Abstract
BACKGROUND:
Cervical cancer (CC) is the most common form of gynecological malignancy. Long intergenic non-protein coding RNA 858 (LINC00858) has been identified to participate in multiple cancers. However, the role and mechanism of LINC00858 in CC cells are still elusive.
AIM:
The aim of this study is to explore the biological functions and mechanisms of LINC00858 in CC cells.
METHODS:
RT-qPCR analysis was used to examine the expression of LINC00858 in CC cells. EdU and colony formation assay were utilized to assess cell proliferation. TUNEL assay and flow cytometry assay were conducted to assess cell apoptosis. The mechanism regarding LINC00858 was certified through RNA pull down, RIP and luciferase reporter assays.
RESULTS:
The up-regulated LINC00858 was detected in CC cells. Reduction of LINC00858 effectively subdued CC cells proliferation and stimulated cell apoptosis. LINC00858 was determined to bind with miR-3064-5p and up-regulate VMA21 in CC cells. In rescue assays, miR-3064-5p down-regulation and VMA21 up-regulation were able to counteract the effect caused by LINC00858 decrease on CC cell proliferation and apoptosis.
CONCLUSION:
LINC00858 enhances cell proliferation, while restraining cell apoptosis in CC through targeting miR-3064-5p/VMA21 axis, implying that LINC00858 may serve as a promising therapeutic target for CC.
Introduction
Cervical cancer (CC) is the most common form gynecological malignancy [1]. Viral infection and high risk type HPV persistent infection are the main risk factors for developing CC [2, 3]. Thus, HPV vaccine becomes a key therapeutic method for CC [4]. Chemotherapy, immunotherapy and the targeted therapy combining noncoding RNAs, especially lncRNAs and miRNAs, are commonly used in recent years, except for surgery, radiation and platinum-based therapy [5, 6, 7]. However, the incidence of CC is rising steadily and the patients are getting younger, thus new and other potential treatment methods are still in need. In this study, we tried to discover a novel lncRNA that has an association with CC.
Long non-coding RNAs (lncRNAs) appear to be the vital modulators for various tumors or diseases such as lung cancer and breast cancer [8, 9, 10]. On this basis, the underlying lncRNAs which affects CC have been explored nowadays [9, 11]. For instance, SNHG1 overexpression is conductive to promote cell proliferation and migration of CC [12]. Both PANDAR and ZFAS1 up-regulations were connected with short survival time of CC patients [13, 14]. Similarly, massive reports indicated that lncRNAs could competitively regulate mRNAs by acting as molecular sponges of miRNAs to impact the development of multiple cancers, including CC. For example, lncRNA XIST has been identified to up-regulate FUS through competitively sponging with miR-200a to further promote the occurrence of CC [15]. LncRNA-CTS/miR-505/ZEB2 axis has been proved by Feng et al. to regulate CC progress [16]. However, the role and molecular mechanism of LINC00858 in CC have not been probed yet.
In terms of this project, we intended to quantify the expression of LINC00858 in CC cells, explore its influence on cell proliferation and apoptosis, as well as delve into its mechanism in CC cells. Our study may be conducive to finding out potential therapy targets for CC treatment.
Materials and methods
Totally, 12 experiments were conducted, including cell culture, real-time quantitative polymerase chain reaction (RT-qPCR), cell transfection, 5-Ethynyl-2’-deoxyuridine (EdU) assay, colony formation, Terminal deoxynucleotidyl transferase (TdT) dUTP Nick-End Labeling (TUNEL) assay, flow cytometry analysis, subcellular fractionation, fluorescence in situ hybridization (FISH), luciferase reporter assay, RNA immunoprecipitation (RIP), and RNA pull down assay. Three bio-repeats of each experiment were run in triplicate. Human normal cervical epithelial cells and four CC cell lines were involved in this study. In terms of the workflow of this research, we intended to dig into the effects of LINC00858 on proliferative and apoptotic abilities of CC cells at the very beginning. Subsequently, this study tried to figure out how LINC00858 exerted its influences on biological behaviors of CC cells. By reviewing previous studies and utilizing bioinformatics tools, the potential mechanism of LINC00858 was inferred, and further testified by mechanism assays. The detailed procedures of different methods were depicted as followed.
Cell culture
Human CC cell lines (CaSki, SiHa, ME180, MS751) and human normal cervical epithelial cells (Ect1/E6E7) in our study, were all procured from ATCC (Manassas, VA, USA). The biosafety level (BSL) of all involved cell lines was 2. 10% fetal bovine serum (FBS) and 1% penicillin/streptavidin served as the supplements of RPMI-1640 medium (all; Gibco, Grant Island, NY, USA). All cells were cultivated in the above medium at 37
RT-qPCR
The total RNA, which was acquired for cDNA synthesis, was first isolated from the cultured cell samples by employing TRIzol Reagent (Invitrogen, Carlsbad, CA, USA), in line with the provided instructions. Then cDNA synthesis was conducted with PrimeScript Reverse Transcriptase Kit (Takara, Kyoto, Japan). RT-qPCR was then followed in the presence of SYBR Green PCR Kit (Takara) for determining gene expression, and 2
Cell transfection
The shRNAs specifically targeting LINC00858 (sh-LINC00858#1 and sh-LINC00858#2), VMA21 (sh-VMA21#1 and sh-VMA21#2), and control-shRNAs (sh-NCs) were acquired commercially from GenePharma (Shanghai, China). The miR-3064-5p mimics, miR-3064-5p inhibitor and the corresponding controls were synthesized by Ribobio (Guangzhou, China). The full-length cDNA sequence of VMA21 was cloned into pcDNA3.1 vector (Invitrogen) for overexpressing VMA21, and the empty vector acted as the negative control. All these plasmids were separately transfected into cells of ME180 and MS751 for 48 hours, with the help of Lipofectamine 3000 (Invitrogen).
EdU assay
The ME180 and MS751 cells acquired from transfection assay were washed in PBS for conducting EdU assay with BeyoClick
Colony formation assay
The transfected ME180 and MS751 cells were reaped and then planted into 6-well plates with 500 cells in each well. Following incubation for 14 days, the resulting colonies were treated with 4% PFA for 30 min and with 0.5% crystal violet solution for 5 min. Finally, the visible colonies in three separately conducted assays were manually counted.
TUNEL assay
The transfected cells of ME180 and MS751 were collected for rinsing with PBS and fixing with 4% PFA. After permeabilization with 0.1% TritonX-100, the apoptotic cells were stained using TUNEL reagent, as per the established method (Merck KGaA, Darmstadt, Germany). The fluorescence microscopy (Olympus) was employed for detecting the stained cells of three individually conducted assays.
Flow cytometry assay
Apoptosis of transfected ME180 and MS751 cells was detected by the use of Annexin V/PI staining kit (Invitrogen) following the standard method. Cells were mixed respectively with the FITC-Annexin V and PI in Binding Buffer for 15 min in the dark. After that, apoptotic cells were analyzed by flow cytometer (BD Biosciences, Franklin Lakes, NJ, USA).
Subcellular fractionation
After being washed in PBS, cells of ME180 and MS751 were lysed in cell fractionation buffer, and then centrifuged to collect the supernatant. The nuclear pallet was lysed in cell disruption buffer and centrifuged. LINC00858 content was examined in cell cytoplasm and cell nucleus using RT-qPCR. GAPDH and U6 worked as the internal reference respectively for cytoplasm and nuclei.
FISH
The ME180 and MS751 cells were fixed using 4% PFA and digested using protease K for 10 min. Then, cell samples were washed twice in PBS for culturing with the specific FISH probe of LINC00858 in hybridization buffer. The DAPI staining was added on the following day. The processed cell samples of three separately performed assays were examined using Olympus fluorescence microscope.
Luciferase reporter assay
LINC00858 or VMA21 fragments which contained the wild-type or mutated miR-3064-5p binding sequences were synthesized and cloned into the pmirGLO dual-luciferase reporter vectors, following the established protocol (Promega, Madison, WI, USA). Thereafter, the constructs were co-transfected with miR-3064-5p mimics or NC mimics into cells of ME180 and MS751. After 48 hours, the dual-luciferase reporter assay system (Promega) was employed for analyzing luciferase activities. Renilla luciferase activity served as internal control.
RIP
Based on the instructions of Magna RIP
RNA pull down
The miR-3064-5p sequences which contained wild-type or mutated VMA21 interacting sites were biotinylated into bio-miR-3064-5p WT/MUT probes. For RNA pull down assay, the cell protein lysates were prepared using RIPA lysis buffer, and then mixed with magnetic beads and bio-NC (control group) or bio-miR-3064-5p WT/MUT probes. Relative RNA enrichment was studied by RT-qPCR.
Statistical analyses
Each data with its associated error bar was the mean
LINC00858 contributes to cell growth in CC. A. LINC00858 expression in CC cell lines (CaSki, SiHa, ME180 and MS751) and normal cervical epithelial cell line (Ect1/E6E7) was analyzed by RT-qPCR. B. Analysis of LINC00858 levels by RT-qPCR in ME180 and MS751 cells transfected with sh-LINC00858. C. The proliferation of ME180 and MS751 cells mediated by sh-LINC00858 was measured by EdU assay. D. Colony formation assay was performed to obverse the colony formation abilities of ME180 and MS751 cells. E and F. Under the knockdown of LINC00858, the cell apoptosis of ME180 and MS751 was detected via TUNEL assay and flow cytometry assay. The length of each bar represented the mean result, and error bar represented SD. At least three independent experiments were carried out. 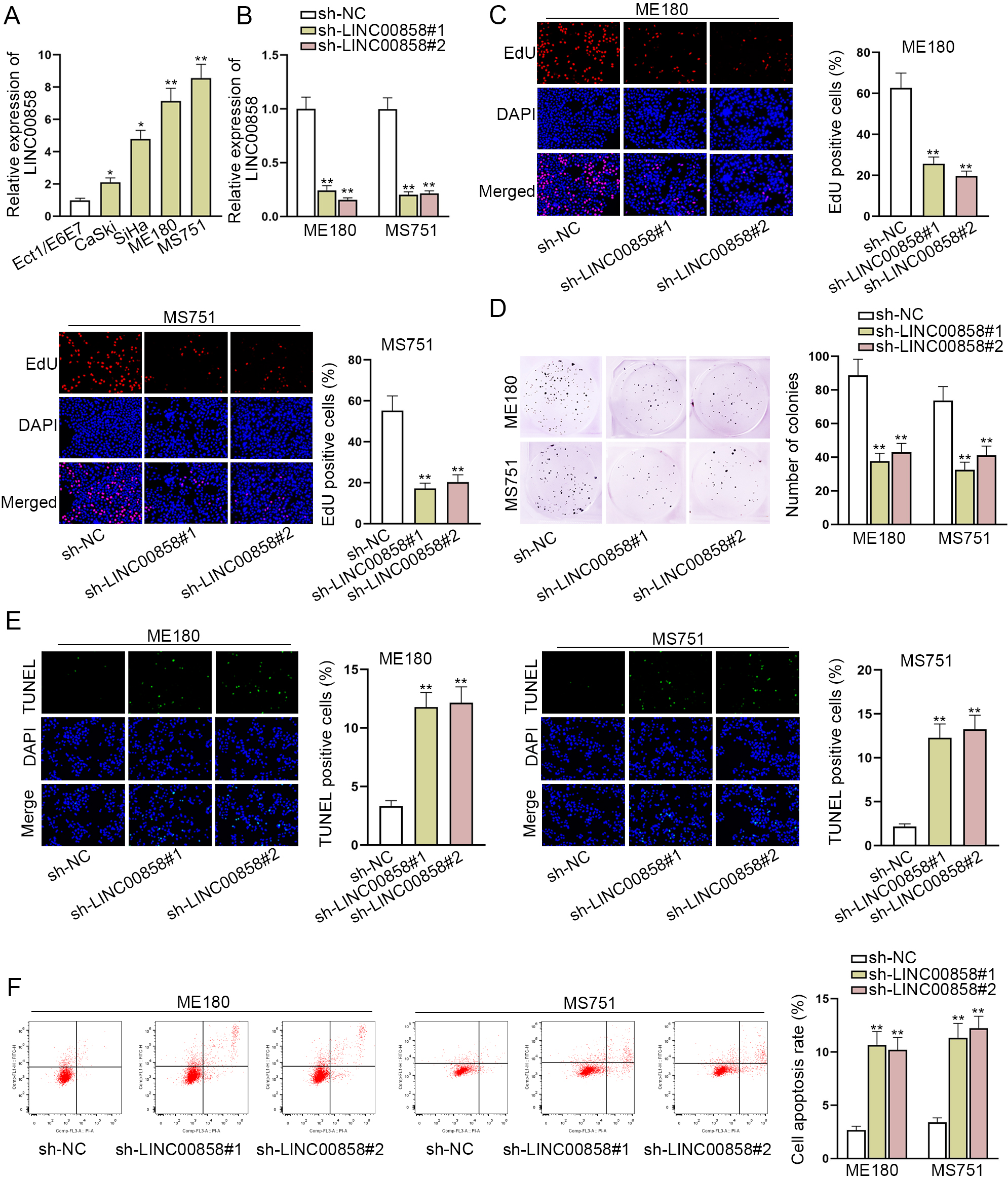
LINC00858 interacts with miR-3064-5p. A and B. Subcellular fraction assay and FISH assay were carried out to locate LINC00858 in CC cells. C. RT-qPCR was used to examine underlying miRNAs expression in CC cell lines and normal cell line. D. The binding sites between LINC00858 and miR-3064-5p were forecasted via starBase. E. The overexpression efficiency of miR-3064-5p mimics was assessed by RT-qPCR. F and G. Luciferase reporter assay and RIP assay were implemented to verify the combination between LINC00858 and miR-3064-5p in ME180 and MS751 cells. The length of each bar represented the mean result, and error bar represented SD. At least three independent experiments were performed. 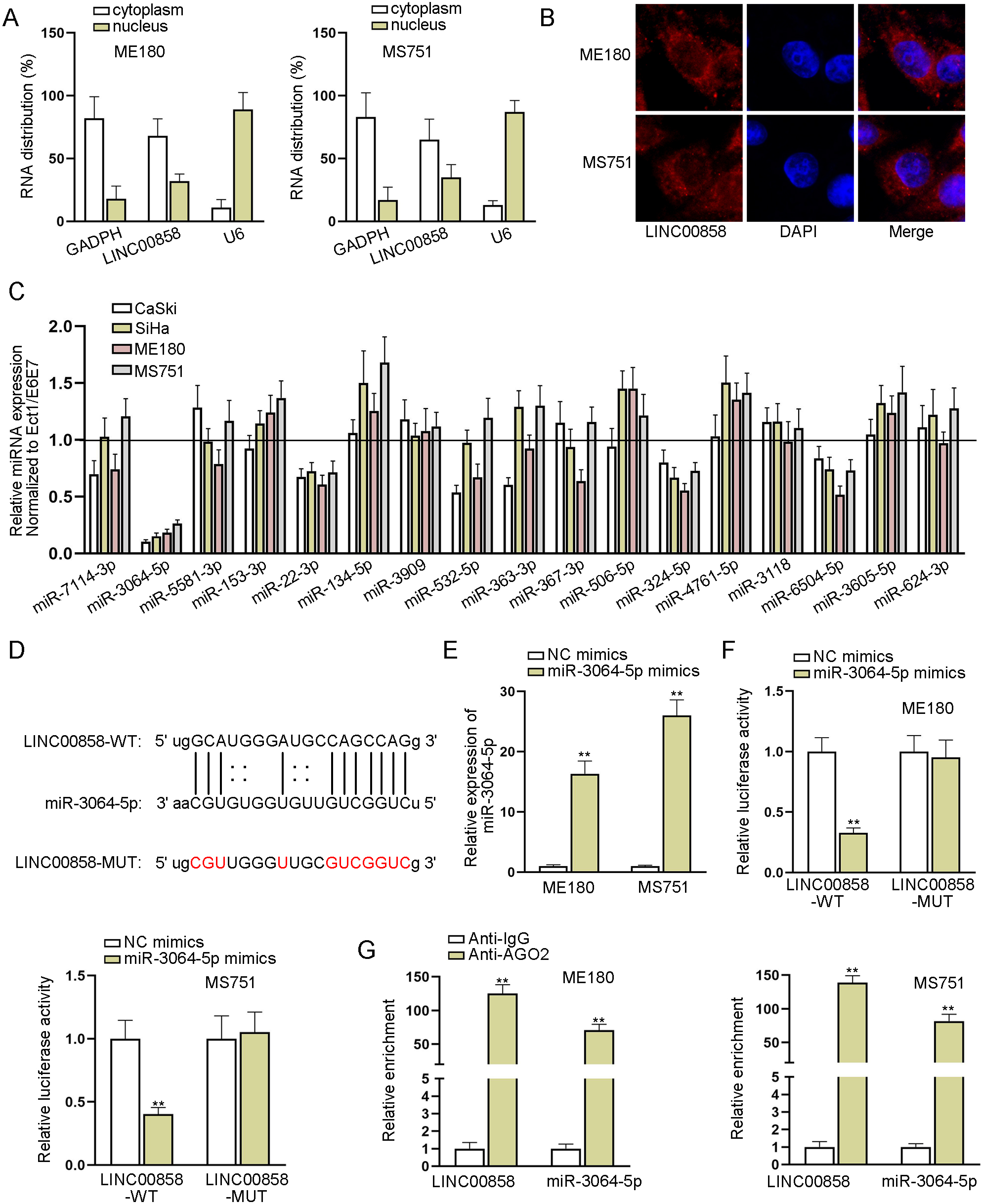
LINC00858 contributes to cell growth in CC
To identify the functional characteristics of LINC00858 in CC cells, we first quantified the expression of LINC00858 in CC cell lines (CaSki, SiHa, ME180 and MS751) and in normal cervical epithelial cell line (Ect1/E6E7) by RT-qPCR. The results showed that CC cell lines, especially ME180 and MS751, exhibited remarkably higher LINC00858 expression, in contrast to normal cervical epithelial cell line (Fig. 1A). To delve into the function of LINC00858 in CC cells, ME180 and MS751 cells were transfected with sh-LINC00858#1 due to their rich expression of LINC00858. RT-qPCR showed that LINC00858 expression was significantly knocked down in ME180 and MS751 cells transfected with sh-LINC00858#1 (Fig. 1B). On the premise of LINC00858 silence, we conducted the loss-of-function assays. The quantitive data from EdU and colony formation assays manifested that lessening LINC00858 expression remarkably restricted the proliferation of CC cells (Fig. 1C and D). The quantitive data from TUNEL assay and flow cytometry assay further indicated that LINC00858 depletion pushed the apoptosis of ME180 and MS751 cells (Fig. 1E and F). Altogether, these results indicated that LINC00858 plays an oncogenic role in CC cell malignant behaviors.
LINC00858 interacts with miR-3064-5p in CC cells
Subcellular fractionation assay and FISH assay determined that LINC00858 was mainly accumulated in the cytoplasm of ME180 and MS751 cells (Fig. 2A and B). Therefore, we assumed LINC00858 might act as a ceRNA in CC cells. To complete the ceRNA model, we used starBase (
MiR-3064-5p-targeted VMA21 prompts cell proliferation and hampers cell apoptosis in CC
To figure out the potential mechanism of miR-3064-5p with its downstream target genes, we took advantage of miRmap, PicTar, TargetScan and microT databases to screen out 54 underlying mRNAs (Fig. 3A). Then we narrowed the range in the case of LINC00858 knockdown and miR-3064-5p overexpression. The results of RT-qPCR demonstrated that VMA21, EIF5, ZBTB4, HECTD1, TXLNA and PHACTR4 were apparently down-regulated (Fig. 3B). Their expression in CC cell lines normalized to that in normal cervical epithelial cell line (Ect1/E6E7) was also examined via RT-qPCR, and the results illustrated that VMA21 was visibly over-expressed in CC cell lines in contrast to Ect1/E6E7 cell line (Fig. 3C). Hence, VMA21 was selected as the target mRNA for further study. In addition, the potential binding sequences between miR-3064-5p and VMA21 were shown via starBase (Fig. 3D). Next, luciferase reporter assay revealed that miR-3064-5p up-regulation led to luciferase activity reduction of VMA21-WT, not that of VMA21-MUT, representing that VMA21 was the direct target of miR-3064-5p (Fig. 3E). RNA pull down assay also suggested that VMA21 could be enriched by bio-miR-3064-5p-WT rather than bio-NC group or bio-miR-3064-5p-MUT group (Fig. 3F). On this basis, we further explored the impact of VMA21 on CC cell lines. Firstly, ME180 and MS751 cells were transfected with sh-VMA21, and RT-qPCR was used to detect the variation of VMA21. VMA21 expression was distinctly inhibited due to knockdown of VMA21 (Fig. 3G). The outcomes of EdU and colony formation assay demonstrated that proliferative ability of ME180 and MS751 was weakened by VMA21 reduction (Fig. 3H and I). Meanwhile, based on TUNEL assay and flow cytometry assay, cell apoptosis of ME180 and MS751 was elevated due to VMA21 silence (Fig. 3J and K). Taken together, VMA21 has the ability of promoting cell proliferation and hampering cell apoptosis.
MiR-3064-5p-targeted VMA21 promotes cell proliferation and hampers cell apoptosis in CC. A. Databases of miRmap, PicTar, TargetScan and microT were used to screen out underlying mRNAs. B. In the case of LINC00858 knockdown and miR-3064-5p overexpression, the relative expression of potential target genes were quantified by RT-qPCR. C. The expression of 6 mRNAs was detected via RT-qPCR in CC cell lines. D. The binding sites between miR-3064-5p and VMA21 were shown via starBase database. E and F. Luciferase reporter assay and RNA pull down assay were used to confirm the binding between miR-3064-5p and VMA21. G. The interference efficiency of sh-VMA21 was evaluated with the help of RT-qPCR. H and I. The cell proliferation and colony formation ability were examined by EdU assay and colony formation assay in ME180 and MS751 cells transfected with sh-VMA21. J and K. The cell apoptosis was assessed through TUNEL assay and flow cytometry assay. The length of each bar represented the mean result, and error bar represented SD. At least three independent experiments were implemented. 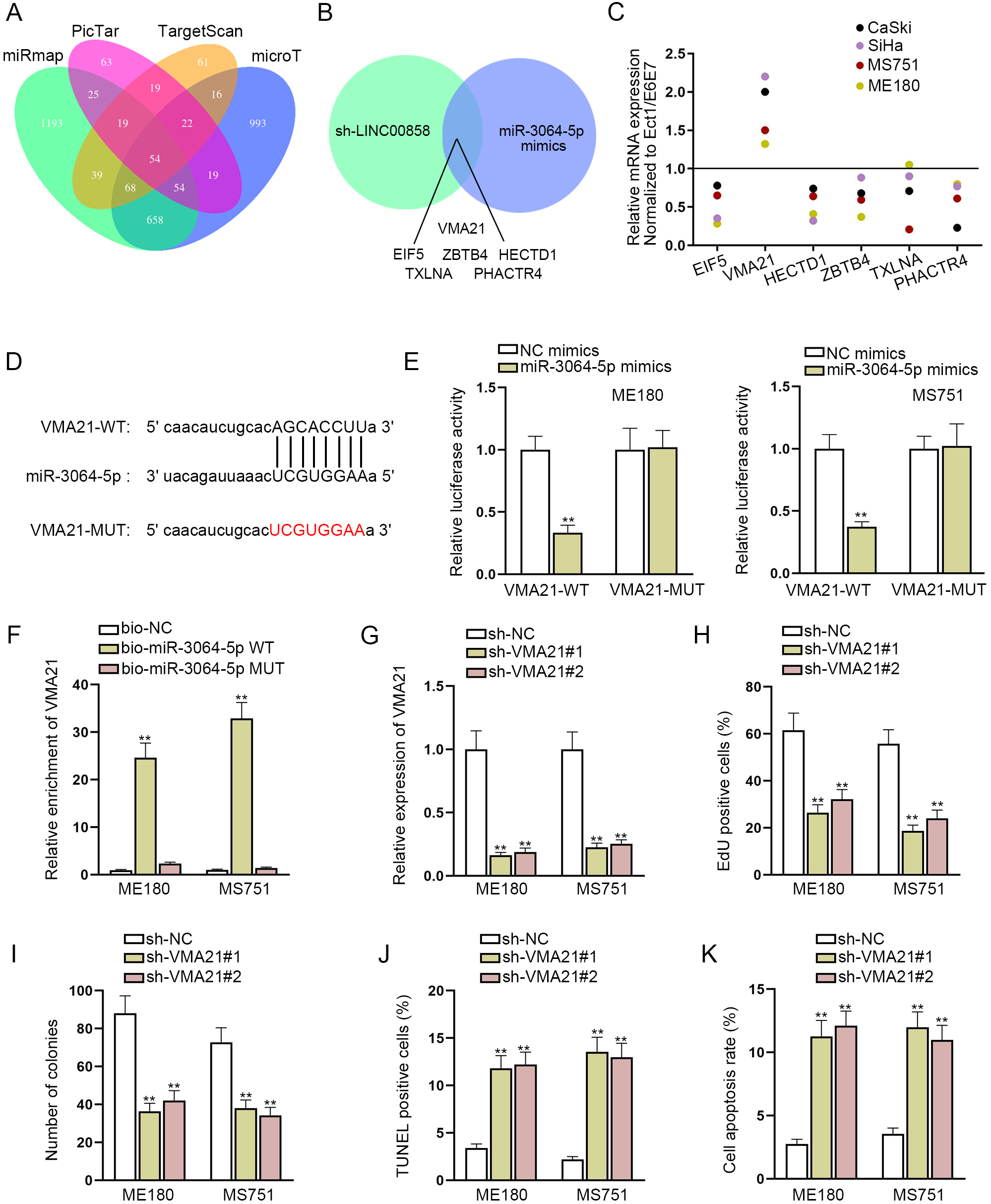
LINC00858 exacerbates CC cell malignancy through regulating miR-3064-5p/VMA21 axis. A and B. RT-qPCR assessed the expression of miR-3064-5p or VMA21 in ME180 and MS751 cells transfected with miR-3064-5p inhibitor and pcDNA3.1-VMA21. C and D. EdU assay and colony formation assay examined cell proliferation in sh-NC, sh-LINC00858, sh-LINC00858+miR-3064-5p inhibitor and LINC00858+pcDNA3.1-VMA21 groups. E and F. TUNEL assay and flow cytometry assay tested ME180 and MS751 cell apoptosis under the same situation. The length of each bar represented the mean result, and error bar represented SD. At least three independent experiments were operated. 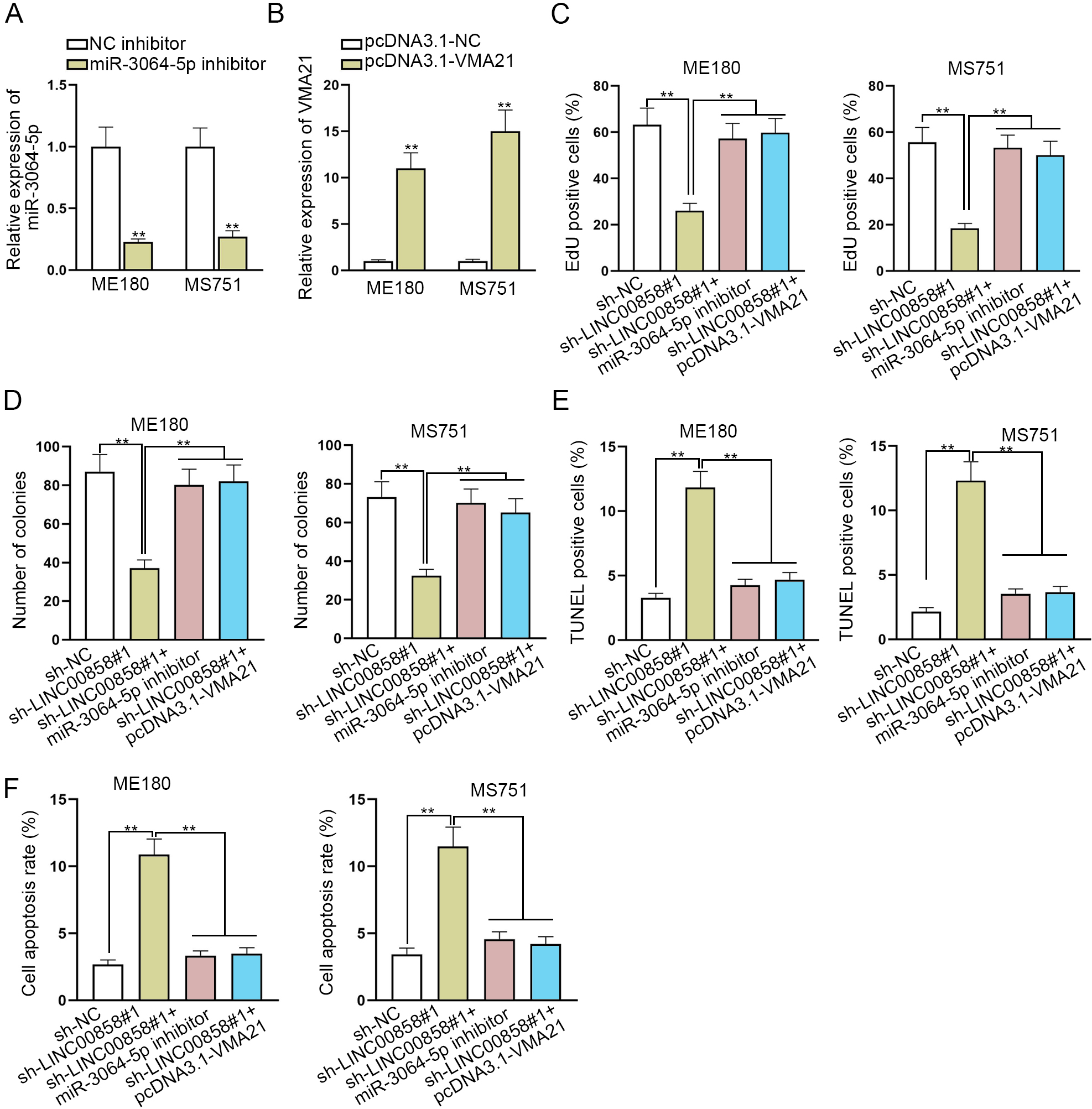
LINC00858 facilitates CC cell malignancy by regulating miR-3064-5p/VMA21 axis.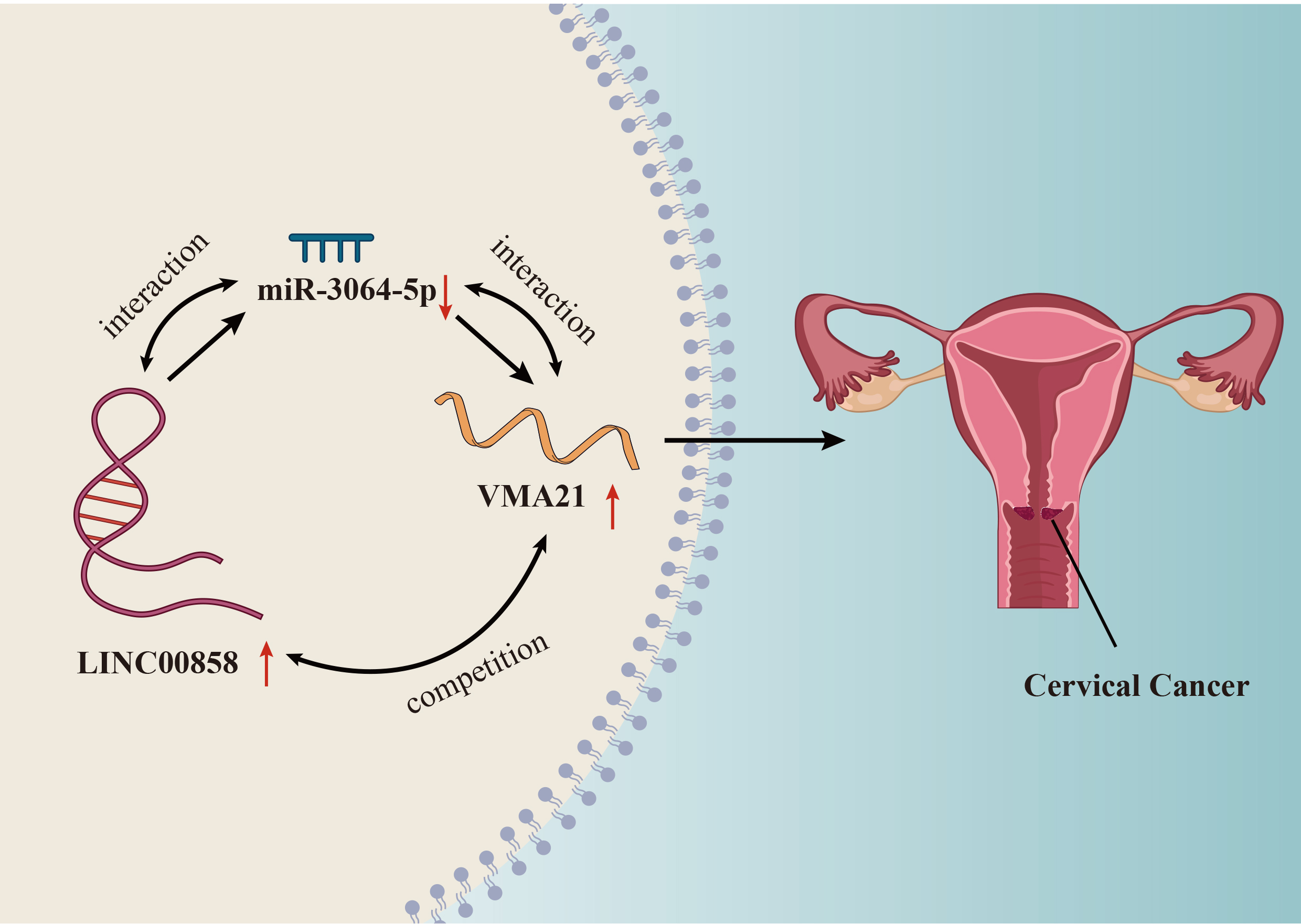
To learn the interplay of LINC00858, miR-3064-5p and VMA21, ME180 and MS751 cells were transfected with miR-3064-5p inhibitor or pcDNA3.1-VMA21. As disclosed in Fig. 4A and B, miR-3064-5p was indeed cut down by miR-3064-5p inhibitor, and VMA21 was effectively increased by pcDNA3.1-VMA21. Then, a chain of rescue experiments were conducted in ME180 and MS751 cells. Knockdown of miR-3064-5p or over-expression of VMA21 was able to reverse the suppressive effect of LINC00858 knockdown on cell proliferative abilities of CC cells, which was illustrated in the quantitive data from EdU and colony formation assay (Fig. 4C and D). Likewise, the results from TUNEL and flow cytometry assay demonstrated that cell apoptosis of CC cells was observably promoted by LINC00858 knockdown, and was almost fully recovered by miR-3064-5p down-regulation or VMA21 overexpression (Fig. 4E and F). Overall, it was proved that LINC00858 promotes CC cell malignancy by regulating miR-3064-5p/VMA21 axis (Fig. 5A).
Discussion
Considering the rapidly increasing incidence of CC in the world in recent years, identifiying more useful and underlying lncRNA-targeted biomarkers is strongly necessary and urgent [17, 18]. At present, several studies have suggested that LINC00858 is positively implicated in the regulation of several tumors. For instance, LINC00858 activates cell growth in non-small cell lung cancer [19]. LINC00858 also affects human osteosarcoma tumorigenesis via miR-139/CDK14 axis [20]. In addition, a new study by Zhan et al. firstly reports that LINC00858 is overtly-overexpressed in colorectal cancer [21]. In like manner, LINC00858 was significantly highly-expressed in CC cells according to RT-qPCR analysis in this study. A strain of loss-of-function assays uncovered that LINC00858 promoted cell proliferation and repressed cell apoptosis, implying that LINC00858 served as an oncogene and impacted biological behaviors of CC cells.
It is known that lncRNAs can act as molecular sponges of miRNAs to affect the development of tumor [22, 23]. For instance, H19 is apparently up-regulated in NSCLC to push NSCLC development by sponging miRNA-107 [24]. In glioma, CCAT1 accelerates glioma tumorigenesis by binding with miR-181b [25]. In this study, through bioinformatics analysis and experiments, we found that LINC00858 could bind to miR-3064-5p, which has been reported to be involved in the regulation of gastric cancer in previous studies [26]. Additionally, miR-3064-5p was proved to suppress angiogenesis in human hepatocellular carcinoma via the FOXA1/CD24/Src pathway [27]. Meanwhile, its suppressing role in breast cancer glycolysis was also verified [28]. Herein, we validated that LINC00858 directly targeted miR-3064-5p by luciferase reporter assay and RIP assay. In rescue assays, miR-3064-5p silence could wipe the inhibition or promotion effect caused by LINC00858 silence on cell proliferation or apoptosis, indicating the participation of miR-3064-5p in LINC00858 regulation. These data demonstrated that LINC00858 modulated CC cellular processes by binding to miR-3064-5p.
VMA21 has been reported to involve in lung adenocarcinoma by interplaying with miR-18b-5p [29]. In addition, VMA21 was also proved to participate in the regulation of ovarian cancer by interacting with miR-18b-5p [30]. In this study, the results of luciferase reporter assay confirmed that VMA21 was a direct target of miR-3064-5p. Silencing VMA21 induced proliferation inhibition and apoptosis increase of CC cells. Furthermore, recuse assays demonstrated that VMA21 overexpression offset the effect of LINC00858 silence-mediated proliferation and apoptosis of CC cells.
In conclusion, our findings indicated that LINC00858 promotes proliferation and restrains apoptosis of CC cells through miR-3064-5p/VMA21 axis. This finding enriched the understanding of CC, which might benefit for developing effective therapeutic strategy for CC. However, herein, only the effect of LINC00858 on CC cell proliferation and apoptosis was studied, and whether LINC00858 will affect other biological behaviors of CC cells such as cell migration, invasion and EMT still needs further study.
Footnotes
Acknowledgments
We appreciate the support of laboratory.
Conflict of interest
The authors declare that there are no competing interests in this study.
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-200033.
