Abstract
BACKGROUND:
The interest in plasma biomarkers has increased recently. Plasma exosome-derived microRNA-532 is aberrantly expressed in a variety of human cancers and has the prognostic value in many solid tumors. However, the prognostic impact of the expression value on AML remains unclear.
OBJECTIVE:
The aim of this study is to investigate the prognostic value of exosome-derived microRNA-532 in AML patients.
METHODS:
We performed the real-time PCR to quantify exosome-derived microRNA-532 in plasma of 198 AML patients. To assess the prognostic value, we performed Cox regression analyses in the context of well-established clinical and molecular markers. Cellular metabolic profile was conducted to help us understand the biological insight of its expression.
RESULTS:
The expression level was not associated with white blood cell counts, age, FAB subtypes, cytogenetic risk groups and genes of FLT3-ITD, NPM1, CEBPA and DNMT3A mutations. Interestingly, high expressers had a favorable overall survival in the univariate analysis. This prognostic value was testified in the multivariate analysis. Moreover, up-regulation of miR-532 was negatively associated with cellular energy like fructose and glutamine.
CONCLUSION:
We found plasma exosome-derived microRNA-532 can be used as a novel survival predictor for acute myeloid leukemia.
Background
Acute myeloid leukemia (AML) is a clinically and molecularly heterogeneous disease. Approximately 30% of newly diagnosed patients did not achieve complete remission and about 50% of them experienced the relapsed disease after intensive chemotherapy [1]. Therefore, the robust predictors are still of importance. To date, cytogenetic abnormalities like complex karyotypes and genes of NPM1, FLT3-ITD and CEBPA mutations were the well-established predictors in the clinical practice [2]. But these biomarkers are derived from the bone marrow blasts and require bone marrow biopsy and aspiration. Recent years observed an increased interest in the plasma biomarkers as potential diagnostic and prognostic biomarkers.
We have demonstrated high expression of huntingtin interacting protein 1 (HIP1) is associated with poor clinical outcome in AML patients [3]. But the exact role of HIP1 remains unclear. It involves in the cell-to-cell communication via the clathrin-mediated endocytosis, which was regarded as the mode of exosome internalization. Specifically, exosomes are membrane vesicles which could shelter micro-ribonucleic acids (miRNAs) from degradation, and then played important pathogenesis of tumors [4]. For example, it was reported that exosomes play a critical role in the lymphomagenesis by transferring the oncogenes between tumor cells and its microenvironment [4]. Moreover, utilities of exosomes as vehicles have now been proposed as a promising treatment strategy. The therapeutic mechanism of miRNA was either by restoring tumor-suppressive miRNAs or by inhibiting the oncogenic miRNAs. As far as we know, there were a lot of miRNAs derived from blasts with the biological insights or clinical significance in AML such as miR-181a/b, miR-124, miR-320, miR-362 and so on [5, 6, 7]. Similarly, plasma miRNAs are of interest because they will be used as a non-invasive tool to predict the survival of leukemia patients. For example, plasma miR-328, miR-339, miR-342 and miR-150 were down-regulated and miR-523 and let-7b were up-regulated in AML with diagnostic potentials [8, 9].
Recently, numeric studies demonstrated miR-532 is aberrantly expressed in a variety of cancers, such as ovarian cancer [10], gastric cancer [11], hepatocellular carcinoma [12, 13], bladder cancer [14] and chronic lymphoblast leukemia [15] and so on. Additionally, miR-532 plays an important role in cancer growth, metastasis, drug resistance and disease progression [4, 10, 11, 13, 14, 16, 17]. Taken together, it is reasonable for us to believe that exosome-derived microRNA-532 might provide new insight into the pathogenesis and disease progression of AML.
Methods
Patients
Clinical data were extracted from medical records of AML patients at the department of hematology in our hospital. Between 2014 and 2018, 198 patients with detailed diagnoses and treatment information were included in this study. Chromosomal abnormalities were described according to the International System for Human Cytogenetic Nomenclature [18]. Cytogenetic groups of patients were classified as favorable, intermediate, and unfavorable risks according to the NCCN guideline [19]. Cytogenetically normal AML (CN-AML) was defined as AML with the karyotype 46, XY [20] or 46 XX [20] in all 20 metaphase cells analyzed. Gene mutations of NPM1, FLT3-ITD, CEBPA and DNMT3A were analyzed by whole-gene sequencing as previously described [20]. Patients were treated with IA (idarubicin and cytorabine) based intensive induction chemotherapy as previous reported [21, 22]. In the consolidation therapy, younger patients were treated with a high-dose cytarabine-based chemotherapy [21]. The chemotherapy consolidation for elderly patients was decided by the physicians in an individualized way [21]. No patients in our study received allogeneic transplantation in the first induction remission. Patients with secondary AML or acute promyelocytic leukemia were excluded. All the study methods were carried out in accordance with the approved guidelines. The study was approved by the Institutional Review Boards of the First Affiliated Hospital of Zhejiang University, and performed in accordance with the Declaration of Helsinki.
Plasma samples collection and quantitative reverse transcriptase-PCR
Blood was drawn into ethylene diamine tetra acetic acid (EDTA) tubes from AML patients at diagnosis. We conducted the quality analysis control for the exosome isolation according to previous published methods [23]. Total RNA was extracted from 1 ml plasma with the exoRNeasy Serum/Plasma Midi Kit (Qiagen, CA, USA) according to the manufacturer’s instructions for isolation of total RNA. Total RNA from each sample were converted into cDNA using All-in-one miRNAsqPCR-PCR Detection Kit. The expression of miR-532 was evaluated by real-time polymerase chain reaction (PCR) analysis according to the manufacturer’s protocol. U6 snRNA was used as the endogenous control. Quantitative PCR was performed in triplicate using SYBR-Green PCR Master Mix kit (Takara, Japan) on an IQ5 real time PCR instrument (Bio-Rad, USA). Quantitative PCR was performed in triplicate using standard settings: 95
Cellular metabolomic profiles
Metabolomic profiling of bone marrow samples was performed on XploreMET platforms (Metabo-Profile, Shanghai, China) as previously described [20, 24]. A total of 132 metabolites were identified by the comparison with the internal library built with the standard reference compounds. The intensities of these metabolites were used to further analyze. Detailed descriptions of sample preparation and GC-TOFMS analysis methods are provided in the supplemental Appendix.
Statistical analysis
Patient characteristics were summarized using descriptive statistics, which included frequency counts, median, and range. The main objective of this study was to evaluate the prognostic impacts of miR-532 expression on Overall survival (OS) of AML patients. OS was defined as time from date of diagnosis until death due to any cause or the last follow-up. Relapse free survival was defined as time from date of complete remission until relapse, death and the last follow-up. The proportional-hazards assumption was checked for each variable before fitting Cox models. The multivariable analysis was carried out to determine the independent effect of miR-532 expression in the context of clinical and molecular factors. The “multtest” package was used to test for the difference of the cellular metabolite signatures between high and low miR-532 expressers. Hierarchical clustering based on expression levels of these metabolites was performed and visualized by heatmap. All statistical analyses were conducted with R statistic packages, version 3.6.1 (www.r-project.org). The two-sided level of significance was set at
Clinical characteristics of patients with different expression of exosome-derived miR-532
Clinical characteristics of patients with different expression of exosome-derived miR-532
Abbreviations: EsmiR-532, exosome-derived microRNA-532;
Survival curves of AML patients. Kaplan-Meier estimates of OS (A) by high and low miR-532 expression for our patients according to the three quartiles (Q1: 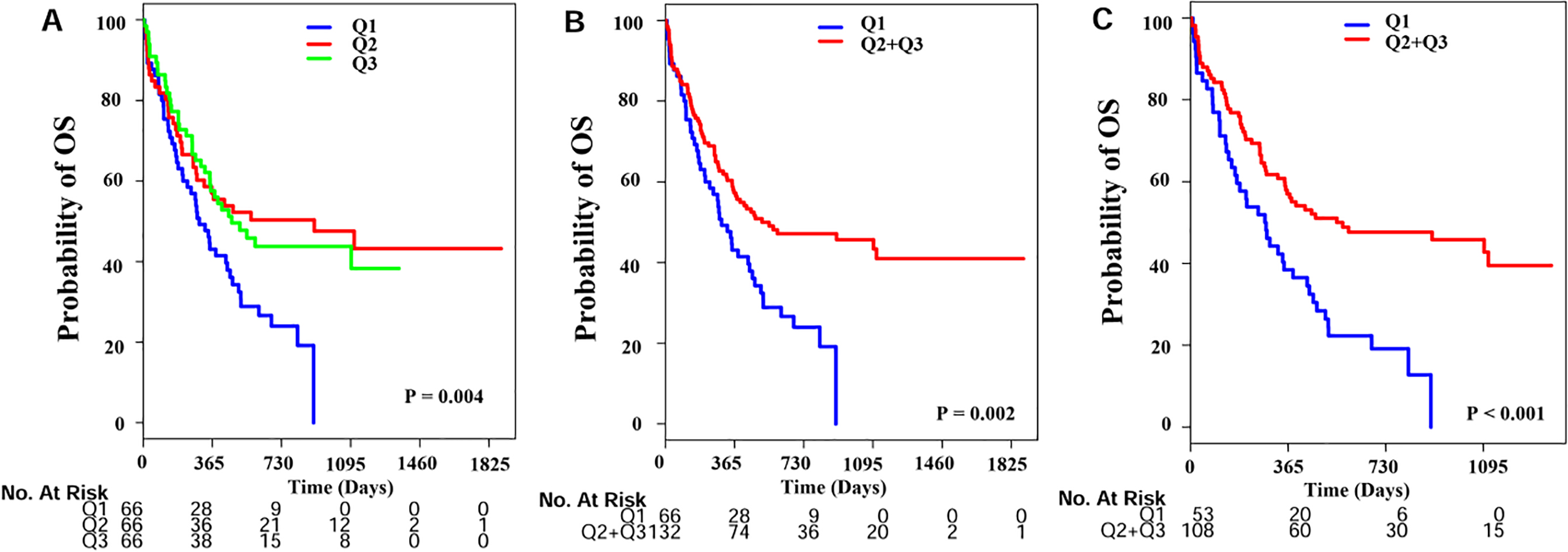
Characteristics of patients with high exosome-derived miR-532
Exosomal particles was isolated and observed by transmission electron microscopy (Fig. S1). CD81 and TSG101 protein as markers of the extracted exosomes were detect by Western Blot (Fig. S2). Exosome-derived miR-532 (esmiR-532) expressions were analyzed in 198 serum samples from AML patients at diagnosis. The non-normal distributions are illustrated in Fig. S3. The expression level of miR-532 in AML patients was similar to healthy controls (
Prognostic value of esmiR-532 expression in AML
The median follow-up of our living patients was 777 days. Three year survival rate for these patients was 36% with 95% confidence interval from 30% to 45%. The main objective of this study was to assess the prognostic value of esmiR-532 expression on overall survival (OS). High expressers (
Unvariate and multivariable analysis for OS in AML patients
Unvariate and multivariable analysis for OS in AML patients
Abbreviations were seen in Table 1.
Heatmap plot illustrating the different cellular metabolites between high and low miR-532 expression.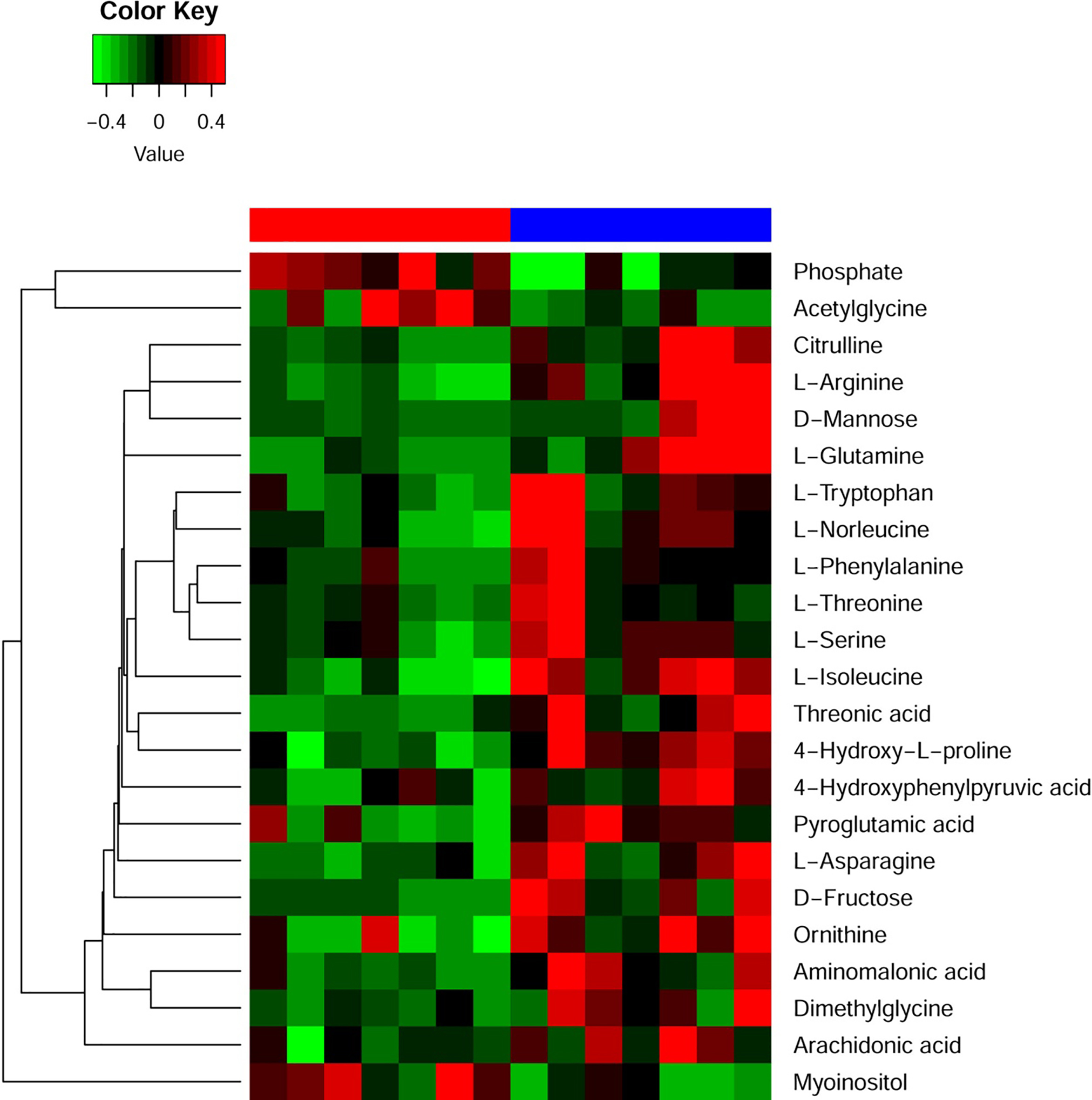
We analyzed the cellular metabolic profiles of 7 CN-AML patients with high and 7 with low expressers (Fig. 2). Resultingly, we found that 3 highly and 20 lowly expressed metabolites in the high esmiR-532 expression group. Of note, these metabolites enriched in eight metabolic pathways including aspartate metabolism, arginine and proline metabolism, glycine and serine metabolism, urea cycle, galactose metabolism, ammonina recycling, fructose and mannose degradation and amino sugar metabolism. It is worth noting that the major energy substrate for leukemia growth like cellular fructose and glutamine were low in high expression groups (Fig. 3).
The significantly expressed metabolites and the enriched pathways in high and low group. Red colors represent high and blue low levels of the metabolites.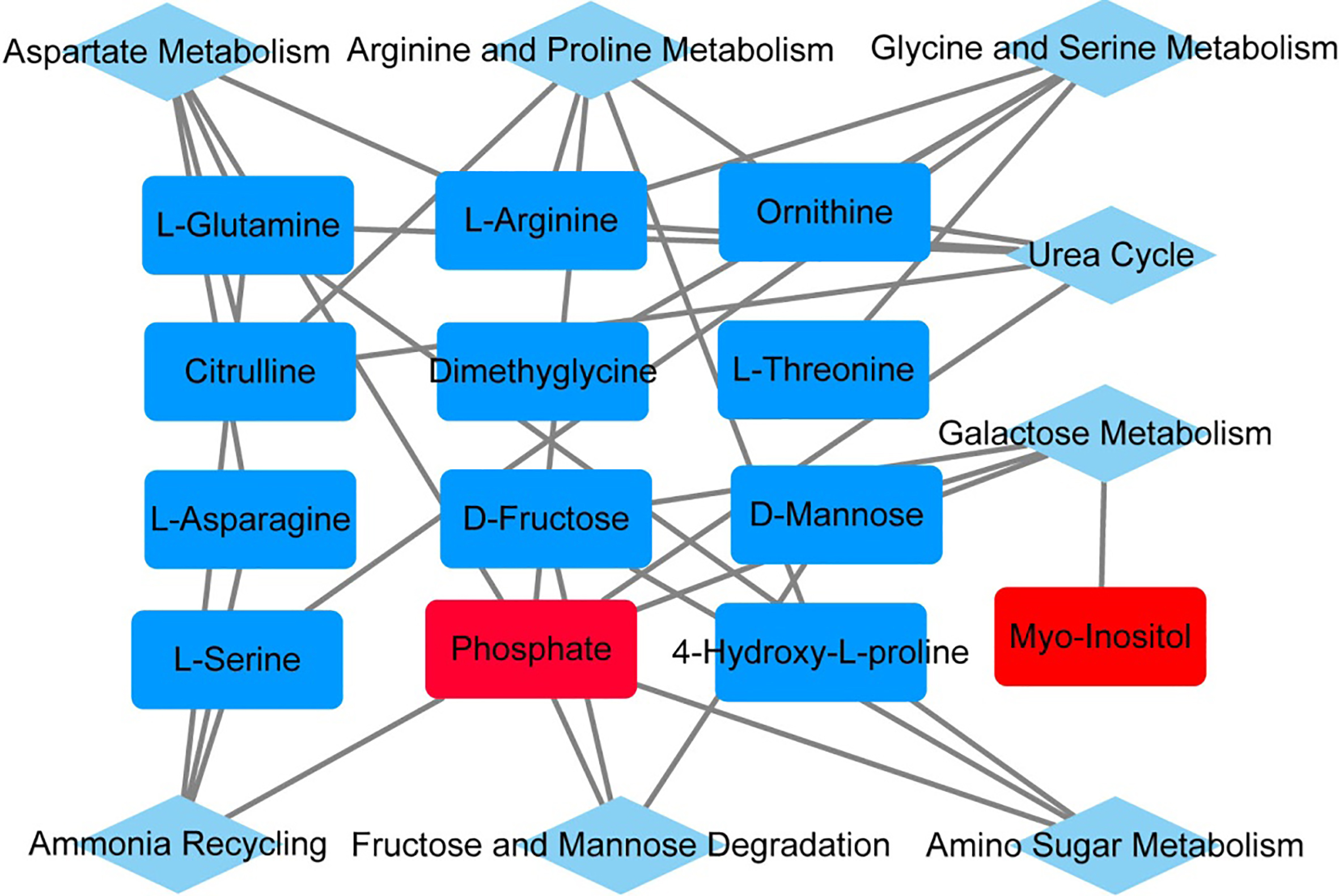
In order to improve clinical outcomes in AML patients, the identification of biomarkers for effective treatment is critical. Recent years observed an increased interest in the significance of microRNAs carried by exosomes as potential diagnostic and prognostic biomarkers. Here, we found that high expression of exosome-derived miR-532 (esmiR-532) in CN-AML had a favorable overall survival. And the benefit of esmiR-532 for overall survival might be associated with the lower cellular energy substrate like glutamine and fructose for leukemia growths.
MircroRNA-532 (miR-532) has diagnostic and prognostic value in solid tumors. Currently, there is no consensus about the biological function and clinical significance of miR-532 in cancer patients. Some researchers suggest miR-532 functions as an oncogenic miRNA in gastric cancer [11], colorectal cancer [16] and hepatocellular carcinoma [12]. However, other studies demonstrate miR-532 acts as a tumor suppressor in the bladder cancer [14] and chronic lymphocytic leukemia [15, 17]. But its clinical impact on AML patients remains unclear. With the rapid advancement of miRNA technology, the identification of the prognostic value of exosome-derived miRNA may be leveraged to develop new therapies to improve patient outcomes in the future. In this study, we found high expression of plasma exosome-derived miR-532 was positive associated with overall survival in univariate analysis. But, the expression level was not associated with the well-established clinical and molecular predictors such as age, white blood cell counts and cytogenetic risk group as well as genes of FLT3-ITD, NPM1 and DNMT3A mutations in AML patients. Even if we took these factors as the potential confounders, plasma exosome-derived miR-532 was still as an independent favorable predictor in multivariate analysis. Recently, miRNAs are proposed as key plays in cancer cell metabolism [9]. Cancer cells are characterized by rewiring the energy production networks to support rapid growth. In order to further understand the poor survival of plasma exosome-derived miR-532 expressed in AML, we conducted a cellular metabolomics study. The increased uptake of glucose, fructose and glutamine has been extensively described as a hallmark of cancer [25, 26]. Consistently, we found several aberrant expressed metabolic pathways existed in high expression group. Notably, fructose and glutamine concentrations were lower in high group than in low group, which implied miR-532 might inhibit the utilities of cellular energy substrate like fructose and glutamine, and then translating into favorable clinical outcome.
There are still some limitations in this study. Firstly, we only examine genes of FLT3-ITD, NPM1, CEBPA and DNMT3A mutations, thus we could not exclude other genes like IDH1/2, TET2, ASXL1 mutations those will confound the prognostic value of miR-532 expression in AML patients. Secondly, the association of miR-532 and the metabolites uncover the important regulatory networks, but the underlying mechanisms are required to further study in the future. Finally, functional study is still required to investigate the exact role of miR-532 in leukemia cell lines in vitro and in vivo models. Therefore, caution in application of our findings is still warranted.
Conclusion
In this study, we find elevated plasma exosome-derived miR-532 was associated with favorable overall survival, and negative associated with low levels of energy production like glutamine and fructose in AML.
Footnotes
Acknowledgments
This work is supported by Zhejiang Provincial Natural Science Foundation of China (LY19H080009, LY16H080004, LY20H080008), University Science and Technology Innovation Talent Support Program of Henan Province (17HASTIT046). We are very thankful to patients who enrolled in this study We are very thankful to Dr Yasmin Elhaimoussa at City of Hope/Biomedical Research Center for carefully polishing this article.
Conflict of interest
The authors declare no potential conflicts of interest.
