Abstract
BACKGROUND:
The role of microRNAs (miRs) as biomarkers to predict outcome in hepatoblastoma (HB), the most common malignant liver tumor in childhood, has still to be determined. Recently, the so-called four-miR signature has been described to efficiently stratify HB patients according to their prognosis.
OBJECTIVE:
We examined the recently described four-miR signature for its clinical relevance in an independent validation cohort of HB patients and tried to optimize its predictive value by analyzing four additional miRs involved in HB biology.
METHODS:
Expression of eight miR was determined in 29 tumor and 10 normal liver samples by TaqMan assays and association studies and Kaplan-Meier estimators determined their clinical relevance.
RESULTS:
Stratifying HB patients by the four-miR signature showed no difference in patients’ outcome, which was also reflected by the lack of association with any clinical risk parameter. Adding miR-23b-5p and miR-23b-3p did also not increase its discriminating power. However, the integration of miR-483-5p and miR-483-3p into the four-miR signature could predict patients with poor outcome that were associated with large tumors and vessel invasive growth with high accuracy.
CONCLUSIONS:
The expansion of the four-miR signature by miR-483 serves as a useful biomarker to predict outcome of HB patients.
Introduction
The discovery of microRNAs (miRs) in 1993 has opened an entirely new level of regulation of gene expression [1]. These short, non-coding RNA molecules of 17–23 nucleotides length are regulators of most physiological and pathological aspects of cell biology by binding to the 3’-untranslated regions of target messenger RNAs (mRNAs) via imperfect base pairing, inducing translational repression or mRNA degradation [2]. One miR is able to regulate a set of several mRNAs, while one mRNA can be regulated by several miRs [3]. The importance of miRs in various malignancies has emerged, since they can act either as oncogenes [4, 5] or as tumor suppressors [6]. Besides its gene regulatory function miRs can also serve as diagnostic and therapeutic tools in oncology [7]. Tong et al. were able to identify a 16-miR signature to predict cancer relapse in prostate cancer by measuring miR expression in the tumor [8]. Moreover, circulating miRs can be predictors for relapse in prostate cancer [9]. Clusters of circulating miRs were described to be able to predict the occurrence of hepatocellular carcinoma (HCC) in cirrhotic patients [10] or the risk of disease progression in patients with chronic hepatitis B [11]. Zhang et al. established a 3-miR signature that was able to identify early stage and low-level alpha feto protein (AFP) HCC patients [12].
Hepatoblastoma (HB) is the most common malignant childhood liver tumor with a yearly incidence of 1.5 per 1,000,000. The clinical outcome of HB patients strongly depends on the age and the serum AFP level at diagnosis, local extend of the disease, and the presence of metastases [13]. Moreover, defining the tumor extend in the liver before any treatment by the radiology-based pretreatment extent of disease (PRETEXT) staging system is of clinical relevance [14]. Standard risk patients with no clinical risk factors have an excellent prognosis of 90% overall survival (OS) [15]. However, high risk patients with advanced age (
In the present study, we examined the clinical relevance of the four-miR signature in an independent cohort of HB patients and determined the prognostic role of miR-23b and miR-483 expression in HB.
Materials and methods
Patient samples
We recruited 29 patients diagnosed with HB undergoing surgical resection in our department. Ten normal liver samples were obtained from HB patients during the surgical tumor resection. Fresh frozen samples were stored in liquid nitrogen until examination. Diagnosis of HB was confirmed by the reference pathology center in Kiel, Germany. Clinical data were retrieved from the German Pediatric Liver Tumor Register in Munich. Experiments were undertaken with the understanding and written consent of each subject. The study conforms with the code of ethics of the World Medical Association (Declaration of Helsinki) and was approved by the ethics board of the University of Munich (no. 431-11).
MicroRNA analysis by real-time PCR
Isolation of total RNA of HB and liver tissue was performed using the Tri Reagent
Statistical analysis
For comparison of miR expression in different samples we used unpaired
Results
Patients’ characteristics
A total of 29 HB specimens and 10 normal healthy liver tissue samples (NL) were collected. Of the 29 HB cases, 14 were from female and 15 from male patients (Table 1). They were on average 34.5 months old at the time of diagnosis. The average serum alpha fetoprotein (AFP) at diagnosis was 348,161 ng/mL. The diagnosis was HB in 25 patients, and transitional liver cell tumor (TLCT) according to Prokurat et al. [29] in 4 patients. Twenty were classified epithelial, whereas 9 were mixed type. PRETEXT (pretreatment extent of disease) staging showed 3 patients with PRETEXT I, 5 with PRETEXT II, 13 with PRETEXT III and 8 with PRETEXT IV. Multifocal tumor growth was found in 11, vessel invasive in 9, extrahepatic in 2, and metastatic in 11 out of 29 HB patients. Seven out of 29 HB patients died of disease to date, and 13 patients had the adverse C2 subtype of the 16-gene signature according to Cairo and colleagues [18].
Study population characteristics (
29)
Study population characteristics (
We conducted an expression analysis to assess the capacity of the four miRs let-7a, miR-100, miR-371, and miR-373 to stratify HB patients according to their prognosis. Expression analysis of let-7a (Fig. 1A) and miR-100 (Fig. 1B) showed no differences in expression levels between NL and tumor samples, but correlated mildly to each other (Fig. 1C). In contrast, expression levels of the two miRs miR-371 (Fig. 1D) and miR-373 (Fig. 1E) revealed a significant upregulation in HB tumor samples compared to NL, with a very strong correlation of both miR expression values (Fig. 1F).
Relative expression of (a) let-7a, (b) miR-100, (d) miR-371, and (e) miR-373 in normal liver and liver tumor samples, as determined by quantitative RT-PCR and normalized to the expression of house-keeping miR RNU43. Mean expression is given as a horizontal line. Significances were determined by unpaired 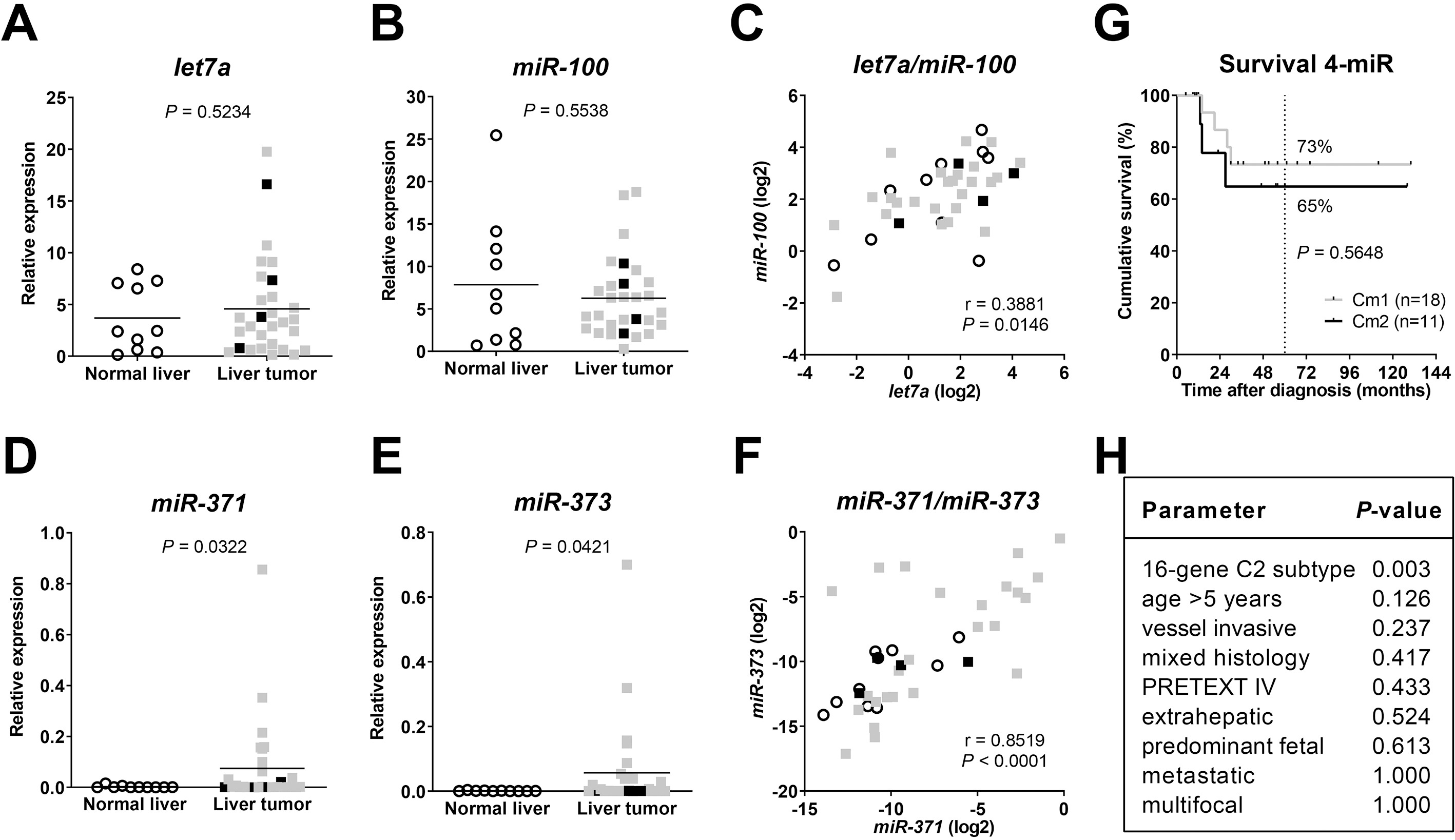
Based on the four-miR expression profile, tumor samples were then classified as either Cm1 or Cm2 according to Cairo and colleagues [19]. Kaplan-Meier estimator comparing both groups of patients displayed no difference in overall survival (Fig. 1G). Along this line, Cm1 and Cm2 grouping was not associated with clinicopathological risk factors such as age at diagnosis
Next, we investigated the role of miR-23b and miR-483 in HB. As their role in HB patient samples was completely unknown, we examined both the 5’ transcript (-5p) and its 3’ (-3p) counterpart for both miRs. The relative expression of miR-23b-5p (Fig. 2A) and miR-23b-3p (Fig. 2B) in HBs showed no significant difference to NL tissues, but their expression values correlated positively (Fig. 2C). This was to be expected, since they are processed from the same hairpin precursor [30]. Of note, the mean relative expression of miR-23b-5p in all samples was approximately 200-times higher than the one of miR-23b-3p.
Relative expression of (a) miR-23b-5p, (b) miR-23b-3p, (d) miR-483-5p, and (e) miR-483-3p in normal liver and liver tumor samples, as determined by quantitative RT-PCR and normalized to the expression of house-keeping miR RNU43. Mean expression is given as a horizontal line. Significances were determined by unpaired 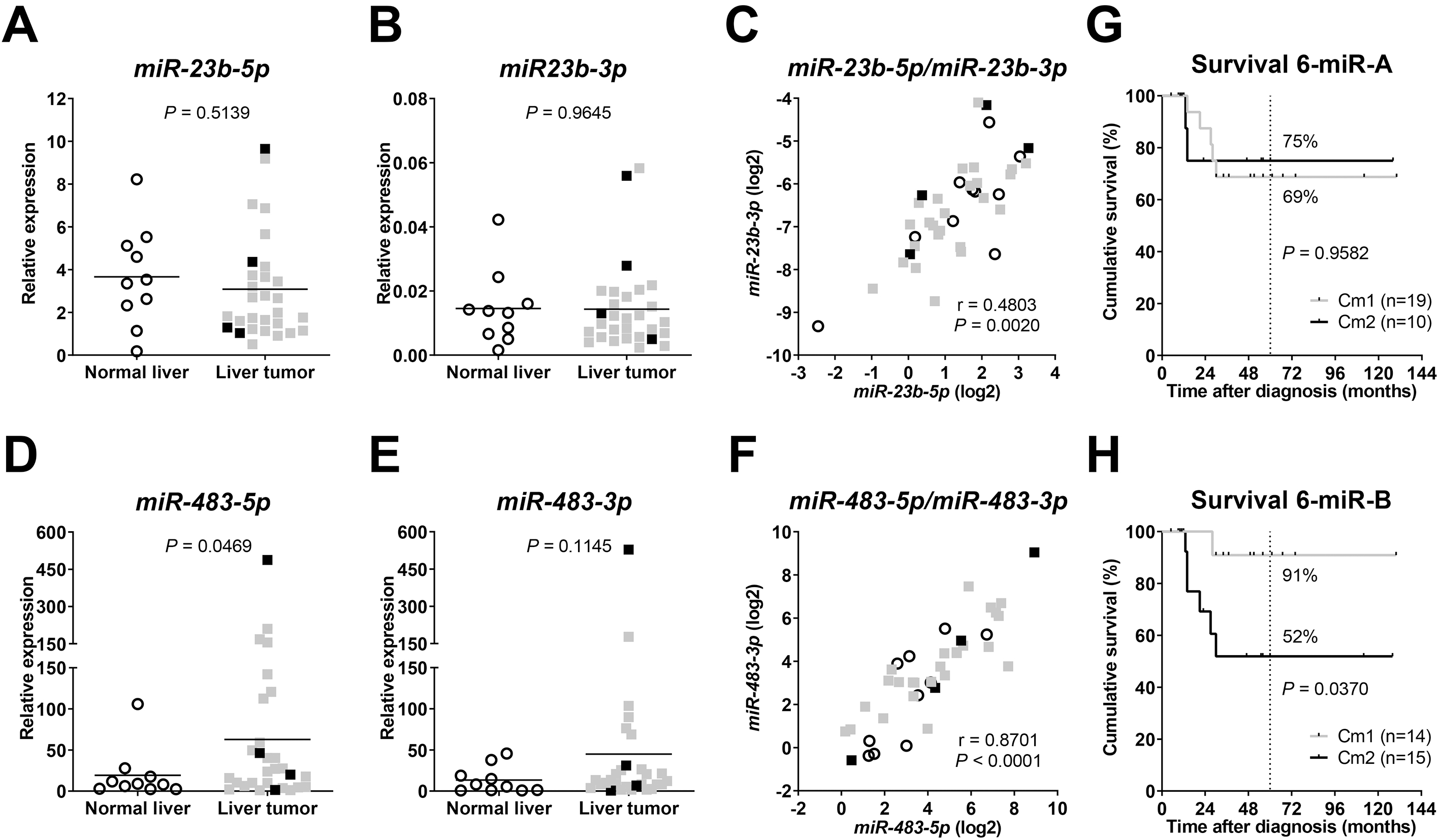
Then, we analyzed the expression of the two mature transcript variants -5p and -3p of miR-483 in HB patients. Significantly higher expression levels could be detected for miR-483-5p in tumor compared to NL samples (Fig. 2D). Although miR-483-3p followed the same trend, this was statistically not significant (Fig. 2E). As anticipated, a strong and highly significant correlation of both variants was found (Fig. 2F).
One of the main findings of Cairo et al. was that the 4 miR signature is able to predict outcome of patients [19]. As we were not able to validate this finding in our cohort (Fig. 1G), we tried to refine the set of miRs and first integrated miR23b-5p and miR23b-3p into the original set of the four-miR signature. However, there was again no difference in overall survival (Fig. 2G). Then, by integrating both variants of miR-483 into the four-miR signature, Kaplan-Meier-survival analysis revealed distinct groups with significantly different outcome (Fig. 2H).
We then investigated if the new six-miR signature is associated with any clinical risk factors [31]. We found that Cm2 patients that have a poor outcome (Fig. 2H) are significantly associated having large PRETEXT IV and vessel invasive tumors (Fig. 3A and B). All other clinical features were not significant.
(a) Association analysis of the six-miR signature with clinicopathological characteristics using Fisher’s exact test with 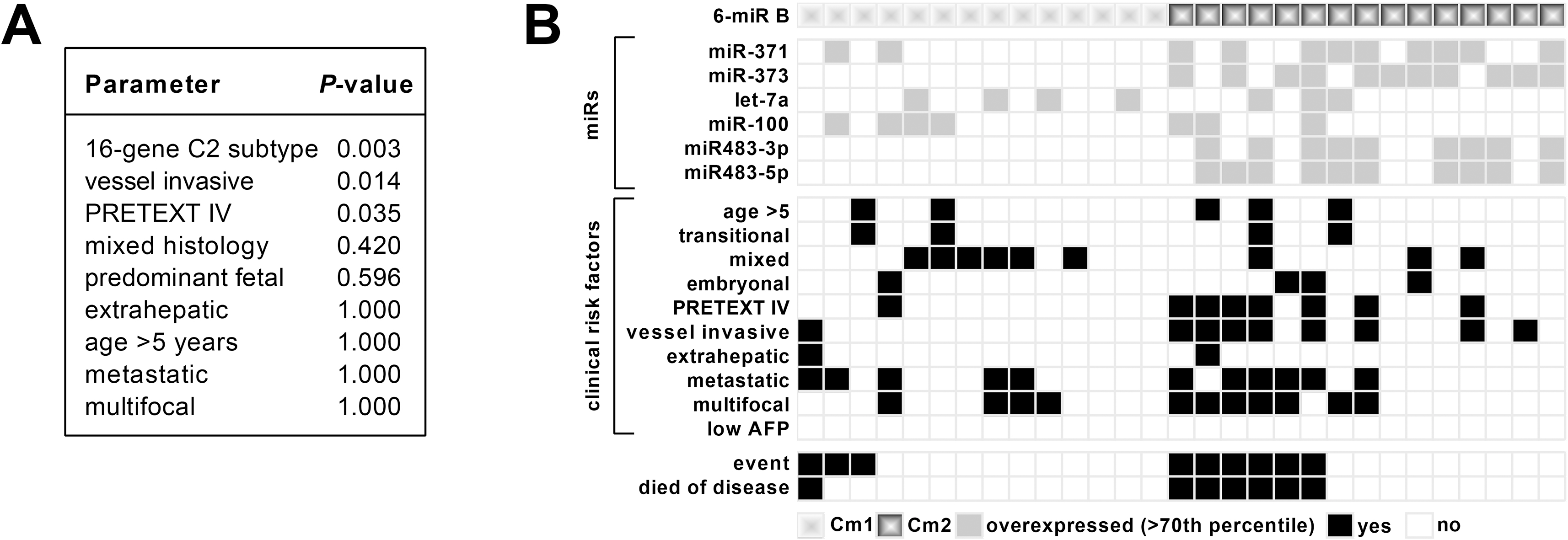
Standard risk HB patients have an excellent outcome of 90% OS, if treated according to the guidelines of SIOPEL [13]. However, high-risk patients still face a poor outcome of 50% OS [17]. In the last decades, numerous studies have identified the involvement of miRs in the initiation and progression of many cancer types by acting as tumor suppressors or oncogenes [32, 33]. As in other types of cancer, several reports proved the importance of miRs in HB development and metastasis [34, 35, 36]. Thus, the present study aimed at defining miRs that can be used as prognostic biomarkers in HB.
As described by Cairo et al., the four-miR-signature consisting of miR-373, miR-371, miR-100 and let-7a can be used to predict outcome of HB patients [19]. To evaluate the robustness of the four-miR signature in an independent cohort of HB patients, we first determined the expression levels of the four miRs and then stratified patients into the Cm1 and Cm2 group according to Cairo et al. [19]. However, we were not able to reproduce the finding of Cairo et al., as patients stratified according to the four-miR signature had no difference in OS. One possible explanation for this discrepancy could be the expression pattern of the individual miRs, which showed only for miR-371 and miR-373 a significant upregulation in the tumors compared to normal liver tissues, whereas let-7a and miR-100 exhibited comparable expression levels. It is worth mentioning that miR-371 and miR-373 as well as let-7a and miR-100 are located next to each other in clusters on chromosome 19q13.4 [37] and 11q24.1 [38], respectively, and are usually co-expressed. Consistent with this, we found a significant correlation of miR-371 and miR-373 expression as well as let-7a and miR-100 expression in our study, but this was much stronger for the former two miRs. Altogether, this first set of data suggests that the predictive power of the four-miR signature, presumably due to the ambiguity of the let-7a and miR-100 cluster, could not be validated in our cohort of 29 HB patients and should be improved to be used for further prognostication.
Thus, we wanted to investigate the prognostic power of two new miRs in HB patients, namely miR-23b and miR-483. Although the 5’ mature products (-5p) of the pre-miRs are generally considered the dominant miR, the 3’ (-3p) can still be functional, which led us to investigate both variants [39].
MiR-23b was previously described as a tumor suppressor miR in various cancers [20, 21] and as a biomarker for HCC, whereby low expression of miR-23b is indicative of metastases, vascular infiltration and poor survival [23]. Of note, miR-23b was one of the co-regulated miRs to let-7a and miR-100 in the initial study by Cairo et al., which all showed downregulation in the adverse HB subtype [19]. In our study, we found no significant differences for both variants of miR-23b in HB compared to normal liver tissue and expression levels were not associated with any clinical parameters. Consistently, integration of miR23b-5p and miR23b-3p into the four-miR signature only led to minor changes in the group assignment and therefore could not improve the discriminating power using the Kaplan-Meier survival estimator. As miR-23b together with let-7a and miR-100 are repressed by the MYC oncogene that contributes to the biology of immature HBs [40], classified as Cm2 [19], it is safe to predict that all three miRs show a similar transcriptional regulation and thus are unsuitable for improving the four-miR signature in HB.
The miR-483 gene is located on chromosome 11p15.5 in the second intron of the IGF2 gene and is implicated in several pathologies [24, 26, 41, 42]. So far, the co-regulation with IGF2 and the transcriptional induction by
Collectively, our results clearly show that the inclusion of miR-483 into the four-miR signature discriminates patients with good and poor outcome and might help to improve biomarker-assisted stratification of HB patients into different risk groups. However, further studies with a larger population size are necessary to substantiate our findings.
Footnotes
Acknowledgments
We acknowledge the assistance of Fatemeh Promoli for technical support. The study was funded by the Friedrich Baur Foundation (Munich, Germany) and the Weigand Foundation (Munich, Germany).
