Abstract
BACKGROUND:
Among patients diagnosed with non-muscle invasive bladder cancer (NMIBC), 30% to 70% experience recurrences within 6 to 12 years of diagnosis. The need to screen for these events every 3 to 6 months and ultimately annually by cystoscopy makes bladder cancer one of the most expensive malignancies to manage.
OBJECTIVE:
The purpose of this study was to identify reproducible prognostic microRNAs in resected non-muscle invasive bladder tumor tissue that are predictive of the recurrent tumor phenotype as potential biomarkers and molecular therapeutic targets.
METHODS:
Two independent cohorts of NMIBC patients were analyzed using a biomarker discovery and validation approach, respectively.
RESULTS:
miRNA Let-7f-5p showed the strongest association with recurrence across both cohorts. Let-7f-5p levels in urine and plasma were both found to be significantly correlated with levels in tumor tissue. We assessed the therapeutic potential of targeting Lin28, a negative regulator of Let-7f-5p, with small-molecule inhibitor C1632. Lin28 inhibition significantly increased levels of Let-7f-5p expression and led to significant inhibition of viability and migration of HTB-2 cells.
CONCLUSIONS:
We have identified Let-7f-5p as a miRNA biomarker of recurrence in NMIBC tumors. We further demonstrate that targeting Lin28, a negative regulator of Let-7f-5p, represents a novel potential therapeutic opportunity in NMIBC.
Introduction
Currently, the U.S. has an estimated 500,000 patients with a history of urothelial carcinoma [1]. The high rate of disease recurrence and progression is a major challenge in patient management [2]. Of patients diagnosed with non-muscle invasive bladder cancer (NMIBC), 30–70% experience recurrence within 6–12 years of diagnosis, and 2–30% of tumors progress to muscle-invasive disease [3]. The need to screen for these events (every 3–6 months and ultimately annually by the invasive cystoscopy procedure) makes bladder cancer one of the most expensive malignancies, costing the U.S. an estimated $3.7 billion in 2001 [4, 5].
Recurrence rates vary considerably [6], even within a single histopathologic group [7]. Primary tumor clinicopathologic characteristics predictive of recurrence include multiplicity, tumor size, T category (depth of invasion), presence of carcinoma in situ, tumor grade, and patient gender [8]. Patients with low stage, low grade (LG) tumors can remain disease free for many years, but poorly differentiated tumors with a high grade (HG) often recur within one year and frequently progress to muscle invasive disease [2]. Management of these patients with TaHG, CIS or T1 tumors is clinically challenging. Bladder removal by cystectomy is performed in the subset of patients experiencing recurrent or progressing tumors that are refractory to treatment [9]. Early identification of those NMIBC patients likely to have progressing and refractory phenotypes would enable individualized treatment and surveillance recommendations, reducing patient burden and disease mortality.
Non-coding RNAs, particularly the microRNAs (miRNAs) have emerged as useful prognostic biomarkers in cancer in part because their small size makes them stable to degradation and thus robust to variations in sample handling [10]. The miRNAs regulate their target genes by binding to specific sites, usually in the 3’ untranslated region (UTR) of the target gene. The miRNA then modifies the target gene via translational repression, cleavage, degradation, or sequestration [11]. The objective of this project was to identify reproducible prognostic miRNAs in resected NMIBC tissue that are predictive of the recurrent tumor phenotype as potential biomarkers and molecular therapeutic targets.
Methods
All study procedures were approved by the Committee for the Protection of Human Subjects at Dartmouth College, Federal Wide Assurance (FWA#3095).
Patients
We utilized patients diagnosed with primary NMIBC in two independent cohorts for a biomarker discovery/ validation approach.
The New Hampshire (NH) Population-based Cohort was comprised of bladder cancer patients diagnosed in the state of NH between 2002–2004. Eligible cases were residents of the State of NH at the time of diagnosis identified using the State Cancer Registry, hospital pathology departments, and hospital cancer registries, as described. Information on bladder tumor clinicopathological features and recurrences was obtained from medical records, or provided by the treating hospital(s) (both in and outpatient records, including any pathology reports) covering the 2-year follow-up period (
For the Dartmouth-Hitchcock Medical Center (DHMC) Cohort, we retrospectively selected a sequential set of bladder cancer patients identified through the hospital tumor registry diagnosed in the years 2008–2014. For this miRNA expression assessment project, we identified a subset of those patients with the clinically challenging non-muscle invasive histologic types: primary Ta High-Grade and T1 tumors (TaHG/T1) who had archived formalin-fixed, paraffin-embedded (FFPE) tissue blocks. We ensured that all tissue samples utilized represent the tissue that was removed prior to the administration of any intravesical immunotherapy or chemotherapy. We reviewed the patient medical records to ascertain clinical information related to recurrence and progression events. The study pathologists reviewed the hematoxylin and eosin (H&E)-stained slide and circled the non-cauterized tumor portion. For each tumor, we performed macrodissection on several of the matching 10 micron unstained tissue sections to select the circled portions. RNA was isolated from this portion using Qiagen Deparaffinization Reagent followed by the Qiagen AllPrep FFPE tissue kit (Qiagen Inc, Germantown, MD, USA) (
Tissue-matched bio-fluid samples
On a subset of DHMC bladder cancer patients with tumor tissue samples, we also collected blood and urine samples post-diagnosis (
Total RNA was extracted from (3) 20-
miRNA expression levels
The Dartmouth Genomics Shared Resource used the Nanostring human v3 microRNA expression assay (NanoString Technologies, Seattle, WA, USA) to quantify the levels of 800 miRNAs. Specific tags are ligated to the 3’ end of each miRNA. miRNA is then hybridized to a panel of miRNA:tagspecific nCounter capture and barcoded reporter probes. The nCounter Digital Analyzer counted individual fluorescent barcodes and quantified the target RNA molecules present in each sample. We used Nanostring Nsolver 3.0/4.0 software to normalize the count data to the positive controls and to average geometric mean of the top 100 detected miRNAs. We estimated the background level using the counts in the negative controls (mean
Cell lines and reagents
The NMIBC cell line HTB-2 (RT4) was obtained from ATCC (Manassas, VA, USA). Cells were cultured in McCoy’s growth medium (Sigma-Aldrich, Burlington, MA, USA) with 10% fetal bovine serum and 1% Penicillin-Streptomycin.
RT-qPCR assays
HTB-2 cells (10
Cell growth assays
HTB-2 cells were seeded in triplicate at 8,000/well in 96-well plates. The next day, cells were treated with dose ranges of C1632 for 5 d. Relative numbers of adherent cells were determined by sulforhodamine B (SRB) staining as described [15], and staining intensity was quantified by spectrophotometry (Abs
Non-muscle invasive bladder cancer patient characteristics by cohort
Non-muscle invasive bladder cancer patient characteristics by cohort
NH population cohort top-ranking miRNAs for recurrence
Among 331 detectable miRNAs.
Patient and tumor characteristics stratified by let-7f-5p
HTB-2 cells were plated in triplicate and grown to 90% confluency in flat bottom cell culture 12-well plates (Corning, NY, USA). Monolayers were scratched using a 1-mL pipette tip, and dead cells were washed off. Cells were grown for an addition 72 h, after which adherent cells were fixed and stained with 0.5% crystal violet in 20% methanol. Plates were scanned using an Epson Perfection V600, and migration distance was measured using the ruler function in Adobe Photoshop software.
Target gene analysis
To assess a priori target genes, we performed a secondary analysis of data from bladder tumors assayed as part of the National Cancer Institute’s Genomic Data Commons (GDC) project, restricting to
Statistical analyses
We defined the first recurrent tumor as any tumor identified following a disease-free remission period, more than 90 d after the date of initial primary bladder tumor diagnosis. These recurrent tumors include subsequent tumors of the same level of invasiveness, as well as those progressing to a higher stage/grade. Persistent primary tumors that did not have a remission period were excluded from the analysis of recurrence. Time to recurrence was calculated as the time between the initial diagnosis date and the date of the first recurrence event. For progression, the event was the diagnosis of a tumor with a greater stage or grade than the initial primary bladder tumor. If no events were reported, the date the patient was last seen documented in the medical record was used for censoring.
Median times to first recurrence, progression or survival were calculated using the Kaplan-Meier method. Multivariable analysis of time to the first bladder tumor recurrence, progression and survival analyses were performed using Cox-proportional hazards regression analysis. We tested the Cox model proportional hazards assumption by checking the independence of the scaled Schoenfeld residuals with time, and that the deviance residuals were symmetric. The standard prognostic model included adjustment for age at diagnosis of first bladder tumor, gender, tumor size (
Recurrence-free survival prognosis of NH and DHMC cohort by Let-7f-5p levels in tumor tissue. Kaplan-Meier plots depict the probability of recurrence stratified by let-7f-5p levels 
Correlation of Lef-7f-5p levels in urine and plasma versus tumor tissue. Tissue-matched bio-fluid specimens from the DHMC cohort were assayed for let-7f-5b levels relative to tumor tissue. Let-7f-5p expression levels in both urine (
The effect of Lin28 inhibition on Let-7f-5p expression, cell viability, and cell migration in non-muscle invasive bladder cancer cells. (A) Let-7f-5p levels in HTB-2 cells were analyzed by qPCR following treatment with Lin28 inhibitor C1632 for 48 hours at the indicated concentrations (x-axis, 
As shown in Table 1, the NMIBC patient cohorts assessed for miRNAs included a majority of male patients (72–76%) diagnosed with bladder cancer in their 60’s. Multiple tumors were present in a subset of patients (26–30%), and less than half of patients had large tumors (
We used the NH population cohort as a ‘discovery’ set to identify tumor tissue miRNAs associated with time-to-first recurrence in a multivariable model adjusted for sex, age, multiplicity, tumor size, stage, and grade. miRNA expression was stratified by the median value. Out of the 331 detectable miRNAs ranked by p-value, Let-7f-5p had the strongest association with recurrence in the NH population cohort (
To establish if blood and urine could also be used for minimally-invasive assessment of prognostic miRNA levels, we used a set of blood (
Because post-transcriptional downregulation of let-7 miRs can be attributed to Lin28 activation [17], we asked if inhibition of Lin28 with small molecule Lin28 inhibitor C1632 would result in increased let-7f-5p levels in NMIBC cell line HTB-2. Indeed, treatment of HTB-2 cells with Lin28i induced a dose-dependent increase in let-7f-5p (Benjamini Hochberg FDR-adjusted
We also used publically available data to perform a secondary analysis of Let-7f target genes in bladder tumor tissue, restricting to
Discussion
In the present study, we sought to identify prognostic miRNAs in NMIBC tumor tissue. We used patient cohorts representing two different clinical settings: population-based cohort representing patients who were residents of New Hampshire, including those treated in community hospitals, and a referral hospital cohort representing patients seen at Dartmouth-Hitchcock Medical Center. We identified let-7f-5p as a prognostic miRNA that reliably corresponds with probability of recurrence-free survival in both cohorts examined. Furthermore, we found that let-7f-5p levels in tumor tissue correlate with levels in plasma and urine, which can be collected without biopsy. Thus, Let-7f-5p expression may have clinical utility as a biomarker of recurrence.
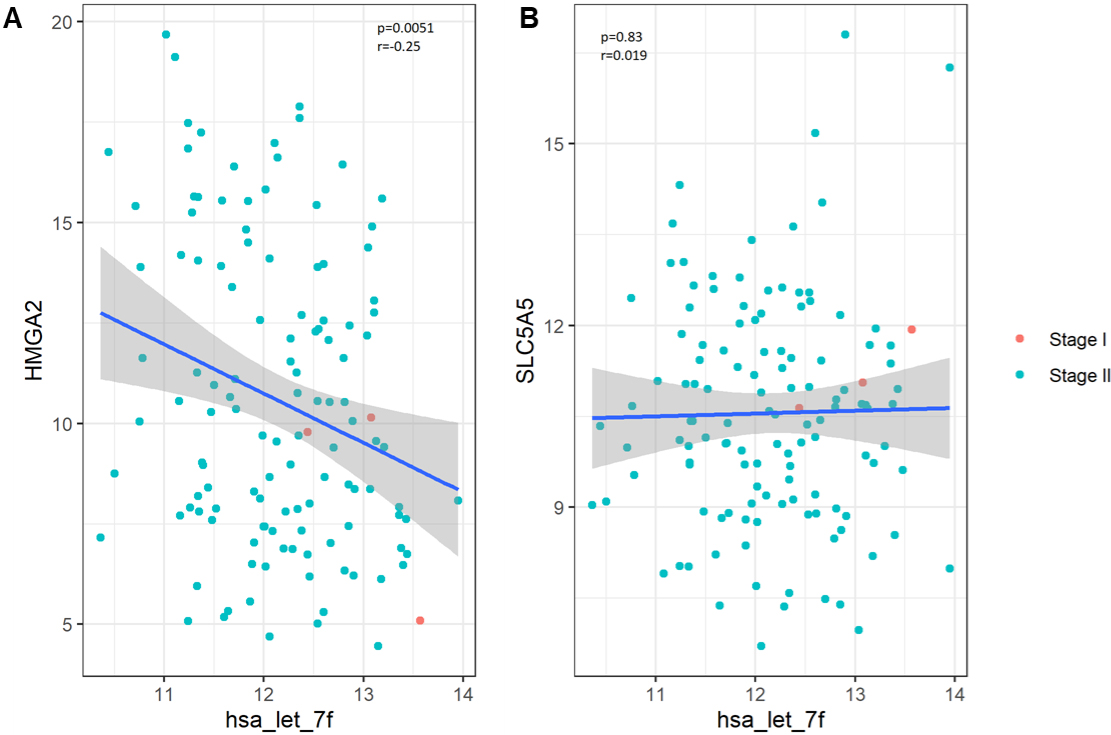
The let-7 family of miRNAs is comprised of thirteen different members found on nine different chromosomes, and expression of let7 miRNAs are frequently downregulated in multiple tumor types [18]. Importantly, let7 miRNAs are thought to regulate tumorigenesis by targeting multiple oncogenic factors, including c-MYC, Ras, high-mobility group A (HMGA), and others, by targeting transcripts of these proteins directly or by targeting their downstream effectors [19]. Manipulating Let-7f levels in cell lines from the thyroid and the lung modified levels of HMGA2, and of the sodium iodide symporter (NIS; SLC5A5) [20, 21, 22, 23, 24]. Our analysis of levels of these genes in bladder tumors showed Let-7f has a statistically significant inverse correlation with HMGA2, but not with SLC5A5.
Because of its action as a tumor suppressor miRNA, reduction of let-7 levels may enable the oncogenic activity of these factors, leading to tumor formation and progression. Due to the importance of let-7 miRNAs in tumorigenesis, these miRNAs may be useful biomarkers with prognostic value. Indeed, let-7f has been identified as an indicator of early stage ovarian cancer, and is also downregulated in the tumor tissue and plasma of colorectal carcinoma patients [25, 26]. To our knowledge, the prognostic utility of let-7 miRNAs assessed in urine and plasma biofluids in the context of NMIBC has not been explored. Urinary exosomal Let-7f levels were 1.73-fold lower in high-grade bladder cancer patients (
There is currently a critical need for new therapeutic strategies to prevent recurrence in NMIBC. Our findings suggest that Lin28, a negative regulator of let-7 miRNAs, may be a therapeutic target for NMIBC. Lin28 is an RNA binding protein whose oncogenic activity is attributed to its role in selectively targeting let-7 for degradation by DIS3L2 exonuclease [31]. Inhibition of Lin28 by the small molecule C1632 (Lin28i) can inhibit stemness and induce differentiation of mouse embryonic stem cells, as well as inhibit proliferation of human cancer cells [32]. Accordingly, we treated NMIBC cell line HTB-2 with Lin28i and observed an increase in let-7f-5p, suggesting Lin28 acts as a negative regulator of let-7f-5p in these cells (Fig. 3). Further analysis showed that Lin28i reduced HTB-2 cell viability, colony formation, and migration (Supplemental Figure). Taken together, these observations suggest that Lin28 inhibition may reactivate the tumor-suppressive functions of let-7f-5p and could serve as a novel therapeutic opportunity in NMIBC.
A limitation of our study is that the cohorts may not be generalizable to populations with other demographics. New Hampshire has proportionally higher rates of bladder cancer diagnosis compared to the rest of the country, found to be due at least in part to higher levels of arsenic in drinking water [33]. Additional national and international studies are thus required to generalize these findings to the broader population. An additional limitation is sample size, particularly in the DHMC cohort, which may have limited interpretations of significance in our validation analysis. However, despite these limitations, our study suggests that determination of let-7f-5p expression may highlight patients at elevated risk for recurrence, that let-7f-5p represents an optimal biomarker because expression in tumor tissue correlates with expression inin urine and plasma, and that targeting the Lin28-Let-7 axis may represent a novel therapeutic opportunity in NMIBC. These findings offer potential precision medicine opportunities with important clinical implications, and warrant further study.
Footnotes
Acknowledgments
This publication was funded in part by grant numbers F30CA216966, R21CA182659, K07CA102327, R03CA121382, R03CA099500, P42ES07373, P20GM 104416, and P30CA023108 from the National Cancer Institute, NIH, from the National Institute of Environmental Health Sciences, NIH, the National Center for Research Resources, NIH, and the National Institute of General Medical Sciences, NIH. The New Hampshire State Cancer Registry is supported by the Centers for Disease Control and Prevention’s National Program of Cancer Registries through cooperative agreement U58/ DP000798 awarded to the New Hampshire Department of Health and Human Services, Division of Public Health Services, Bureau of Public Health Statistics & Informatics, Health Statistics and Data Management Section.
Conflict of interest
The authors have no conflicts of interest to disclose.
Supplemental Figure
The effect of Lin28 inhibition on Lin28 expression, cell viability, and cell migration in non-muscle invasive bladder cancer cells. (A)–(C) Lin28A (A), Lin28B (B), and Actin (C) levels in HTB-2 cells were analyzed by qPCR following treatment with Lin28 inhibitor C1632 for 48 hours at the indicated concentrations (x-axis, 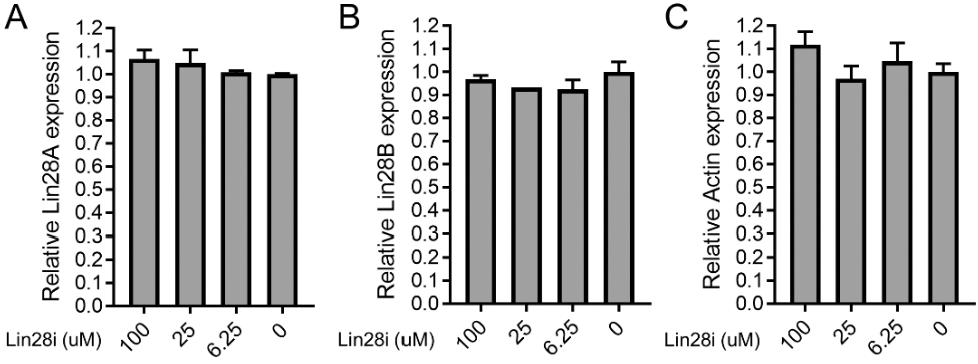
The effect of Lin28 inhibition on cell viability, and cell migration in non-muscle invasive bladder cancer cells. (A) Representative wells from Sulforhodamine B (SRB) assay with HTB-2 cells following treatment with C1632. Experiment was performed in triplicate wells. Quantification is shown in manuscript Fig. 3B. (B) Representative wells from scratch assay with HTB-2 cells following treatment with C1632. Quantification is shown in manuscript Fig. 3C.
The relationship between Let-7f and putative target genes in bladder tumor tissue. We performed a secondary analysis of Let-7f target genes in bladder tumor tissue data from the National Cancer Institute’s Genomic Data Commons (GDC) project, restricting to