Abstract
BACKGROUND:
MicroRNA (miRNA) expression has been implicated in leukaemia. In recent years, miRNAs have been under investigation for their potential as non-invasive biomarkers in acute promyelocytic leukaemia (APL). We investigated whether miR-638 in circulating leukaemia cells is a non-invasive biomarker in diagnosis, assessment of the treatment response and minimal residual disease (MRD) surveillance of APL.
METHODS:
Sixty cases of acute myeloid leukaemia (AML), including 30 cases of APL and 30 cases of non-APL AML, were selected. Thirty healthy controls were also selected. Bone marrow (BM) and peripheral blood (PB) samples were collected from APL patients at diagnosis and post-induction. Microarray analysis and quantitative real-time PCR (qRT-PCR) were performed for miRNA profiling and miR-638 expression analysis, respectively. For statistical analysis, Mann-Whitney U test, Wilcoxon Signed Rank test, receiver operating characteristic (ROC) curve analysis and Spearman’s rho correlation test were used.
RESULTS:
Both microarray and qRT-PCR data showed that miR-638 was significantly upregulated in BM after APL patients received induction therapy. Moreover, miR-638, which is specifically downregulated in APL cell lines, was upregulated after all-trans retinoic acid (ATRA)-induced myeloid differentiation. Receiver operating characteristic (ROC) curve analyses revealed that miR-638 could serve as a valuable biomarker for differentiating APL from controls or non-APL AML. Furthermore, miR-638 expression was sharply increased after induction therapy and complete remission (CR). An inverse correlation was observed between miR-638 and PML-RAR
CONCLUSIONS:
Our study suggested that miR-638 may serve as a potential APL biomarker for diagnosis and assessment of the response to targeted therapy, and PB miR-638 could be used for non-invasive MRD surveillance in APL.
Keywords
Introduction
Acute promyelocytic leukaemia (APL) is characterized by specific chromosomal translocations (15; 17) (q22; q12), leading to the fusion of the promyelocytic (PML) gene with the retinoic acid receptor
Studies have shown that the circulating tumor nucleic acid, circulating tumor cells and exosomes can be used in liquid biopsy and are really to become part of future clinical practice [9, 10]. MRD is a valuable marker for evaluating the treatment response, and MRD surveillance is recommended as part of the standard of care for acute myeloid leukaemia (AML) patients [11]. MicroRNAs (miRNAs) are small non-coding RNAs (
The identification of dysregulated miRNAs may provide new targets for the diagnosis and treatment of APL [18, 21]. miR-638, which is downregulated in various solid tumors [22, 23], might act as a sensitizer in cancer chemotherapy and accompany chemotherapy drugs to enhance chemotherapeutic efficacy and to improve the chance of recovery from cancer [24]. Furthermore, miR-638 has been proposed to play roles in haematopoiesis or leukemogenesis [25, 26]. Recently, Lin et al. reported that miR-638 was reduced in primary AML samples vs. cells undergoing normal haematopoiesis [27]. miR-638 may promote ATRA-induced myeloid differentiation in leukaemic cells, the downregulation of which may promote proliferation and contribute to the transformation of leukaemic cells [27]. The miR-638/cyclin-dependent kinase 2 (CDK2) axis may serve as a marker for prognosis or treatment response in AML. Nevertheless, whether miR-638 can be applied for the non-invasive monitoring of APL remains unknown.
Characteristics of 60 patients with AML
Characteristics of 60 patients with AML
AML, acute myeloid leukemia; APL, acute promyelocytic leukemia.
In this study, we first investigated the expression of miR-638 in cell lines and clinical samples. We then explored the role of miR-638 as a potentially new biomarker for diagnosis in APL. Furthermore, we found that miR-638 could be used to predict treatment response and MRD surveillance, especially in non-invasive monitoring of APL. We aim at provide a new target for APL diagnosis and treatment.
Patient profiles
The present study enrolled 60 leukaemia patients who were from The Second Affiliated Hospital of Wenzhou Medical University from January 2015 to October 2017 (Table 1). Patient diagnosis was established according to the 2016 World Health Organization criteria [1], including morphology, immunophenotyping, cytogenetics and molecular examination. BM specimens were collected from 60 AML patients (including 30 APL patients) at the time of diagnosis, 30 matched-APL patients post-induction and at complete remission (CR), 4 APL patients at relapse time, and 30 normal controls. Moreover, 30 peripheral blood (PB) samples were also taken from APL patients at the time of diagnosis and matched-APL patients after induction. The normal controls were patients with matched age and sex not suffering from any haematologic or other types of malignancy. No chemotherapy was administered prior to participation in the study. All patients with APL received substantial therapy according to the latest National Comprehensive Cancer Network (NCCN) guidelines [28]. Haematologic CR was defined according to standard criteria [29]. All samples were collected in accordance with clinical standard procedures. Approximately 2-ml EDTA-K2 anticoagulated BM samples from BM aspirates or 2-ml EDTA-K2 anticoagulated PB samples were isolated using Density Reagemt (QuantoBio, China) according to the manufacturer’s instructions. Subsequently, the mononuclear BM or PB cells were resuspended in 1 ml TRIzol (Invitrogen, USA), and temporarily stored at
Cell line, cell culture and induced myeloid differentiation
The human leukaemia cell lines NB4, HL-60, U937, THP-1, K562, Kasumi-6 and HEL were purchased from the Shanghai Institution of Hematology, and maintained in RPMI 1640 medium containing 10% foetal bovine serum (GIBCO BRL, USA) and cultured at 37
MiRNA microarray
MiRNA expression profiling was conducted using a GeneChip miRNA 4.0 Array (Affymetrix, USA) that consisted of probes for 2006 human miRNAs based on Sanger miRBase release 19.0. BM samples were taken from three patients with APL before and after induction therapy. Total RNA was extracted from BM samples using TRIzol (Invitrogen, USA). The arrays were washed and scanned according to the manufacturer’s instructions (Shanghai OE Biotech Co., China). The fluorescent intensities were calculated using the Feature extraction software (Agilent Technologies, USA). The results were identified by arbitrarily setting the threshold at a fold change (FC) of 2.0 or above combined with
RNA extraction, reverse transcription and qRT-PCR
Total RNA was isolated from BM samples or cell samples (including NB4 and HL-60 cells) with TRIzol according to the manufacturer’s instructions and eluted with 50
MRD analysis
MRD analysis was conducted on 30 BM samples obtained from patients with APL before and after induction therapy. MRD assessment was performed using qRT-PCR to detect PML-RAR
Statistical analysis
The microarray data were uploaded to the R software package and analysed using Bioconductor’s Oligo package. The non-parametric Mann-Whitney U test and Wilcoxon Signed Rank test were performed to evaluate the differences in miR-638 expression between leukaemia and control samples and between BM samples at diagnosis and follow-up, respectively. The discriminatory ability of miR-638 for APL was examined via receiver operating characteristic (ROC) curve analysis. Correlation analyses were performed using Spearman’s rho correlation test.
Results
General expression profiles of dysregulated miRNAs in APL with targeted therapy
To investigate and verify the potential involvement of miR-638 in APL therapy, we first scanned the global miRNA expression profiles of BM mononuclear cells (BMMCs) from 3 patients with APL before and after induction therapy. The microarray data were filtered through a volcano plot to determine the dysregulated miRNAs in patients with APL post-induction vs. matched controls at diagnosis (Fig. 1a). The heat map showed the results of the unsupervised hierarchical clustering based on the significantly differentially expressed miRNAs (Fig. 1b). After filtering, we found that 5 miRNAs were downregulated and 25 miRNAs were upregulated in patients with APL post-induction compared with the matched controls at diagnosis (Fig. 1b). Moreover, 83% (25/30) of the dysregulated miRNAs were upregulated, and miR-638 was the most significantly upregulated (Fig. 1c). Subsequent qRT-PCR experiment (including miR-638) in the enrolled patients also verified the microarray results (Fig. 1d). This finding raised the possibility that miR-638 might be associated with APL therapy.
General expression profiles of dysregulated miRNAs in BM samples from patients with APL post-induction and corresponding BM samples from patients with APL at diagnosis. (a) Volcano plot of the dysregulated miRNAs for patients with APL post-induction vs. matched controls at diagnosis. The red dots on the right represent upregulated miRNAs and the red dots on the left represent downregulated miRNAs. (b) Heat map of dysregulated miRNAs. Unsupervised hierarchical clustering analysis of miRNA expression profiles. (c) Component diagram for dysregulated miRNAs. (d) RTqPCR verification of 5 selected miRNAs from an additional 30 samples from patients with APL, including 20 postinduction BM samples and 10 preinduction BM samples.
To assess the specific expression of miR-638 in haematopoietic cell development, we measured the expression of miR-638 in different leukaemia cell lineages. We found that miR-638 was significantly downregulated in promyelocytic leukaemia cell lines, including HL-60 and NB4, compared with the non-promyelocytic leukaemia cell lines U937, THP-1, K562, Kasumi-6, and HEL (Fig. 2a). Furthermore, we used NB4 and HL-60 cells as models of myeloid differentiation. As a result, qRT-PCR confirmed that miR-638 was upregulated in HL-60 (Fig. 2b) and NB4 cells (Fig. 2c) treated with ATRA compared to DMSO-treated cells (control). Furthermore, upregulation of miR-638 coincided with increased myeloid-specific surface marker CD11b in NB4 (Fig. 2d) and HL-60 (data not shown) cells induced by ATRA, as measured by flow cytometry. Taken together, these findings suggested that miR-638 is specifically downregulated in APL cell lines and ATRA-induced myeloid differentiation might promote miR-638 dysregulation.
Differential expression of miR-638 in myeloid lineage, and miR-638 dysregulation may contribute to ATRA-treated APL. (a) Lineage-specific expression of miR-638. miR-638 expression levels in MNCs, NB4, HL-60, U937, THP-1, K562, Kasumi-6, and HEL cells were analysed by qRT-PCR. (b) and (c) HL-60 and NB4 cells were treated with ATRA as indicated. miR-638 expression was measured by qRT-PCR. (d) NB4 cells were treated with ATRA as indicated and CD11b expression was measured by flow cytometry. 
The representative miR-638 and U6 amplification plots by qRT-PCR analysis. (a) Amplification plot of miR-638 from a patient with APL at diagnosis. (b) Amplification plot of U6 from a patient with APL at diagnosis. (c) Amplification plot of miR-638 from a patient with APL post-induction. (d) Amplification plot of U6 from a patient with APL post-induction. The x-axis represents the cycle number, and the y-axis represents the relative change in the fluorescence values. The threshold of amplification plot is set at 0.158009.
Evaluation of miR-638 expression levels in APL patients. (a) Box plots presenting miR-638 levels in APL BM samples and non-APL patients for disease diagnosis as well as that of normal controls. 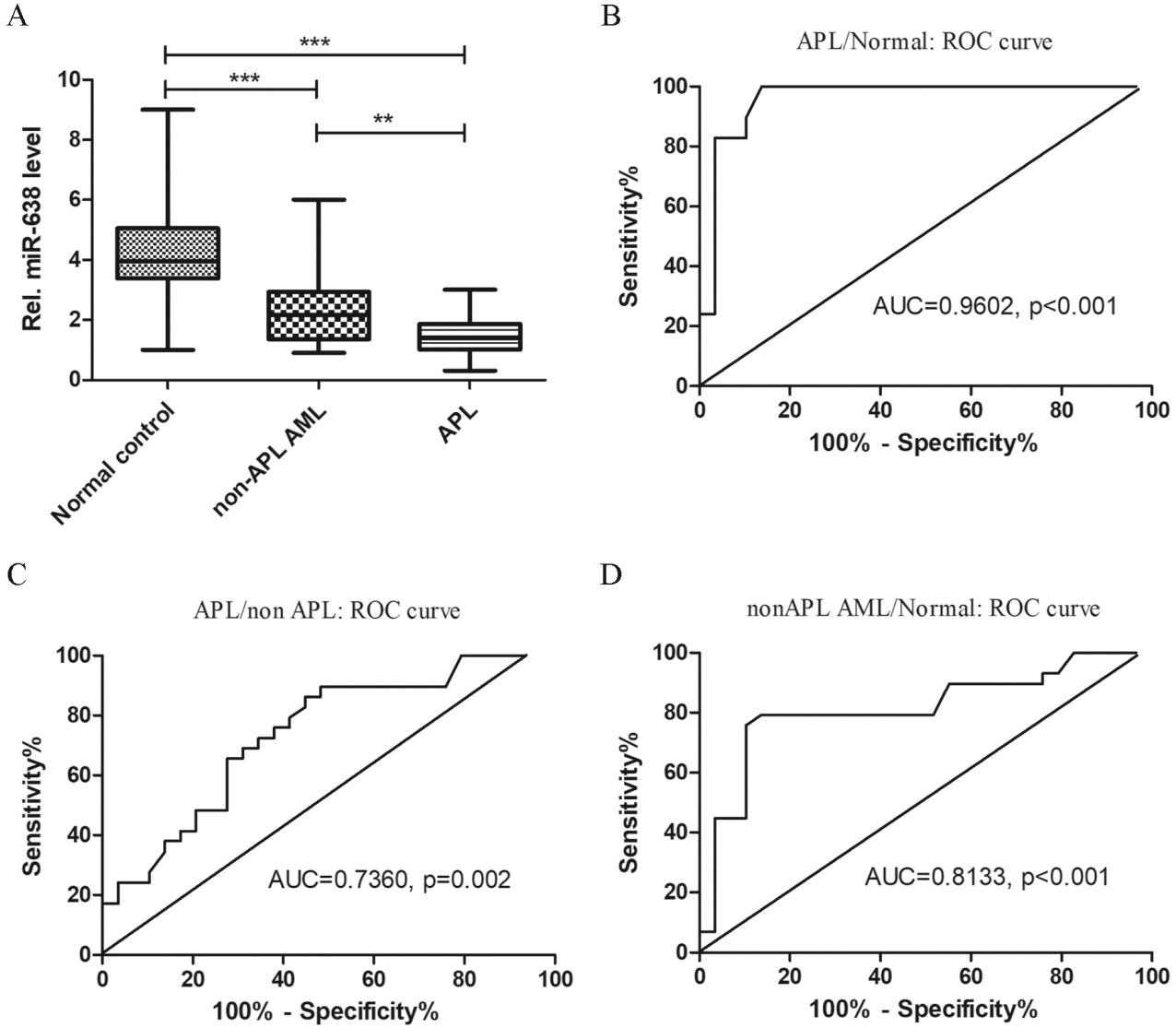
To confirm the accuracy and universality of our findings, we performed qRT-PCR assays on BM samples from 60 primary AML patients, including 30 patients with APL and 30 non-APL AML patients, and 30 normal controls. The representative amplification plots of U6 and miR-638 by qRT-PCR are shown on Fig. 3. The significant downregulation of miR-638 (
miR-638 expression varied in the different APL therapeutic response groups. qRT-PCR was repeated three times, and similar results were obtained. (a) miR-638 expression levels in 30 APL patients at new diagnosis, post-induction and CR by qRT-PCR. (b) miR-638 expression levels in APL patients at new diagnosis, CR and relapse by qRT-PCR.
BM morphological changes in APL patients during targeted therapy. (a) BM morphology in APL patients at diagnosis. (b) BM morphology in APL patients after induction. (c) CR morphology from BM in APL patients. (d) BM morphology after relapse in APL patients. The black arrows indicate the accumulation of abnormal promyelocytes in BM, while the red arrows indicate mature macrophages or segmented neutrophils. BM cell morphology was examined by staining cells with May-Grünwald-Giemsa and the representative images are shown at 
miR-638 as a non-invasive biomarker for MRD surveillance by RT-qPCR analysis. (a) miR-638 levels were inversely correlated with PML-RAR
To determine the clinical significance of miR-638 in APL patients receiving targeted therapy, we collected 30 paired BM samples from APL patients at diagnosis, after induction therapy, and after three years of clinical follow-up. For comparison, we also collected samples from 4 relapsed patients. The results showed that miR-638 expression was low in patients with APL at new diagnosis, but increased sharply after induction therapy and CR (Fig. 5a and Supplemental Table 1). Moreover, there were no differences between post-induction and CR miR-638 expression (
MiR-638 may be used in MRD monitoring
To assess whether miR-638 can be used for MRD monitoring, we analysed the levels of PML-RAR
PB miR-638 as a potential biomarker for non-invasive treatment surveillance
To explore the role of miR-638 in the early detection of residual disease after targeted APL therapy, we collected 30 paired PB and BM samples from patients with APL before and after targeted therapy. The levels of miR-638 in PB and BM were analysed. Spearman’s rho analysis displayed a significantly positive correlation between PB and BM miR-638 expression levels (
Discussion
In this study, we first compared the miRNA expression profiles of BMMCs from three APL patients before and after induction therapy by miRNA array and identified 30 dysregulated miRNAs. Due to methodological differences, the results of microarray-based high-throughput screening must be verified by qRT-PCR. The differential expression of miR-638 was most significant among the upregulated miRNAs using qRT-PCR. This phenomenon led us to consider whether miR-638 in circulating leukaemia cells is a non-invasive biomarker in APL diagnosis, treatment response and MRD surveillance. miR-638 expression in both cell lines and clinical samples was analysed by qRT-PCR. The results showed that miR-638 can be used to effectively diagnose APL and non-APL AML by ROC curve analysis. An inverse correlation between miR-638 and PML-RAR
Circulating miRNA have recently implicated with several cancers and become the biomarker potential of tumour-derived miRNAs. Serum miR-638 with other three miRNAs could efficiently differentiate patients with esophageal squamous dysplasia from the controls [35]. Similarly, the combined-miR-panel including miR-638 could effectively discriminate non-small cell lung carcinoma [36]. Moreover, exosomal miRNAs (including miR-638) could strongly predict prognosis of solid tumor patients [37]. Furthermore, the dysregulation of miR-638 in APL holds a diagnostic relevance. Mi’s group reported that as few as two miRNAs could be used to discriminate between ALL or AML at an accuracy of
miR-638 plays a role in the differentiation of haematopoiesis. A number of groups have reported that miRNA expression is correlated with the differentiation of APL cells treated with ATRA, and this adds relevant prognostic information in APL patients [18, 19, 20, 39]. Similarly, the upregulation of miR-638 was observed in both ATRA-induced APL cell lines and APL patients who received induction therapy. miR-638 acts as a differentiation factor in AML cells through enhancer-binding protein-alpha (CEBPA) [40], which is a master regulator within normal haematopoiesis by direct regulation of specific miRNAs [41]. miR-638 expression was upregulated during cellular terminal differentiation and is involved in mediating DNA damage repair processes [42]. We speculated that miR-638 dysregulation might contribute to APL.
Since several miRNAs are regulated by PML-RAR
PB represents an attractive specimen source for MRD surveillance, allowing for frequent sampling due to the ease of sample collection. Although droplet digital PCR protocols have been established for the quantification of PML-RAR
In conclusion, we identified the differential expression of miR-638 and its clinical significance in patients with APL. Some limitations can be attributed to individual heterogeneity, relatively small sample size, and shorter observation duration. To the best of our knowledge, this is the first to study the role of miR-638 in APL’s clinical evaluation, including diagnosis, assessment of the response to treatment, and MRD surveillance. Moreover, our results suggest that miR-638 may be used as a potentially effective noninvasive biomarker for diagnosis, assessment of the treatment response and MRD surveillance in APL.
Footnotes
Acknowledgments
This work was supported by the Medical and Health Research Science and Technology Plan Project of Zhejiang Province (2017KY112), the Basic Scientific Research Project of Wenzhou City (Y20190090), the Lin He’s New Medicine and Clinical Translation Academician Workstation Research Fund (18331203), and the Basic Public Welfare Technology Research Project of Zhejiang Province (LGF20H200005).
Conflict of interest
All authors declare that they have no conflict of interest.
Abbreviations
Supplemental Table
Fold changes of miR-638 expression in different therapeutic response groups of APL FC, fold change;
Groups
Median FC
Range
Newly diagnosis
1.40
0.30–3.00
–
Post induction
12.01
5.40–18.63
0.001
Complete induction
10.30
5.56–19.17
0.001
