Abstract
BACKGROUND:
Zinc finger MYND (Myeloid, Nervy and DEAF-1)-type containing 8 (ZMYND8) is closely correlated with tumor proliferation and invasiveness. However, its prognostic value has not been estimated in colorectal cancer (CRC).
OBJECTIVE:
We aimed to elucidate the prognostic significance of ZMYND8 expression and the pN and pM classification supplemented by its expression in CRCs.
METHODS:
The candidate gene ZMYND8 is identified by TCGA database and GEO database, and then we retrospectively evaluated the status and prognostic significance of ZMYND8 expression of 174 patients with CRC.
RESULTS:
Online data showed high expression of ZMYND8 is closely correlated with worse overall survival. Our study revealed high expression of ZMYND8 in CRC patients was significantly associated with worse overall and disease-free survival (
CONCLUSION:
The combined prognostic model could improve the ability to determine the clinical outcome of patients with CRC.
Introduction
Colorectal cancer (CRC) is one of the most prevalent carcinomas and it is thought to be the fourth leading cause of cancer-related deaths worldwide [1]. Patients with CRC have the pessimistic prognosis due to predominantly lack of an effective method for early diagnosis in clinical practice. Despite the recent therapeutic strategies were improved for CRC therapy, the prognosis of patients with CRC remains to be unfavorable for the low overall 5-year survival of approximate 60% [2]. In view of the increased incidence and poor clinical outcome of CRC, it is extraordinarily important to understand the development of CRC and identify the related prognostic factors. A work reported that quite a few clinicopathological parameters including tumor histology, depth of invasion, and perineural invasion have been put forward as independent prognostic factor for CRC [3]. To more precisely improve the predictive accuracy of traditional and time-honored classifications including anatomic stage, histological type, and tumor grade, Eighth Edition of the AJCC cancer staging manual enrolled into the status of BRAF, KRAS and NRAS mutations as the prognostic indicator for CRC to have allow us to better understand the molecular underpinnings of CRC in terms of oncogenesis and drug resistance [4, 5]. Therefore, although traditionally proposed TNM stage has the high prognostic accuracy, there remains to be need to improve the accuracy of existing prognosis predictive system and especially combining molecular classification of cancer accepted as an imminent reality with clinical traditional classification.
Although various signaling pathways and abnormally expressed genes involving in CRC have been hitherto reported [6, 7, 8, 9], a plenty of biomarkers still have not been used to precisely evaluate the prognosis of patients with CRC. Therefore, it is imperative to find the novel signal pathway linking to molecular and biological changes of CRC, and further generate new prognostic biomarkers to more precisely predict prognosis of patients, which could be helpful of individual treatments for patients with CRC. Thus it is necessary to explore the novel biomarkers for predicting prognosis, and investigate new effective therapeutic target for CRC.
The bromodomain protein zinc finger MYND (Myeloid, Nervy and DEAF-1)-type containing 8 (ZMYND8) functioned as epigenetic reader, which was initially identified as an activated protein-kinase-C-binding protein (also called PRKCBP1 or RACK7) [10], and its domain architecture included a PHD finger, BROMO domain, and PWWP (also called triple PHD-BRD-PWWP) domain, which located at its N terminus involvement with Histone modification and DNA binding, and a C terminus MYND domain associated with protein-protein interactions [11, 12, 13, 14]. ZMYND8 is recognized as an interacting element of several complexes involving in transcription regulation [13, 14, 15, 16, 17] or acted as a crucial factor linking chromatin with DNA damage repair activities [11, 13, 14, 18, 19]. Impressively, ZMYND8 inhibits tumorigenicity in the study of cervical cancer, prostate cancer and breast cancer [20, 21, 22]. Above mentioned likely explains the critical role of ZMYND8 in chromatin regulation, DNA damage and tumor formation. It is also showed that ZMYND8 is required for efficient somatic hypermutation of the IgH variable regions of B cells, suggesting its association with immunity regulation [23]. A recent study described that ZMYND8 was regarded as a component of transcription coregulator complexes, which can play a role of activation or repression of transcription by interacting with different signaling-dependent program [24]. It could recognize transcription activation signatures including H3.1K36Me2/H4K16Ac to activate the expression of genes and exerts its function by interacting with several lysine demethylase (KDM) components, such as KDM5A or KDM5C [16, 17]. Therefore, ZMYND8 has the complicated transcription networks and could be served as the activator or repressor in different context. More recently, a study showed that ZMYND8 acetylation by P300 mediates hypoxia-induced factors (HIF)-dependent breast cancer progression and metastasis [25] and another study reported that overexpression of ZMYND8 via a positive feedback loop of the estrogen receptor (ER) pathway was more common in Luminal B subtypes and these partial patients with breast cancer have the worse survival [26]. The Cancer Genome Atlas (TCGA) Research Network found that ZMYND8 have a recurrent somatic copy number alteration in high-grade serous ovarian cancer [27]. Overexpression of ZMYND8 increases tumor angiogenesis in prostate cancer [28]. However, there are no relevant reports on the prognostic value of ZMYND8 and the traditional pN and pM classification supplemented by its expression in CRC. In this study, we investigated the expression status of ZMYND8 protein in CRC tissues by tissue-microarray-based immunohistochemistry and the combined prognostic model containing the pN, pM classification supplemented by the status of ZMYND8 expression whether could improve the ability to determine the clinical outcome of patients with CRC.
Materials and methods
Bioinformatics analysis of TCGA data and GEO data
Oncomine analysis found candidate genes that were significantly upregulated in colon adenocarcinoma according to the below method. The candidate genes were selected using the conditions as follow: firstly, the genes are significant upregulation in samples of colon adenocarcinoma compared with that of normal colon tissues, (false discovery rate [FDR]
Patients and tissue specimens
We collected paraffin samples from September 2012 to November 2014 in Department of Pathology, Sun Yat-sen University Cancer Center. In this study, the diagnostic criterion of all tissue samples was based on the 2004 WHO criteria for tumor classification, and tumor staging was on the basis of UICC (International Union against Cancer) and the AJCC (American Joint Committee on Cancer). The clinicopathological features of patients with CRC are shown in Table 1. The CRC cohort is composed of 103 (59.2%) men and 71 (40.8%) women with median age of 58 years. The follow-up period ranges from 9.0 to 69.0 months (median, 53.9 months). Forty-four patients (25.3%) with late stages (III and IV), and other 130 patients (74.7%) were identified as early stages (I and II), and 22 of 174 patients metastasize to distant organs when were diagnosed.
Correlation between the ZMYND8 expression and clinicopathological variables in CRC patients
Correlation between the ZMYND8 expression and clinicopathological variables in CRC patients
Age (1): median; Tumor size (2): median; MSI status (3): Microsatellite instability.
ZMYND8 protein was stained by IHC according to standard EnVision
Online databases analysis of ZMYND8 expression in CRC. Oncomine analysis of ZMYND8 expression in human cancers compared to normal cells. 10% gene bank; 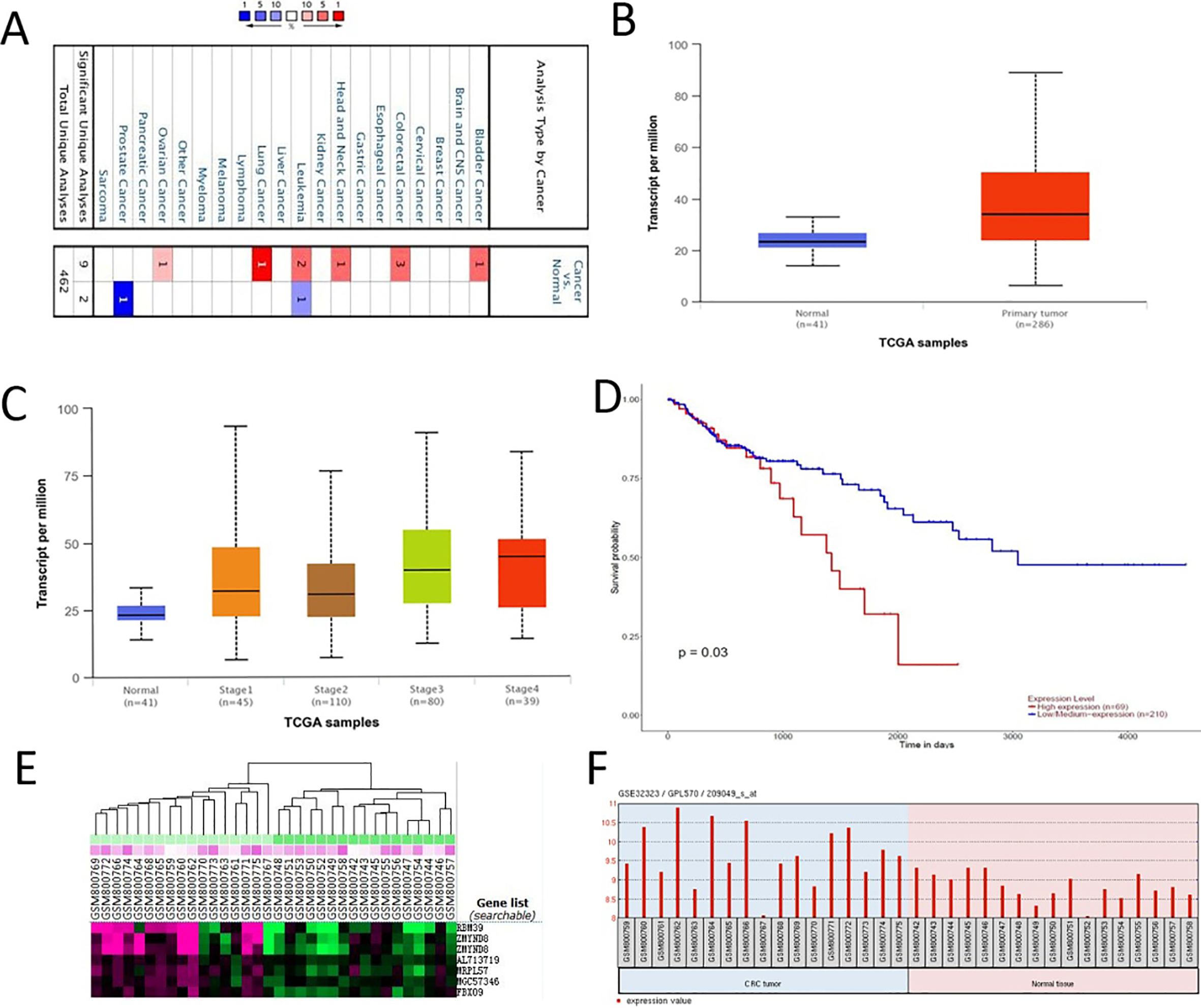
Expression of ZMYND8 protein in CRC tissues. A (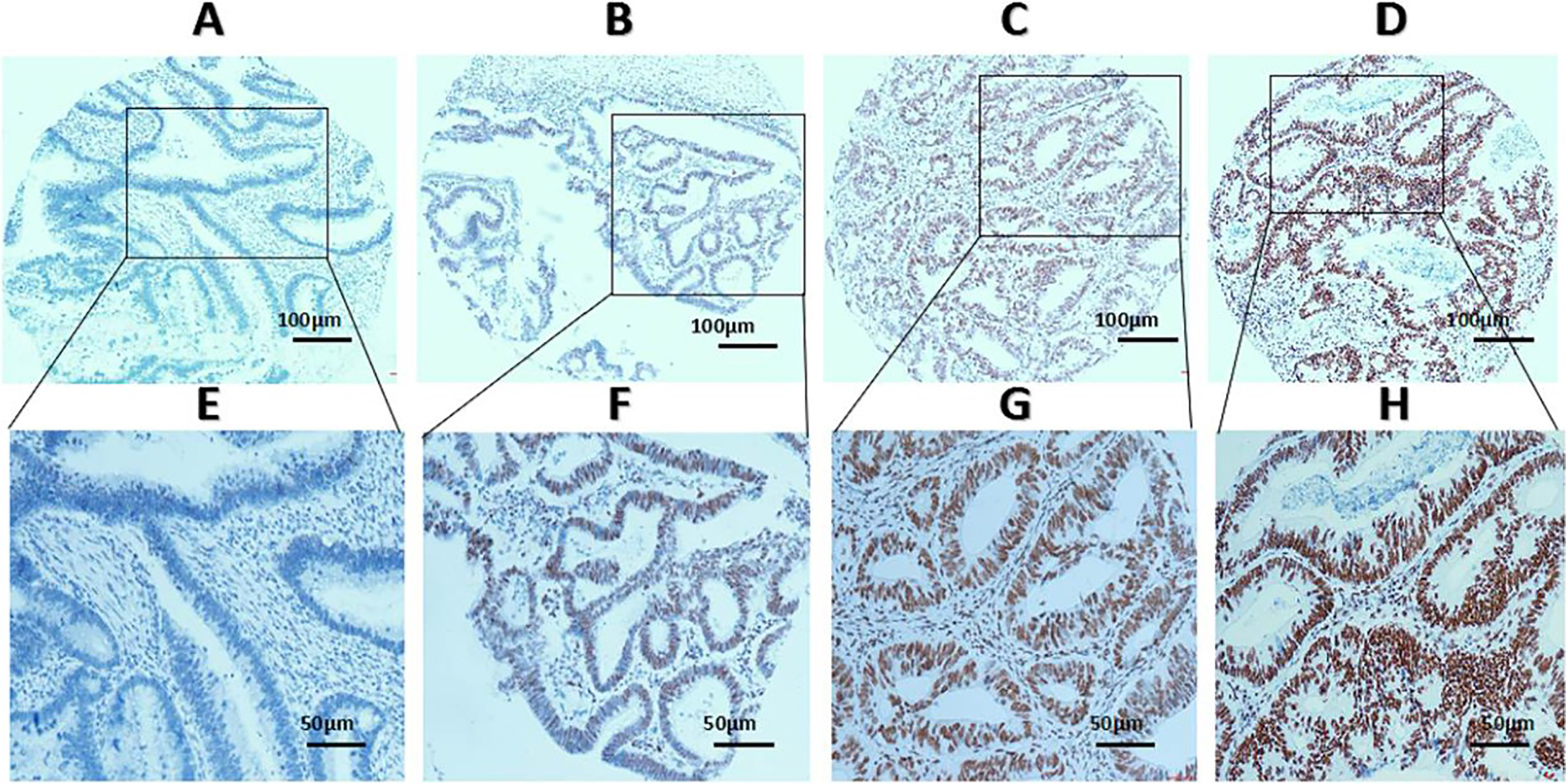
The status of ZMYND8 protein expression was assessed according to the method mentioned below. The presence of nuclei dark brown was identified to be positive for ZMYND8 expression, and scoring criteria as follow: each sample tissue harbored an intensity score (I score) from 0–3 (such as I0, I1, I2 and I3: I0-negative expression, I1-weak expression, I2-moderate expression, and I3-strong expression). Subsequently, ZMYND8 was elucidated according to the percentage of positively stained cells in 5% increments ranging from 0% to 100%, which performed a percentage score (P score). The total H score (range from 0 to 300) was calculated by multiplying each I score and P score. (Such as H score
Selection of cutoff value
The plot of TPF (sensitivity) versus FPF (1-speci- ficity) by variable cutoff values formed a curve which named an receiver operating characteristic (ROC) curve in the unit square, ROC curve analysis could calculate the optimal cutoffs by judging the closest distances from the point in the different curves to the point (0.0, 1.0) or (1.0, 0.0) [30, 31]. The sensitivity and specificity for each clinicopathological variable was plotted for determining the status of ZMYND8 expression in the study, and thus generating the corresponding ROC curves for the ZMYND8 score. The score was selected as the cutoff value that had both maximum sensitivity and specificity for evaluating the status of ZMYND8 expression. The score less than or equal to the cutoff value was regarded as low expression of ZMYND8, and more than the cutoff value was recognized as that of high expression. The clinicopathological characteristics were enrolled in the ROC curve analysis, including survival outcome, clinical stage, pT status, pM status, pN status and tumor relapse.
Statistical analysis
Statistical analyses were performed using SPSS software, version 16.0 (SPSS, Chicago, USA) and R, version 3.3.1 (
ROC curve analysis was employed to determine the cutoff value for the status of ZMYND8 expression in CRC. The sensitivity and specificity for each outcome was plotted: Survival outcome (A), Clinical stage (B), pT stage (C), pM stage (D), pN stage (E), Relapse (F).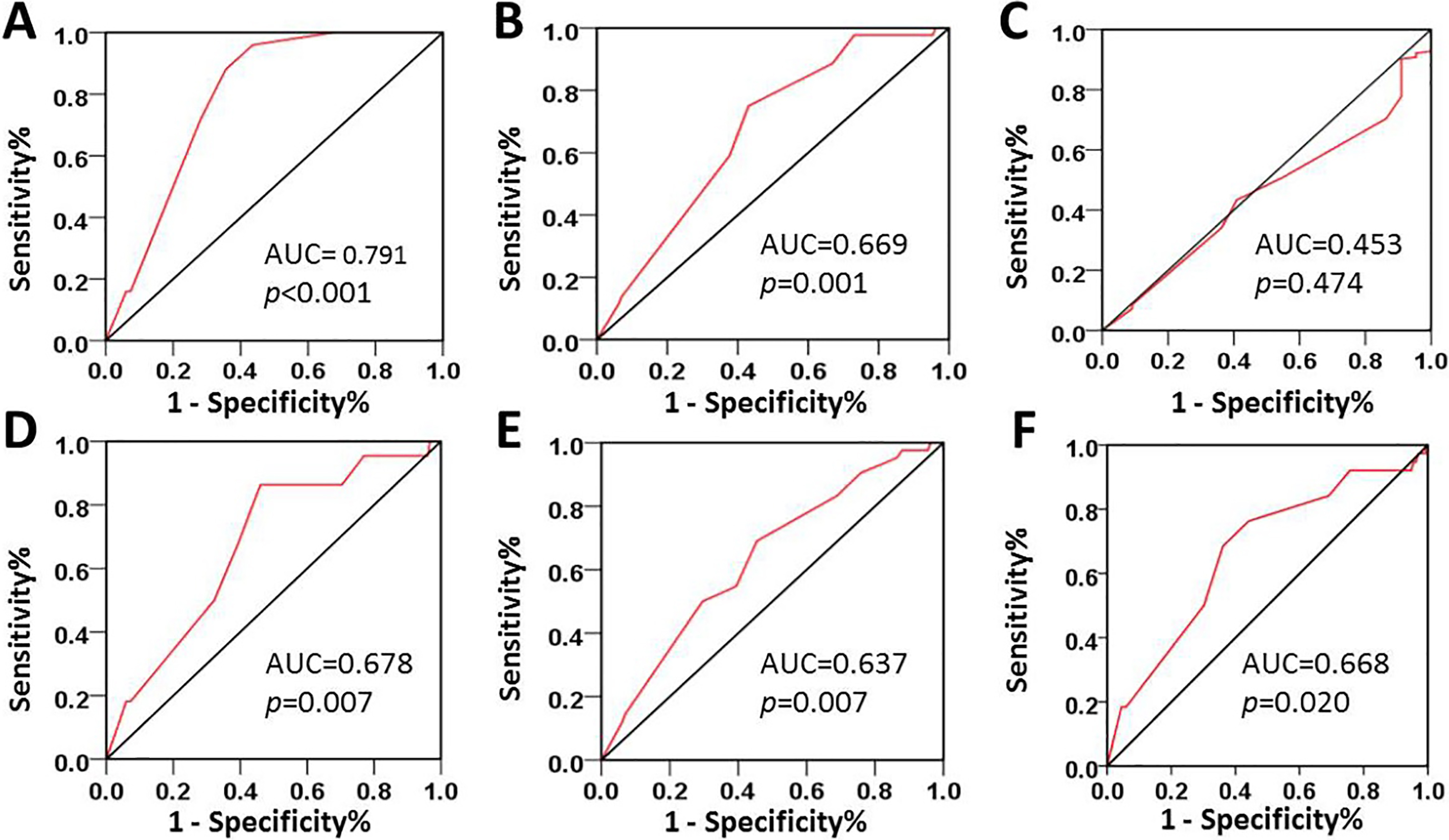
Kaplan-Meier survival analysis of ZMYND8 expression in total CRC patients and subsets with N0 and M0 CRC patients and comparison of overall survival and disease-free survival. Survival curve of ZMYND8 expression in total CRC patients for OS (A). Survival curve of ZMYND8 expression in CRC patients with N0 stage for OS (B). Survival curve of ZMYND8 expression in CRC patients with M0 stage for OS (C). Survival curve of ZMYND8 expression in total CRC patients for DFS (D). Survival curve of ZMYND8 expression in CRC patients with N0 stage for DFS (E). Survival curve of ZMYND8 expression in CRC patients with M0 stage for DFS (F).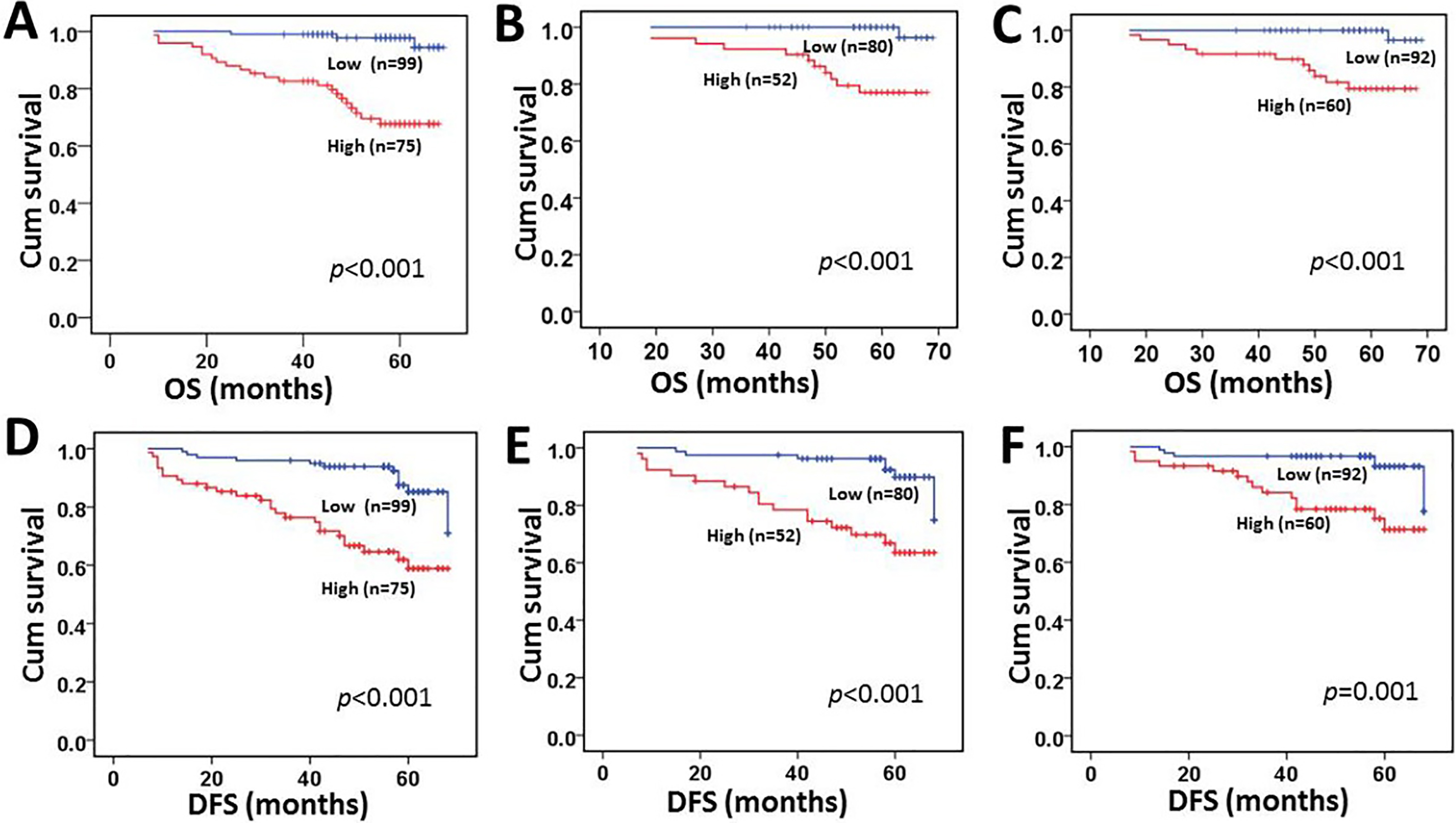
Identification of ZMYND8 in colon adenocarcinoma on the basis of the analysis of the online databases
We analyzed publicly available online databases of the gene expression, and selected genes over-expressed in colon adenocarcinoma compared with normal colon tissues. We chose ZMYND8 gene as a candidate potentially correlated with intestinal tumorigenesis. Oncomine analysis showed that ZMYND8 was markedly upregulated in colon adenocarcinoma and lung cancer (Fig. 1A). The results from the UALCAN analysis based on TCGA database indicated that ZMYND8 expression was higher in colon adenocarcinoma tissues (Fig. 1B) and samples of patients with different clinical stages (Fig. 1C) compared with normal colon tissue, and the high expression of ZMYND8 is closely correlated with worse overall survival (Fig. 1D). GEO database reveal ZMYND8 mRNA expression heatmap (Fig. 1E) and its expression was significantly upregulated in CRC tumors (
The selection of the cutoff score for ZMYND8 expression
The expression status of ZMYND8 protein is demonstrated in Fig. 2. To select a suitable cutoff score of ZMYND8 expression for further analysis, there is a point in the ROC curves of each clinicopathological characteristics closest to the point (0.0, 1.0) in Fig. 3, which maximizes both the sensitivity and specificity for the result according to the part of materials and methods mentioned. Tumor tissues with score more than the obtained cutoff value were regarded as high expression of ZMYND8. As it was shown in Fig. 3, a point in the corresponding ROC curve of the survival status is closest to the point (0.0, 1.0). On the basis of this outcome, we selected 220 score as cutoff value of ZMYND8 expression by the survival status for survival analysis. According to the ROC curve analysis, low expression of ZMYND8 is identified in 99/174 (56.9%) of CRC patients, and 75/174 (43.1%) with high expression.
The relationship between ZMYND8 expression and the clinicopathological variables of patients with CRC
The results demonstrated that high expression of ZMYND8 was significantly correlated with presence of nerve invasion, elevated serum CA199 level, pM1 stage, clinical stage and tumor relapse (
Univariate analysis of different prognostic factors in 174 patients with CRC
Univariate analysis of different prognostic factors in 174 patients with CRC
Univariate analysis showed a significant impact of well-known clinicopathological prognostic factors on CRC patients’ survival rates such as N classification, M classification, clinical stage and tumor relapse (
Multivariate analysis of prognostic factors for overall survival and disease-free survival in 174 patients with CRC
Multivariate analysis of prognostic factors for overall survival and disease-free survival in 174 patients with CRC
According to the results of our multivariate analysis, we proposed a new clinicopathologic prognostic model with 3 prognostic factors, including pN classification, pM classification and ZMYND8 expression. We designated a high-risk group harboring 2 or 3 prognostic factors among the presence of the advanced pN classification, pM1 classification and high expression of ZMYND8, an intermediate-risk group as the presence of 1 factor (the presence of the advanced pN classification or pM1 classification or high expression of ZMYND8), and a low-risk group as the presence of none (the pN0 classification, pM0 classification and low expression of ZMYND8). Our results revealed that the proposed model could significantly stratify the risk (low, intermediate and high) for overall survival (Fig. 5A,
The new combined model (including pN, pM classification and the status of ZMYND8 expression) could significantly stratify the risk (low, intermediate and high) for prognosis of CRC patients. Overall survival (A). Disease-free survival (B).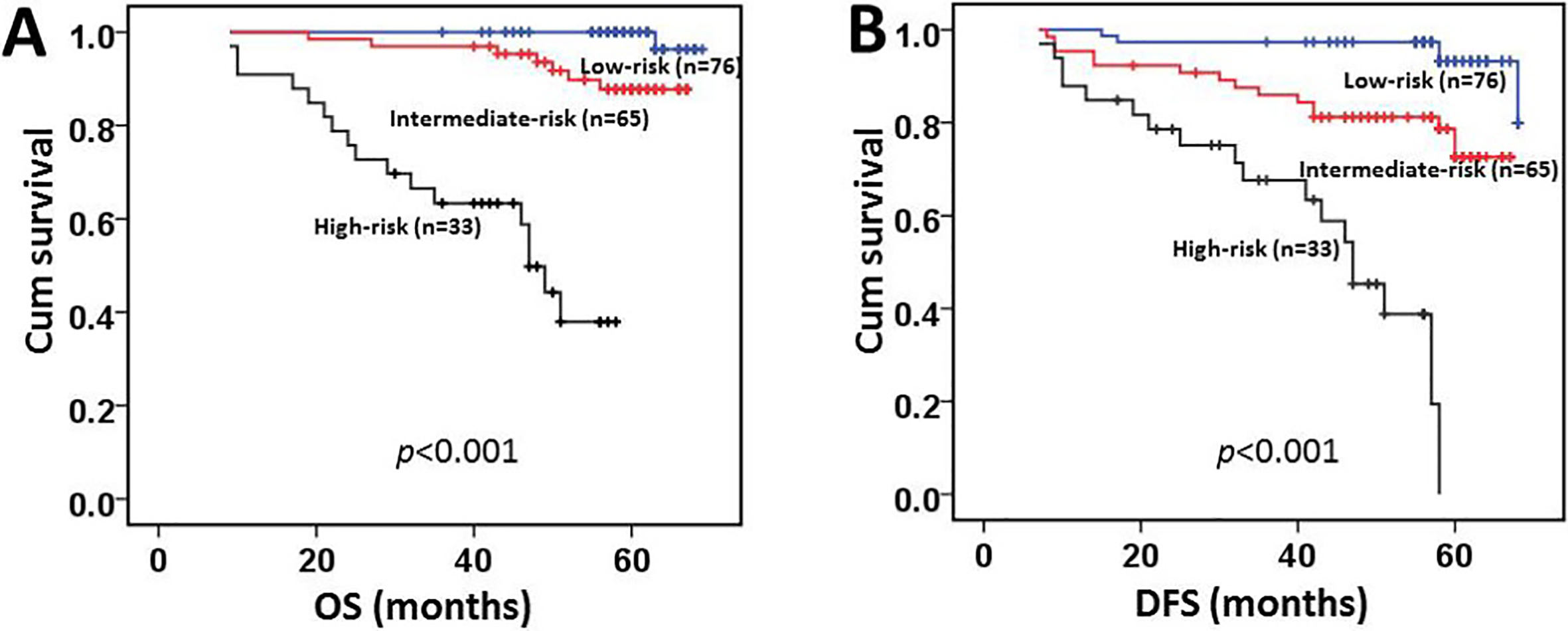
In this study, application of C-index to the proposed new prognostic model showed improved predictive ability when compared with the traditional pN and pM classification model (C-index of 0.597 vs 0.545, respectively).
CRC is one of the most common and lethal malignancies worldwide, accounting for approximately 1 in 10 cancer associated deaths [32], and thereby it is indispensable to improve the prognosis of patients with CRC by exploring ideal and individual management method. TNM stage is acted as the most conventionally prognostic factor of CRC. However, it is difficult to predict patients’ prognosis with these traditional factors due to the heterogeneity of this tumor [33].
In this study, we retrospectively evaluated the data containing 174 patients with CRC to determine the prognostic accuracy of the pN and pM classification compared with this classification supplemented by the status of ZMYND8 protein expression. Our findings demonstrated that the high expression of ZMYND8 in CRCs was significantly associated with pM1 stage, late clinical stage and the presence of tumor relapse. Nevertheless, multivariate analyses had the conclusion that high expression of ZMYND8 was a prognostic factor independent of well-known clinical factors, including pT stage, pN stage and pM stage and etc. The C-index analysis revealed that the proposed new prognostic model (combination pN classification, pM classification with the status of ZMYND8 expression) could improve the predictive ability when comparing with traditional pN and pM classification.
Our results of the status of ZMYND8 expression and the correlation between its expression and the outcome of patients with CRCs are consistent with that other studies provided. Our conclusions identified that high expression of ZMYND8 was significantly associated with worse overall survival and disease-free survival by univariate analysis. A group of data showed that high expression of ZMYND8 was more common in patients with Luminal B subtypes of breast cancer, which patients have the worse survival [26]. Another data found that ZMYND8 is overexpressed in prostate cancer xenografts and prostate cancer samples from patients and promotes their tumor angiogenesis [28]. ZMYND8 is induced by HIF-1 and HIF-2 and upregulated in human breast cells, and is correlated with poor survival of patients with breast cancer [25], supporting its potential roles of tumorigenesis, which is agreement with results our study showed, therefore, these above mechanisms also could involve in the CRC. However, in contrast to this conclusion, we previously reported the low expression of ZMYND8 based-immunohistochemistry was related with aggressive features and poor prognosis in nasopharyngeal carcinoma [34], which is consistent with other researches showed ZMYND8 regulates transcription and suppress cancer associated phenotypes by combining with members of the H3K4me3-specific KDM5 family, such as KDM5C and KDM5D [20, 21], the recent study also highlighted in nasopharyngeal epithelial cells, the association of EBV infection with aberrant modifications in histone bivalent marks, including H3K4me3 and H3K27me3 [35], therefore, we speculated that ZMYND8 could play the role of suppresser in nasopharyngeal carcinoma by parting histone modification members and the detailed mechanism remain to be investigated. In a word, ZMYND8 played the dual role in tumor regulation including tumorgenesis and tumor inhibition. Accordingly, the same gene may exert different molecular mechanisms on the various tumors. A comprehensive molecular characterization of human colorectal cancers showed that a low mutation rate (2/54; 3.7%) of ZMYND8 in a study harboring the 54 samples with mismatch repair deficient of 195 patients with colorectal cancers through TCGA network, and the research also manifested that no significant associations were found between ZMYND8 mutation and ZMYND8 mRNA, overall survival time [36]. Nevertheless, a prior research reported a mutation frequency (17/91; 19%) of ZMYND8 was found in mismatch repair deficient colorectal cancers [37], our results suggested that there has no strong relationship was showed between ZMYND8 expression and microsatellite instability (MSI) status. Therefore, the association of ZMYND8 mutation and MSI status and their roles in CRCs remain to be investigated due to the entirely relative low mutation rate in our data.
In addition, our findings support the idea that the pN and pM classification supplemented by the status of ZMYND8 expression could improve the predictive ability to determine the outcome of patients with CRCs, especially for the patients without lymph node metastasis or distant metastasis. To the best of our knowledge, the well-established factors including pTNM stage and the degree of tumor differentiation are the common risk factors influencing the prognosis of patients with CRCs. However, both of variables might have exposed their limitations in providing pivotal information affecting the prognosis and therapeutic strategies on the basis of specific clinicopathologic features and the extent of the tumor. Furthermore, several patients with same clinical stage has different clinical outcome, the reason why has not been well investigated [38]. Therefore, there is an urgently need for novel and objective method that can effectively differentiate the patients with different clinical outcome. In our study, the results support the content that the status of ZMYND8 expression, as detected by IHC, can distinguish whether patients with CRCs have the aggressive clinical behavior and/or unfavorable outcome. Thus, evaluation of the status of ZMYND8 expression could be regarded as a cost and effective factor for better predicting prognosis. The newly combined model can reflect the aggressively biological behavior of CRC and there are also strong efficacies improve their predictive ability by integrating the novel biomarkers into established clinicopathologic models [39]. Thereby the combined model could become meaningful for predicting the prognosis of patients with CRC.
Our study was a retrospective analysis, but all the clinicopathologic variables involved in the study were re-evaluated by two pathologists. Therefore, our results remained to be strengthened by the fact above mentioned. Finally, we proposed the new prognostic model in combination with pN, pM classification and the status of ZMYND8 expression by IHC that may be easily judged by the pathologists and more accurately predict the biological phenotype of patients with CRC. As far as we know, it is initially investigate the prognostic ability of the pN and pM classification supplemented by the status of ZMYND8 expression by IHC. However, it is necessary to be performed by multicenter works for the external validation of this combined model as well.
In summary, our data evidenced by univariate and multivariate analysis demonstrated that high expression of ZMYND8 was an independent biomarker for predicting the prognosis of CRC patients. The proposed new prognostic model in combination with the pN, pM classification and the status of ZMYND8 expression by IHC might improve the predictive ability to the prognosis of CRCs. Therefore, the status of ZMYND8 expression by IHC could be considered as an additional effective instrument to discriminate those CRC patients with high risk of tumor progression. The instrument might be of help for the clinician to select the ideal therapeutic strategies for the individuals.
Footnotes
Acknowledgments
This research is supported by grants from the Key Foundation of National Natural Science Foundation of China (81730072), the Youth Foundation of National Natural Science Foundation of China (81902420) and Guangdong Esophageal Cancer Institute Science and Technology Program Project (Q201903).
Conflict of interest
The authors declare no competing interests.
