Abstract
BACKGROUND:
The breast cancer subtype deficient in estrogen receptor and human epidermal growth factor receptor-2 (ER-/HER2-) displays enhanced aggressiveness, metastasis and disease relapse due to chemoresistance. ER-/HER2- patients lack molecularly targeted treatment hence, new therapeutic and prognostic biomarkers are required for better patient management.
OBJECTIVES:
To investigate the prognostic role of protein tyrosine phosphatase genes in Breast Cancer and their relevance as predictive markers for chemoresistance.
METHODS:
We examined the expression of 114 protein tyrosine phosphatase (PTP) genes in 1700 breast cancer patient’s tumor samples with respect to ER-/HER2- subtype. Correlation of relevant candidates with chemoresistance was analyzed in breast cancer cells resistant to taxane/anthracycline based drugs. The prognostic value of key candidates was assessed using Kaplan Meier plots and Nottingham prognostic index and expression pattern was confirmed using qRT-PCR. The epigenetic regulation was analyzed using ChIP-Seq datasets. By plotting ROC plots, clinical outcome after treatment with taxane and anthracycline was established.
RESULTS:
Overexpression of CDC25A and CDC25C and under-expression of DUSP16 was observed in tumor samples of ER-/HER2- patients and breast cancer cells. Similar expression patterns of these candidate genes were observed in MCF7 cells resistant to paclitaxel and adriamycin and also correlated with poor prognosis of breast cancer patients. Increased CDC25A and CDC25C in ER-/HER2- cells was found to be regulated epigenetically by histone H3K4 methylation. Overall, the present study establishes increased expression of protein tyrosine phosphatase CDC25C as a poor prognostic marker for breast cancer.
CONCLUSION:
Our study highlights the role of CDC25C in chemoresistance to taxane and anthracycline based therapy and proposes CDC25C as a potential predictive marker for these cancer therapies.
Introduction
Breast cancer is one of the most common cancers in female accounting for 15% of all cancer deaths among women [1]. One of the most aggressive and metastatic subtype of breast cancer is Triple Negative Breast Cancer (TNBC), accounting for 10–20% of cancer incidences [2]. TNBC lacks clinical expression of estrogen receptor (ER), progesterone receptor (PR), and does not undergo amplification of human epidermal growth factor receptor 2 (HER2), while other subtypes overexpress at least one of the receptors [3, 4]. The presence and/or absence of these hormone receptors not only facilitate molecular sub-categorization of breast cancer but also are deciding factors for therapy regimens.
Various classification models of breast cancer specifically assign a tumor sample to a certain subtype. Appropriate categorization of tumor provide better molecular understanding and facilitate effective treatment strategies. Gene expression profiling of multiple tumor samples and identification of key genes is used to construct sub-classification. Some of the classifications are single sample predictors (SSPs) based on 50 to 500 genes while others are subtype classification models (SCMs) based on ER, HER2 and the proliferation phenotypes. Among SCMs, 3-gene classifier has been established as robust method to classify breast cancers [5]. 3-gene classifier model is based on the expression of ER, HER2 and Aurora Kinase A and divides Breast Cancer into ER-/HER2-, HER2+ and ER+/HER2- low/high proliferation subtypes each corresponding to basal-like, HER2 enriched and luminal A/B subtypes of PAM-50 SSP classification, respectively. Identifying molecular signatures associated with these subtypes would better stratify patients and also provide therapeutic, prognostics and predictive biomarkers for better patient management and treatment outcome.
The 3-gene classifier system captures main features of ER signaling and thus is a robust way of classification also facilitating decisions of therapeutic approaches. For example, patients positive for ER
In this study, we focussed on elucidating the role of protein tyrosine phosphatases (PTPs) in breast cancer specifically in ER-/HER2- subtype. PTPs function both as positive and negative regulators of cellular signaling pathways and work in unison with protein tyrosine kinases (PTK) to control the cellular homeostasis. Signaling defects due to alterations in kinases and phosphatases may succumb cells to abnormal growth, proliferation, differentiation and ultimately tumorigenesis. While the role of kinases such as receptor tyrosine kinases, MAP kinases etc. in breast cancer tumorigenesis is well established [17], the pathological contribution of protein tyrosine phosphatases in breast cancer have received less attention. The PTP gene family comprises of 125 proteins with phosphatase enzymatic activity on specific substrates and mediate diverse functions [18, 19]. One study identified PTP4A3 as a key therapeutic target in breast cancer [20]. PTPN12 is also implicated as a poor prognostic marker for triple negative breast cancer [21]. However, till date the potential role of PTPs in acquiring resistance to chemotherapeutic drugs in breast cancer is largely unstudied except for a report demonstrating overexpression of PTP1B as a chemoresistance feature to tamoxifen in breast cancer cells [22]. Chemoresistance is a major reason for cancer recurrence and high mortality rates of ER-/HER2- breast cancer patients, therefore identifying new molecular biomarkers predicting resistance of breast cancer to chemotherapeutic drugs is pertinent.
Hence, in our study, we investigated the transcriptional landscape of protein tyrosine phosphatases in ER-/HER2- aggressive, metastatic subtype of breast cancer to identify relevant PTPs, which could be associated with prognosis, chemoresistance and predict the response to drug treatment in breast cancer. We explored specific clusters of up/downregulated PTP genes employing RNA-Seq data of ER-/HER2- patient tumor samples from The Cancer Genome Atlas (TCGA) database and compared it with other subtypes. Then we evaluated the association of these gene clusters with chemoresistance to paclitaxel and adriamycin, relapse free patient survival and assessed their predictive significance. We identify CDC25C as a key PTP gene, whose overexpression is correlated with poor prognosis in breast cancer. Further, potential role of these genes in acquiring chemoresistance to paclitaxel and adriamycin and their predictive value for relapse free survival of patients’ post-chemotherapy, was explored. We find CDC25C as a predictive biomarker for taxane and anthracycline based therapies and propose its potential role in acquiring chemoresistance and metastatic recurrence. We further observe that expression of CDC25C is epigenetically regulated by histone modification, putting forth a new avenue of targeting ER-/HER2- breast cancers with inhibitors of epigenetic modulators regulating CDC25C transcription.
Materials and methods
Cell line and cell culture
The ER+/HER2- cell line MCF7 (Luminal-A) and the ER-/HER2- cell line MDA-MB-231 (Triple negative claudin low [23] were used in the study. MCF7 cells were cultured in RPMI 1640 media (Himedia laboratories) supplemented with 10% (v/v) fetal bovine serum (Gibco), 100 U/ml Penicillin, 0.1 mg/ml streptomycin and 2 mM glutamine. MDA-MB-231 cells were cultured in DMEM (Himedia laboratories) with the same additives as described above.
RNA isolation and real time qpcr
Total RNA was isolated using RNeasy
qRT-PCR was carried out with the real-time PCR detection system (The StepOnePlus
Kaplan-Meier survival analysis
Prognostic association of gene expression with relapse free survival of breast cancer patients was evaluated using online database (
Bcgenexminer analysis
To evaluate the prognostic value of selected genes, published annotated transcriptomic data from Breast Cancer Gene-Expression Miner v4.4 (
Receiver operating characteristic (ROC) plot analysis
To validate predictive value of selected genes, their expression was correlated with 5-year relapse free survival of breast cancer patients in response to chemotherapeutic drugs taxane and anthracycline using the online validation tool ROC plotter which links gene expression and response to therapy using transcriptome of breast cancer patients [29]. Patients were segregated into response cohorts with the outcome of survival without disease (responders) and disease relapse before 5 years after therapy (Non-responders). Area Under Curve (AUC) between 0.6–0.7 for a gene represents a predictive marker of potential clinical utility. The
Bioinformatic in silico analysis
The mRNA expression data from 1700 breast cancer samples was downloaded from cBioportal.org [30]. The z score (i.e. a value of 0 corresponds to the mean gene expression of that gene and
ChIP-Seq analysis
For ChIP-Seq analysis data was downloaded from NCBI-SRA database (PRJNA311514). After pre-processing, the raw reads were aligned to the human genome (hg19) with Bowtie2 using default options. Peak calling was performed with MACS2 using only unique mapping and non-duplicated reads from each ChIP sample and input control.
Results
Absence of expression of estrogen receptor and non-amplification of HER2 receptor is associated with the aggressiveness and chemoresistance of breast cancer [14, 15]. To investigate the role of protein tyrosine phosphatases in breast cancer aggressiveness and chemoresistance, we first conducted an expression analysis of 114 PTP genes in tumor samples of 1700 breast cancer patients derived from TCGA dataset [30]. Expression profiles were plotted as heatmap for PTP expression in the patients classified into four categories on the basis of the 3-gene classifier model: (1) ER-/HER2- (which included mostly the basal and claudin-low subtypes) (2) ER+/HER2- high proliferating and (3) ER+/HER2- low proliferating (both of which included mostly patients with luminal A or Luminal B subtypes) and (4) HER2+ (which included mostly HER2 enriched and normal-like subtypes). The number of patients belonging to these categories and their corresponding PAM-50 classifiers is enlisted in Supplementary Table 2. The heatmap showed 3 signature clusters with PTP genes that were specifically either under or over-expressed in ER-/HER2- subtype in stark contrast to other subtypes of breast cancer patients (Fig. 1). We called these signature clusters as either DR-cluster or UR-cluster depending on whether the genes were downregulated or upregulated, respectively. Twelve genes i.e. DUSP4, DUSP5, DUSP16, FIG4, GAK, INPP4B, PTEN, PTP4A2, PTPN13, PTPRT, SSH3 and SYNJ2 were found to be specifically downregulated in ER-/HER2- tumor samples of Breast Cancer patients while 11 genes i.e. CDC25A, CDC25B, CDC25C, CDKN3, EYA3, INPP4A, PTP4A3, PTPN2, PTPN14, SSU72 and UBASH3B were found to be upregulated. Notably, DR-cluster included PTEN, which is a well-established tumor suppressor gene. Also, the UR-cluster presented certain cell cycle regulatory genes CDC25A, CDC25B and CDC25C suggesting an oncogenic potential of upregulated UR-cluster genes. The expression of each of the genes of DR-cluster and UR-cluster in tumor samples of ER-/HER2- breast cancer patients was compared with the other 3 subtypes using violin plots (Fig. 2). By performing ANOVA statistical analysis, we confirmed that these genes showed statistically significant differential expression in ER-/HER2- breast cancer patients compared to other subtypes (Supplementary Table 3).
Gene expression analyses of protein tyrosine phosphatases (PTP) in tumor samples of breast cancer patients pinpoint 3 gene-clusters specifically mis-expressed in ER-/HER2- subtype. Heatmap of PTP mRNA expression in tumor samples of breast cancer patients categorized as ER-/HER2-, ER+/HER2- HP, ER+/HER2- LP and HER2+ subtypes based on 3-gene classifier. The patient dataset is derived from METABRIC, TCGA, cbioportal and their categorization in molecular subtypes is provided in Supplementary Table 2. Genes falling in cluster 1 (6 genes) and cluster 2 (6 genes) show reduced expression in ER-/HER2- subtype, designated as DR-cluster genes while genes of cluster 3 (11 genes) display enhanced expression in ER-/HER2- subtype, designated as UR-cluster genes.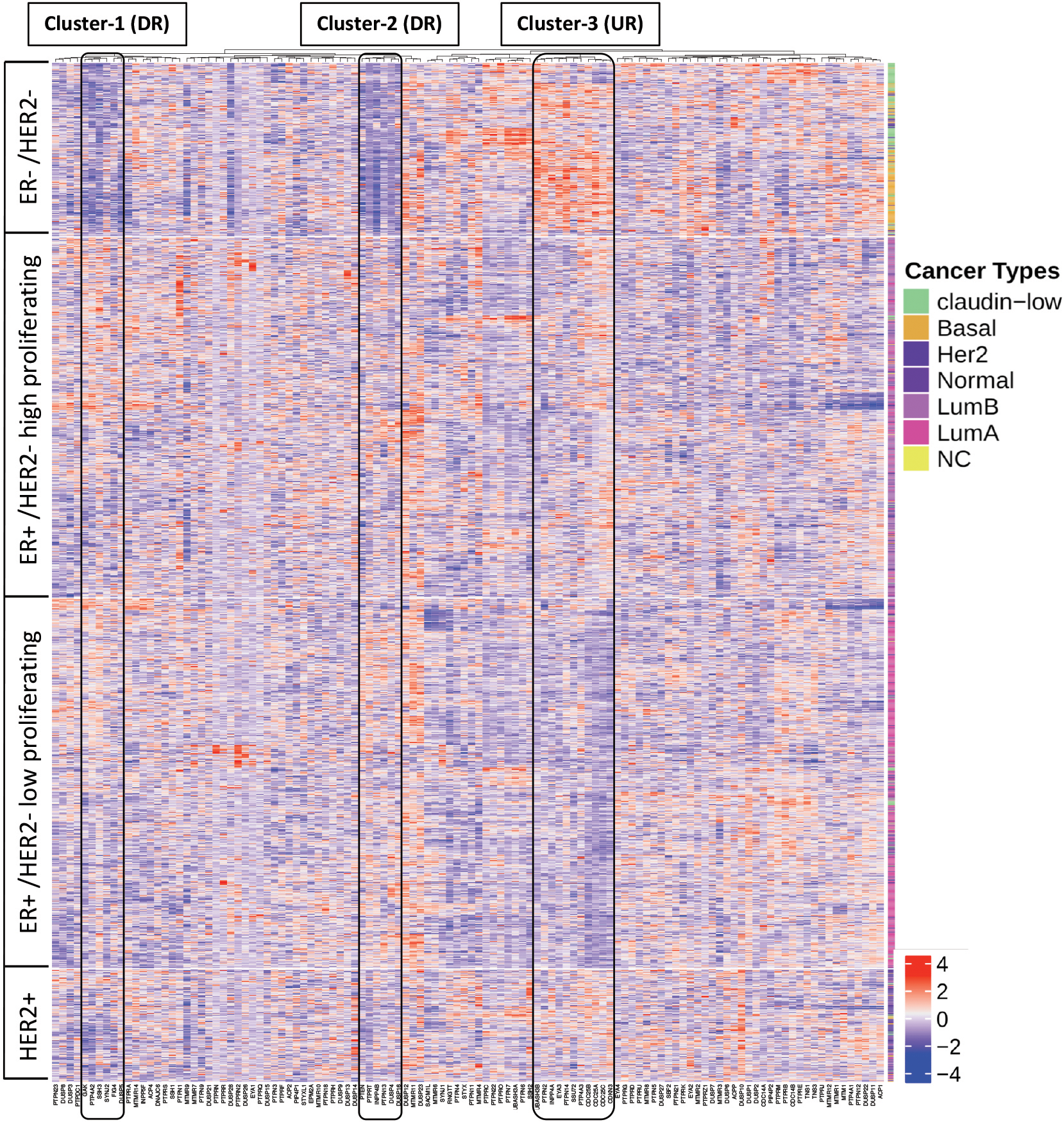
Violin plots demonstrate comparison of expression values (z-score) of DR-cluster and UR-cluster genes in tumor samples of four molecular subtypes of breast cancer patients. a. Significant reduced expression of twelve genes of DR-cluster in ER-/HER2- tumor samples compared to other categories is observed. b. Eleven genes of UR-cluster show significant increased expression in ER-/HER2- breast cancer patients in comparison to other subtypes. The statistical significance among various groups is determined by ANOVA test using Tukey multiple comparisons of mean and 95% family-wise confidence level and the 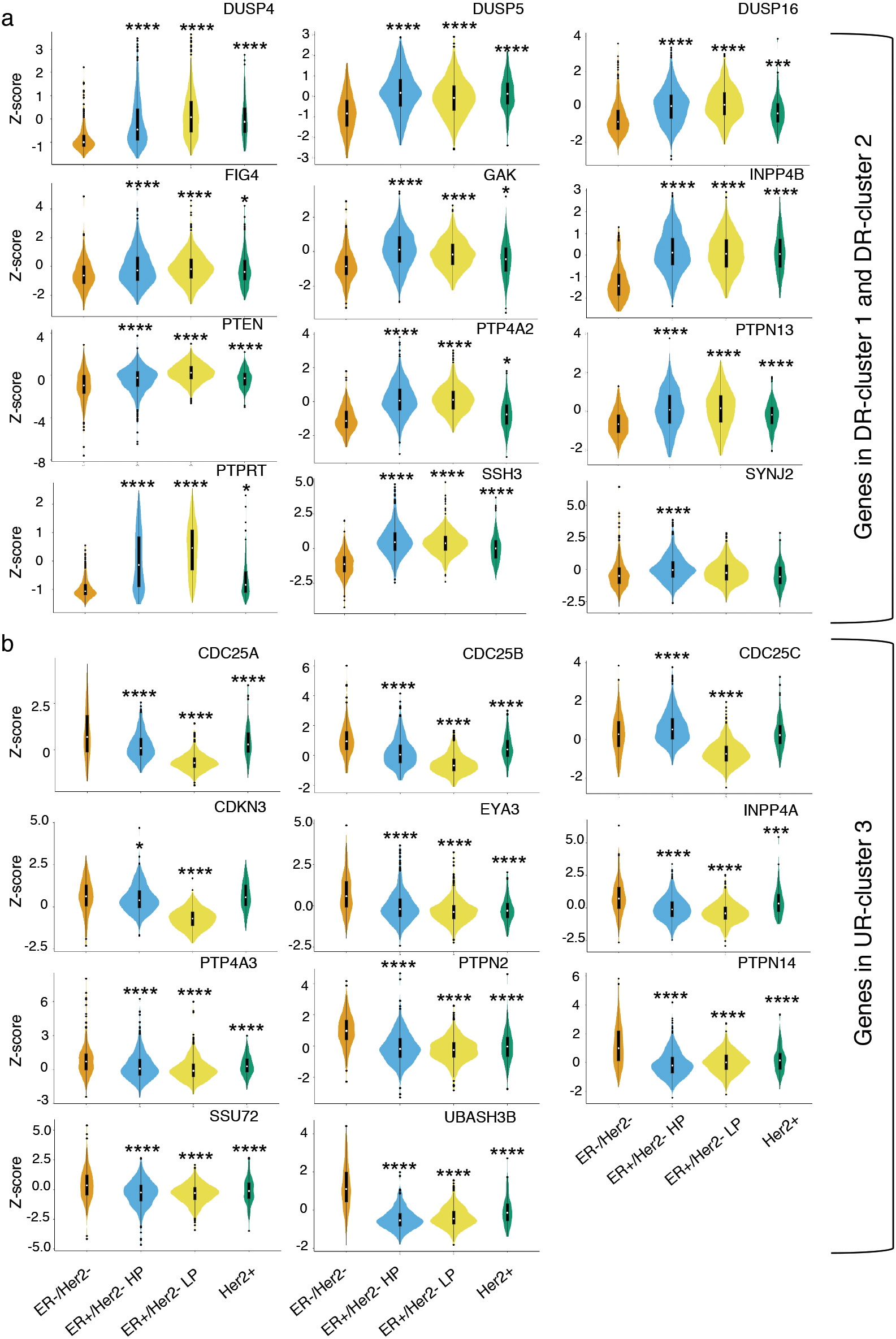
It is well established that hormone independent breast cancer shows enhanced chemoresistance [14, 15]. Hence, we next investigated potential correlation of DR/UR-cluster genes with chemoresistance by evaluating their expression in breast cancer cells resistance to chemotherapeutic agents. We chose a publicly available GEO dataset “GSE68815” in which MCF7 cells were intermittently treated either with paclitaxel or adriamycin, drugs commonly used for treating hormone independent breast cancer, to generate paclitaxel and adriamycin chemoresistant MCF7 cells [31]. We analyzed RNA-sequencing data of these cells and generated a PTP specific expression heatmap, comparing paclitaxel-resistant MCF7 (MCF7-PTX) and adriamycin-resistant MCF7 (MCF7-ADM) cells with untreated chemosensitive MCF7 (MCF7-untr) (Fig. 3a). A similarly altered expression pattern of a gene in both paclitaxel and adriamycin resistant cells in contrast to the untreated cells will suggest that the particular gene may have a role in chemoresistance pathways. Even though these analyses highlighted multiple PTP genes, we focused on ER-/HER2- specific DR/UR gene clusters. By comparing read counts of DR/UR cluster genes in chemoresistant versus sensitive cells, we found that 6 genes (CDC25A, CDC25C, EYA3, INPP4A, PTPN14, UBASH3B) of the UR-cluster showed consistent overexpression in both the paclitaxel-resistant MCF7-PTX and adriamycin-resistant MCF7-ADM cells in comparison to sensitive untreated MCF7(Supplementary Fig. 1a). While only 3 genes (DUSP16, FIG4, SSH3) from the DR-cluster showed a consistent under-expression in both chemoresistant cells(Supplementary Fig. 1b). Thus, 9 PTP genes were narrowed down which showed differential expression in ER-/HER2- patient samples and their differential expression patterns correlated with chemoresistance of breast cancer cells to paclitaxel and adriamycin. We next questioned the clinical utility of these genes as prognostic biomarkers for disease recurrence in breast cancers. High/low expression of these genes above/below a median value was correlated with 5-year relapse free survival (RFS) of breast cancer patients using Kaplan Meier survival curves (Fig. 3b). A hazard ratio cut-off of less than 0.7 or more than 1.3 was selected as a significant threshold. We found that downregulation of DUSP16 and upregulation of CDC25A and CDC25C significantly associated with poor relapse free survival, identifying them as potential prognostic markers for disease relapse/recurrence in breast cancers.
Identification of PTP genes of DR/UR clusters that are associated with chemoresistance to paclitaxel and adriamycin and show poor prognosis in breast cancer patients. a. Heatmap of PTP mRNA expression (z-score) in chemoresistant MCF7 cells resistant to paclitaxel (PTX) and adriamycin (ADM) compared to sensitive untreated MCF7 cells. Nine PTP cluster genes showed similar expression patterns in chemoresistant MCF7 cells contrasting to chemosensitive MCF7. b. Kaplan Meier survival plots analyzing prognostic value of nine candidate PTP genes by correlating their gene expression with the probability of relapse free survival of patient groups expressing high or low levels of these genes. A hazard ratio (HR) cut-off value of less than 0.7 correlates lowly expressed gene DUSP16 with poor prognosis, whereas HR 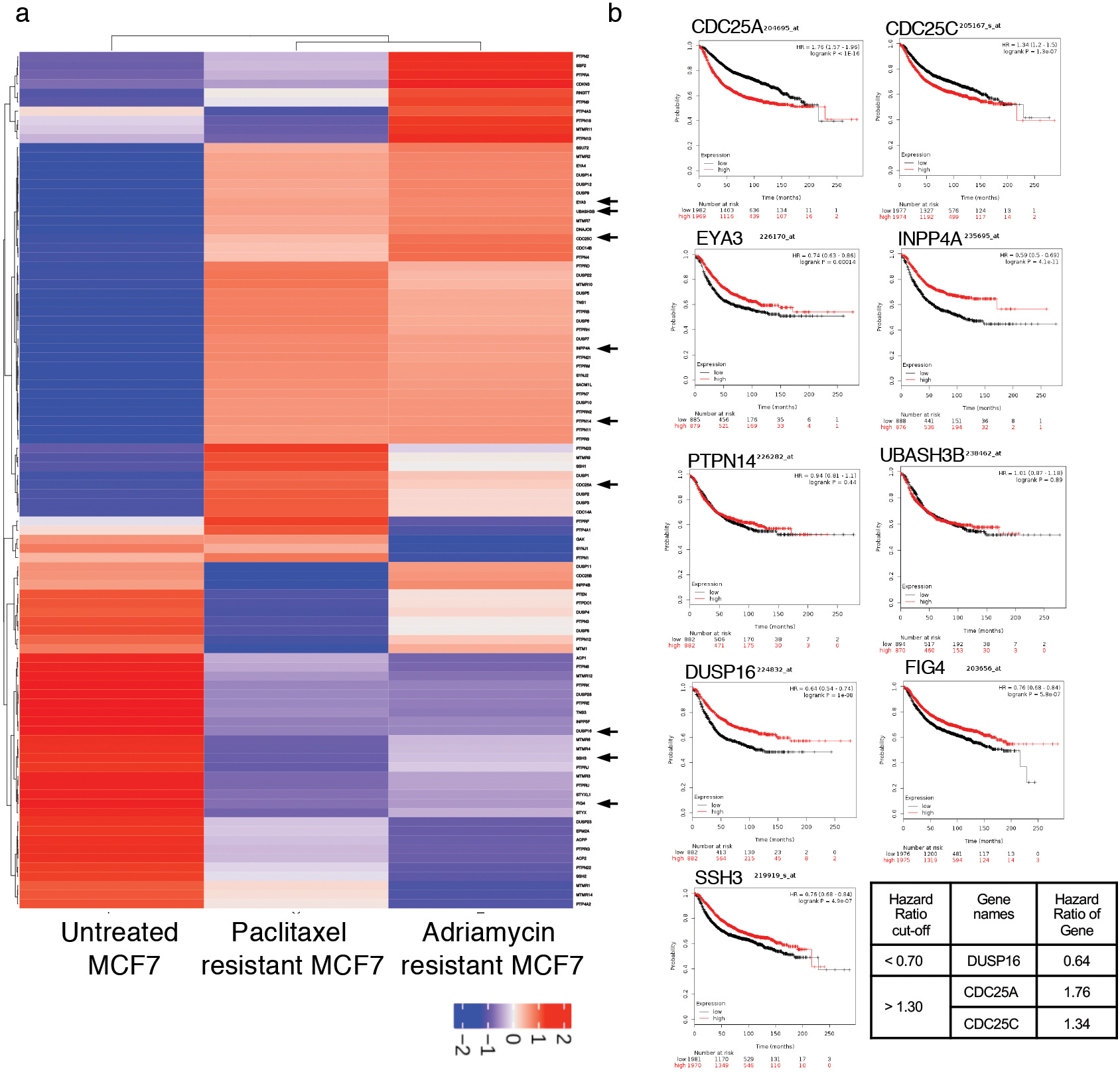
CDC25C is a robust prognostic marker for metastatic relapse free survival of breast cancer patients as evaluated by Nottingham prognostic index. a–c. Kaplan Meier survival plots of metastatic relapse free survival associates with mRNA expression of CDC25C, CDC25A and DUSP16. Adjusted multivariate Cox analysis with respect to Nottingham prognostic index identifies CDC25C as a significant prognostic marker (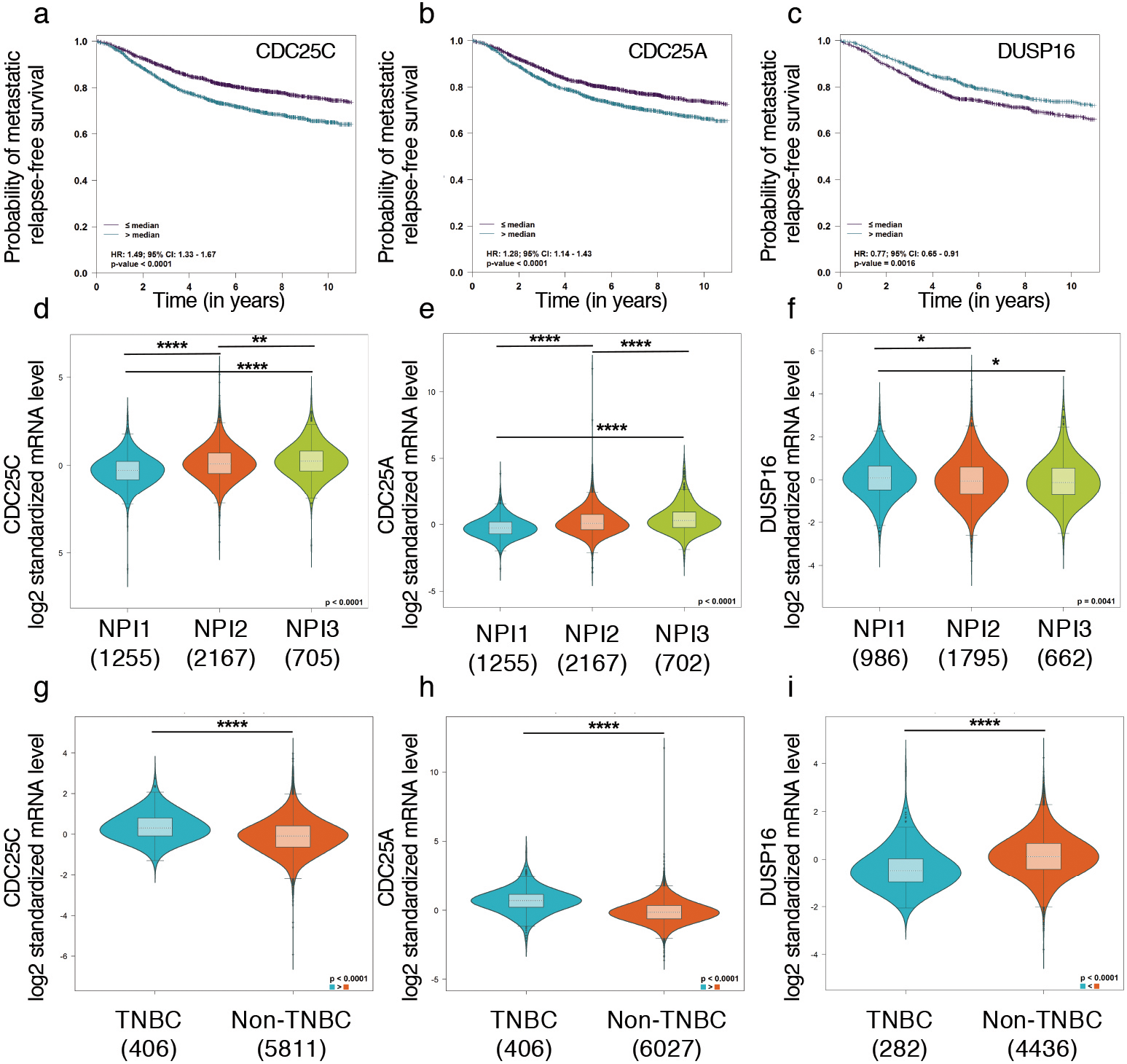
Increased expression of CDC25A and CDC25C in ER-/HER2- cells is mediated by enrichment of histone H3K4me3 active marks at respective promoter sites. a–c. Represent fold change in mRNA expression of CDC25A, CDC25C and DUSP16, respectively, in MDA-MB-231 (ER-/HER2-, TNBC claudin low) cells using MCF7 (ER+HER2-LP, Luminal A) cells as control. GAPDH was used as internal loading control and the expression values are plotted as mean 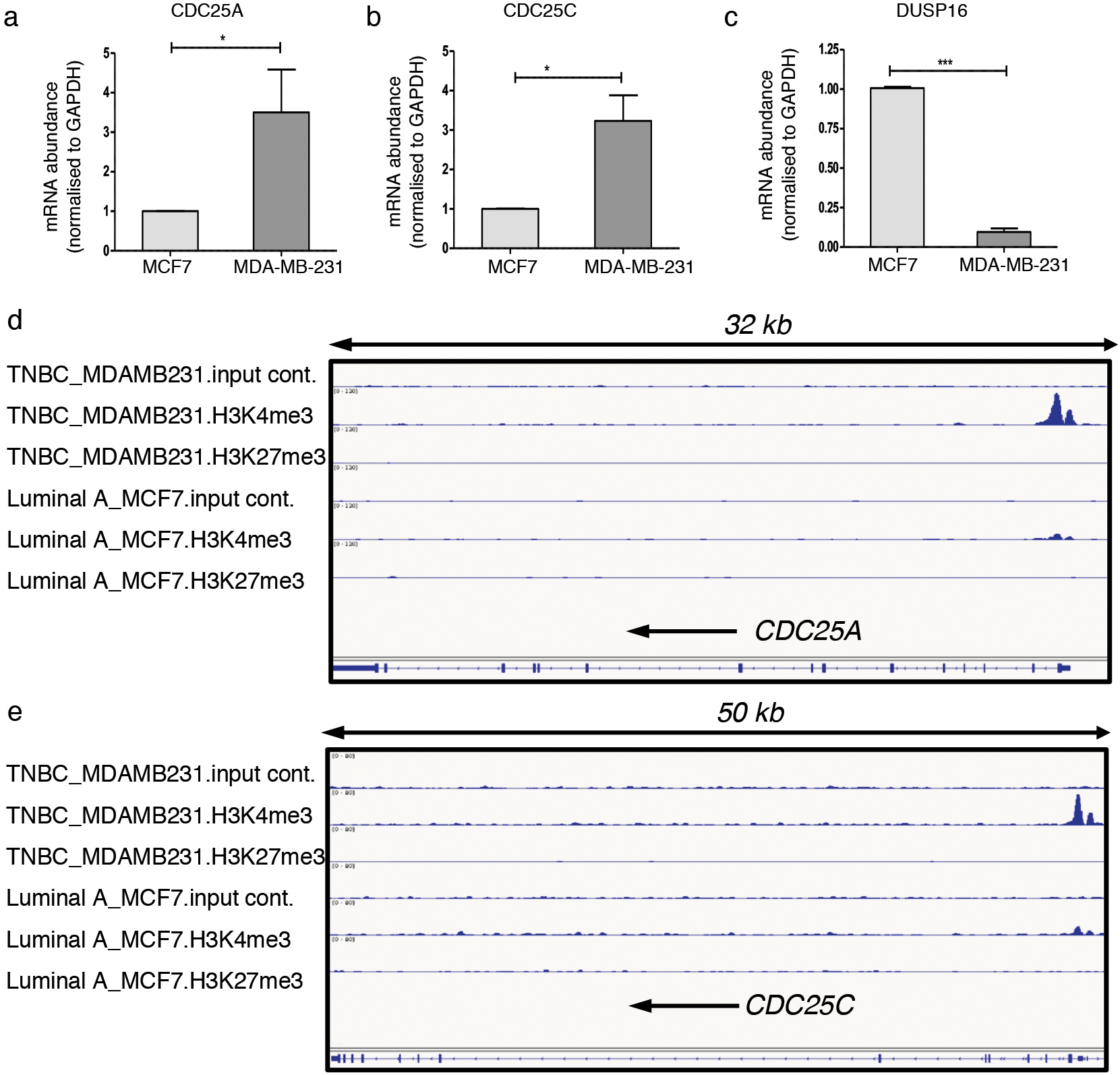
Next, we validated the prognostic significance of these 3 potential markers in large pooled patient cohort by BC Gene-Expression Miner (bc-GenExMiner v4.0). Using DNA microarray data of pooled cohort of patient samples, expression of CDC25C, CDC25A and DUSP16 was correlated with the probability of metastasis relapse free patient survival. Univariate Cox proportional hazard model analysis demonstrated that increased expression of CDC25C and CDC25A above the median value reduced the probability of metastasis relapse free survival and thus showed poor prognosis with respect to metastatic relapse (Fig. 4a and b). Expression of DUSP16 lower than median value also showed poor prognosis and associated with increased metastatic relapse in patients (Fig. 4c). To have a better estimate of prognostic impact of these markers relative to well-established clinical prognostic index “The Nottingham Prognostic Index” (NPI), adjusted multivariate cox analysis on pooled patient cohorts with available NPI data was performed. The NPI index takes into consideration the tumor size, lymph node involvement and the grade of breast tumor. This analysis established CDC25C as a robust prognostic marker for metastatic relapse-free patient survival based on Nottingham prognostic index (
We next assessed the mechanism regulating transcription of these genes in ER-/HER2- cells. Gene transcription is regulated either by transcriptional machineries along with the RNA polymerase activity or by epigenetic regulation such as histone methylation at the chromatin. We tested epigenetic-meditated transcriptional regulation by analyzing enrichment of H3K4me3 and H3K27me3 histone modifications at the promoter and gene body region of selected genes. Enriched methylation at the lysine 4 of histone H3 activates gene transcription while enrichment of H3K27me3 mediates transcriptional repression. We analyzed publicly available chromatin immunoprecipitation-sequencing (ChIP-seq) data (GEO accession no. GSE77772) that used H3K4me3 and H3K27me3 antibodies in MDA-MB-231 (TNBC; ER-/HER2-) cells and compared with MCF7 (Luminal A; ER+/HER2-) cells. We observed that the activating mark H3K4me3 was highly enriched at the promoter and gene body of CDC25A and CDC25C in MDA-MB-231 cells in comparison to MCF7 cells (Fig. 5d and e). No enrichment of H3K27me3 above the input control on the gene promoter of CDC25A and CDC25C was observed both in MDA-MB-231 and MCF7 (Fig. 5d and e). This suggests that transcription of CDC25A and CDC25C is regulated epigenetically by histone methylation at the lysine 4 of Histone H3. We did not observe any histone methylation mediated epigenetic regulation of DUSP16 in the tested cell lines. We thus ascertain that CDC25A and CDC25C are epigenetically regulated at transcriptional level in breast cancer cells and with a higher degree in ER-/HER2- MDA-MB-231 cells.
Patient data and statistical analysis of ROC plots as plotted in Fig. 6, evaluating predictive value of CDC25A, CDC25C and DUSP16 in response to taxane and anthracycline therapy in breast cancer
CDC25C is a predictive molecular marker for poor clinical outcome in breast cancer patients after treatment with taxane and anthracycline. The patient cohorts were treated with chemotherapeutic drugs and the response outcomes were determined by relapse free survival over 5 years (ROCplot transcriptomic dataset). (I) CDC25A (a) and CDC25C (b) expression was significantly higher in non-responders to taxane when compared to responders. Low expression of DUSP16 (c) was observed in non-responders but was not statistically significant. d–f. Represent ROC plots illustrating the predictive biomarker value of CDC25A, CDC25C and DUSP16 in response to taxane therapy. (II) CDC25C (h) expression was significantly higher in non-responders when compared to responders to anthracycline whereas was insignificant for CDC25A (g) and DUSP16 (i). j–l. Represent ROC plots illustrating the predictive biomarker value of CDC25A, CDC25C and DUSP16 in response to anthracycline therapy. Statistical significance was calculated by Mann-Whitney test and is provided in Table 1.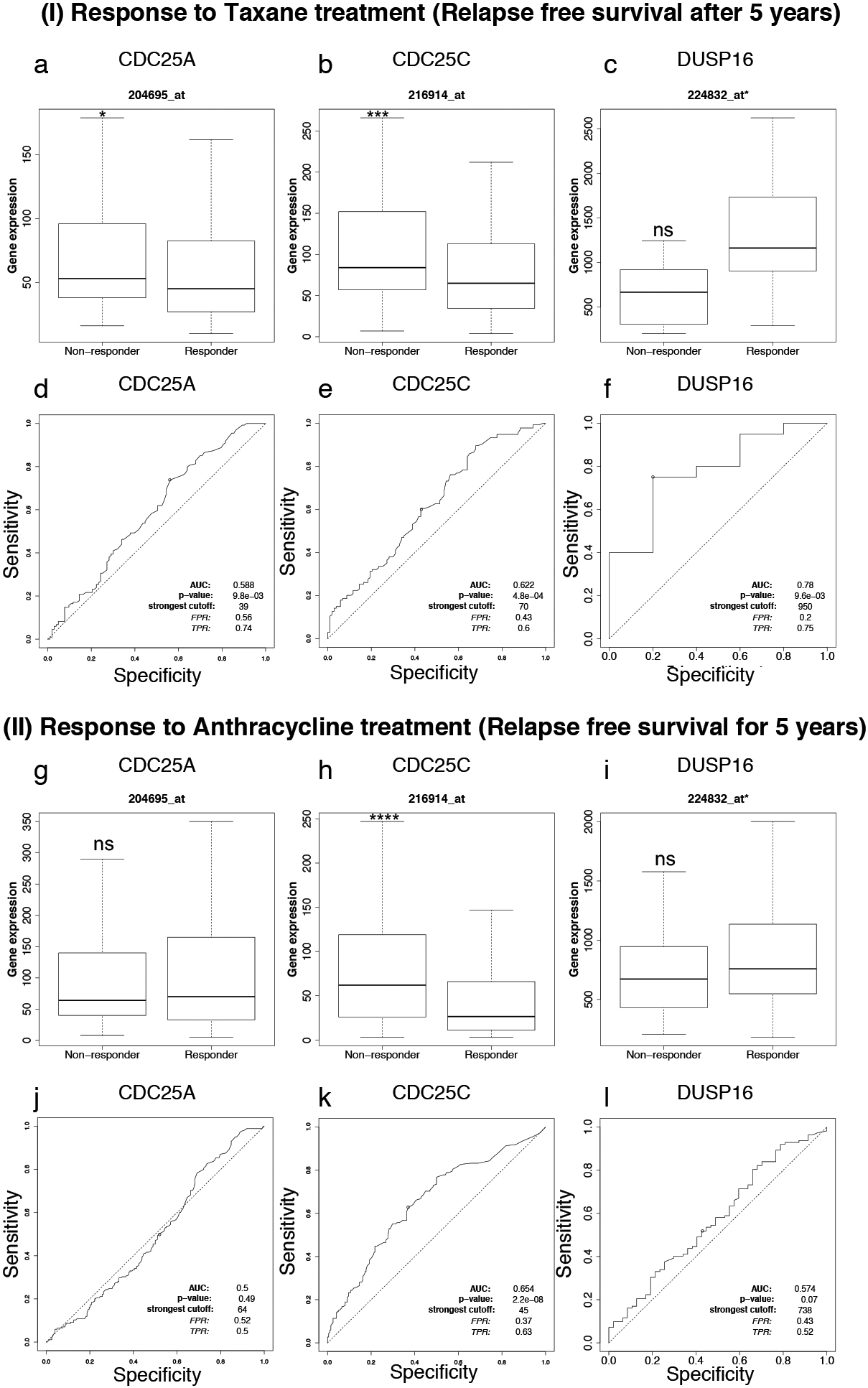
Our study identified CDC25A, CDC25C and DUSP16 as prognostic markers in breast cancer and also showed their altered expression in ER-/HER2- breast cancer subtype and correlated it with chemoresistance to taxane based (Paclitaxel) and anthracycline based (Adriamycin) drugs. Next, we questioned the predictive value of these prognostic markers in response to drug treatment to explore their clinical utility. To do so, a largescale study of validating predictive markers in breast cancer after various therapies was used as the resource database [29]. The patient cohorts were treated with taxane or anthracycline and subdivided into two groups of responders and non-responders. The response outcome was based on the relapse free survival (RFS) within a recovery period of 5 years for responders whereas, patients whose disease relapsed after the drug treatment within a period of 5 years were designated as non-responders. The expression of these genes was evaluated in responders and non-responders and plotted as boxplots (Fig. 6a–c, g–i). The fold change and Mann-Whitney
In this study we identify CDC25C as a prognostic and predictive biomarker for aggressiveness, chemoresistance to taxane and anthracycline drugs and associated disease relapse in breast cancers.
Protein tyrosine phosphatases are major regulators of signalling pathways that control the essential cellular processes such as survival, proliferation, apoptosis and division. Mis-regulation of these key regulators contributes to cancer development and progression. In breast cancer, the role of protein tyrosine phosphatases and their correlation with aggressiveness and therapy resistance is sparsely studied. It is reported that the aggressiveness and development of resistance to chemotherapeutic drugs in breast cancer partially depends on the status of hormone receptors ER and HER2. Keeping this in view, we performed an integrative in silico genomic analysis of protein tyrosine phosphatases in ER-/HER2- subtype of breast cancers and compared it with other subtypes to identify potential candidates that might have prognostic and predictive potential. By constructing an expression heatmap of PTP genes, we identified 3 clusters that were specifically mis-regulated in ER-/HER2- subtype of breast cancer. A detailed analysis of the effect of the expression of these cluster genes on development of chemoresistance and relapse free survival of patients identified 3 candidate genes namely CDC25C, CDC25A and DUSP16.
CDC25A and CDC25C are threonine and tyrosine dual specificity phosphatases that regulate G2/M transition by acting on CDKs/cyclins [32]. Notably, overexpression of CDC25A has been described in eleven different cancers and it is also associated with aggressiveness and poor prognosis [32, 33]. In our study, we also observe enhanced expression of CDC25A specifically in ER-/HER2- breast cancer cells and patient tumor samples. Even though enhanced expression of CDC25A correlated with poor relapse free survival of patients, still it was identified as low potential NPI prognostic and predictive marker for chemotherapy in patients. Although previous studies discuss the overexpression of CDC25A in a subpopulation of breast cancer cells, association of CDC25C with breast cancer was not reported. Here, by performing focussed analysis of protein tyrosine phosphatases in hormone receptor deficient, most aggressive subtype of breast cancer i.e. ER-/HER2- patients, we identified specific overexpression of CDC25C in this subtype compared to hormone dependent subtypes. Consistent with our study, another report in vulvar carcinomas associated CDC25C overexpression with increased metastasis and aggressiveness of the disease [34]. We further established CDC25C as a poor prognostic marker for breast cancer that may predict response to chemotherapy such as therapy with taxane and anthracycline. Thus, our study opens up a new avenue for exploring clinical utility of CDC25C as a predictive marker for 5-year relapse free survival of patients after the completion of chemotherapy. Apart from chemotherapeutic drugs, target specific therapeutic agents such as HER2-targeted antibody-drug conjugate immunotherapy for HER2 positive BCs, PARP inhibitors for BRCA1/2 mutated BCs and CDK4/6 inhibitors for hormone receptor-positive, HER2-negative metastatic breast cancers have been approved by Food and Drug Administration for specific subtypes of breast cancers. In this study, we also analysed biomarker value of CDC25C to predict response to Trastuzumab treatment, however due to small sample size, its role in Trastuzumab treatment could not be established at this point. A larger patient cohort will be required to clearly understand its predictive value upon Trastuzumab treatment. Also, further studies are required to investigate the predictive value and clinical utility of CDC25C for other targeted therapies for metastatic breast cancer.
Mechanistically, CDC25C has been implicated in increasing genomic instability in breast cancer. Phosphorylation of serine 214 of CDC25C activates the protein prematurely and leads to faster mitotic entry without appropriate DNA repair [35]. This will lead to accumulation of genetic instability, which is one of the characteristic features of cancer cells. Thus, specific catalytic inhibitors for CDC25C may have a potential therapeutic role in breast cancer and more specifically towards basal and claudin-low types, which are mostly ER-/HER2-. Another way to target CDC25C is to identify how is it transcriptionally regulated. We identify that overexpression of CDC25C in ER-/HER2- cells is epigenetically regulated. We find enrichment of H3K4me3 histone marks on the promoter of CDC25C, a positive regulator of its transcription. Moreover, we also find that CDC25A is also regulated epigenetically by enrichment of transcriptional activation histone mark H3K4me3. This highlights that inhibiting H3K4 methylation by way of small molecule epigenetic inhibitors could be a potential therapeutic approach in case of ER-/HER2- breast cancers, in which a potential targeted therapy is still missing. Thus, to summarise, our study identifies CDC25A and CDC25C as markers for aggressiveness and chemoresistance in breast cancer.
Out of the identified DR-cluster genes, downregulation of DUSP16 was seen in ER-/HER2- tumor samples of breast cancer patients and also correlated with chemoresistance. However, due to small sample size, its value as a predictive biomarker in response to taxane drug was not clear. As such, expression of this gene did not show any correlation with the relapse free survival of patients that are being treated with anthracycline drug. DUSP16 is a critical component of mitogen-activated protein kinase (MAPK) pathway and functions by dephosphorylating p38 [36]. In coherence with our study, a recent report showed that repression of DUSP16 is observed in chemotherapy resistance to paclitaxel, a taxane based drug [37]. They further show that this repression is mediated by hypoxia induced transcriptional repressor REST. In this study, we explored the possibility of epigenetic mediated transcriptional repression of DUSP16 by analysing H3K27me3 enrichment on its promoter or gene body, however no epigenetic transcriptional regulation of DUSP16 gene in breast cancer cells was observed. Thus, it is likely that transcriptional downregulation of DUSP16 in breast cancer is majorly governed by transcription factors such as REST and others.
Thus, with this study, we suggest that re-analysing large scale patient genomic, transcriptomics datasets raising a specific question is a great tool that could pave ways to new discoveries and identify novel molecular therapeutic targets and prognostic/predictive biomarkers for cancers.
Author contribution
P.A. and M.K. conceptualized and supervised the study. R.T., N.N. and P.K. performed and analyzed the experiments. R.T., N.N., M.K. and P.A. prepared the manuscript. M.K. and P.A. revised the final manuscript with inputs from all authors. All authors have approved the final version of manuscript submitted for publication.
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-200858.
Footnotes
Acknowledgments
We are thankful to Prof. Bhudev C. Das and Prof. Simran Tandon for sharing cell lines and their continued support. This work was partly supported by Department of Biotechnology “DBT-BioCare fellowship” (BT/PR18554/BIC/101/259/2016), “DBT-RLS fellowship” (BT/RLF/Re-entry/16/2016) and Department of Science and Technology “DST-SERB ECR Award” (ECR/2017/000857) to P.A. and “DST-SERB YSS” (YSS/2015/000207) award to M.K.
Conflict of interest
The authors declare no conflict of interests.
