Abstract
This article has been retracted, and the online PDF replaced with this retraction notice.
Keywords
Introduction
Lung cancer is one type of malignant cancer with increasing morbidity and mortality, and accounts for approximately a quarter of cancer-caused death worldwide [1, 2]. Besides, the occurrence rate of lung cancer ranks the first in males and second in females [3]. Non-small cell lung cancer (NSCLC) is commonly diagnosed in lung cancer patients, and lung squamous cell carcinoma (LUSC) is a subtype of NSCLC [4]. Many factors, such as smoking and air pollution, were considered to be closely associated with the carcinogenesis of LUSC [5]. Despite the improvement in therapeutic level, LUSC patients still present a terrible prognosis with dismal overall survival rate in 5-years. Thus, further exploration in LUSC treatment is urgently required.
Long non-coding RNA (lncRNA) belongs to a member of non-coding RNAs, which possesses no less than 200 nucleotides in length and is unable to encode proteins [6, 7, 8]. Emerging reports confirmed that the dysregulation of lncRNA was related to the initiation and development of numerous cancers. As examples, lncRNA SNHG10 drives hepatocarcinogenesis and metastasis via a positive feedback loop [9]. Additionally, LINC00707 is identified as a tumor facilitator and enhances cell proliferation and metastasis in gastric cancer [10]. In mechanism aspect, lncRNA has been widely reported to function as competitive endogenous RNA (ceRNA) to regulate messenger RNA (target gene) expression by sponging microRNA (miRNA) [11, 12], and thereby to regulate the progression of tumors [13]. For instance, MNX1-AS1 boosted esophageal squamous cell carcinoma progression through miR-34a/SIRT1 axis [14]. NR2F1-AS1 boosts proliferation and migration and inhibits apoptosis of thyroid cancer cells by sponging miR-338-3p and regulating CCND1 [15]. MFI2-AS1 improves the proliferation, migration and invasion capacities of colorectal cancer cells via miR-574-5p/MYCBP axis [16]. Recently, long intergenic non-protein coding RNA 01116 (LINC001116) was reported to drive oral squamous cell carcinoma progression, and boost cell proliferation in prostate cancer [17, 18]. However, the biological role and mechanism of LINC01116 in LUSC was still unclear.
In the current research, we uncovered the high expression of LINC01116 in LUSC tissues and cells. Furthermore, LINC01116 exacerbated LUSC progression by targeting miR-744-5p/SCN1B axis, implying an underlying therapeutic marker for LUSC patients.
Materials and methods
Tissue samples
The LUSC samples (
Cell lines
Human normal bronchial epithelial cells (HBE) and LUSC cell lines (SK-MES-1, A549, H2170 and H1299) were bought from the Chinese Academy of Science Cell Bank (Shanghai, China). All the cells were cultivated in Dulbecco’s Modified Eagle Medium (DMEM, Gibco, USA) with 10% fetal bovine serum (FBS) in a damp atmosphere of 5% CO
Cell transfection
The short hairpin RNAs (shRNAs) targeting LINC01116 (sh-LINC01116#1/2/3) with negative control (sh-NC) was employed to knockdown LINC01116 expression. MiR-744-5p mimics with negative control (NC mimics) was used to overexpress miR-744-5p. The pcDNA3.1 vector cloned with the full length of SCN1B was utilized to construct pcDNA3.1/SCN1B for overexpression of SCN1B expression, with empty pcDNA3.1 as control. All vectors synthesized by GenePharma (Shanghai, China) were separately transfected into LUSC cells through Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA) obeying the manufacturer’s instruction.
Real-time reverse-transcription polymerase chain reaction (RT-qPCR)
According to manufacturer’s instruction, TRIzol reagent (Invitrogen, Carlsbad, CA, USA) was adopted to extract all RNAs from cells. A Reverse Transcription Kit (Takara, Dalian, China) reversely transcribed lncRNA and mRNA into cDNA. As for miRNA, a microRNA First-Strand cDNA Synthesis Kit (Sangon Biotech, Shanghai, China) was applied. LINC01116 and SCN1B were amplified via SYBR-Green PCR Master Mix kit (Applied Biosystems, Foster City, CA, USA), and miR-744-5p was amplified via TaqMan miRNA assay kit (Applied Biosystems). Foregoing procedures were implemented on a 7900HT Fast Real-Time System (Applied Biosystems). The relative expression levels of genes were dissected by the 2
CCK-8 assay
Cell Counting Kit-8 (CCK-8) was employed to determine cell viability. At first, 1000 cells were seeded into per well of 96-well plates. Cell viability was evaluated using the Cell Counting Kit-8 (Dojindo Laboratories) at 24, 48, 72 and 96 hours after incubation. Cell viability curves were plotted according to the absorbance values at 450 nm at each time point.
5-ethynyl-2’-deoxyuridine (EdU) assay
EdU incorporation experiment was carried out with the Cell-Light
Transwell assay
Transwell chambers (BD Biosciences, San Jose, CA, USA) with or without matrigel (BD Biosciences) were used to estimate the migration and invasion of LUSC cells. The bottom compartment of the chamber was added with DMEM (Gibco) containing 10% FBS, whereas, the upper compartment was seeded with LUSC cells and DMEM (Gibco) with free FBS. After incubation of LUSC cells for 24 h at 37
Western blot
BCA Protein Assay (Thermo Fisher Scientific, Waltham, MA, USA) was utilized to test the protein density of extracted proteins. Dissolved proteins by 10% sodium dodecyl sulfate-polyacrylamide gel electrophoresis were transferred to nitrocellulose membranes (Millipore, Billerica, MA, USA). Post electrophoresis, proteins were moved onto polyvinylidene difluoride (PVDF) membrane. The membranes blocked in 5% nonfat milk were in mixture with primary antibodies (1:1000) for one night at 4
Nuclear-cytoplasmic fractionation
The location of LINC01116 was segregated with the application of a PARIS kit (Life Technologies, MA, USA). Briefly, collected LUSC cells were lysed on ice. After centrifugation, the supernatant was harvested. GAPDH and U2 were respectively utilized as the cytoplasm and nucleus controls. The extracted RNAs were detected by RT-qPCR.
Bioinformatics analysis
GEPIA (
Luciferase reporter assay
LINC01116-WT (or SCN1B 3’UTR-WT) and LINC01116-Mut (or SCN1B 3’UTR-Mut) were constructed into pmirGLO plasmids (Promega, Madison, WI, USA). MiR-744-5p mimics or NC mimics were co-transfected with the constructed plasmids into A549 and H1299 cells for 48 hours. Luciferase reporter assay system (Promega, Madison, WI, USA) was adopted to examine relative luciferase activities.
RNA Pull down assay
MiR-744-5p-Wt (sense) and miR-744-5p-Mut (antisense) were transcribed employing Transcript Aid T7 High Yield Transcription Kit (ThermoFisher Scientific, USA). Biotinylated miR-744-5p-Wt, miR-744-5p-Mut and negative control (Bio-NC) were co-cultivated with cell lysate (A549 and H1299) at 4
Statistical analyses
SPSS 20 software (Abbott Laboratories, Chicago, IL, USA) was applied for statistical analyses. And data were dissected through unpaired Student’s
LINC01116 was up-regulated and boosted LUSC progression. (A) Data from GEPIA database showed the expression of LINC01116 in 486 LUSC tumor tissues (red box) and 338 normal adjacent tissues (black box). (B–C) RT-qPCR was performed to detect LINC01116 expression in LUSC tissues (or cells) and normal tissues (or cells). (D) The knockdown efficacy of sh-LINC01116 (sh-LINC01116#1/2/3) was examined by RT-qPCR. (E–F) The proliferation of sh-LINC01116 transfected cells was assessed by CCK-8 and EdU assays. (G–H) Transwell assay was utilized for the evaluation of migration and invasion in cells upon LINC01116 knockdown. (I) Western blot assay measured the protein levels of E-cadherin, N-cadherin and Vimentin. 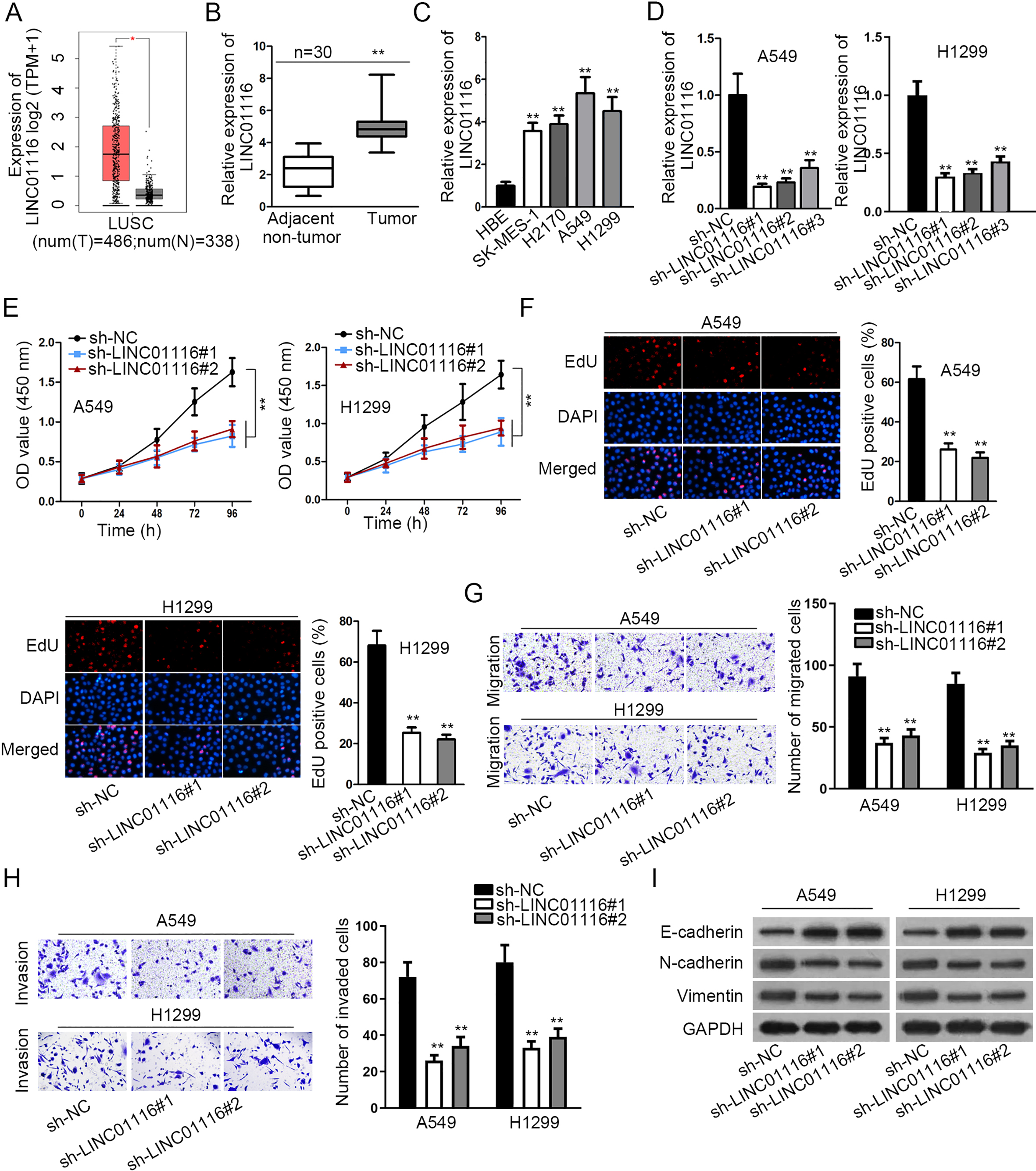
LINC01116 directly sponged miR-744-5p in LUSC. (A) Nuclear-cytoplasmic fractionation assay was employed to determine LINC01116 localization. (B) The miRNAs with binding site on LINC01116 were predicted by starBase. (C) The expressions of these miRNAs in LUSC cells and HBE cells were tested by RT-qPCR. (D) MiR-744-5p expression in LUSC tissues and adjacent non-tumor tissues was tested. (E) The transfection efficiency of miR-744-5p mimics was confirmed. (F) The expression of LINC01116 or miR-744-5p was detected after transfecting the indicated plasmids. (G) The expression relevance between LINC01116 and miR-744-5p was verified by Pearson correlation analysis. (H) RNA pull down assay was used for verifying the interaction between LINC01116 and miR-744-5p (I) Binding sequence between LINC01116 and miR-744-5p were hypothesized. (J) Luciferase reporter experiments confirmed the luciferase activity of LINC01116-WT/Mut with the transfection of miR-744-5p mimics or NC mimics. 
SCN1B served as a target of miR-744-5p in LUSC. (A) Venn diagram showed the possible target genes. (B) The expression of target genes in LUSC tissues was monitored by RT-qPCR. (C) SCN1B expression in LUSC cell lines and HBE cell line. (D–E) Effect of miR-744-5p mimics on the expressions of SCN1B mRNA and protein was analyzed by RT-qPCR and western blot assays. (F) Pearson correlation analysis was applied to analyze expression correlation between SCN1B and miR-744-5p. (G) The enrichment of SCN1B in biotinylated miR-744-5p-Wt/Mut pellet was validated by RNA pull down assay. (H) The binding sequence of miR-744-5p on SCN1B 3’UTR were depicted. (I) Luciferase reporter assay verified the binding affinity between SCN1B and miR-744-5p. 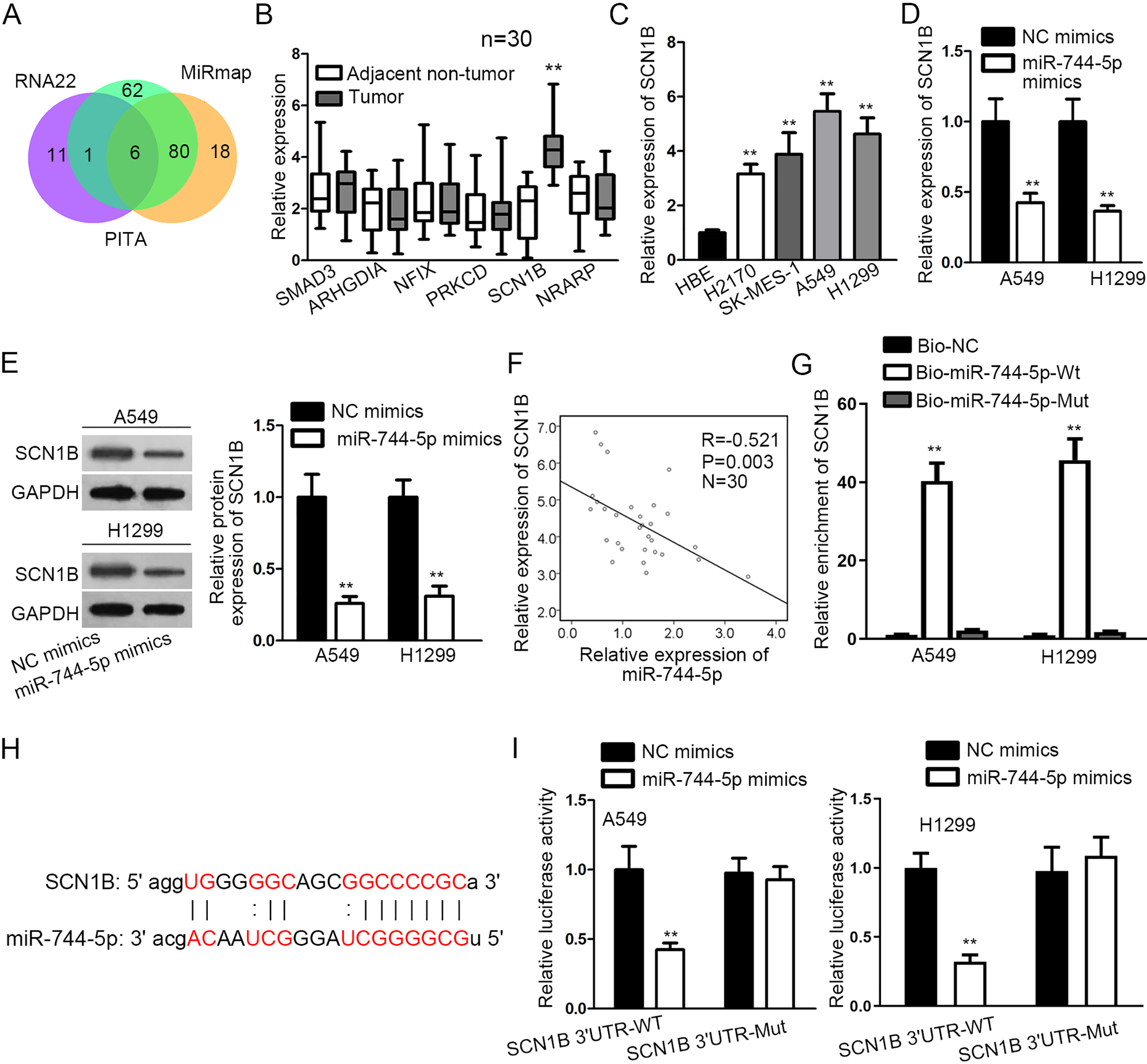
LINC01116 exacerbated LUSC progression by increasing SCN1B expression. (A–B) The detection of SCN1B mRNA and protein levels was implemented by western blot assay after transfecting the indicated plasmids. (C–D) Cell proliferation in each group was evaluated through CCK-8 and EdU assays. (E–F) The migratory and invasive abilities of LUSC cells were assessed by transwell assay with the transfection of indicated plasmids. (G) EMT-relevant protein expressions in each group were investigated by western blot. 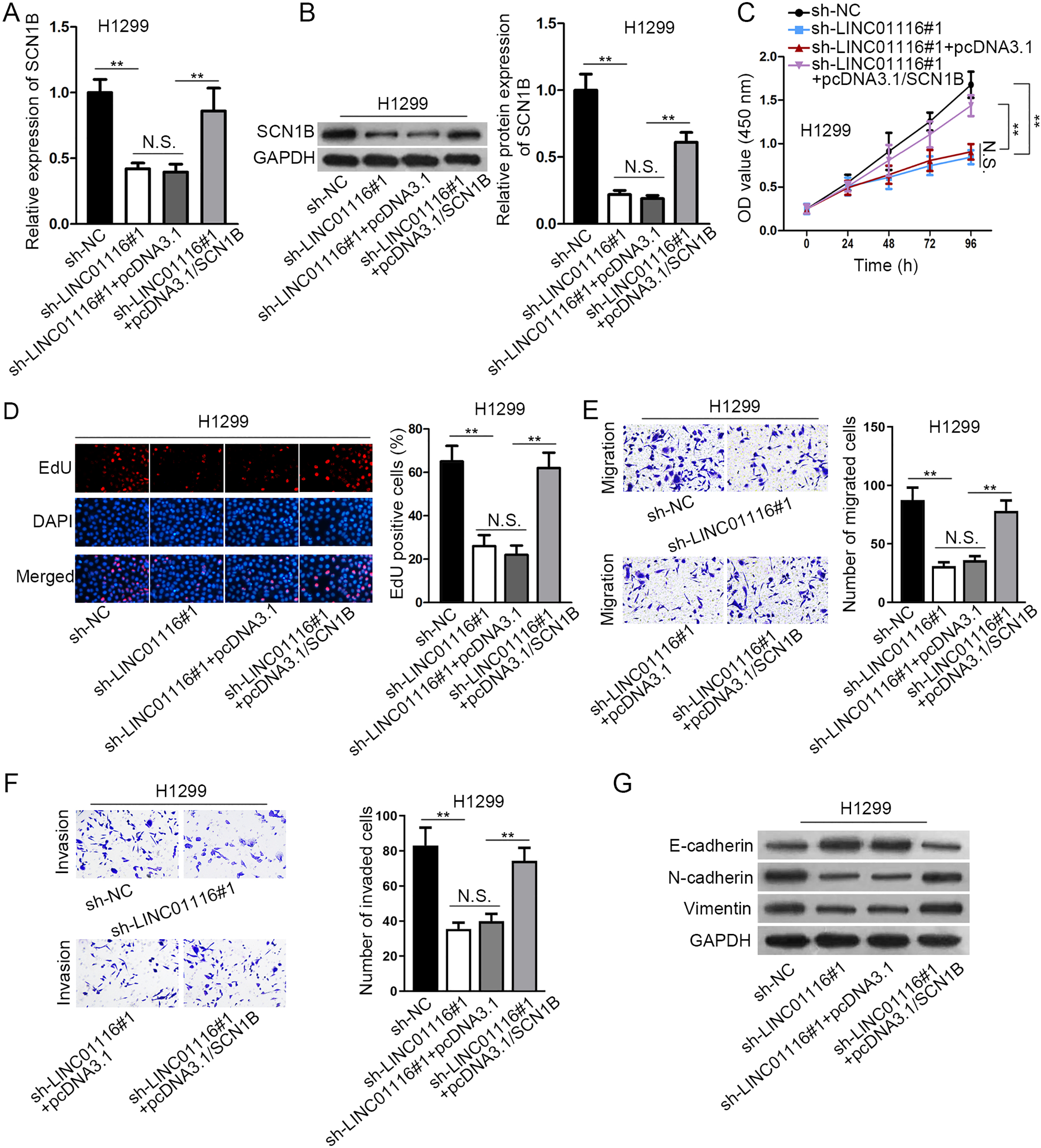
LINC01116 was up-regulated and boosted LUSC progression
Several studies have identified that LINC01116 functioned as an oncogene in cancers, but no further exploration of LINC01116 in LUSC has been reported. According to GEPIA database, we found that the expression of LINC01116 was up-regulated in LUSC (Fig. 1A). To confirm the expression pattern of LINC01116 in LUSC, we detected LINC01116 expression in LUSC tissues and cell lines (SK-MES-1, A549, H2170 and H1299) by RT-qPCR. The adjacent non-tumor tissues and human normal bronchial epithelial cell line (HBE) were separately seen as a control. Results revealed that LINC01116 expression in LUSC tissues or cells was markedly higher than that in matched normal tissues or HBE cell line (Fig. 1B and C). Then, LINC01116 expression was markedly reduced by specific LINC01116 shRNAs (sh-LINC01116#1/2/3) (Fig. 1D). With better knockdown efficacy, sh-LINC01116#1 and sh-LINC01116#2 were adopted in loss-of-function assay. At the beginning, CCK-8 assay uncovered that cell viability was obviously hindered upon LINC01116 knockdown (Fig. 1E). Besides, LINC01116 suppression also caused an evident decline of proliferative ability through EdU assay (Fig. 1F). Subsequently, transwell assay demonstrated that both migration and invasion of A549 and H1299 cells were inhibited by transfecting sh-LINC01116 (Fig. 1G and H). Moreover, western blot assay suggested that the protein expression of E-cadherin was increased whereas that of N-cadherin and Vimentin plummeted by LINC01116 silencing (Fig. 1I). To sum up, LINC01116 was up-regulated and boosted LUSC progression.
LINC01116 directly sponged miR-744-5p in LUSC
In mechanism, a multitude of researches have claimed that lncRNA could act as a ceRNA in cytoplasm to regulate gene expression [19, 20]. To explore the mechanism of LINC01116 in LUSC, we firstly determined its localization in A549 and H1299 cells. As expected, LINC01116 was mainly distributed in the cytoplasm of LUSC cells via nuclear-cytoplasmic fractionation assay (Fig. 2A). Through starBase, five miRNAs possessing putative binding site on LINC01116 were predicted (Fig. 2B). Then, the expressions of five potential miRNAs were detected in LUSC cell lines. The result of RT-qPCR disclosed that miR-744-5p was down-regulated in LUSC cells whilst the rest displayed no expression difference (Fig. 2C). Likewise, miR-744-5p presented lower expression in LUSC tissues compared with adjacent normal tissues (Fig. 2D). To evaluate the interaction between LINC01116 and miR-744-5p, a series of experiments were designed and carried out. Before the experiments, miR-744-5p expression was dramatically enhanced by transfecting miR-744-5p mimics (Fig. 2E). As we observed, miR-744-5p amplification decreased LINC01116 expression and LINC01116 inhibition increased miR-744-5p expression (Fig. 2F). Furthermore, the negative expression correlation was found between LINC01116 and miR-744-5p (Fig. 2G). In addition, LINC01116 was pulled down by biotinylated miR-744-5p-Wt through RNA pull down assay (Fig. 2H). As illustrated in Fig. 2I, the binding sequences between LINC01116 and miR-744-5p were hypothesized. Luciferase reporter assay manifested that the luciferase activity of LINC01116-WT vector was conspicuously weakened with the transfection of miR-744-5p mimics, whereas no evident changes were observed in that of LINC01116-Mut vector (Fig. 2J). Taken together, LINC01116 directly sponged miR-744-5p in LUSC.
SCN1B served as a target of miR-744-5p in LUSC
Next, we sought the targets of miR-744-5p in LUSC and six potential target genes were predicted and shown as a Venn diagram (Fig. 3A). For selecting the optimal target of miR-744-5p, RT-qPCR was utilized to detect expression levels of the targets, and we discovered that only SCN1B were up-regulated LUSC tissues (Fig. 3B). Meanwhile, the high expression of SCN1B was also detected in LUSC cells (Fig. 3C). Furthermore, SCN1B mRNA and protein expressions were both decreased in cells transfected with miR-744-5p mimics (Fig. 3D and E). As exhibited in Fig. 3F, SCN1B expression was negatively correlated with miR-744-5p expression. RNA pull down confirmed the considerable enrichment of SCN1B in biotinylated miR-744-5p-Wt pellet (Fig. 3G). Additionally, the 3’UTR of SCN1B was postulated to contain a binding sequence of miR-744-5p (Fig. 3H). Luciferase reporter assay uncovered that miR-744-5p augmentation resulted in an alleviated luciferase activity of SCN1B 3’UTR-WT but such alteration was not observed in SCN1B 3’UTR-Mut (Fig. 3I). Collectively, SCN1B served as a target of miR-744-5p in LUSC.
LINC01116 exacerbated LUSC progression by increasing SCN1B expression
To confirm the role of LINC01116/miR-744-5p/ SCN1B axis in LUSC progression, some rescue assays were performed. Firstly, we found that the repressive effect of LINC01116 depletion on SCN1B mRNA and protein expressions was recovered by SCN1B overexpression (Fig. 4A and B). Besides, the transfection of pcDNA3.1/SCN1B restored the suppressive role of LINC01116 knockdown in cell proliferation via CCK-8 and EdU assays (Fig. 4C and D). Furthermore, transwell assays elucidated that the restrained migration and invasion resulted from LINC01116 deficiency was counteracted by SCN1B amplification (Fig. 4E and F). Afterwards, SCN1B up-regulation abolished sh-LINC01116-mediated promotion in E-cadherin protein level and inhibition in N-cadherin and Vimentin protein levels (Fig. 4G). In conclusion, LINC01116 exacerbated LUSC progression by increasing SCN1B expression.
Discussion
With the highest death rate, lung cancer has taken away millions of lives [21]. As a kind of lung cancer, LUSC was hard to be cured because of high recurrence and distant metastasis [22]. Increasing evidence has proved that aberrant lncRNAs were capable to affect diverse cellular processes in cancers [23, 24]. The up-regulation of SNHG8 improved cell migration in hepatocellular carcinoma [25]. MALAT1 overexpression contributed to colorectal cancer cell growth and metastasis [26]. Although several lncRNAs have been identified in LUSC [27, 28, 29], the underlying carcinogenesis of LUSC remains unknown. LINC001116 is involved in the modulation of cell proliferation and migration in glioma, osteosarcoma and breast cancer [30, 31, 32]. In our research, LINC01116 was up-regulated in lung squamous cell carcinoma tissues and cells, and LINC01116 repression restrained LUSC cell proliferation, migration, invasion, and EMT process. These data demonstrated the oncogenic property of LINC01116 in LUSC.
MiRNAs are also reported as an important regulator in cancer progression. For instance, miR-4516 functions as a novel oncogene and predicts poor prognosis via targeting PTPN14 in human glioblastoma [33]. MiR-142-3p acts as a tumor suppressor in colorectal cancer and impairs cell growth by targeting CDK4 [34]. Mechanistically, ceRNA pattern has attracted considerable attentions globally [35]. In this pattern, lncRNA could up-regulate target gene expression by interacting with miRNA [36]. Based on this theory, we detected LINC01116 distribution in LUSC cells and found that LINC01116 was a cytoplasmic RNA. Afterwards, miR-744-5p was verified to directly interact with LINC01116. MiR-744-5p was reported to regulate HNRNPC and NFIX expressions and induce cell apoptosis in ovarian cancer [37]. MiR-744-5p was also sponged by lncRNA MAFG-AS1 in lung adenocarcinoma [38]. Our study confirmed the low level of miR-744-5p in LUSC cells and tissues. Moreover, LINC01116 expression was negatively associated with miR-744-5p expression. Collectively, LINC01116 served as a sponge of miR-744-5p in LUSC.
According to the result of bioinformatics analysis and experiments, sodium voltage-gated channel beta subunit 1 (SCN1B) was predicted to serve as a target of miR-744-5p. In our exploration, SCN1B expression was up-regulated and negatively associated with miR-744 expression. Importantly, the combination between miR-744-5p and SCN1B was testified. Last but not least, rescue assays using H1299 cells validated that SCN1B overexpression restored the prohibitive effect of LINC01116 silencing on the proliferation, migration, invasion, and EMT process.
In conclusion, our study was the first to confirm that LINC01116 was overexpressed in LUSC and exacerbate LUSC progression by regulating miR-744-5p/SCN1B axis, which could provide a meaningful revelation for exploring the potential therapeutic methods for LUSC patients.
Footnotes
Acknowledgments
We appreciate the support of lab members.
Conflict of interest
The authors declare no competing interests in this work.
