Abstract
BACKGROUND:
Glioblastoma (GBM) is the most common and aggressive primary malignant brain tumor with a high mortality rate. Aberrant activation of signal transducers and activators of transcription (STAT) signaling results in tumor pathogenesis and progression by regulating cell cycle, cell survival and immune response.
METHODS:
Therapeutic targets and prognostic biomarkers within the STAT family in GBM were explored using web applications and databases.
RESULTS:
High levels of STAT1/3/5A/5B/6 and low levels of STAT4 were observed in GBM patients. GBM patients expressing high STAT1/2/3/5A/6 and low STAT4/5B levels had the worse overall survival. Among the STAT family, STAT4 and STAT6 were the most frequently mutated genes. A low to moderate correlation among members of the STAT family was observed. Additionally, the STATs were involved in activation or inhibition of cancer related pathways. Analysis of immune infiltrates showed STAT5A levels to be significantly correlated with abundance of immune cells and levels of immune gene biomarkers. Gene ontology (GO) functions and KEGG pathway analysis indicated that STAT5A is involved in immune response-regulating signaling pathway, neutrophil and lymphocyte mediated immunity, single-stranded DNA binding, cytokine-cytokine receptor interaction, NOD-like receptor signaling pathway, NF-kappa B signaling pathway and TNF signaling pathway. Moreover, several kinase and transcription factor targets of STAT5A in GBM were identified.
CONCLUSION:
We report here therapeutic targets, prognostic biomarkers and regulation network of STAT family in GBM. These findings lay a foundation for further studies on the role of STAT family in therapy and prognosis of GBM. Further studies are required to verify our results.
Background
Glioblastoma (GBM) is the most common and aggressive primary malignant brain tumor, accounting for about 45.6% of all brain malignancies [1]. In 2020, the estimated number of new cases of brain malignancies in America alone is 23,890 [2]. In the past two decades, no significant progress has been made in the standard of care for GBM patients [3]. Moreover, prognosis of GBM patients is poor, with a median overall survival less than 20 months and a 5-year-survival of 5% [4, 5]. Therefore, identification and use of novel biomarkers for prognosis and therapy, and a combination of current treatments are the most promising strategies in management of GBM.
Signal transducers and activators of transcription (STAT) signaling pathways are crucial in the function, proliferation and apoptosis of immune cells such as CD8+ cells and dendritic cells [6]. Aberrant activation of STAT signaling results in tumor pathogenesis and progression by regulating cell cycle, cell survival and immune response [7, 8]. The STAT family comprises of STAT1-4, STAT5A/5B and STAT6. Previous studies have shown STAT1 to function as an anti-oncogene in GBM under hypoxia and inhibited angiogenesis [9]. Interaction between STAT3 and Forkhead box protein M1 (FOXM1) confers radio-resistance in GBM cells [10]. These findings demonstrate a vital function played by STATs in occurrence and progression of GBM. However, potential applications and mechanisms of the STAT family in GBM therapy and prognosis are still unclear.
This study explored expression levels and prognostic value of the STAT family in GBM. The study evaluated single nucleotide variation (SNV), correlation, cancer-related pathways, drug sensitivity, immune infiltrates and functional analysis of the STAT family in GBM. Our findings highlight potential application value and mechanisms of the STAT family in therapy and prognosis of GBM.
Materials and methods
Oncomine
Oncomine (
GEPIA
Gene Expression Profiling Interactive Analysis (GEPIA –
PrognoScan
PrognoScan (
TIMER
Tumor IMmune Estimation Response (TIMER,
GSCALite
Gene Set Cancer Analysis (GSCALite,
Transcriptional expression of STAT family in 20 different types of cancer diseases from Oncomine database. Difference in transcriptional expression compared by Student’s 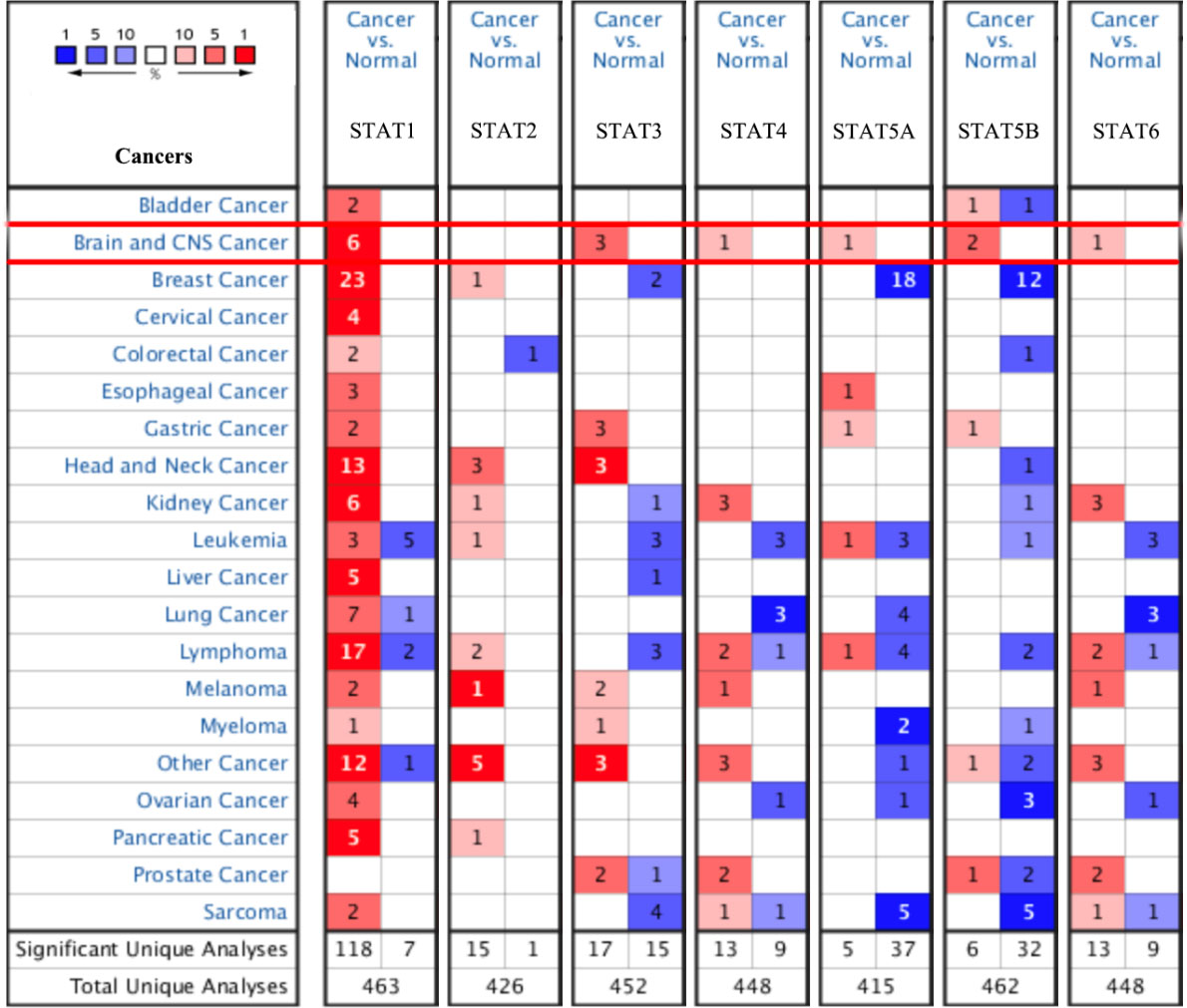
LinkedOmics (
GeneMANIA
GeneMANIA (
Results
Expression and prognosis analysis of STAT family in GBM
To assess the potential involvement of STAT family in GBM, their differential expression at mRNA and protein level was analyzed using multiple datasets hosted on Oncomine and GEPIA. Analysis of STAT family mRNA expression in 20 cancer types (Oncomine) showed a significant upregulation and downregulation in several cancers including GBM (Fig. 1). Particularly, STAT1 was significantly upregulated in tumor tissues, with a fold change of 3.136 and a
Expression and prognosis analysis of STAT family in GBM (GEPIA). (A) Box plots showing gene expression using data derived from GEPIA comparing expression of specific STAT family members in GBM and normal tissues. Difference in STAT family expression compared by Student’s 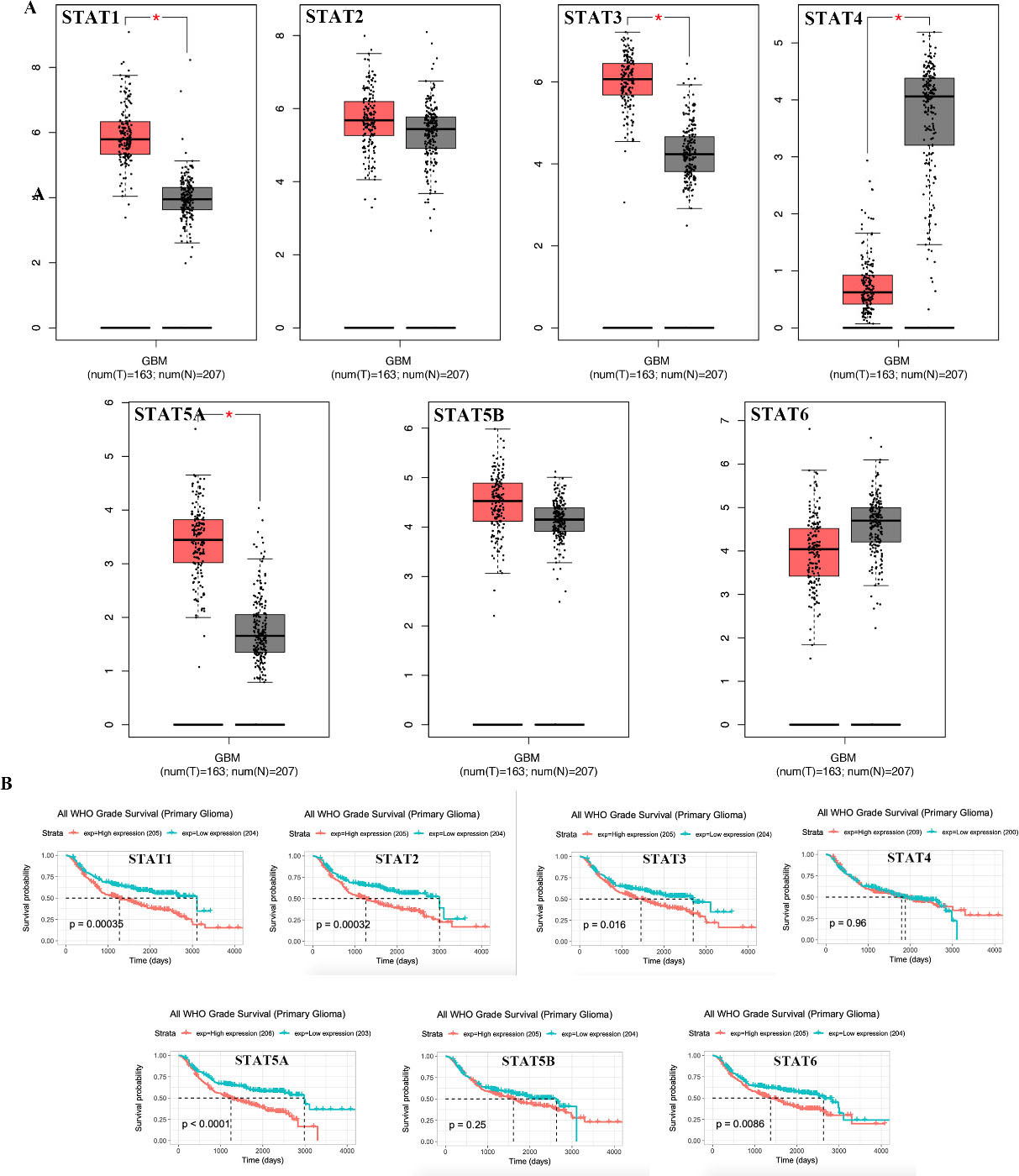
mRNA levels of STAT family in glioblastoma (ONCOMINE)
Difference in expression of members of the STAT family determined using Student’s
Single Nucleotide Variation (SNV), co-expression and cancer-related pathway analysis of STAT family in GBM (GSCALite). (A) SNV analysis of STAT family in GBM. (B) Correlation heat map of STAT family in GBM; Pearson correlation coefficient, 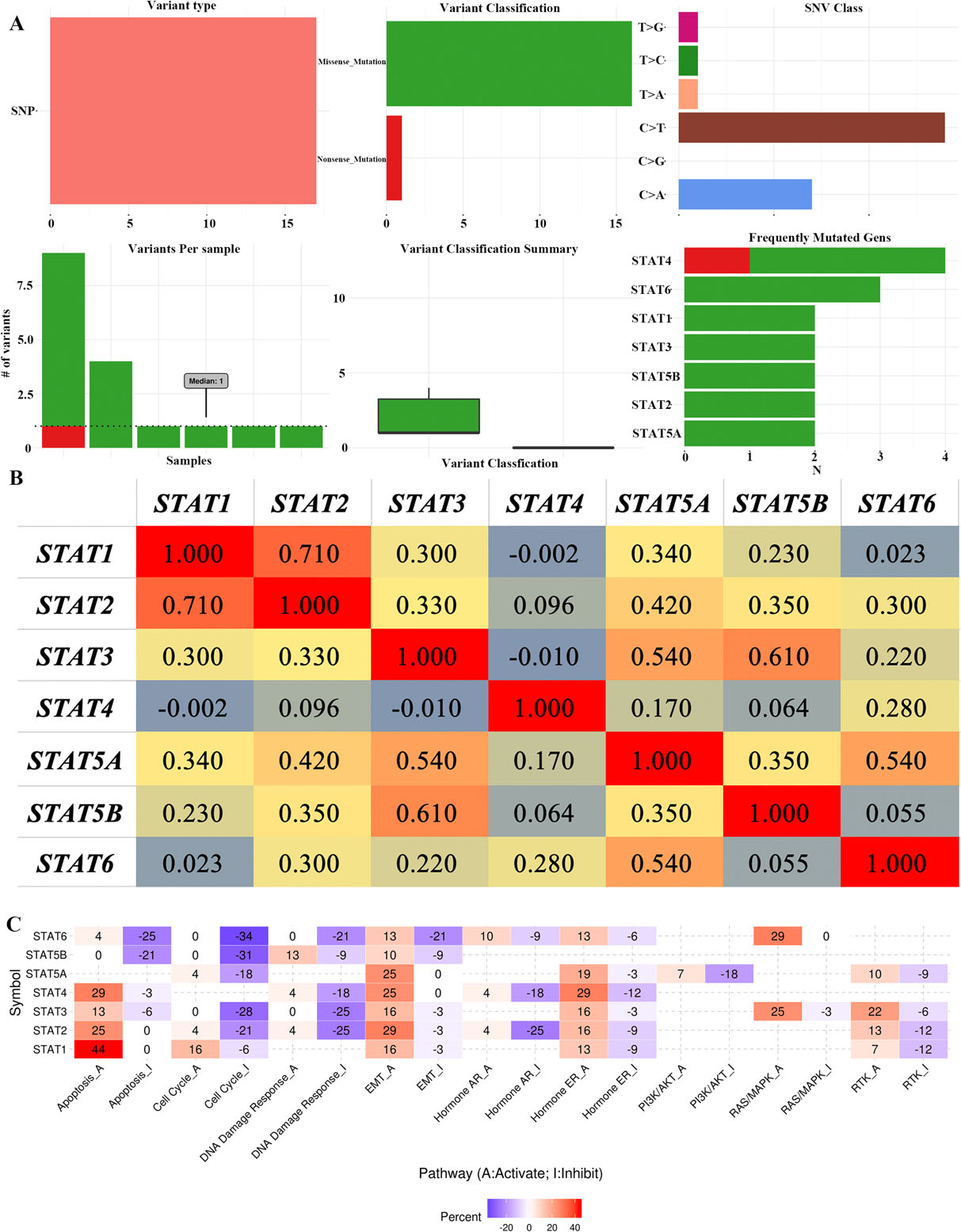
The results of prognosis analysis of STAT family in GBM using PrognoScan are shown in Table 2. Notably, GBM patients with high expression of STAT1 (HR [95% CI]: 1.55 [1.11–2.16],
Overall Survival of STAT family in glioblastoma (PrognoScan)
Analysis; Kaplan-Meier. N; numbers of patients.
Drug sensitivity of STAT family in GBM (GSCALite). Negative correlation indicate that low expression of gene is resistant to the drug and vice versa. Spearman’s correlation analysis conducted with TCGA GBM dataset (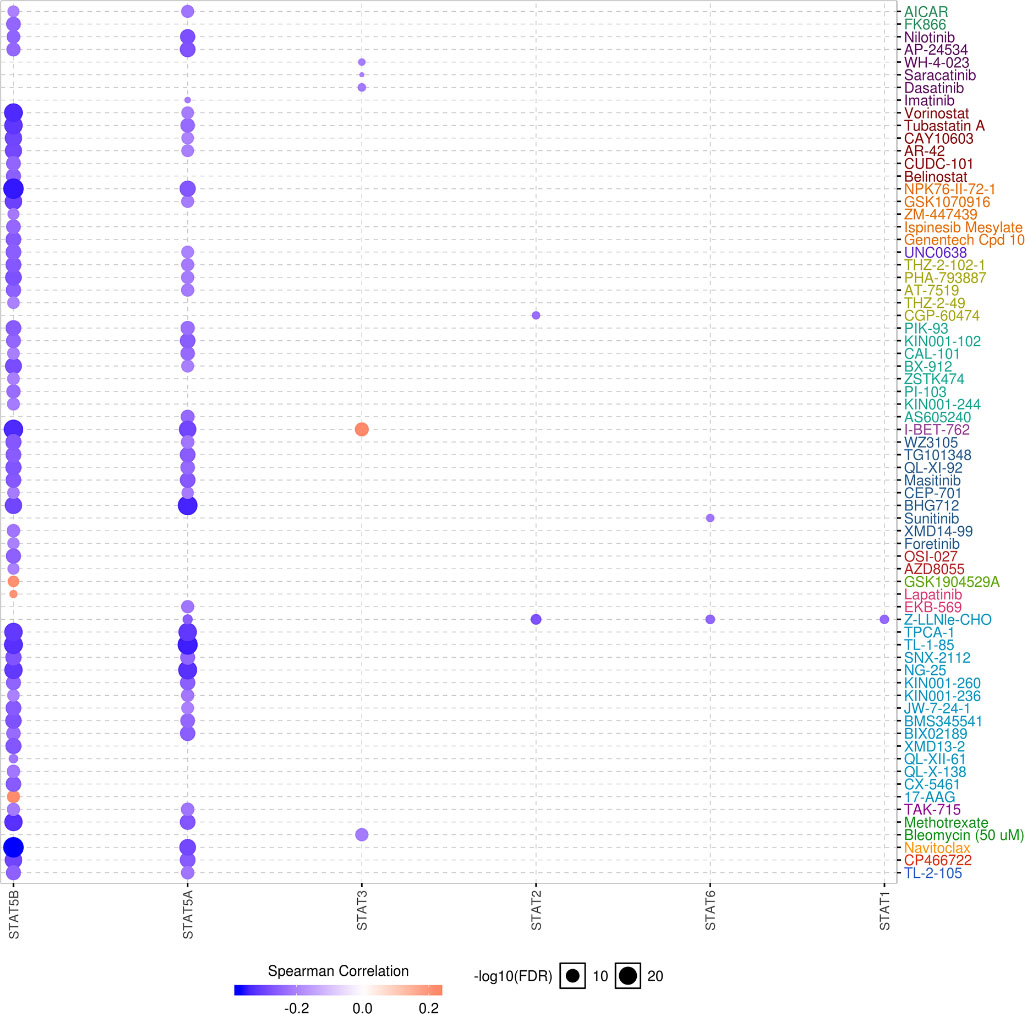
Missense mutation was the most common variant classification and C
Immune cell infiltration of STAT5A in GBM (TIMER). STAT5A expression is significantly positively associated with infiltrating levels of B cells, CD8+ T cells, CD4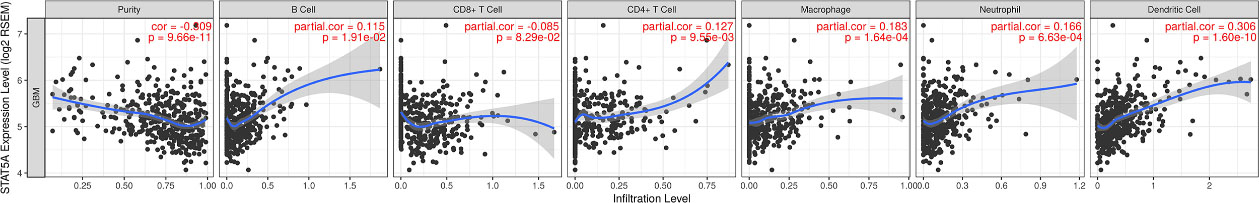
Expression of STAT5A was high in GBM, making it a potential prognostic biomarker. Moreover, STAT5A was involved in activation and inhibition of key cancer related pathways and drug resistance. This implies that STAT5A plays significant roles in GBM pathogenesis and progression. Under this backdrop, STAT5A was selected for further analysis. Correlation between STAT5A and immune infiltrates in GBM was analyzed owing to the important roles the STAT family play in immune response [25, 26]. The levels of STAT5A were significantly correlated with abundance of B cell (
Expression of STAT5A was also strongly correlated with expression of gene biomarkers in GBM presented in Table 3. In reference to gene biomarkers of CD8+ T cell, data obtained from TIMER showed CD8B expression to be positively associated with STAT5A expression while data from CGGA showed the expression of both CD8A and CD8B to be positively associated with expression of STAT5A. Data obtained from both TIMER and CGGA indicated a strong correlation between expression of STAT5A and the expression of T cell (CD3D, CD3E, CD2), Monocyte (CD86 and CD115), TAM (CCL2, CD68, IL10) and M2 Macrophage (CD163, VSIG4, MS4A4A) gene biomarkers. From the analysis of gene biomarkers of Natural killer cell in TIMER, the levels of KIR2DL3, KIR2DL4, and KIR3DL1 were positively correlated with STAT5A expression. In both TIMER and CGGA, eight gene biomarkers of Dendritic cell were strongly associated with STAT5A expression. Similarly, the levels of STAT4, STAT1, TNF, GATA3, STAT6 and IL13 were positively correlatively with STAT5A expression. Two gene biomarkers (FOXP3 and TGFB1) of Treg and three gene biomarkers (PD-1, CTLA4 and GZMB) of exhibited a strong correlation with STAT5A expression. Therefore, STAT5A presents a suitable target for immunotherapy in GBM.
Correlation between STAT5A and gene biomarkers of immune cells in GBM (TIMER)
Correlation between STAT5A and gene biomarkers of immune cells in GBM (TIMER)
*
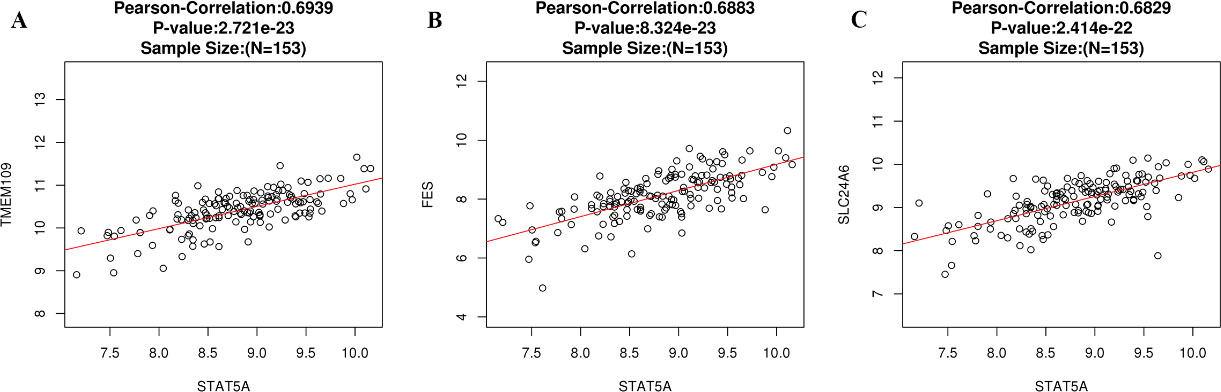
Figure 6A showed the genes significantly correlated with STAT5A in GBM. Among these genes, the top 50 most significant genes positively (Fig. 6B) and negatively (Fig. 6C) associated with STAT5A were also extracted. Moreover, Supplementary Fig. 1 showed the top three genes positively associated with STAT5A, which were TMEM199 (cor
Enrichment of STAT5A in GBM (LinkedOmics). (A) Genes associated with STAT5A in GBM based on TCGA GBM dataset (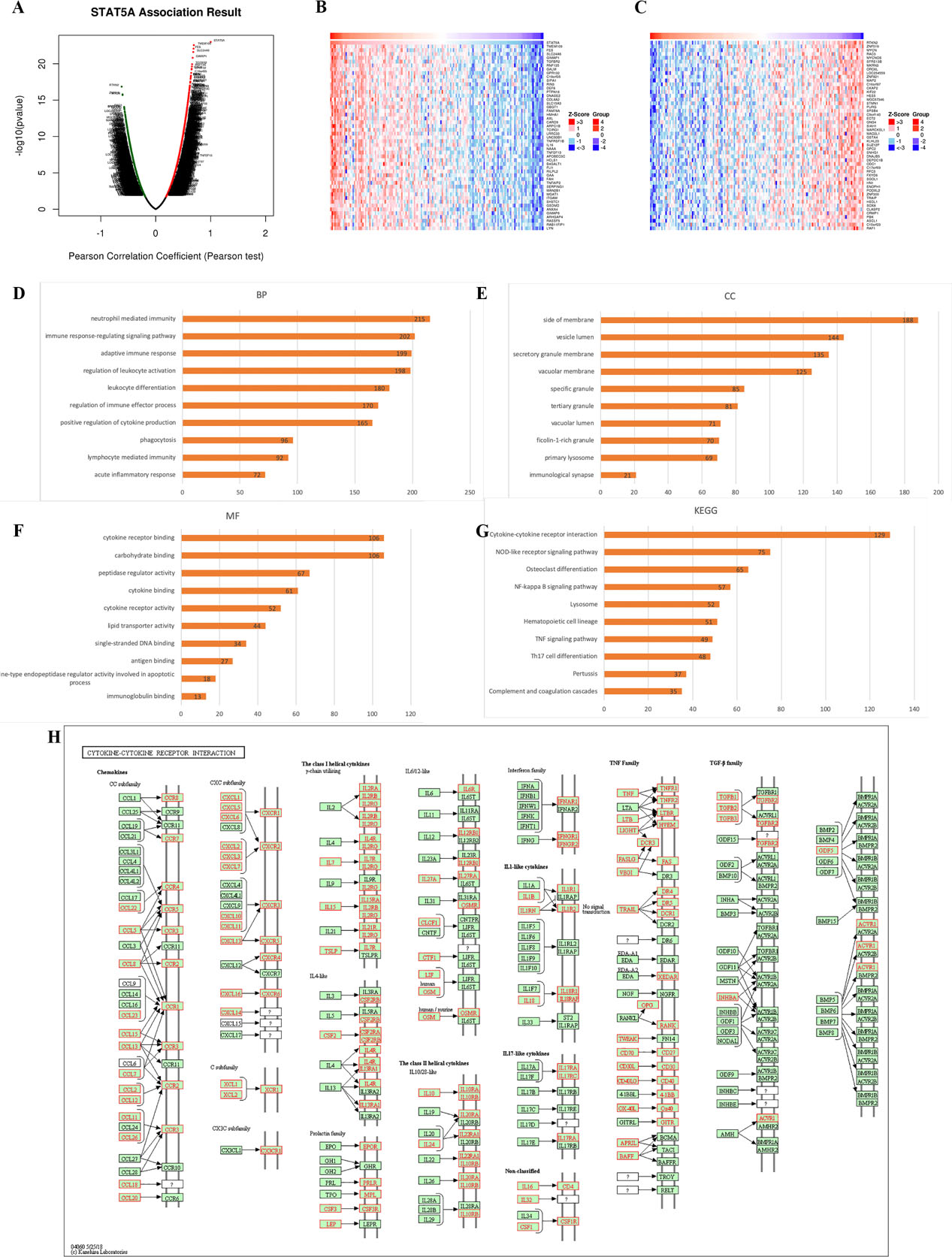
Kinase and transcription factor target networks of STAT5A in GBM were also explored (Table 4). Here, kinases LCK, SYK, LYN, HCK, and TTK were identified as the five most significant kinase network targets. The PPI network showed kinase LCK target networks to be involved in regulation of immune response, antigen receptor-mediated signaling pathway, T cell activation and receptor signaling pathway (Fig. 7A). The five most significant transcription factor targets identified were V$ELF1_Q6, V$IRF_Q6, V$PEA3_Q6, V$PU1_Q6, and V$NERF_Q2. PPI network indicated that transcription factor ELF1 networks are involved in regulation of immune response, lymphocyte, leukocyte and B cell activation (Fig. 7B).
PPI networks of kinase LCK and transcription factor ELF1 networks (GeneMANIA). (A) The applied bioinformatics methods and the biological functions of gene sets of kinase LCK networks. (B) The applied bioinformatics methods and the biological functions of gene sets of transcription factor ELF1 networks.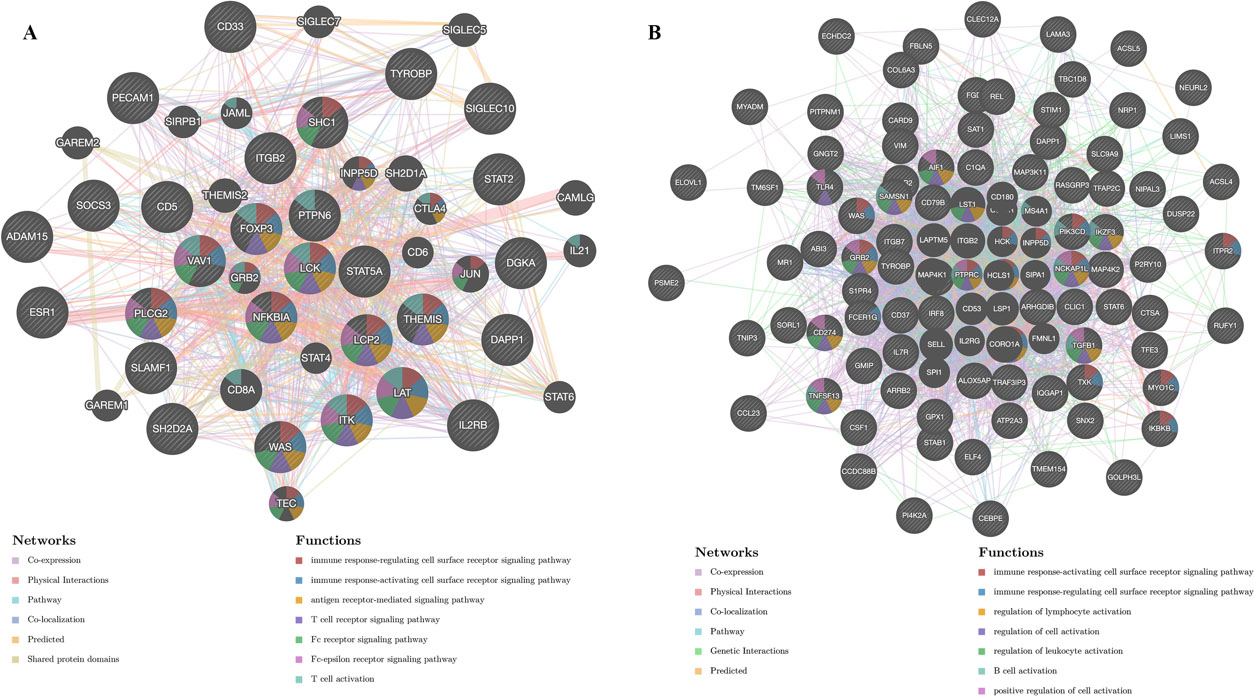
Kinase and transcription factor targets of STAT5A in GBM (LinkedOmics)
Enrichment analyses performed using GSEA.
The STAT family mediates various biological processes including cellular immunity, invasion, differentiation and apoptosis [27]. Activated by diverse cytokines, STAT signaling play a central role in immunity, cell death, cancer pathogenesis and progression [8]. Studies have shown that the STAT family exert important functions in cancers such as hepatocellular carcinoma. However, the role of the STAT family in GBM has not been studied.
From the expression analysis, the levels of STAT1/3/ 5A/5B/6 were high in GBM, while the levels of STAT4 were significantly low. GBM patients with high expressions of STAT1/2/3/5A/6 and low expressions of STAT4/5B had a worse overall survival. These findings are consistent with results from previous studies. For instance, Balaram et al reported STAT1 to be upregulated in GBM and correlated with a worse prognosis [28]. Another study fronted STAT3 as a potential biomarker for prognosis of GBM [29]. These findings corroborate our recommendation of the STAT family as potential prognostic biomarkers in GBM.
Cancer-related pathway analysis showed that the STAT family play significant roles in GBM mainly by activating; the apoptosis pathway, EMT pathway, hormone ER pathway, RAS/MAPK pathway, RTK pathway, and inhibiting; the cell cycle pathway, DNA damage response pathway, hormone AR pathway and P13K/AKT pathway. In an earlier study, Adetola et al reported that STAT3 inhibition induced apoptosis in cancer cells [30]. In hepatocellular carcinoma, STAT1 has been reported as a tumor-suppressor, inducing G0/G1 cell cycle arrest and apoptosis of tumor cells [31]. Therefore, the STAT family may regulate pathogenesis and progression of GBM via these pathways.
In the present study, we observed the levels of STAT5A to be significantly correlated with abundance of immune cells and the levels of immune gene biomarkers. These immune cells and immune gene biomarkers have previously been reported as therapy targets and play a significant role in various types of cancer. Immune cells, including dendritic cells and CD8+ cytotoxic T lymphocytes are promising therapeutic targets in bone metastasis [32]. In breast cancer, CD8+ T cells and regulatory T cells have been reported as reliable prognostic biomarkers [33]. Similarly, CTLA4 and PD-1 have been proposed as immune checkpoint for cancer immunotherapy [34]. Here, we present STAT5A as a potential therapeutic target for cancer immunotherapy.
Correlation between STAT5A expression and several drugs or small molecules, including I-BET 762 and Nilotinib was evaluated. I-BET 762 has been shown to inhibit tumor cell proliferation, serving as a potential therapy for pancreatic cancer. Another study demonstrated the potential use of I-BET 762 in preventing breast and lung cancer. Nilotinib, a second-generation tyrosine kinase inhibitor, is widely regarded as a therapy target for chronic myeloid leukemia. In GBM, Nilotinib could regulate cell invasion. I-BET 762 and Nilotinib could also serve as therapy targets or play significant roles in GBM progression through STAT5A-associated signing. Further studies are required to explore this possibility.
Gene Ontology (GO) functions and KEGG pathways analysis indicated that STAT5A is involved in immune response-regulating signaling pathway, neutrophil and lymphocyte mediated immunity, single-stranded DNA binding, cytokine-cytokine receptor interaction, NOD-like receptor signaling pathway, NF-kappa B signaling pathway and TNF signaling pathway. These functions and pathways are associated with immune response, carcinogenesis and progression. Fernando et al. reported NOD-like receptors as important players and targets in the interface between innate immunity and cancer [35]. A previous study has also shown TNF signaling pathway to be involved in regulation of proliferation and invasion in cervical carcinoma [36]. Single-stranded DNA-binding proteins are crucial in maintaining the integrity of a genome [37]. Furthermore, single-stranded DNA-binding proteins are involved in cell-cycle checkpoint activation following DNA damage in cancer [37]. Therefore, STAT5A may affect pathogenesis and progression of GBM through these signaling pathways.
This study had some limitations. First, the sample size was relatively small, and we recommend a larger cohort study to be performed. Moreover, our study was short of clinicopathological features analysis. Functional analysis should be done to validate the role of STAT family in GBM both in vivo and vitro.
Conclusion
This study outlines expression and demonstrates the prognostic value of STAT family in GBM. Expression levels of STAT5A are significantly correlated with abundance of immune cells and the level of immune gene biomarkers. Moreover, STAT5A is involved in immune response-regulating signaling pathway, cytokine-cytokine receptor interaction, NOD-like receptor signaling pathway, and TNF signaling pathway. These results underscore the clinical significance of and regulation networks of STAT family in GBM. The findings lay a foundation for further studies on potential roles of the STAT family in therapy and prognosis of GBM.
List of abbreviations
GBM: Glioblastoma, STAT: Signal transducer and activator of transcription, SNV: single nucleotide variation, TCGA: The Cancer Genome Atlas, GO: Gene Ontology, KEGG: Kyoto Encyclopedia of Genes and Genomes
Footnotes
Funding
This study was funded by High-level Hospital Construction Research Project of Maoming People’s Hospital.
Supplementary data
The three most significant genes correlated with STAT5A in GBM (LinkedOmics).