Abstract
Colorectal cancer (CRC) remains one of the most commonly diagnosed malignancies worldwide. Circular RNAs (circRNAs) are being found to play crucial roles in human cancer, including CRC. The purpose of this study was to explore the function and mechanism of circ_0007031 on CRC progression and 5-fluorouracil (5-FU) resistance. The levels of circ_0007031, ATP-binding cassette subfamily C member 5 (ABCC5) and miR-133b were assessed by quantitative real-time polymerase chain reaction (qRT-PCR) or western blot. Cell survival and proliferation were detected by the 3-(4,5-dimethylthiazol-2yl)-5-(3-carboxymethoxyphenyl)-2-(4-sulfophenyl)-2H-tetrazolium (MTS) assay. Cell colony formation was evaluated using a standard colony formation assay. Transwell assays were performed to determine cell migration and invasion. Targeted correlations among circ_0007031, miR-133b and ABCC5 were verified by dual-luciferase reporter, RNA immunoprecipitation (RIP) and RNA pulldown assays. Animal experiments were performed to observe the role of circ_0007031 in vivo. Our data indicated that circ_0007031 up-regulation was associated with CRC resistance to 5-FU. Circ_0007031 knockdown repressed CRC cell proliferation, migration and invasion and enhanced 5-FU sensitivity. Circ_0007031 directly interacted with miR-133b. Moreover, circ_0007031 knockdown regulated CRC cell progression and 5-FU sensitivity by miR-133b. ABCC5 was a direct target of miR-133b, and circ_0007031 mediated ABCC5 expression via acting as a miR-133b sponge. Furthermore, miR-133b overexpression regulated CRC cell progression and sensitivity to 5-FU by down-regulating ABCC5. Additionally, circ_0007031 knockdown suppressed tumor growth in vivo. Our current work had led to the identification of circ_0007031 knockdown that repressed CRC cell malignant progression and enhanced 5-FU sensitivity via regulating ABCC5 expression by sponging miR-133b.
Keywords
Introduction
Globally, colorectal cancer (CRC) is the third most commonly diagnosed malignancy in men and the second in women, accounting for above 1.8 million new cases and almost 881,000 deaths in 2018 [1]. The basic therapies against CRC cover surgery, radiotherapy and adjuvant chemotherapy, which have shown clinical benefit [2]. 5-fluorouracil (5-FU), a potent anti-tumor agent, has been widely used for CRC chemotherapy and 5-FU-based combination therapy has greatly improved CRC survival [3]. Recently, the development of drug resistance has become a severe limitation to 5-FU treatment for CRC patients [4, 5]. Hence, it is indispensable to better understand the mechanisms of CRC progression and chemoresistance and to identify new approaches for improving 5-FU clinical benefits in CRC patients.
Circular RNAs (circRNAs) are a special class of unusually stable noncoding RNAs with covalently closed-loop structures, which play potential regulatory roles in gene expression through acting as microRNA (miRNA) sponges [6, 7]. Intensive research in the past two decades has demonstrated circRNAs to be crucial players in human cancers, including CRC [8]. For example, Li et al. reported that circRNA vesicle-associated membrane protein associated protein A (circVAPA, has_circ_0006990) was highly expressed in CRC, and it promoted CRC progression via sponging miR-101 [9]. Bian and colleagues uncovered that circRNA zinc finger RNA binding protein (circZFR, has_circ_103809) was down-regulated in CRC, and it modulated CRC cell growth and migration through targeting the miR-532-3p/forkhead transcription factor AFX (FOXO4) axis [10]. Moreover, Ren et al. underscored that circRNA p72 RNA helicase (circDDX17) was down-regulated in CRC, and the elevated circDDX17 level repressed CRC progression and promoted CRC cell sensitivity to 5-FU by functioning as a miR-31-5p sponge [11]. As for circRNA_0007031, derived from back-spliced exons of tubulin-gamma complex protein 3 (TUBGCP3) gene, it was discovered as one of the top 10 up-regulated circRNAs in 5-FU-based chemoradiation resistant CRC cells [12]. This report also considered that circ_0007031 might be one of the most important circRNAs in the development of CRC chemoradiation resistance. Nevertheless, the function and mechanism of circ_0007031 on CRC progression and 5-FU resistance are still undefined.
MiRNAs have recently emerged as essential regulators in the carcinogenesis of CRC [13]. MiR-133b has been demonstrated as a tumor suppressor in CRC tumorigenesis and progression [14, 15]. Moreover, Lv et al. reported that the up-regulation of miR-133b weakened CRC cell stemness and enhanced cell sensitivity to 5-FU through targeting telomeric silencing 1-like [16]. However, whether miR-133b was involved in the regulatory network of circ_0007031 on CRC progression and chemoresistance remains unknown.
In the present work, we firstly demonstrated that the up-regulation of circ_0007031 was associated with CRC 5-FU resistance. Consequently, we determined the impact of circ_0007031 on CRC cell proliferation, migration, invasion and 5-FU sensitivity and the mechanism governing it.
Materials and methods
Tissue specimens and ethics statement
Sixty CRC tissue specimens from 30 primary patients and 30 recurrent patients after treatment with 5-FU-based chemotherapy were evaluated in this project, which were obtained from Liuzhou People’s Hospital. Patients who met the following criteria were selected: (1) no history of other malignant tumors; (2) no preoperative chemoradiotherapy or other treatment; (3) postoperative standard treatment based on 5-FU; (4) complete clinical and follow-up recorded information. These specimens were stored at
Cell culture and transfection
Two CRC cell lines HCT116 and SW480 (European Collection of Cell Cultures, Wiltshire, UK) were grown in Dulbecco’s Modified Eagle’s medium (DMEM, PAA Laboratories, Cölbe, Germany) plus 10% fetal bovine serum (FBS, PAA Laboratories) and 1% penicillin/streptomycin (PAA Laboratories) in a humidified atmosphere with 5% CO
For in vitro circ_0007031 depletion assays, HCT116/ 5-FU and SW480/5-FU cells (1.0
Quantitative real-time polymerase chain reaction (qRT-PCR)
The RNA STAT-60 reagent (Biogenesis, Poole, UK) was used for RNA extraction from tissues and cells, based on the recommendation of producers. Levels of the indicated human genes and miR-133b were tested by qRT-PCR using One Step PrimeScript
The list of primers used in the study
The list of primers used in the study
HCT116/5-FU and SW480/5-FU cells were implemented with the indicated transfections as described above. In cell survival assays, transfected cells (1
Transwell migration and invasion assay
Transfected cells in FBS-free media were seeded into the top chamber with the non-coated membrane (8
Western blot
Cell lysates were prepared and fractioned by SDS polyacrylamide gels as previously reported [17]. The separated proteins were transferred to polyvinylidene fluoride (PVDF, Alfa Aesar, Karlsruhe, Germany) membranes and subsequently probed with primary antibodies against Cyclin D1 (ab40754), Ki67 (ab92742), matrix metalloproteinase 9 (MMP9, ab137867), Snail (ab53519), ABCC5 (ab180724) and
Dual-luciferase reporter, RNA immunoprecipitation (RIP) and RNA pulldown assays
The circ_0007031 fragment harboring the miR-133b-binding sites and the 3’UTR of ABCC5 were severally inserted into a Promega pGL3 vector to create luciferase reporter plasmids (circ_0007031-WT and ABCC5 3’UTR-WT). The mutant-type reporters (circ_ 0007031-MUT and ABCC5 3’UTR-MUT) were produced using the TaKaRa MutanBEST Kit following the manufacturer’s suggestion. Each reporter plasmid (100 ng) was transiently introduced into HCT116/ 5-FU and SW480/5-FU cells (1.0
The up-regulation of circ_0007031 was associated with 5-FU resistance of CRC. Circ_0007031 expression was detected by qRT-PCR in 60 clinical samples from 30 primary patients and 30 recurrent patients with 5-FU-based chemotherapy (A), HCT116, HCT116/5-FU, SW480 and SW480/5-FU cells (B). (C) and (D) HCT116, HCT116/5-FU, SW480 and SW480/5-FU cells were exposed to various concentrations of 5-FU for 24 h, followed by the detection of cell survival by MTS assay. 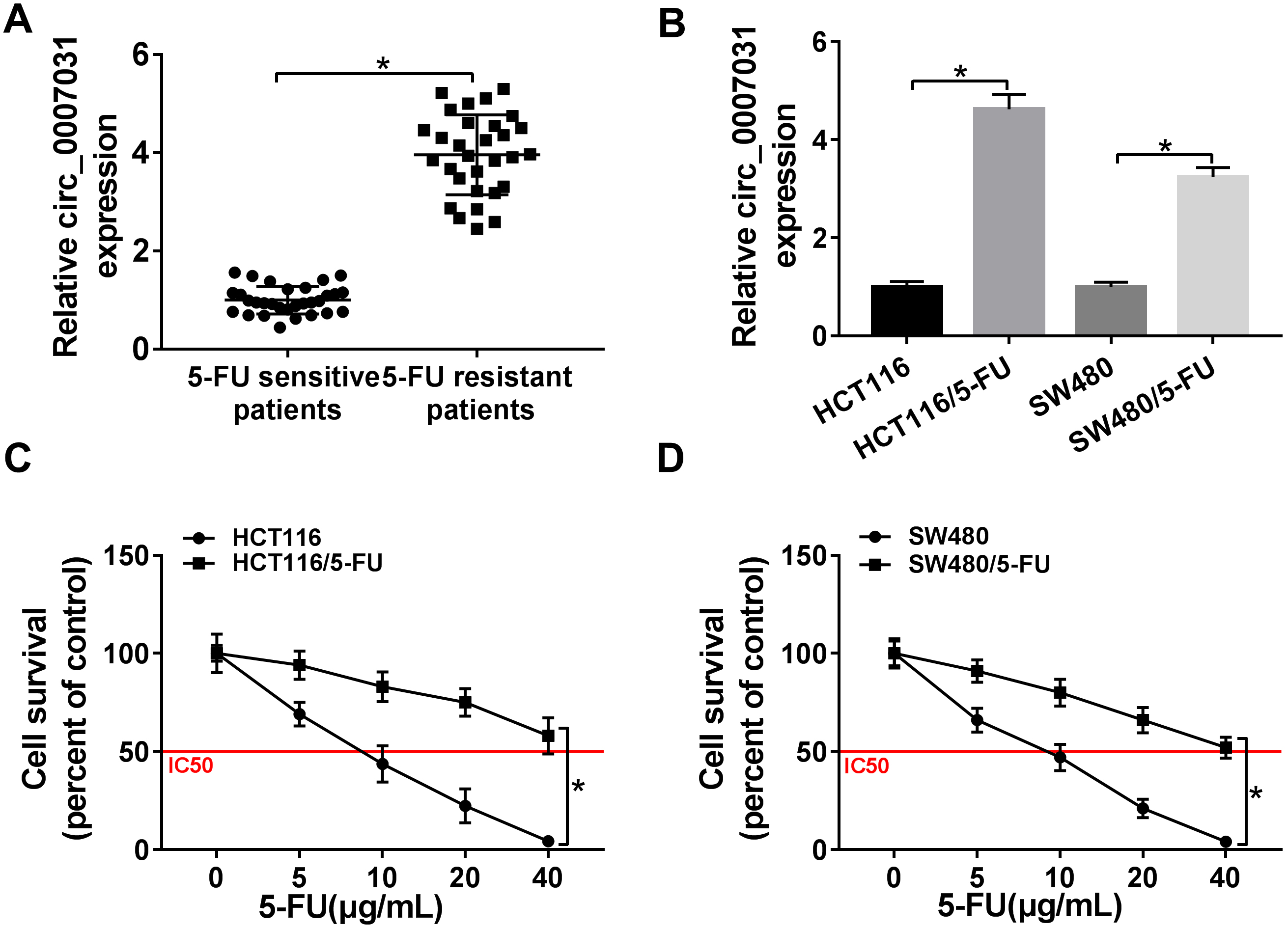
Circ_0007031 silencing hampered CRC cell proliferation, migration and invasion and enhanced 5-FU sensitivity. (A) and (B) Circ_0007031 expression was assessed by qRT-PCR in HCT116/5-FU and SW480/5-FU cells transfected with si-NC or si-circ_0007031. (C) and (D) HCT116/5-FU and SW480/5-FU cells were transfected with si-NC or si-circ_0007031 before 5-FU exposure, and then cell survival was detected by MTS assay. HCT116/5-FU and SW480/5-FU cells were transfected with si-NC or si-circ_0007031, followed by the measurement of cell proliferation by MTS (E) and (F), cell colony formation using a colony formation assay (G) and (H), cell migration (I) and (J) and invasion (K)–(N) by transwell assay, the levels of Cyclin D1, Ki67, MMP9 and Snail by western blot (O) and (P). 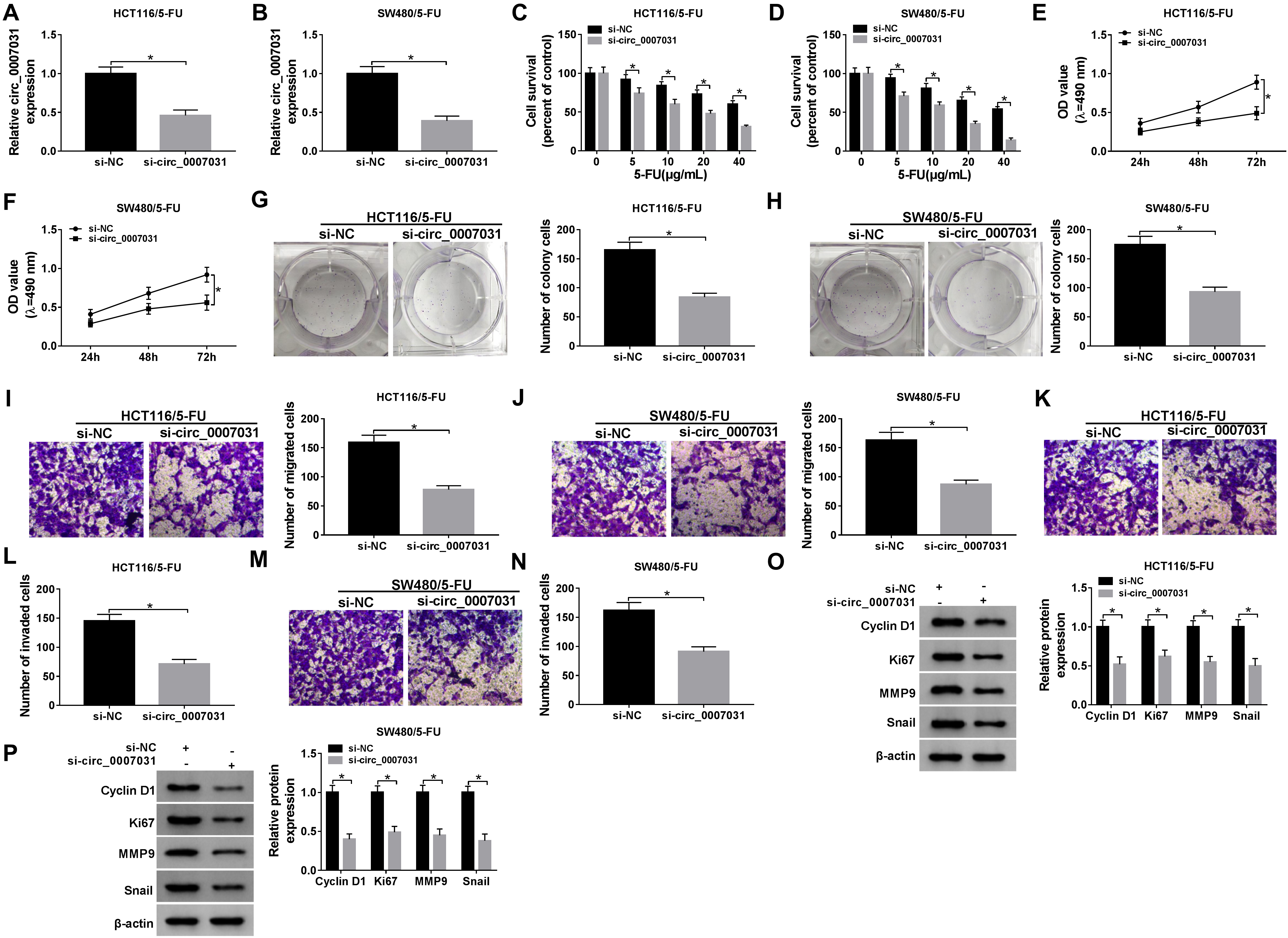
Circ_0007031 acted as a sponge of miR-133b. (A) Schematic model of the target sequence for miR-133b within circ_0007031 and the mutation of the target sequence. (B) and (C) The luciferase activity was determined in both HCT116/5-FU and SW480/5-FU cells cotransfected with circ_0007031-WT or circ_0007031-MUT and miR-133b mimic or miR-NC mimic. Cellular lysates of HCT116/5-FU and SW480/5-FU cells were incubated with anti-Ago2 or anti-IgG antibody (D) and (E) or Bio-miR-133b or Bio-NC (F), and then the expression of circ_0007031 or miR-133b was assessed by qRT-PCR. MiR-133b expression was evaluated by qRT-PCR in HCT116/5-FU and SW480/5-FU cells transfected with si-NC or si-circ_0007031 (G), in 60 clinical samples from 30 primary patients and 30 recurrent patients with 5-FU-based chemotherapy (H), HCT116, HCT116/5-FU, SW480 and SW480/5-FU cells (I). (J) The Spearman test was performed to determine the correlation between miR-133b expression and circ_0007031 level in the resistant CRC tissues. 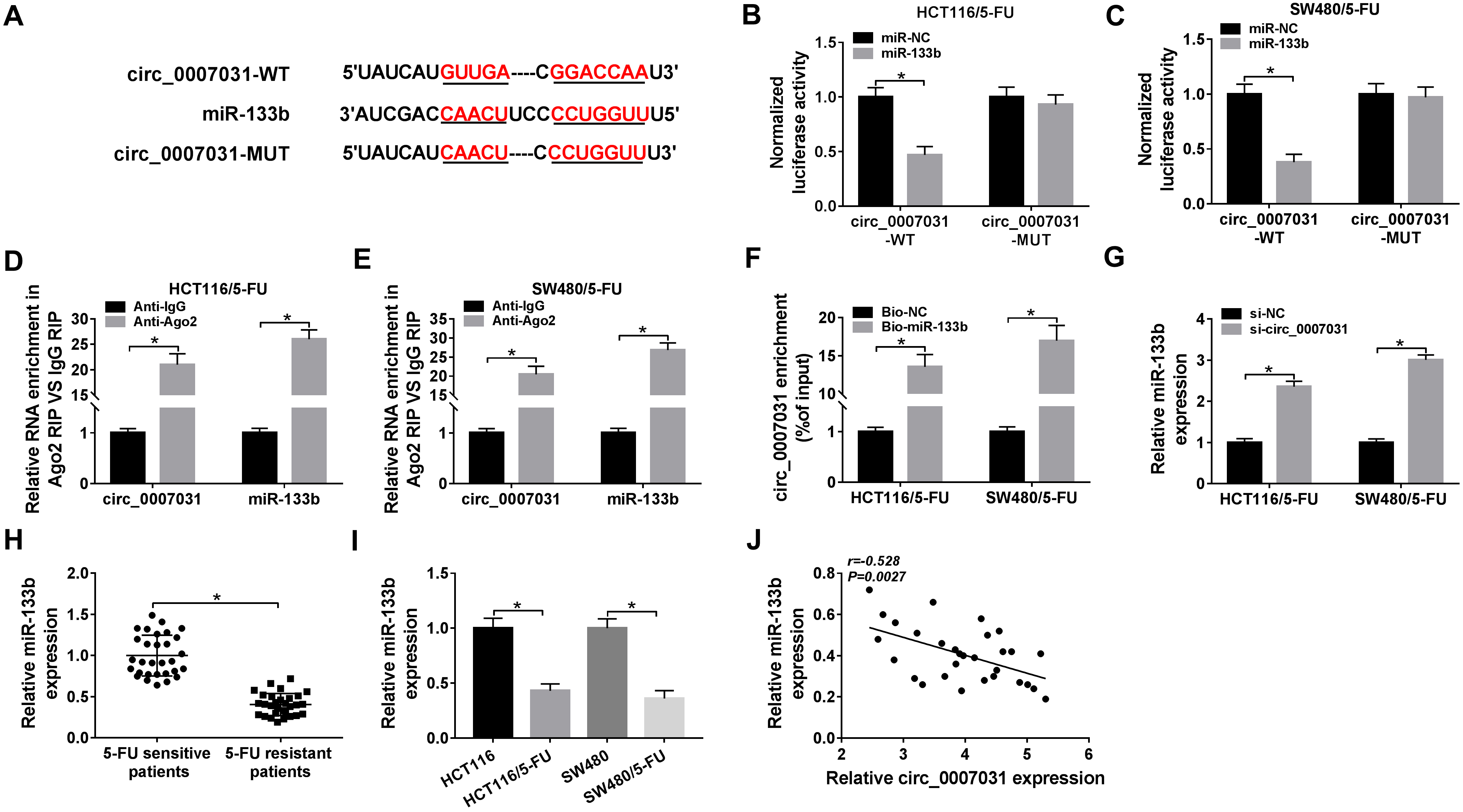
Circ_0007031 regulated CRC progression and 5-FU resistance by miR-133b. MiR-133b expression was tested in HCT116/5-FU and SW480/5-FU cells transfected with anti-miR-NC or anti-miR-133b (A), and si-NC, si-circ_0007031, si-circ_0007031+anti-miR-NC or si-circ_0007031+anti-miR-133b (B). (C) and (D) The two resistant cells were performed the indicated transfections before 5-FU exposure, and then cell survival was determined by MTS assay. The two resistant cells were transfected with si-NC, si-circ_0007031, si-circ_0007031+anti-miR-NC or si-circ_0007031+anti-miR-133b, followed by the determination of cell proliferation by MTS (E) and (F), cell colony formation using a colony formation assay (G), cell migration (H) and invasion (I) by transwell assay, the levels of Cyclin D1, Ki67, MMP9 and Snail by western blot (J) and (K). 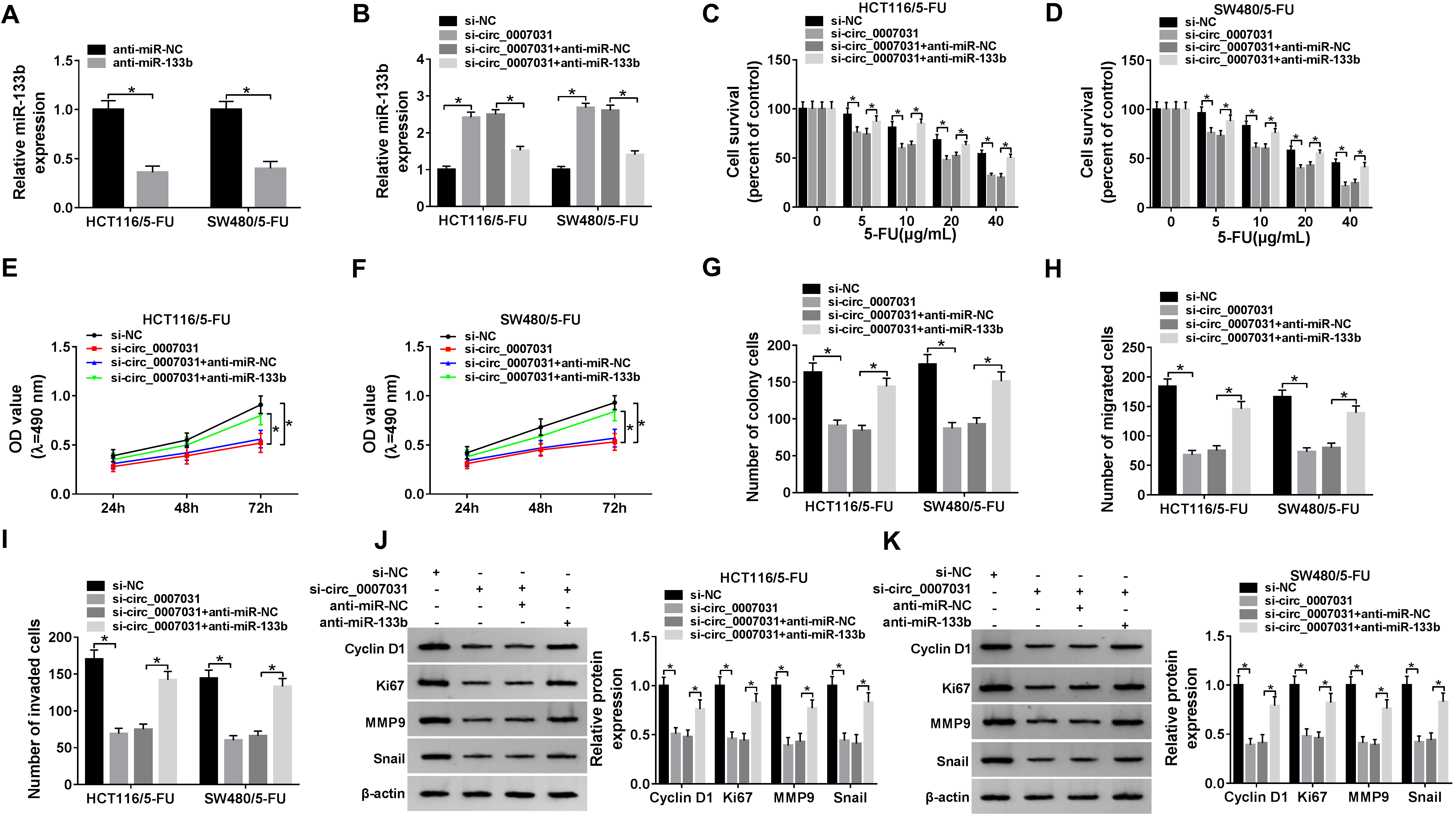
ABCC5 was a direct target of miR-133b. (A) Schematic of the putative complementary sequence for miR-133b within ABCC5 3’UTR and mutated target sequence. (B) and (C) Relative luciferase activity was monitored in the two resistant cells cotransfected with ABCC5 3’UTR-WT or ABCC5 3’UTR-MUT and miR-133b mimic or miR-NC mimic. (D) and (E) The enrichment of miR-133b and ABCC5 were detected by qRT-PCR in cell lysates using anti-Ago2 or anti-IgG antibody. ABCC5 mRNA and protein levels by qRT-PCR and western blot in HCT116/5-FU and SW480/5-FU cells transfected with miR-133b mimic or miR-NC mimic (F) and (G), clinical samples from primary patients and recurrent patients with 5-FU-based chemotherapy (H) and (I), HCT116, HCT116/5-FU, SW480 and SW480/5-FU cells (J)–(L). (M) and (N) Correlation between ABCC5 mRNA expression and miR-133b or circ_0007031 level was assessed by the Spearman test. 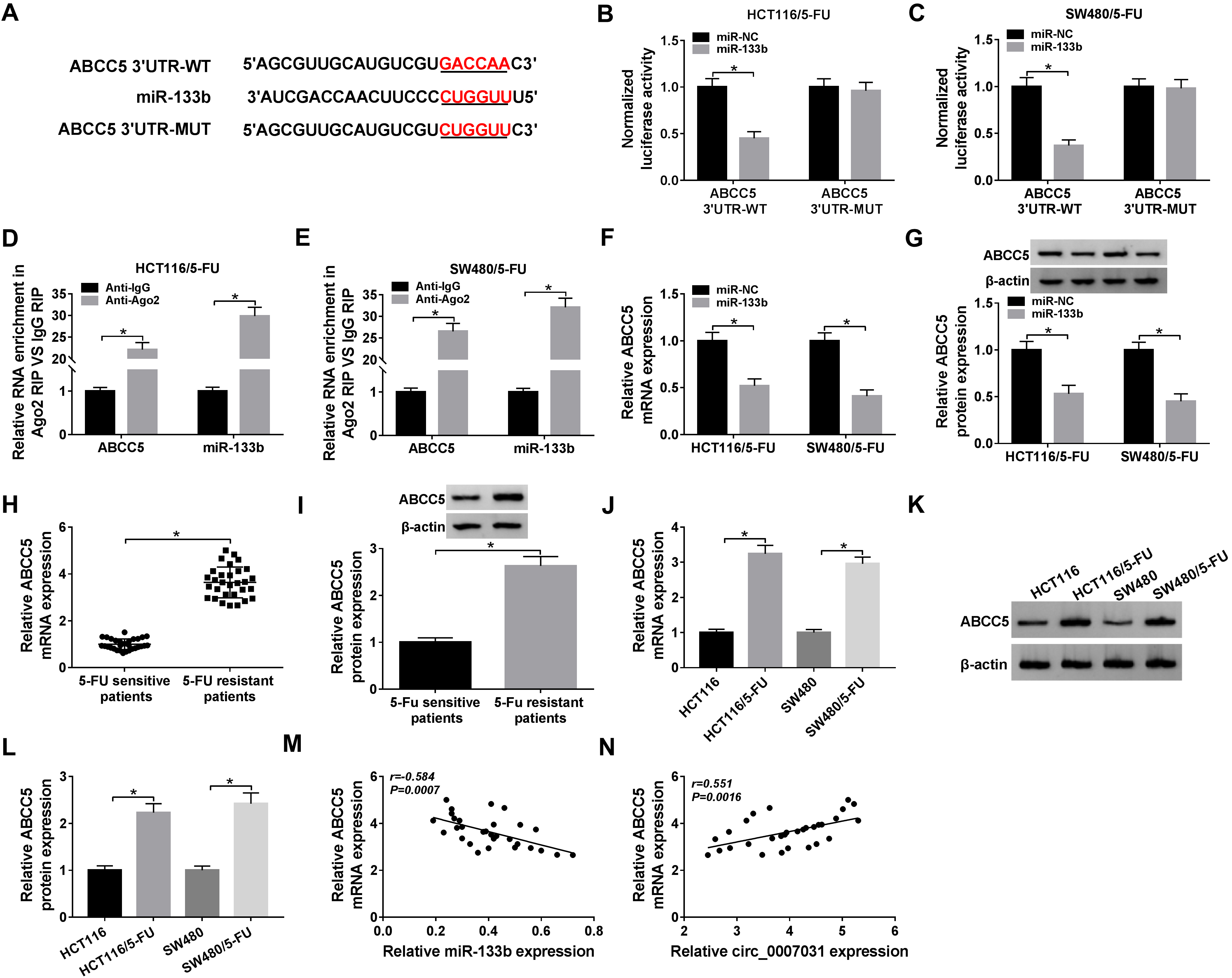
MiR-133b overexpression hampered CRC cell proliferation, migration and invasion and enhanced 5-FU sensitivity by ABCC5. ABCC5 mRNA and protein levels by qRT-PCR and western blot in HCT116/5-FU and SW480/5-FU cells transfected with pcDNA or pcDNA-ABCC5 (A) and (B), and miR-133b mimic, miR-NC mimic, miR-133b mimic+pcDNA or miR-133b mimic+pcDNA-ABCC5 (C) and (D). (E) and (F) The two resistant cells were conducted the indicated transfections prior to 5-FU exposure, and cell survival was detected by MTS assay. The two resistant cells were performed the indicated transfections, followed by the measurement of cell proliferation by MTS assay (G) and (H), cell colony formation using a colony formation assay (I), cell migration (J) and invasion (K) by transwell assay, the levels of Cyclin D1, Ki67, MMP9 and Snail by western blot (L) and (M). 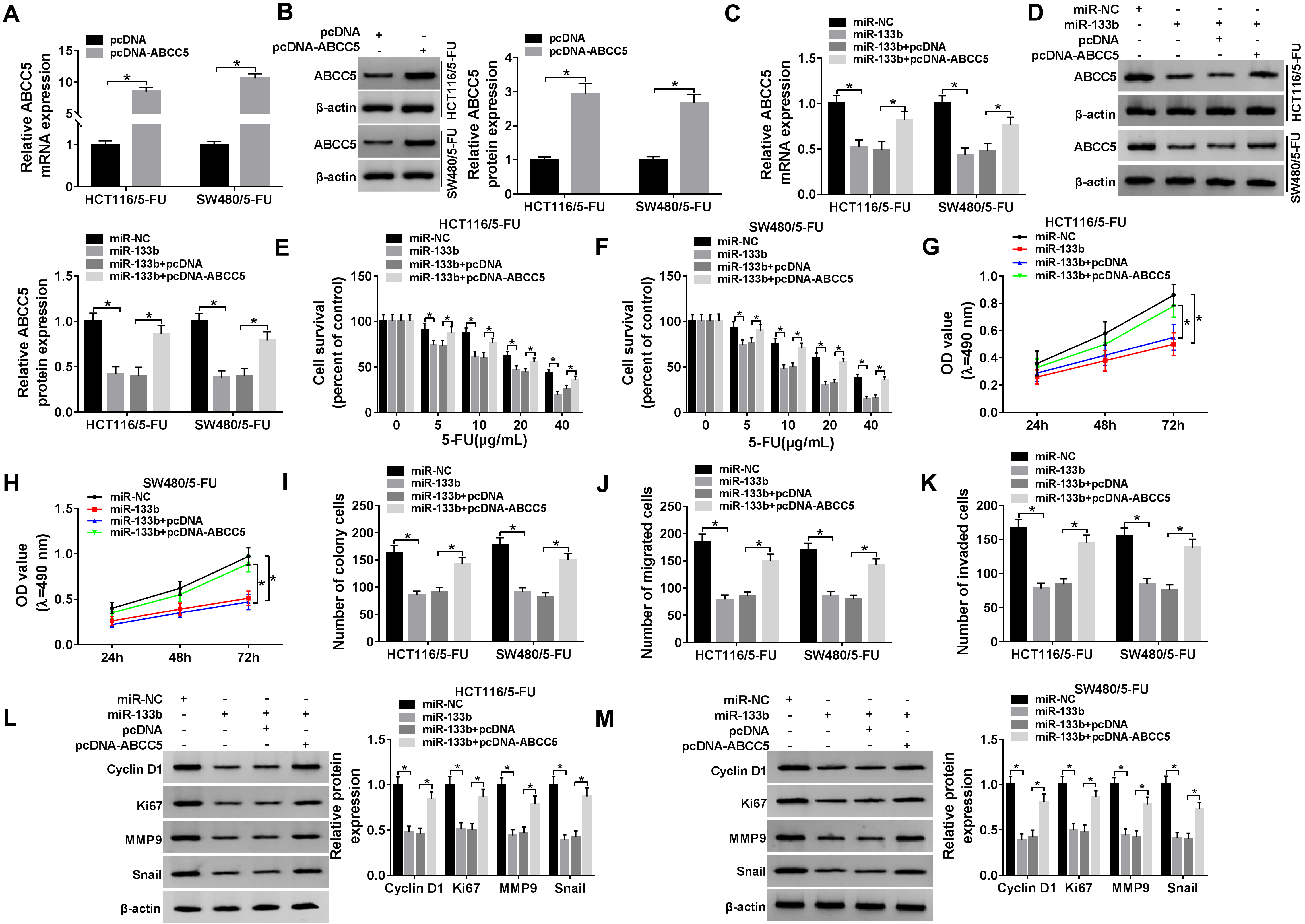
Circ_0007031 regulated ABCC5 expression through sponging miR-133b. HCT116/5-FU and SW480/5-FU cells were transfected with si-NC, si-circ_0007031, si-circ_0007031+anti-miR-NC or si-circ_0007031+anti-miR-133b, and then ABCC5 mRNA expression was detected by qRT-PCR assay (A), and ABCC5 protein expression was assessed by western blot (B). 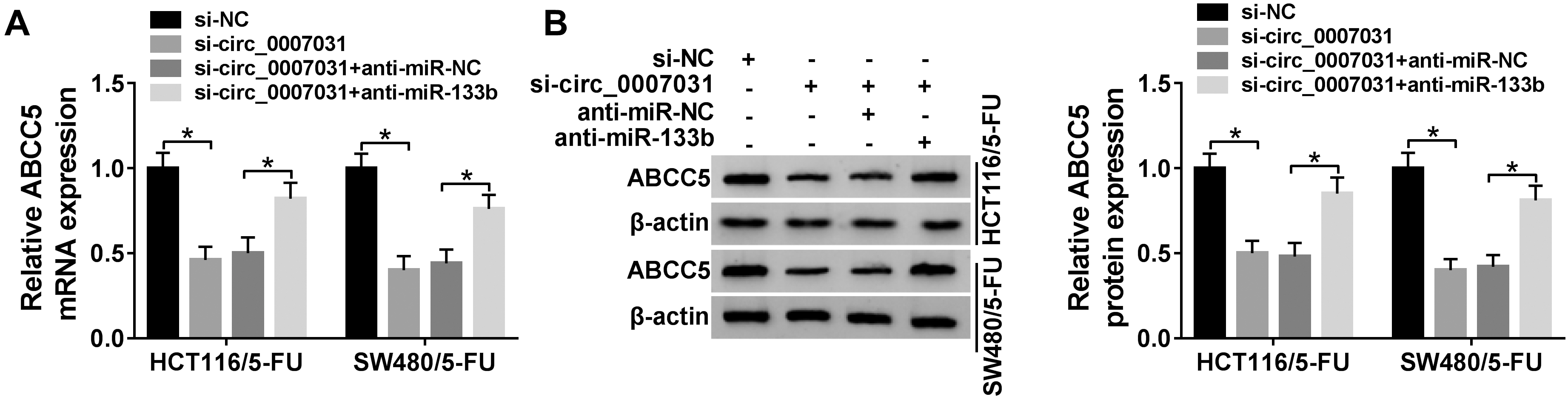
Circ_0007031 silencing mitigated tumor growth in vivo. sh-NC- or sh-circ_0007031-transducing SW480/5-FU cells were subcutaneously inoculated into the nude mice (n 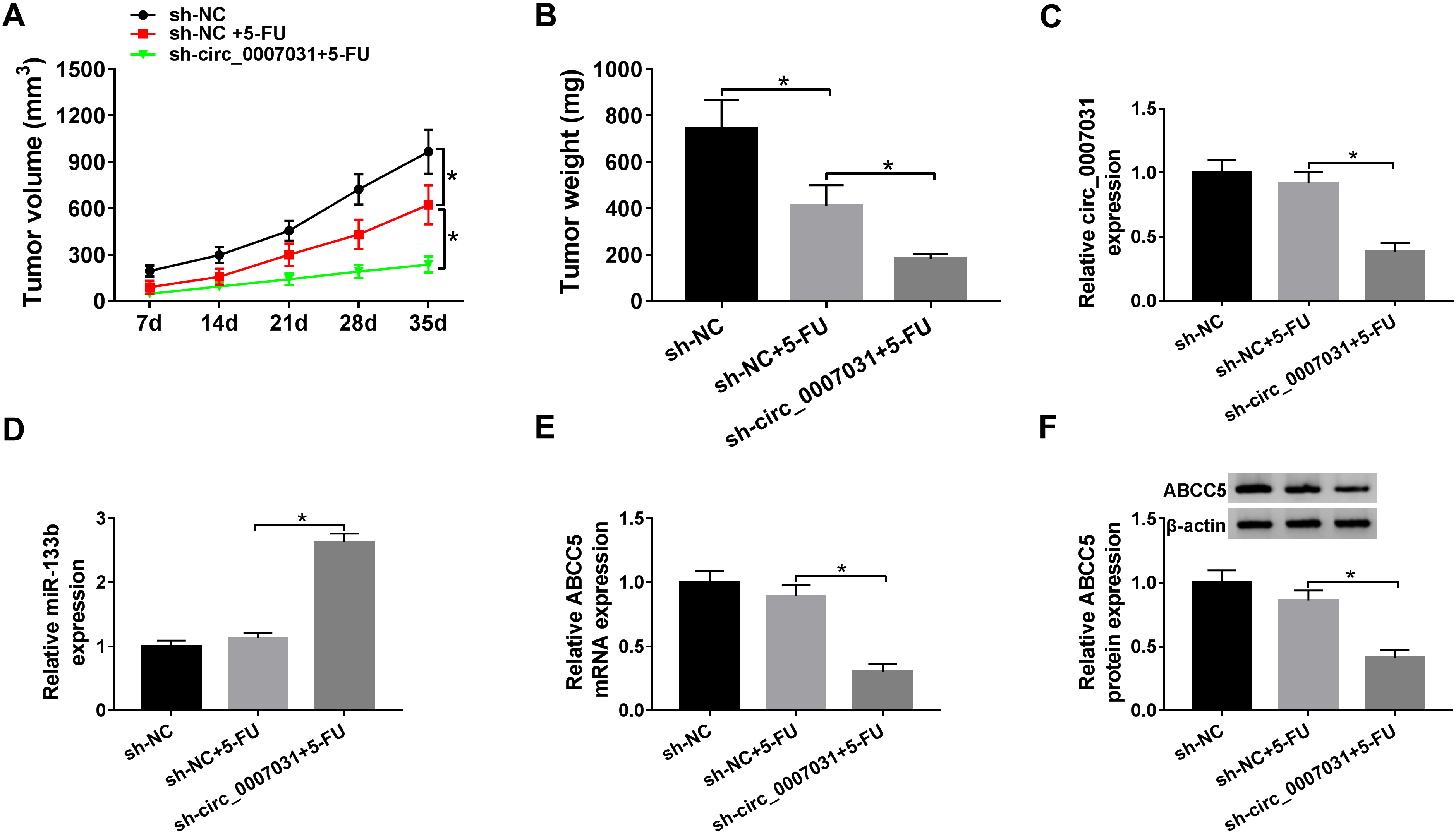
For RIP assays, cell lysates were incubated with magnetic beads-coated antibody against Argonaute 2 (Ago2, ab186733) or isotype IgG (ab172730, all from Abcam) at 4
The animal procedures were in line with the National Guidance for the Care and Use of Laboratory Animals, and the animal study was approved by the Ethics Committee of Liuzhou People’s Hospital. Eighteen 6-week-old BALB/c nude mice (Shanghai Animal Laboratory Center, Shanghai, China) were used for the establishment of the xenograft model. sh-NC- or sh-circ_0007031-transducing SW480/5-FU cells (1
Statistical analysis
Data were presented as the mean values (
Results
Up-regulation of circ_0007031 was associated with CRC resistance to 5-FU
To preliminarily observe the involvement of circ_ 00007031 in CRC chemoresistance, firstly, we detected its expression in clinical tissues from primary patients and recurrent patients with 5-FU-based chemotherapy. As shown by qRT-PCR, circ_0007031 level was higher in 5-FU resistant group than that in the sensitive group (Fig. 1A). Subsequently, we established two 5-FU resistant CRC cells (HCT116/5-FU and SW480/5-FU). In comparison to their parental cells, circ_0007031 expression was significantly elevated in the resistant cells (Fig. 1B). Moreover, 5-FU exposure dose-dependently led to a distinct reduction in cell survival in the two resistant cells and their parents (Fig. 1C and D). Moreover, the IC50 values for 5-FU in HCT116/5-FU (43.92
Knockdown of circ_0007031 suppressed CRC cell proliferation, migration and invasion and promoted sensitivity to 5-FU
To investigate the role of circ_0007031 in CRC, we manipulated its expression using si-circ_0007031. In contrast to the negative control, circ_0007031 expression was prominently down-regulated by si-circ_ 0007031 in the two resistant CRC cells (Fig. 2A and B). MTS data revealed that circ_0007031 silencing led to a significant reduction in cell survival under 5-FU exposure (Fig. 2C and D), demonstrating the promotion of circ_0007031 knockdown on cell sensitivity to 5-FU. Moreover, circ_0007031 silencing triggered a striking repression in cell proliferation (Fig. 2E and F), colony formation (Fig. 2G and H), migration (Fig. 2I and J) and invasion (Fig. 2K–N) in the two resistant cells. Additionally, circ_0007031 knockdown resulted in decreased levels of Cyclin D1, Ki67, MMP9 and Snail in both HCT116/5-FU and SW480/5-FU cells (Fig. 2O and P).
Circ_0007031 directly interacted with miR-133b through binding to miR-133b
Then, we investigated the mechanism by which circ_0007031 regulated CRC progression and 5-FU resistance. Using the starBase v.3 software (
The regulatory effects of circ_0007031 knockdown on CRC cell proliferation, migration, invasion and 5-FU sensitivity were mediated by miR-133b
We further explored whether circ_0007031 regulated CRC progression and chemoresistance by miR-133b. In contrast to the control group, qRT-PCR data revealed that miR-133b expression was remarkably reduced by anti-miR-133b in the two resistant cells (Fig. 4A), and si-circ_0007031-mediated miR-133b up-regulation was significantly reversed by anti-miR-133b (Fig. 4B). Moreover, miR-133b expression restoration led to a prominent augment in cell survival under 5-FU exposure, which was strongly reduced by circ_0007031 silencing (Fig. 4C and D), demonstrating that the enhancement impact of circ_0007031 knockdown on CRC cell 5-FU sensitivity was strikingly abrogated by the restored expression of miR-133b. Further functional experiments showed that si-circ_0007031-mediated anti-proliferation (Fig. 4E and F), anti-colony formation (Fig. 4G), anti-migration (Fig. 4H) and anti-invasion (Fig. 4I) were prominently abolished by miR-133b expression restoration. Additionally, miR-133b expression restoration significantly increased the levels of Cyclin D1, Ki67, MMP9 and Snail, which were highly decreased by circ_0007031 knockdown (Fig. 4J and K).
ABCC5 was directly targeted and inhibited by miR-133b
Using the online software starBase v.3 (
Up-regulation of miR-133b weakened CRC cell proliferation, migration and invasion and enhanced 5-FU sensitivity by down-regulating ABCC5
In contrast to the negative control, ABCC5 mRNA and protein levels were prominently increased by pcDNA-ABCC5 introduction in the two resistant cells (Fig. 6A and B). In addition to the repressive impact on ABCC5 expression (Fig. 6C and D), miR-133b overexpression led to a striking enhancement in 5-FU sensitivity (Fig. 6E and F) and a significant suppression in cell proliferation (Fig. 6G and H), colony formation (Fig. 6I), as well as a prominent inhibition in cell migration (Fig. 6J) and invasion (Fig. 6K). Moreover, the elevated expression of miR-133b resulted in reduced levels of Cyclin D1, Ki67, MMP9 and Snail in the two resistant cells when comparing to the negative control (Fig. 6L and M). To provide further understand in the link between miR-133b and ABCC5 on CRC progression and chemoresistance, the two resistant cells were cotransfected with miR-133b mimic and pcDNA-ABCC5. These data revealed that miR-133b overexpression-mediated ABCC5 reduction (Fig. 6C and D) and 5-FU sensitivity enhancement (Fig. 6E and F) were remarkably abolished by ABCC5 expression restoration. Furthermore, the suppressive effects of miR-133b up-regulation on cell proliferation (Fig. 6G and H), colony formation (Fig. 6I), migration (Fig. 6J) and invasion (Fig. 6K) were markedly reversed by pcDNA-ABCC5 introduction. Besides, ABCC5 expression restoration significantly recovered the levels of Cyclin D1, Ki67, MMP9 and Snail, which were strongly reduced by miR-133b overexpression (Fig. 6L and M).
Circ_0007031 sponged miR-133b to regulate ABCC5 expression
CircRNAs are widely known to mediate gene expression through acting as miRNA sponges. Next, we observed whether circ_0007031 regulated ABCC5 expression in CRC cells. The data of qRT-PCR and western blot showed that in comparison to the negative control, ABCC5 mRNA and protein levels were prominently decreased when circ_0007031 silencing in the two resistant cells, and these effects were remarkably abrogated by anti-miR-133b cotransfection (Fig. 7A and B).
Knockdown of circ_0007031 repressed tumor growth in vivo
Lastly, we evaluated the influence of circ_0007031 on tumor growth in vivo using the xenograft mouse model derived from sh-circ_0007031-transducing SW480/5-FU cells. These data validated the anti-cancer activity of 5-FU in CRC in vivo, as presented by the decreased tumor volume (Fig. 8A) and weight (Fig. 8B). Moreover, in comparison to the negative control, the transduction of sh-circ_0007031 led to a significant repression in tumor growth under 5-FU treatment (Fig. 8A and B). Furthermore, circ_0007031 and ABCC5 levels were strikingly down-regulated, and miR-133b expression was highly up-regulated in tumor tissues derived from sh-circ_0007031-transducing SW480/5-FU cells (Fig. 8C–F).
Discussion
CircRNAs have been continuously discovered to function as critical players in human cancer; however, the underlying mechanisms of the vast majority in cancer biology remain unclear [18]. Up to now, several circRNAs, such as has_circ_001680 and has_circ_0005963, have been demonstrated to be implicated in CRC tumorigenesis and chemoresistance [19, 20]. The focus of this project was to investigate the influence of circ_0007031 on CRC progression and 5-FU resistance and to examine the miR-133b/ABCC5 axis as a molecular mediator of circ_0007031.
Earlier research demonstrated a correlation between circ_0007031 level and 5-FU-based chemoradiation resistance in CRC [12]. Herein, we validated that the elevated expression of circ_0007031 was associated with 5-FU resistance of CRC. To observe the impact of circ_0007031 in CRC, we firstly established two 5-FU resistant CRC cells, and we found a negative role of circ_0007031 knockdown in CRC 5-FU resistance. Similar to our findings, circDDX17 was reported to be involved in 5-FU resistance of CRC [11]. Cyclin D1 is a proto-oncogene that regulates tumor cell proliferation and cell cycle progression [21]. The expression of MMP9 and Snail proteins is associated with cell migration and invasion [22, 23]. For the first time, we uncovered that circ_0007031 knockdown suppressed CRC cell proliferation, migration, and invasion in vitro and hampered tumor growth in vivo.
CircRNAs have gene-regulatory potential in human cancer through functioning as efficient miRNAs [18]. Here, we were first to confirm that circ_0007031 directly interacted with miR-133b through sequestration in CRC cells. MiR-133b is important in cancer pathogenesis, and it has been shown as a tumor-suppressive miRNA in numerous human cancers, including esophageal squamous cell carcinoma, gastric cancer and prostate cancer [24, 25, 26]. Moreover, miR-133b overexpression repressed CRC progression and promoted CRC cell sensitivity to 5-FU [16, 27, 28]. We also validated that the up-regulation of miR-133b hindered CRC malignant progression and enhanced 5-FU sensitivity in CRC cells. More importantly, for the first time, we identified that circ-0007031 knockdown regulated CRC cell progression and 5-FU sensitivity through miR-133b.
Using the starBase v.3 software, we next identified and ascertained that ABCC5 in CRC cells was directly targeted and suppressed by miR-133b. ABCC5 is a member of the multidrug resistance proteins (MRPs) of ATP-binding cassette (ABC) transporters, which contributes to chemotherapeutic resistance in many human tumors, such as breast cancer, nasopharyngeal carcinoma and ovarian cancer [29, 30, 31]. It was also reported that ABCC5 overexpression induced CRC cell resistance to 5-FU through the efflux of monophosphate metabolites [32]. In this study, we were first to highlight that the elevated miR-133b expression hampered CRC cell malignant progression and promoted 5-FU sensitivity through down-regulating ABCC5. Zhang et al. underscored that miR-361 contributed to CRC cell sensitivity to 5-FU through regulating ABCC5 [33]. More interestingly, we first uncovered that circ_0007031 modulated ABCC5 expression via sponging miR-133b through sequestration.
In conclusion, our current work had led to the identification of circ_0007031 knockdown that repressed CRC cell malignant progression and enhanced 5-FU sensitivity through targeting the miR-133b/ABCC5 axis. Our research provided a useful database for further understanding of CRC progression and chemoresistance and identified novel targets to improve 5-FU clinical benefits in CRC patients.
Footnotes
Conflict of interest
The authors declare that they have no competing interests.
