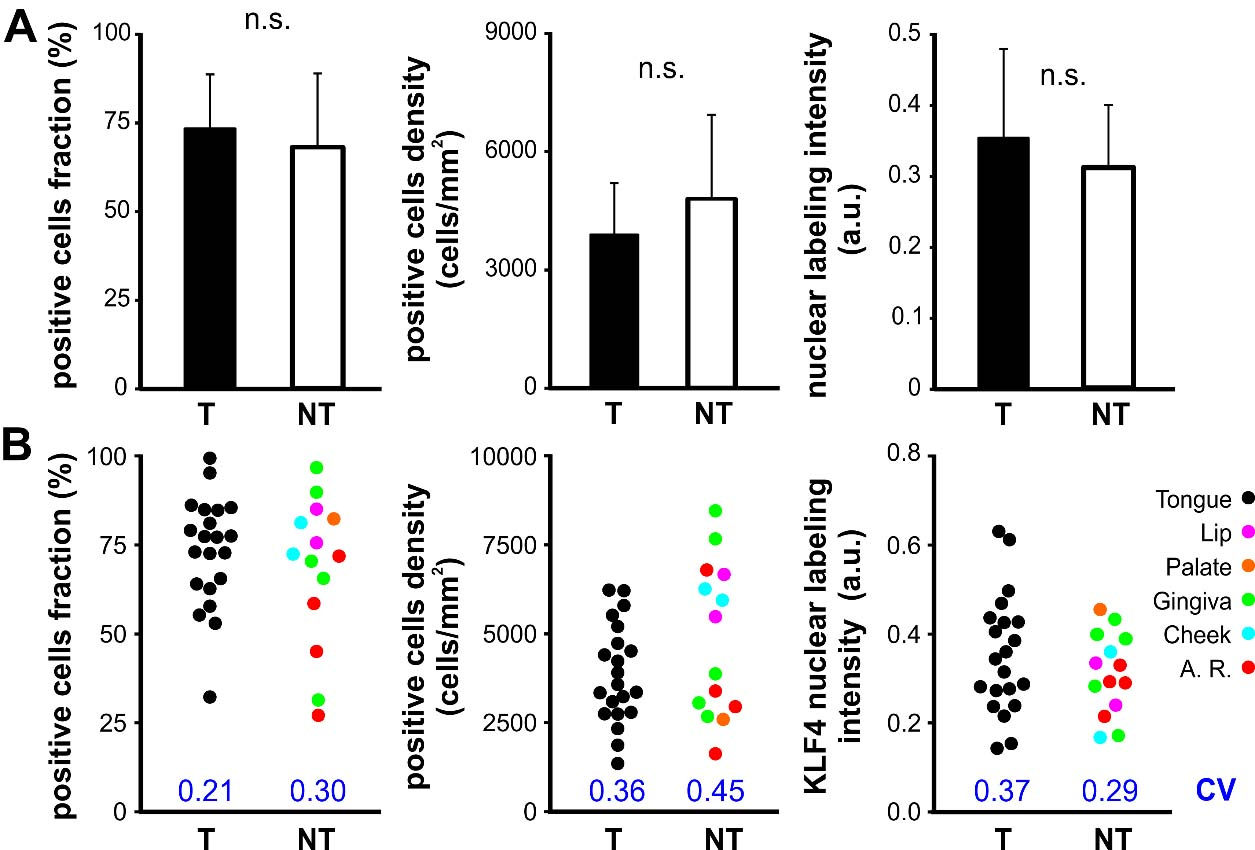
Quantitative analysis of KLF4 expression in OTSCC developed in different tongue subsites. Fraction (A), density (B) and intensity (C) of KLF4 positive cells for OTSCC from the borders of the tongue (tongue border) and the rest of the oral tongue subsites (other tongue site). Bars represent mean 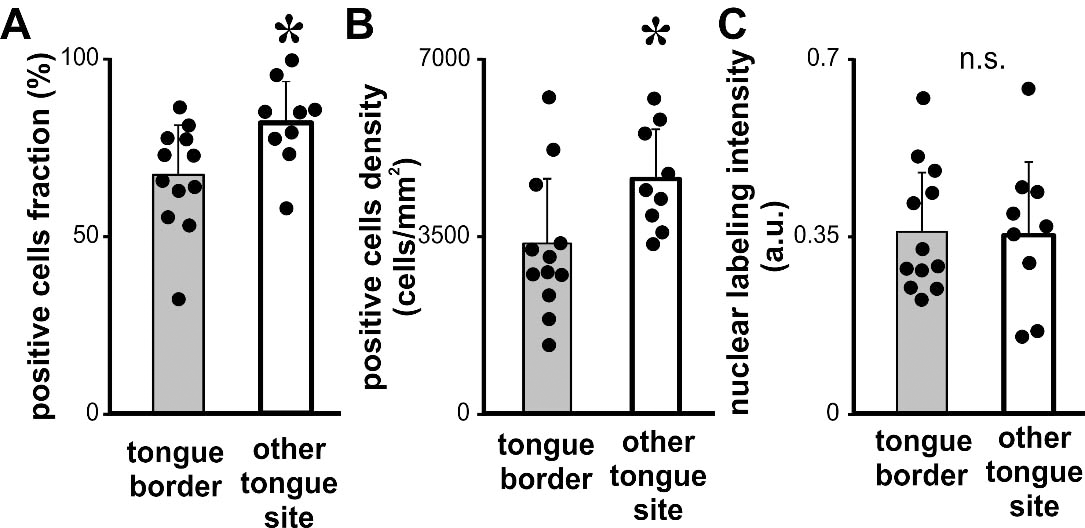
Histologic degree of OSCC and KLF4 expression. Fraction (A), density (B) and intensity (C) of KLF4 positive cells for OSCC grouped according to histologic degree WD, well differentiated; MD, moderately differentiated; PD, poorly differentiated. Bars represent mean 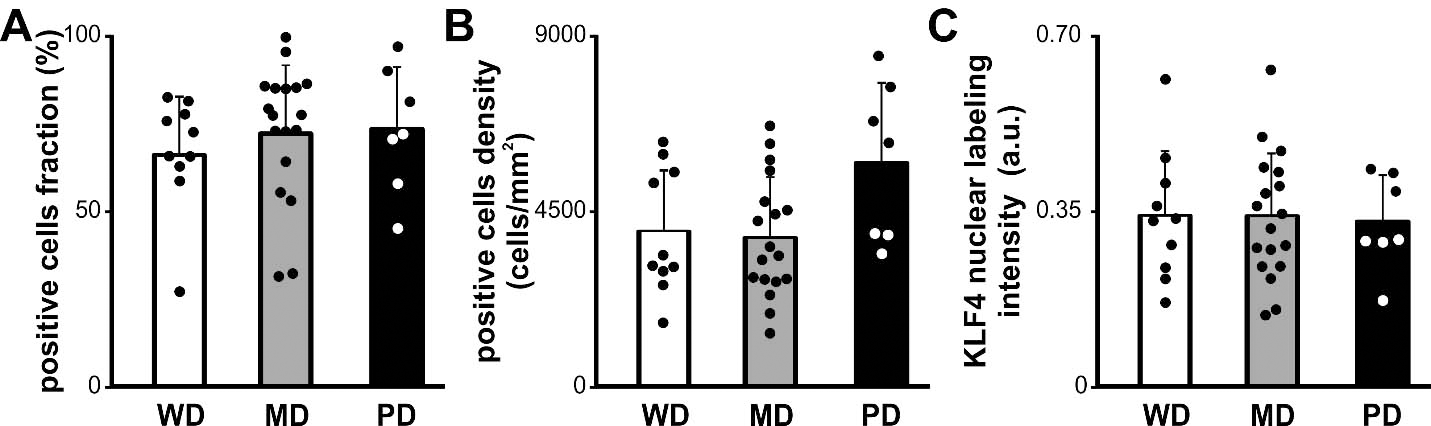
With the aim to characterize KLF4 protein expression in tongue carcinomas and its relation to oral cancer we performed immunohistochemistry in OSCC biopsies from different oral locations (Table 1). Nuclear staining for KLF4 in the intermediate and upper layers of the normal oral mucosa epithelium was observed (Fig. 2A). KLF4 protein was detected at variable levels and clearly stained in the cell nucleus throughout the epithelium in 94.3% (33 out of 35) of the OSCC analyzed by qualitative evaluation (Fig. 2). However the pattern and intensity of expression for KLF4 in OSCCs was very diverse. We described positive cases displaying regions with near full cell labelling intermingled with areas almost lacking of KLF4 staining. Therefore we performed whole slide scanning in all stained biopsies in order to do quantitative analysis on digital images using Qupath software. On average, a ROI area equivalent to 4.72 mm
We analyzed KLF4 immunostaining in OTSCC developed in the borders or in the rest of the tongue (Fig. 4). We found statistically significant differences in KLF4 positive cells fraction in OTSCC from the borders of the tongue when compared with OTSCC from the rest of the tongue (Fig. 4A). The same results were found when we calculated the density of KLF4 positive cells (Fig. 4B). Although we did not observed changes in the KLF4 nuclear labeling intensity (Fig. 4C). The lack of subsite categorization in the available TCGA databases precluded the comparative assessment of KLF4 mRNA levels whiting the tongue. Considering that KLF4 has been reported as a differentiation marker for skin and oral epithelia as well as OSCC [31, 32] we analyzed a possible relation between the histologic degree and KLF4 expression in our group of cases. We did not find any association between the grade of differentiation of the carcinomas and KLF4 expression (Fig. 5). Furthermore no association was found between differentiation degree and KLF4 expression pattern when the analysis was restricted to oral tongue carcinomas (one-way ANOVAs, cell fraction:
SOX2 immunohistochemical expression in oral squamous cell carcinomas. Representative sections of WD, MD and PD oral carcinomas with SOX2 expression. A, B. Image from a SCC developed in the gingiva and with low expression of SOX2. SOX2 immunostaining was localized in the nucleus of the SCC cells (B). C, D. Image from a carcinoma from the lateral border of the tongue presenting a moderate to strong expression of SOX2 (D). E, F. Representative image from a SCC developed in the gingiva showing strong SOX2 expression throughout the parenchyma of the carcinoma (F). Insets in A, C and E show high power view from each carcinoma with different histologic degree studied.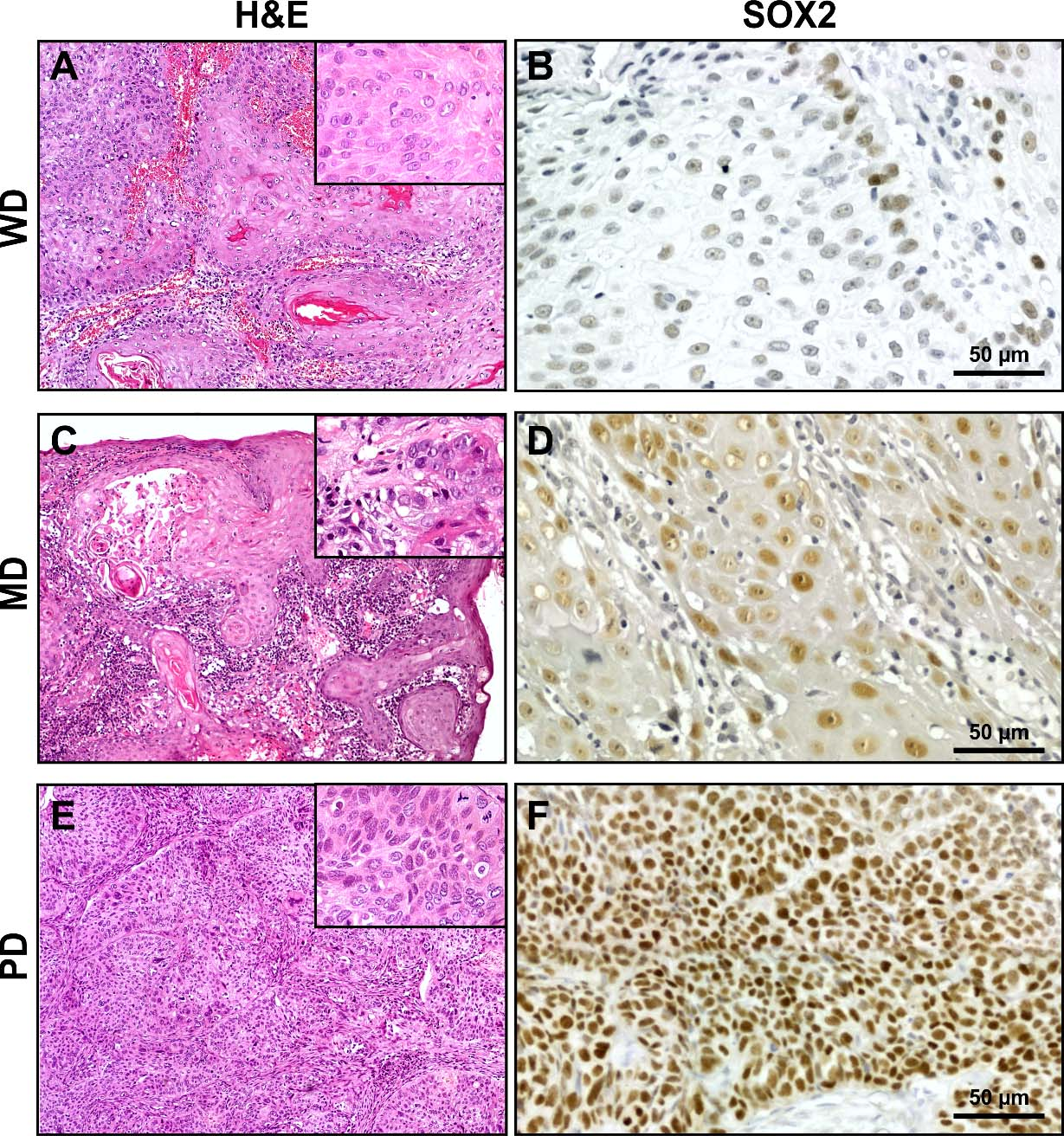
Histologic degree of OSCC and SOX2 expression. Fraction (A), density (B) and intensity (C) of SOX2 positive cells for OSCC grouped according to the histologic degree WD, well differentiated; MD, moderately differentiated; PD, poorly differentiated. Bars represent mean 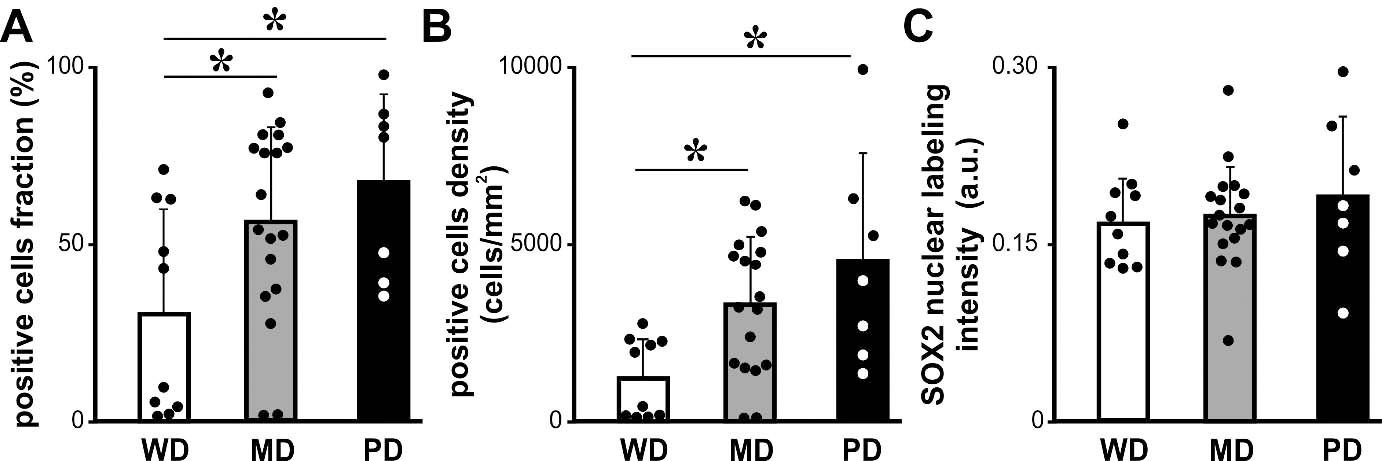
Quantitative analysis of SOX2 expression in OTSCC developed in different tongue subsites. Fraction (A), density (B) and intensity (C) of SOX2 positive cells for OTSCC from the borders of the tongue (tongue border) and the rest of the oral tongue subsites (other tongue site). Bars represent mean 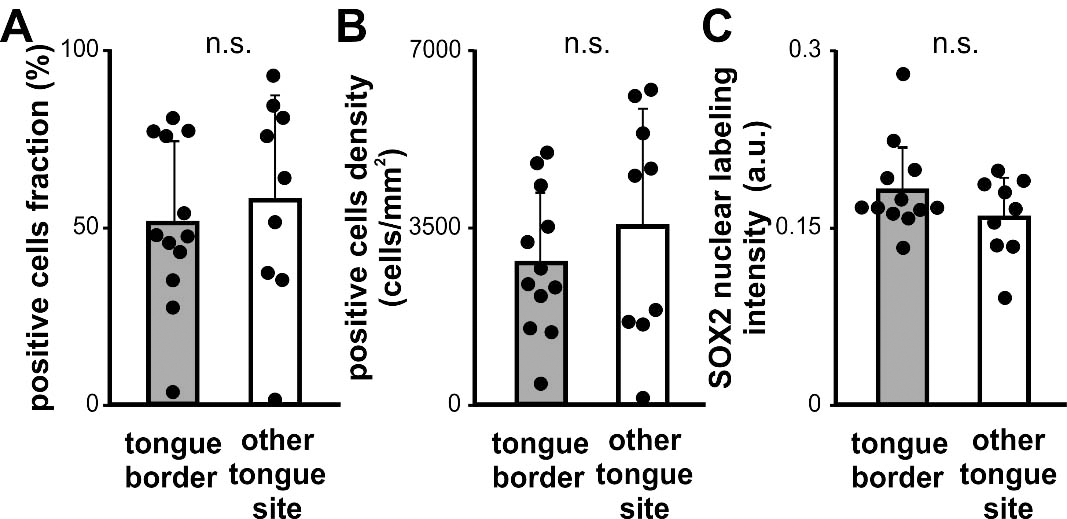
Correlation analysis between SOX2 and KLF4 expression in OSCC and OTSCC. Scatter plots depicting KLF4 and SOX2 positive cells fractions (A) and nuclear labeling intensities (B) for all analyzed cases. Each dot represents an individual case. A. No significant correlation was observed between KLF4 and SOX2 positive cells fraction in OTSCC (T) R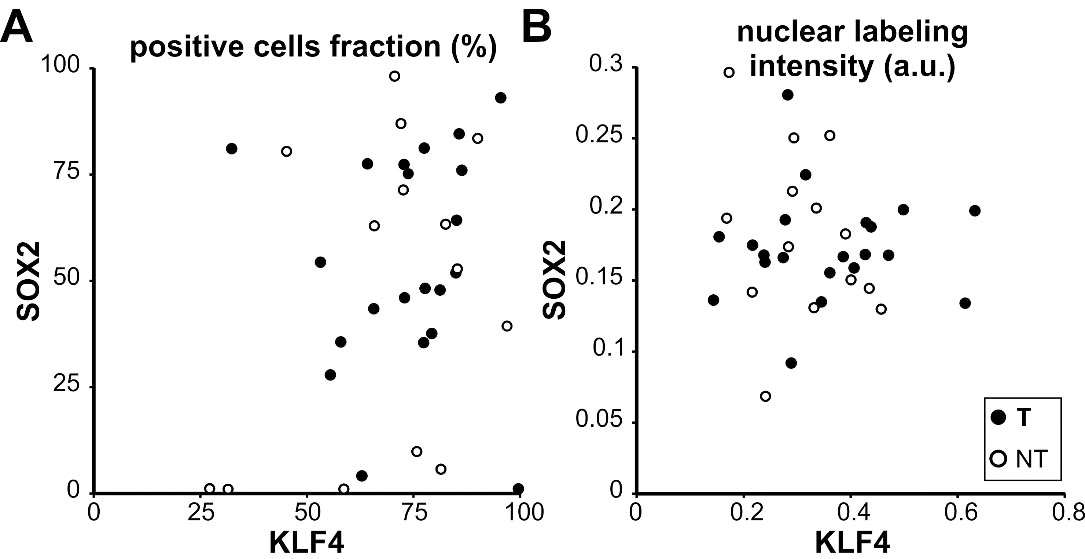
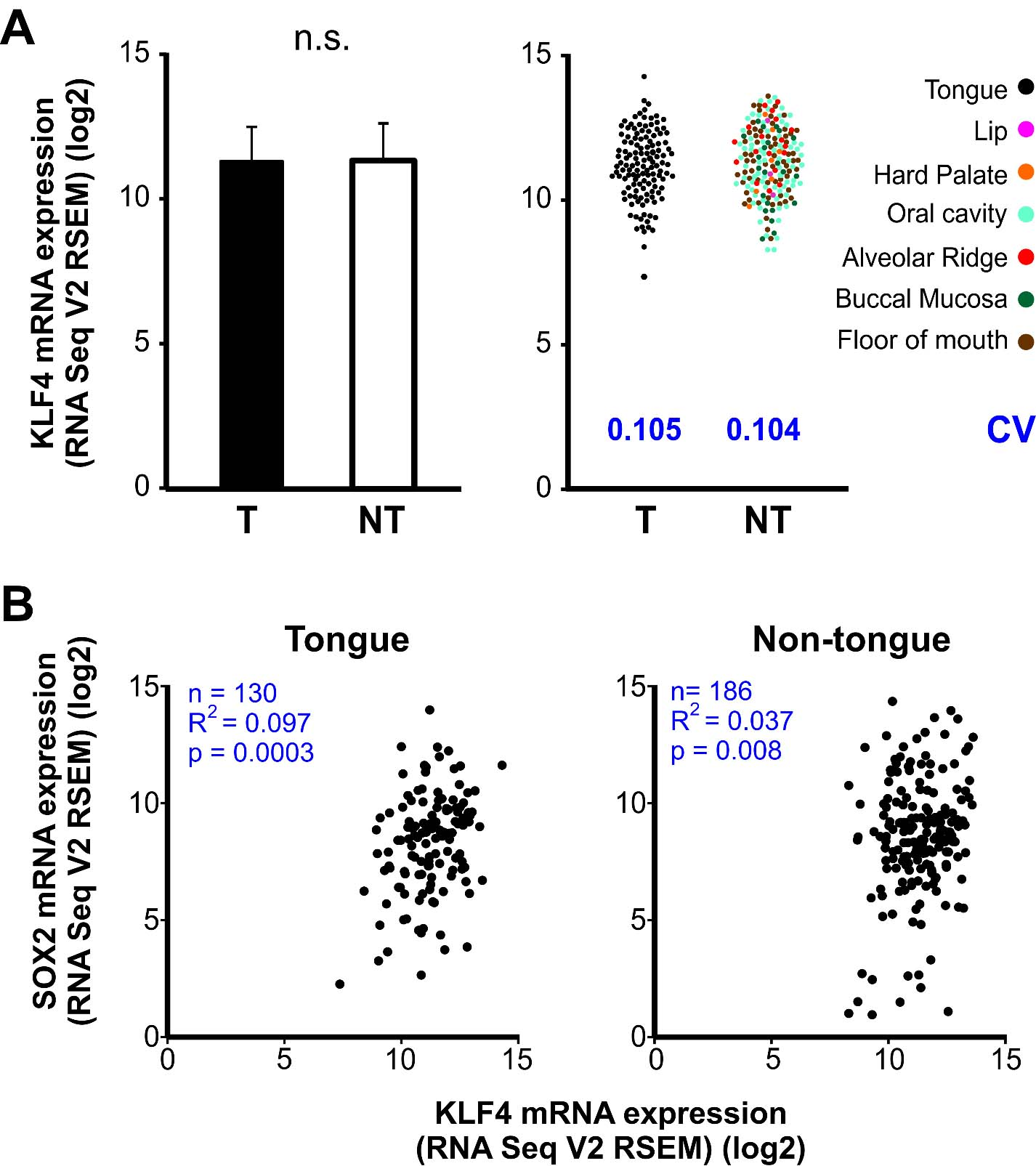
To extend our analysis we analyzed SOX2 expression by immunohistochemistry in the OSCC cases already studied for KLF4 expression. SOX2 was localized in the nucleus of the OSCC cells throughout the parenchyma of the carcinomas at different levels of expression (Fig. 6). The quantitative analysis of the SOX2 immunostaining revealed statistically significant differences between OSCC of different histological grade (Fig. 7). We found increased fraction and density of SOX2 positive cells in moderate and poorly differentiated carcinomas when comparing with well differentiated OSCC. Our data is in concordance with previous ones already reported for OSCC [33]. Therefore we studied if there is a differential expression for SOX2 in OTSCC from different areas of the oral tongue. No statistically significant differences were found regarding SOX2 expression parameters between OTSCC from the border of the tongue versus OTSCC developed in others areas of the tongue (Fig. 8). These results suggest that KLF4 differential expression within tongue subsites is not related to SOX2 differences between these areas. To further evaluate if KLF4 expression is associated with SOX2 labeling we calculated the correlation between these markers using mRNA data from TGCA database and immunolabeling results from our cases. Although KLF4 mRNA expression displayed a significant correlation with SOX2 mRNA levels (Supplementary Fig. 1B) the extremely low R2 obtained indicates that KLF4 expression is not a good predictor of SOX2 mRNA levels for tongue or non-tongue locations. Furthermore, when assessing our immunolabeled samples we found that none of the parameters studied show a significant correlation between KLF4 and SOX2 expression in OSCC (Fig. 9). Limiting the analysis to oral tongue samples produced the same non-significant results supporting the idea that KLF4 and SOX2 can be considered as independent markers in OSCC in general and in tongue SCC in particular.
We describe here for the first time a differential KLF4 pattern of expression in OTSCC from different tongue subsites. We performed a quantitative analysis on digital images and we showed a reduction in KLF4 positives cells in OTSCC from the lateral borders of the tongue when compared with carcinomas developed in the rest of the oral tongue. However OTSCC did not differ in KLF4 expression versus SCC developed in others oral cavity anatomical locations. Previous reports assessing KLF4 expression levels in human cancer support the idea that KLF4 inhibit carcinogenesis in several tissues and organs as mentioned in the introduction. However, when considering squamous epithelium carcinogenesis the evidence is not compelling. While a qualitative assessment of KLF4 expression in skin SCC showed a decrease in its expression [34] other report supports the notion that KLF4 expression was increased and significantly associated with moderately and poorly differentiated skin SCC [35]. Regarding to OSCC the data is also contradictory. For instance, it has been reported that KLF4 expression was related with tumor grade and its expression was much lower in poorly differentiated oral cancers than in well differentiated cancer cells [22]. Conversely, no significant differences between KLF4 expression and the degree of tumor differentiation have been reported elsewhere [21, 36]. These discrepancies may arise from the methodology used since these studies had assessed KLF4 expression in a non-quantitative manner. In most cases a categorical classification based on visual inspection or semi-quantitative methods was used and the analysis was performed on a rather limited number of sampled areas. In our study, using a quantitative large-scale assessment, we could not find any differential expression of KLF4 across tumors with dissimilar differentiation levels confirming that KLF4 is not associated with tumor differentiation at least in OSCC. The lack of association between KLF4 levels and differentiation degree could be attributed to the diverse OSCC localization analyzed, however, when our analysis was restricted to the tongue, the results were similar to the ones obtained combining tumors from all oral locations, supporting our interpretation that KLF4 is not linked to histological degree in OSCC.
In this regard, inspection of mRNA expression data from a public oral TCGA database (42 samples) that includes case differentiation status, suggests an association between mRNA KLF4 levels and differentiation status of the tumor [37]. However, recent studies have shown that discrepancies between mRNA and protein abundance are common [38, 39]. A comprehensive tissue- and gene-specific analysis revealed that across the 14 tissues studied (including normal and cancer samples) correlation between mRNA and protein expression was positive in only 6.1% of the samples [39]. Furthermore, neither KLF4 nor SOX2 were reported in the group of genes and proteins with highly correlated expression, probably as consequence of differences in mRNA/protein in vivo half-life or varied post-transcriptional mechanisms involved in turning mRNA into protein.
In particular, our results in OTSCC show a significant reduction in the fraction and density of KLF4 positives cells in carcinomas developed in the border of the tongue. In this regard, epidemiological evidence suggests that biological differences and distinct clinical behaviors do exist between tumors arising from different oral cavity subsites and should be examined as different entities [40]. Furthermore carcinomas arising from different subsites within mobile tongue present distinct prognosis [41]. For instance, different series of cases reported the lateral side of the tongue as the more affected area of the mobile tongue as well as a high risk oral site distinctive from the rest of the oral cavity [42, 43].
Animal studies also support the oral tongue subsites differences reported in humans regarding the biological behavior from tumors originated in the lateral border or other tongue subsite. Evidence from genetic manipulation of KLF4 expression when it was restricted to a cellular compartment that includes the oral epithelial stem cells, showed oral premalignant lesions development in the tongue margins [44, 45]. Furthermore, the concomitant Klf4 deletion and Ras activation was sufficient to promote the development of SCC in the margins of the tongue [44]. The results presented here constitute a first attempt to uncover the biological basis aimed to explain clinical behavioral differences reported in tumors arising from diverse locations within the tongue, and it might be revealing an anatomical-dependent function during oral tongue carcinogenesis as proposed for KLF4 in others cancer models [20, 46].
The combined evidence obtained from our previous studies using mouse models of KLF4 ablation in a tongue cell compartment containing epithelial stem cells lead us to explore the relationship in human cancer between KLF4 and SOX2, a stem cell marker involved in carcinogenesis. The transcription factor SOX2 it is extensively known for its roles during mammalian development and its critical roles in the self-renewal and maintenance of embryonic and adult normal stem cells as well as in CSC [24]. In particular, SOX2 expression has been reported increased at protein level for glioblastoma, esophageal, breast and prostate cancer and the SOX2 increased expression correlates with increased histologic grade [24, 47]. Regarding to SOX2 role in oral cancer the reported data is controversial. Bayo et al. showed that tumors with high SOX2 levels of expression have a highest progression-free survival when compared to SOX2 expressing tumors at lower levels [48]. However this seems contradictory with the reported results demonstrating that SOX2 amplifications are common in OSCC and SOX2 amplification is significantly associated with the pathological grade [33, 49]. Our SOX2 data reported here is aligned with the later proposal since we found increased fraction and density of SOX2 positive cells in moderate and poorly differentiated carcinomas when comparing with well differentiated OSCC. This discrepancy could be apparent when considering a recent proposal where histological degree does not correlate with survival outcome [50, 51, 52].
Considering that SOX2 expression was linked to tumor differentiation status and KLF4 expression differ across tongue subsites we did not anticipated a strong association between these two markers of stemness. However, previous studies qualitatively assessing their expression have described some degree of association between them in OSCCs [36]. Thus, to further clarify this point we took advantage of our quantitative data and formally tested the relationship between the expression levels of these two markers by calculating the correlation index in a case by case manner. Our result could not identify any significant correlation, neither in OSCC nor in tongue samples, between KLF4 and SOX2 expression. These results reinforce the notion that they might be involved in different stages of the oral carcinogenesis process.
Conclusion
We show here for the first time that OTSCC developed in the borders of the tongue contain fewer tumor cells positive for KLF4 expression than their counterpart from the rest of the tongue. Interestingly this reduction takes place without changes in KLF4 expression level per cell. KLF4 may be useful in understanding the biological behavior of carcinomas originated in different oral tongue subsites. In addition the present work highlights the potential benefits of unbiased quantitative whole slide scanning analysis for diagnostic. Our study prompts future OTSCC prospective studies looking for clinical prognosis to incorporate detailed subsite information in the analysis.
Footnotes
Acknowledgments
We thank German La Iacona from the Microscopy Facility IFIBIO-Houssay and Juan Belforte who kindly provided image analysis technical support. The valuable technical advice for immunostaining of Valeria Sanchez is also acknowledged.
This work was supported by grants from the National Agency for the Promotion of Science and Technology, Argentina (ANPCyT, PICT 2017-0361) and the Argentinean National Research Council (CONICET, PIP11220150100251CO).
Conflict of interest
The authors state that they have no conflicts of interest.
Author’s contributions
Conception: Ana Rosa Raimondi.
Interpretation or analysis of data: Maria Luisa Paparella, Dario Martin Ferri, Krissya Maria Villegas.
Preparation of the manuscript: Maria Luisa Paparella, Ana Rosa Raimondi.
Revision for important intelectual content: Ana Rosa Raimondi.
Supervision: Ana Rosa Raimondi.
Supplementary data
A. Left. KLF4 mRNA expression levels in OTSCC (T) and the rest of the SCC developed in the oral cavity (NT). Bars represent mean