Abstract
MicroRNAs (MiRNAs) have been clarified as crucial regulators of the pathological processes in various carcinomas in the past years. Interestingly, existing evidence has manifested that microRNA-204-5p (miR-204-5p) is engaged in the initiation and progression of multiple carcinomas. However, the potential of miR-204-5p in cervical cancer remains to be disentombed. This study focused on unraveling the detailed role of miR-204-5p in cervical cancer. MiR-204-5p exhibited a low level in cervical cancer cells. The functional assays demonstrated that miR-204-5p upregulation exerted suppressive impact on the functions of cervical cancer cells, including proliferation, invasion, migration and epithelial-mesenchymal transition (EMT) process. Moreover, transcription factor AP-2 alpha (TFAP2A) was screened to be the most affected target gene by miR-204-5p, and TFAP2A was discovered to transcriptionally repress miR-204-5p in cervical cancer. The mutual regulation between TFAP2A and miR-204-5p was testified through molecular mechanism assays. Final rescued-function assays demonstrated that overexpression of TFAP2A could recover the suppressed cellular process caused by miR-204-5p upregulation. In conclusion, miR-204-5p/TFAP2A feedback loop promoted the proliferative and motorial capacities of cervical cancer cells. This finding suggested a novel modulatory loop of miR-204-5p/TFAP2A in cervical cancer, offering promising biomarkers for cervical cancer therapy.
Introduction
Cervical cancer is one kind of malignant tumor in gynecology [1, 2, 3]. Death rate caused by cervical cancer in developing countries and other areas with underserved medical condition remains high [4, 5]. The paucity of effective therapies is noted by a call for accurate diagnosis and better prognosis for cervical cancer patients. Increasing evidence has exposed that non-coding RNAs (ncRNAs) exert pivotal function on the tumorigenesis and progression of human carcinomas [6, 7, 8]. Thus, exploring potent molecules for cervical cancer treatment is the main goal of this study.
MicroRNAs (miRNAs), as endogenous transcripts with almost 22 nucleotides in length, are noted as possible causative agents in cancers [9, 10]. For example, miR-9 regulates BRCA1 and ovarian carcinoma sensitivity to cisplatin and PARP repression [11]. MiR-182 contributes to the leptomeningeal diffusion of non-sonic hedgehog-medulloblastoma [12]. MiR-20a/106a-WTX pathway mediates RhoGDIa/CDC42 axis for colon cancer progression [13]. MiR-224 plays a promotive part in the progression of non-small cell lung cancer [14]. In our research, miR-204-5p was selected. Lee et al. reveals the potential of miR-204-5p in minimal deviation adenocarcinoma of uterine cervix [15]. dbDEMC 2.0 website predicts miR-204-5p is lowly expressed in cervical cancer. As a tumor suppressor, miR-204-5p has been explored in multiple tumors, such as gastric cancer, colorectal cancer, breast cancer, as well as laryngeal squamous cell carcinoma [16, 17, 18, 19]. Nevertheless, the specific mechanism of miR-204-5p in cervical cancer remains to be disentombed.
As is known to all, transcription factors can modulate the expression of target genes, positively or negatively, through interacting with their transcription start site (TSS) [20, 21]. Transcription factor AP-2 alpha (TFAP2A) is identified as a transcription factor and influences the transcription of target genes in various cancers [22, 23, 24]. TFAP2A has also been reported as an oncogene in tumors [25, 26]. University of California, Santa Cruz (UCSC) genome browser assumed the potential role of TFAP2A in regulating miR-204-5p transcription. The biological regulator role of miR-204-5p in TFAP2A-mediated function was also unraveled.
In this paper, gain-of-function assays were conducted to evaluate cellular functions after increased miR-204-5p level. Then, the downstream targets and upstream molecular mechanism of miR-204-5p were analyzed. To summarize, this study revealed the influences of miR-204-5p/TFAP2A feedback loop on cervical cancer cell functions.
Materials and methods
Bioinformatics analysis
The expression pattern of TFAP2A in 306 CESC (Cervical squamous cell carcinoma and endocervical adenocarcinoma) samples and 13 normal samples was obtained from Gene Expression Profiling Interactive Analysis (GEPIA) database (
Cell culture
The normal cervical epithelial cells (H8) and four cervical cancer cells (HeLa, CaSki, SiHa, and C33A) were attained from American Type Culture Collection (ATCC; Manassas, VA, USA). With saturated humidity at 37
Cell transfection
MiR-204-5p mimics, miR-NC, miR-204-5p inhibitor and miR-NC inhibitor were synthesized and obtained from GenePharma (Shanghai, China). Specific short hairpin RNA (shRNA) against TFAP2A (shTFAP2A) and shNC, along with pcDNA3.1/TFAP2A and the empty vector, were obtained from Genechem (Shanghai, China). These plasmids were separately transfected into SiHa or CaSki cells through Lipofectamine 3000 (Invitrogen, Carlsbad, CA, USA).
qRT-PCR
The extraction of total RNAs from SiHa and CaSki cells was achieved by TRIzol reagent (Invitrogen). The gene-specific primers were designed by Primer3 Input (version 0.4.0) and evaluated by Oligo 6 software. Oligo DNA was chemically synthesized by
CCK-8 assay
Transfected SiHa or CaSki cells were cultured in 96-well dishes (2000 cells/well). Absorbance at 450 nm was read by utilizing a microplate reader after treatment for 4 h using 10-
EdU staining
EdU reagents (RIBOBIO, Guangzhou, China) were added for culturing transfected SiHa or CaSki cells in 96-well plates (2000 cells/well). 2 h later, cells were fastened with 4% paraformaldehyde (PFA; Sigma-Aldrich, St. Louis, MO, USA) in PBS (Sigma-Aldrich). Before incubation in 1
Colony formation assay
Transfected SiHa or CaSki cells were first inoculated into 6-well plates (500 cells/well), and incubated for 2 weeks. Upon this, visible colonies were counted via an inverted microscope (Olympus, Tokyo, Japan) after fixation in 4% PFA and dyeing in 0.1% crystal violet.
Wound healing assay
SiHa or CaSki cells were put in 6-well plates (1
Transwell assay
A Matrigel-coated Transwell chamber with 8-
RNA pull-down assay
Cell lysates of SiHa or CaSki cells were incubated with biotinylated miR-204-5p sense/antisense (named as Bio-miR-204-5p sense/antisense), compared with Bio-NC. Subsequently, magnetic beads (Invitrogen) were added. TFAP2A mRNA level was determined via qRT-PCR.
Luciferase reporter assay
The wild-type (WT) or mutant (Mut) sequences of TFAP2A 3’-UTR with or without the binding to miR-204-5p were sub-cloned into pmirGLO luciferase reporter vector (Promega, Madison, WI, USA), which were sequentially co-transfected with miR-204-5p mimics and miR-NC or miR-204-5p inhibitor and miR-NC inhibitor into SiHa or CaSki cells. The pGL3 reporter vector (Promega) containing miR-204-5p promoter was co-transfected with shTFAP2A and shNC or pcDNA3.1/TFAP2A and empty pcDNA3.1 vector into tumor cells, separately. Dual Luciferase Reporter Assay System (Promega) tested the luciferase activities.
Chromatin immunoprecipitation (ChIP)
ChIP was conducted using the Agarose ChIP Kit (Pierce). SiHa or CaSki cells were fixed in 1% formaldehyde to cross-link DNA and protein. Chromatin was digested and sonicated to chromatin fragments (200–1000 bp). The immunoprecipitation was carried out with TFAP2A or IgG antibody (Millipore, Bedford, MA, USA). The precipitated DNA was quantitated via qRT-PCR.
Western blot
Total protein was isolated from transfected SiHa or CaSki cells after treatment with RAPI protein lysate (Beyotime, Shanghai, China) and subsequently quantified using a BCA Protein Assay Kit (Pierce, Rockford, IL, USA). Protein was electro-transferred to PVDF membrane (Invitrogen) at 200 mA for 2 h upon separation by use of SDS-PAGE (Bio-Rad). Next, membranes were blocked for nearly 1 h in TBST (Sigma-Aldrich) with 5% skimmed milk and incubated for one night with primary antibodies against E-cadherin (ab40772, Abcam), Vimentin (ab8978, Abcam), N-cadherin (ab76057, Abcam), TMOD3 (ab157215, Abcam), ZNF282 (PA5-41464, Invitrogen), BIRC6 (ab19609, Abcam), TFAP2A (ab52222, Abcam) and GAPDH (ab9485, Abcam) at 4
Statistical analysis
Values were expressed as means
Results
miR-204-5p exerted anti-cancer function in cervical cancer
At first, miR-204-5p was found to exhibit downregulation in cervical cancer cells (Fig. 1A), especially in SiHa and CaSki cells. Then, miR-204-5p was overexpressed in SiHa and CaSki cells (Fig. 1B) for gain-of-function assays. In CCK-8 assay, cell viability was repressed by enhancing the level of miR-204-5p (Fig. 1C). Data from EdU assay revealed there were lesser EdU positive cells in transfected SiHa and CaSki cells, suggesting that miR-204-5p upregulation could inhibit cell proliferation (Fig. 1D). And data from colony formation assay further revealed that miR-204-5p overexpression hampered cell proliferation (Fig. 1E). Likewise, the result of wound healing analysis revealed that cell migration was suppressed by elevating miR-204-5p expression (Fig. 1F). Meanwhile, we observed through transwell invasion assay that invasive ability was restrained by miR-204-5p mimics (Fig. 1G). Besides, western blot assay was applied to estimate the levels of proteins involved in EMT process. The result showed that overexpressing miR-204-5p increased the level of E-cadherin while had opposite impact on that of N-cadherin and Vimentin (Fig. 1H). Taken together, miR-204-5p upregulation hindered cell functions in cervical cancer.
MiR-204-5p inhibited the proliferation, migration, invasion and EMT process of cervical cancer cells. A. MiR-204-5p level was revealed in normal cell (H8) and cervical cancer cells (HeLa, CaSki, SiHa, C33A) through qRT-PCR. B. The overexpression efficiency of miR-204-5p was assessed by qRT-PCR through transfection with specific miRNA mimics into SiHa and CaSki cells. C and E. CCK-8 (C), EdU staining (D) and colony formation (E) assays were performed in transfected cells for the detection of cell viability, EdU-positive cells and colonies. F. Wound healing for the migratory ability of SiHa and CaSki cells with increased level of miR-204-5p. G. Invasive SiHa and CaSki cells were measured by transwell invasion assay in response to the enhanced level of miR-204-5p. H. Western blot assay for the examination of EMT-related proteins in miR-204-5p-upregulated SiHa and CaSki cells. 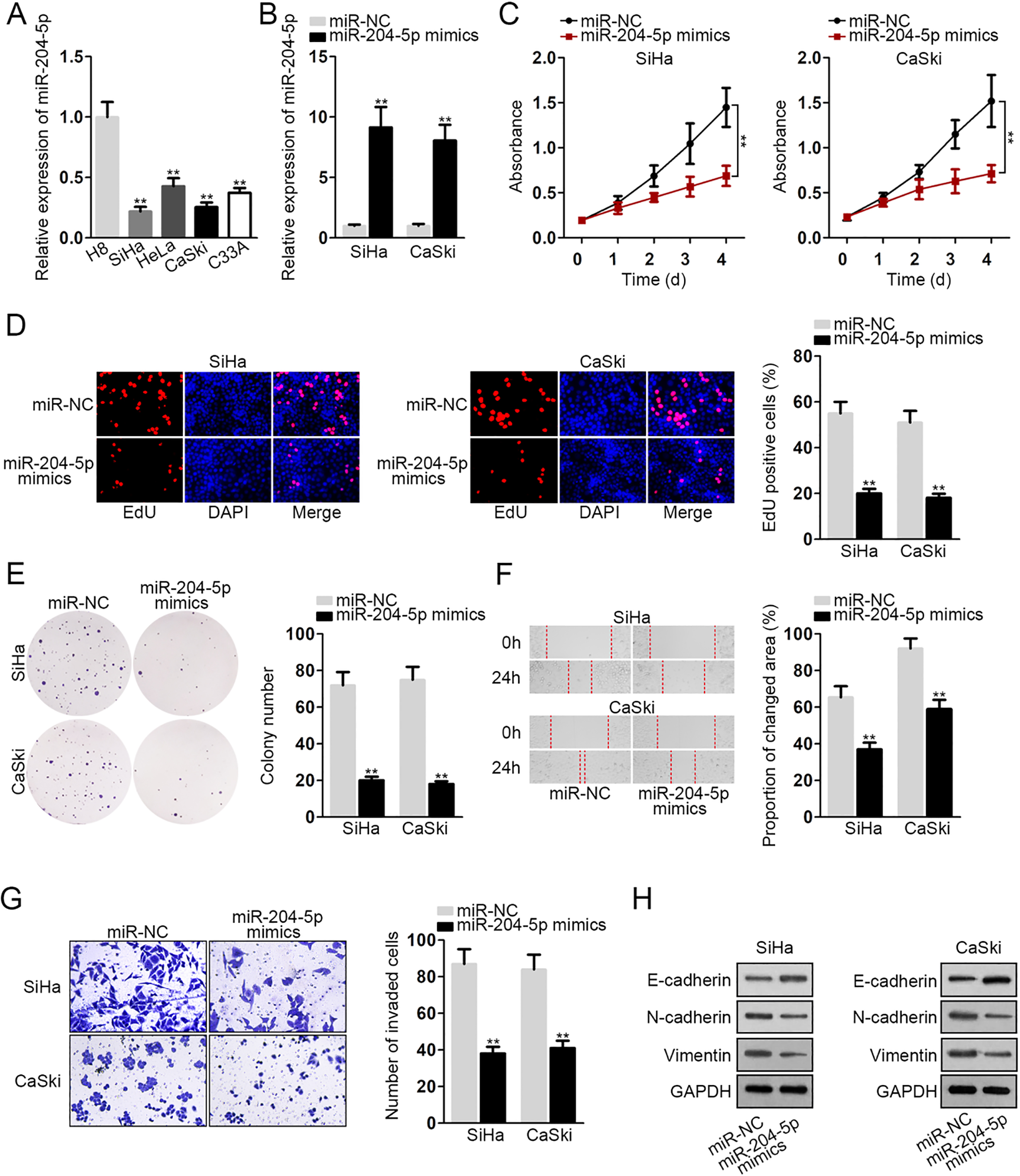
To explore the downstream molecular mechanism of miR-204-5p in cervical cancer, we screened from five online databases (microT, miRanda, PITA, PicTar and TargetScan) and obtained 23 shared potential targets (Fig. 2A). Afterwards, qRT-PCR detected the levels of these mRNAs in cervical cancer cells and the results were plotted into the heatmap. Four mRNAs (ZNF282, BRIC6, TMOD3 and TFAP2A) exhibited the relative highest level in tumor cells (Fig. 2B). Next, the impact of miR-204-5p on these four mRNAs was dissected. After overexpressing miR-204-5p in SiHa and CaSki cells, we detected the distinctly reduced mRNA and protein levels of TMOD3 and TFAP2A (Fig. 2C and D). In addition, the levels of these four mRNAs were also examined in SiHa and CaSki cells after inhibiting miR-204-5p expression. As a result, merely TFAP2A mRNA and protein levels were obviously augmented (Fig. 2E and F). Herein, TFAP2A was chosen for further analysis. Interestingly, TFAP2A was predicted to be highly expressed in TCGA CESC tissues (Fig. 2H). Consistently, high mRNA and protein levels of TFAP2A were observed in cervical cancer cells (Fig. 2I). Subsequently, RNA pull-down assay demonstrated that TFAP2A was only pulled down by biotinylated miR-204-5p sense (Fig. 2J). To further analyze the interaction between miR-204-5p and TFAP2A, the wild-type and mutant sequences of TFAP2A 3’-UTR for interacting with miR-204-5p and the sequences of miR-204-5p for TFAP2A 3’-UTR were predicted and presented in Fig. 2K. Through luciferase reporter assay, we observed that the luciferase activity of TFAP2A (WT) reporter was weakened by miR-204-5p upregulation whereas strengthened by miR-204-5p suppression (Fig. 2L). Altogether, we confirmed that TFAP2A was targeted by miR-204-5p.
MiR-204-5p could bind to TFAP2A in cervical cancer. A and B. 23 shared target genes of miR-204-5p predicted by online datasets were presented as Venn diagram and analyzed by qRT-PCR in cervical cancer cells versus normal H8 cells. C and D. The levels of 4 possible mRNAs and corresponding proteins in SiHa and CaSki cells were evaluated using qRT-PCR and western blot after transfection. E. The inhibition of miR-204-5p expression was confirmed by qRT-PCR, compared with miR-NC group. F and G. qRT-PCR and western blot analyses of the 4 genes in SiHa and CaSki cells responding to miR-204-5p depletion. H. TFAP2A expression in CESC samples and normal samples was predicted by GEPIA database. I. qRT-PCR and western blot for TFAP2A expression in cervical cancer cells and H8 cells. J. RNA pull-down confirmed the binding capacity between TFAP2A and miR-204-5p. K. The predicted wide-type or mutant binding sequences of TFAP2A to miR-204-5p seed region. L. The interaction of TFAP2A with miR-204-5p was testified via luciferase reporter assay. 
Interestingly, TFAP2A is predicted to be a potential transcriptional regulator for miR-204-5p in accordance with the data of UCSC. Therefore, we speculated that TFAP2A and miR-204-5p might form a circuit to modulate each other. The DNA motif of TFAP2A was exhibited in Fig. 3A. ChIP experiment determined that miR-204-5p promoter was enriched in anti-TFAP2A group (Fig. 3B). After TFAP2A expression was downregulated or upregulated, the luciferase activity of miR-204-5p promoter was accordingly enhanced or impaired (Fig. 3C and D). These changes were also observed in relation to the miR-204-5p expression (Fig. 3E). Then, we continued to affirm the impact of TFAP2A and miR-204-5p on each other. MiR-204-5p expression was elevated by miR-204-5p mimics but lowered by TFAP2A upregulation (Fig. 3F). Likewise, TFAP2A levels increased by TFAP2A upregulation were decreased again by miR-204-5p mimics (Fig. 3G and H). In addition, miR-204-5p expression was enhanced by shTFAP2A but regained by miR-204-5p inhibitor (Fig. 3I). These findings illustrated that TFAP2A and miR-204-5p could form a negative feedback loop to regulate each other.
TFAP2A could regulate miR-204-5p expression transcriptionally. A. The DNA motif of transcription factor TFAP2A. B. ChIP assay validated the binding of TFAP2A to miR-204-5p promoter. C. Both levels of TFAP2A mRNA and protein in SiHa and CaSki cells transfected with shTFAP2A or pcDNA3.1/TFAP2A were analyzed via qRT-PCR and western blot, with shNC or pcDNA3.1 as negative control. D. The effect of TFAP2A on miR-204-5p transcription was confirmed by luciferase reporter assay. E. The effect of TFAP2A depletion or TFAP2A overexpression on miR-204-5p expression was detected by qRT-PCR. F. MiR-204-5p level was detected in cells was examined via qRT-PCR responding to TFAP2A overexpression or knockdown. G and H. TFAP2A expression in different groups was examined via qRT-PCR and western blot. I. The expression of miR-204-5p in two cervical cancer cells was detected via qRT-PCR after indicated transfections.
MiR-204-5p/TFAP2A feedback loop regulated cell proliferation, migration, invasion and EMT process in cervical cancer. A–C. The proliferative ability of SiHa cells transfected with different plasmids was assessed via CCK-8, EdU staining and colony formation. D and E. The capabilities of transfected cells to migrate and invade were measured via wound healing and transwell assays. F. Western blot was done to examine the levels of EMT-related proteins in transfected cells. 
Functional rescue assays were conducted to further confirm the role of miR-204-5p/TFAP2A feedback loop in cervical cancer. It was displayed by CCK-8 assay that overexpressed miR-204-5p inhibited cell viability, and this effect was then reversed by increased level of TFAP2A (Fig. 4A). Likewise, EdU and colony forming assays manifested that the proliferation of SiHa cells was restrained by miR-204-5p overexpression, and this phenomenon was counteracted by TFAP2A upregulation (Fig. 4B and C). Based on the results of wound healing assay, overexpression of miR-204-5p repressed cell migratory capacity, which was then neutralized by overexpressing TFAP2A (Fig. 4D). In transwell assay, we observed that TFAP2A overexpression rescued the suppressive influence of miR-204-5p upregulation on cell invasion (Fig. 4E). Moreover, western blot elucidated that E-cadherin expression was augmented by upregulation of miR-204-5p, which was then reversed by TFAP2A overexpression. Conversely, N-cadherin and Vimentin protein levels were lowered by elevating miR-204-5p expression, which was then increased by overexpressing TFAP2A (Fig. 4F). In summary, miR-204-5p/TFAP2A feedback loop contributed to the proliferative and motorial abilities of cervical cancer cells.
Discussion
It is full of challenges to seek for effective therapies for cervical cancer. Therefore, the research for promising biomarkers potentiating the pathology of cervical cancer is significant. In the past decades, the roles of microRNAs (miRNAs) in cervical cancer have been widely explored. For instance, miR-506 suppresses cervical cancer by directly targeting Gli3, a transcription factor of the hedgehog pathway [27]. Up-regulation of B4GALT3 targeted by miR-27a contributes to cervical cancer cells [28]. Repressed miR-126 and upregulated adrenomedullin by cancer-stromal cross talks in the stromal endothelium provoke the angiogenesis of cervical cancer [29]. Suppression of miR-424 and upregulation of Chk1 contribute to cervical cancer progression [30]. As previously reported, miR-204-5p was under-expressed in minimal deviation adenocarcinoma of uterine cervix [15]. What’s more, miR-204-5p was also supposed to be expressed at low levels in cervical cancer. In our current study, we determined that the miR-204-5p exerted anti-oncogenic role in cervical cancer by suppressing cell growth and migration.
Mechanistically, miRNAs usually regulate their downstream mRNA by post-transcriptional regulation [34]. Here, we explored and identified that TFAP2A was the direct target of miR-204-5p. Importantly, past studies have reported that TFAP2A is a tumor promoter by serving as a transcriptional regulator [24, 25]. It is known that miRNAs can be activated or suppressed by their upstream modulators [35, 36]. For example, Cip2a/miR-301a feedback loop facilitates triple-negative breast cancer cell proliferation and invasion [37]. MiR-29a-5p/STAT3 feedback loop mediates the progression of colitis-associated colorectal cancer [38]. TNF-
In summary, this paper disclosed a feedback loop of miR-204-5p with TFAP2A for the modulation of cellular processes in cervical cancer. MiR-204-5p could repress TFAP2A expression at post-transcriptional level, and TFAP2A could in turn inactivate miR-204-5p expression at transcriptional level. These findings suggested that targeting miR-204-5p/TFAP2A might be a prospective therapy for the treatment of cervical cancer patients.
Footnotes
Acknowledgments
All supports in this research were undeniable.
Conflict of interest
None.
Supplementary data
The supplementary files are available to download from http://dx.doi.org/10.3233/CBM-191064.
