Abstract
BACKGROUND:
Oral tongue squamous cell carcinoma (OTSCC) is a highly aggressive malignancy characterized by frequent recurrence, poor survival with relatively few therapeutic options due to the late diagnosis in many cases.
OBJECTIVES:
Understanding the molecular pathways underlying OTSCC tumourigenesis and the discovery of diagnostic and/or prognostic biomarkers.
METHODS:
We performed high-throughput mutational analysis of 44 OTSCC formalin-fixed paraffin-embedded (FFPE) cases using the Cancer Hotspots Panel (CHP) v2 on the Ion Torrent™platform. We determined the frequency of human papilloma virus (HPV) using PCR and Epstein bar virus (EBV) positivity using immunohistochemistry. As a control for EBV infection we screened matched non-tumourous tissues.
RESULTS:
Sequencing analysis identified missense, nonsense and frameshift mutations in TP53 (66%), PIK3CA (27%), CDKN2A (25%), EGFR (18%), and PTEN (14%). Interestingly, no significant associations were found between damaging mutations and clinicopathological data. A total of 10/44 of the OTSCC samples (23%) tested was positive for HPV18 DNA. OTSCC patients with positive HPV infection had worse overall survival compared to HPV-negative cases as determined by Kaplan-Meier survival (
CONCLUSIONS:
Taken together, this study highlights that the two most common events in OTSCC are TP53 mutations and EBV positivity. Helping to understand the contribution of TP53 mutations and EBV infection events could serve as useful biomarkers for OTSCC.
Introduction
Tumors arising from oral cavity, nasal cavity, pharynx and larynx, which are collectively defined as head and neck cancer, are the sixth leading malignancies around the world [1]. Anatomically, these types of tumors are heterogeneous in nature; they differ in incidences, mortalities, as well as in prognoses. Oral Tongue Squamous Cell Carcinoma (OTSCC) is the most common, devastating and aggressive cancer type of oral subsites with around
Human papilloma virus (HPV) infection is a frequent infectious disease that affects oral health. In normal conditions, HPV remains in an episomal state and when active viral DNA can break and integrate into the host genome then induce deregulation of the cell-cycle control [5]. This is believed to be an important mechanism by which HPV induces carcinomas of head and neck with markedly different mutational pattern associated with infection. The oral cavity and pharyngolarynx are considered the primary site for Epstein-Barr virus (EBV) infection, transmission and persistence, since it is rich in lymphoid tissue. Its transmission through the oral cavity usually targets epithelial cells and immature B lymphocytes [6]. The expression of these EBV proteins can disturb the regulatory process of proteins inside the host cells [7]. Extensive genetic and epigenetic alterations of both the host and viral genome were found in EBV-positive neoplasms. These changes lead to cellular modifications by which the virus can persist and propagate inside the host [8]. Epstein-Barr nuclear antigen 1 (EBNA1) is a small subset of EBV oncogenic proteins that is expressed in all forms of latency in proliferating cells and in all EBV-associated tumors [9]. EBNA1 plays a more direct role in cell survival and oncogenesis by altering key cellular proteins and pathways, such as cellular gene transcription, protein turn-over, signaling pathways and apoptosis [9].
Material and methods
Samples and DNA extraction
This retrospective study includes 44 formalin-fixed Paraffin-embedded blocks containing tissue that met the histologic criteria (WHO) for squamous cell carcinoma of the oral tongue. Tissues were retrieved from the department of pathology, King Abdulaziz University Hospital, and from King Abdulaziz Medical City, National Guard Health Affairs, Jeddah, Saudi Arabia, covering the period from 2010 to through 2014. The relevant clinicopathological data (gender, age, grade, and lymph node status and tumor size) were obtained from the patients’ records after obtaining the relevant ethical approvals (Unit of Biomedical Ethics, King Abdulaziz University Hospital No. 1127–13 and King Abdulaziz Medical City No. RJ13/042/J). Genomic DNA was isolated from the FFPE archival tissues utilizing the Qiagen QIAMP FFPE Tissue DNA extraction kit, following the manufacturer’s protocol. Nucleic acid quantity and quality were determined via a NanoDrop 1000 spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA). Qubit DNA high-sensitivity assay kit (Life Technologies, Carlsbad, CA, USA) was utilized to quantify purified DNA. Matched non-tumourous OTSCC tissues (
Library preparation and sequencing
A total of 44 OTSCC specimens were subjected to sequencing on the Ion Torrent Personal Genome Machine (PGM) system as previously described [10]. As low as 10 ng of DNA from each sample were used to construct barcoded Ion Torrent adaptor-ligated libraries utilizing the Ion Ampliseq Library Kit 2.0 (ThermoFisher) as per manufacturer’s protocol. Genomic DNA was amplified using the primer pool from the Ion AmpliSeq™ Cancer Hotspot Panel v2 (ThermoFisher). The panel is designed to cover 2800 hotspots mutations of 50 oncogenes and tumor suppressor genes. The resulting library was purified with AMPure beads (Beckman Coulter), and the concentration of the library was determined by Ion Library AmpliSeq™ Quantitation Kit (ThermoFisher) following the manufacturer’s protocol. 100 pM of DNA from each library was pooled and templated spheres were prepared using the Ion OneTouch 2.0 machine. The enriched template-positive ISPs were loaded onto Ion chips 316 version 2.0 and sequencing was performed using the Ion Sequencing 200 v2 kit following the Ion PGM™ 200 Sequencing protocol (ThermoFisher).
Variants calling and filteration
Sequencing data generated from the PGM runs were processed using Torrent Suite version 4.4.3. The Coverage analysis report, identification of low frequency variants, as well as aligned reads to the reference genome was achieved by the Ampliseq CHP v2 single sample workflow within the Ion Reporter suite v4.6. In order to exclude errors during base calling, a number of filtering steps were followed to generate final variant calling. Within our study, somatic mutations with a total coverage greater than 100
HPV detection
Detection of HPV DNA was performed by PCR using GP5+/GP6+ consensus primers. GP5+ primer sequence is 5’-TTTGTTACTGTGGTAGATACTAC-3’ and GP6+ primer sequence is 5’-GAAAAATAAACTG TAAATCATATTC-3’. GP5+/GP6+ PCR assay was designed to target distinct highly conserved L1 ORF (Open Reading Frame) regions within HPV genome. The final 50
Mutational events identified in OTSCC and the location of pathogenic variants identified in TP53 gene.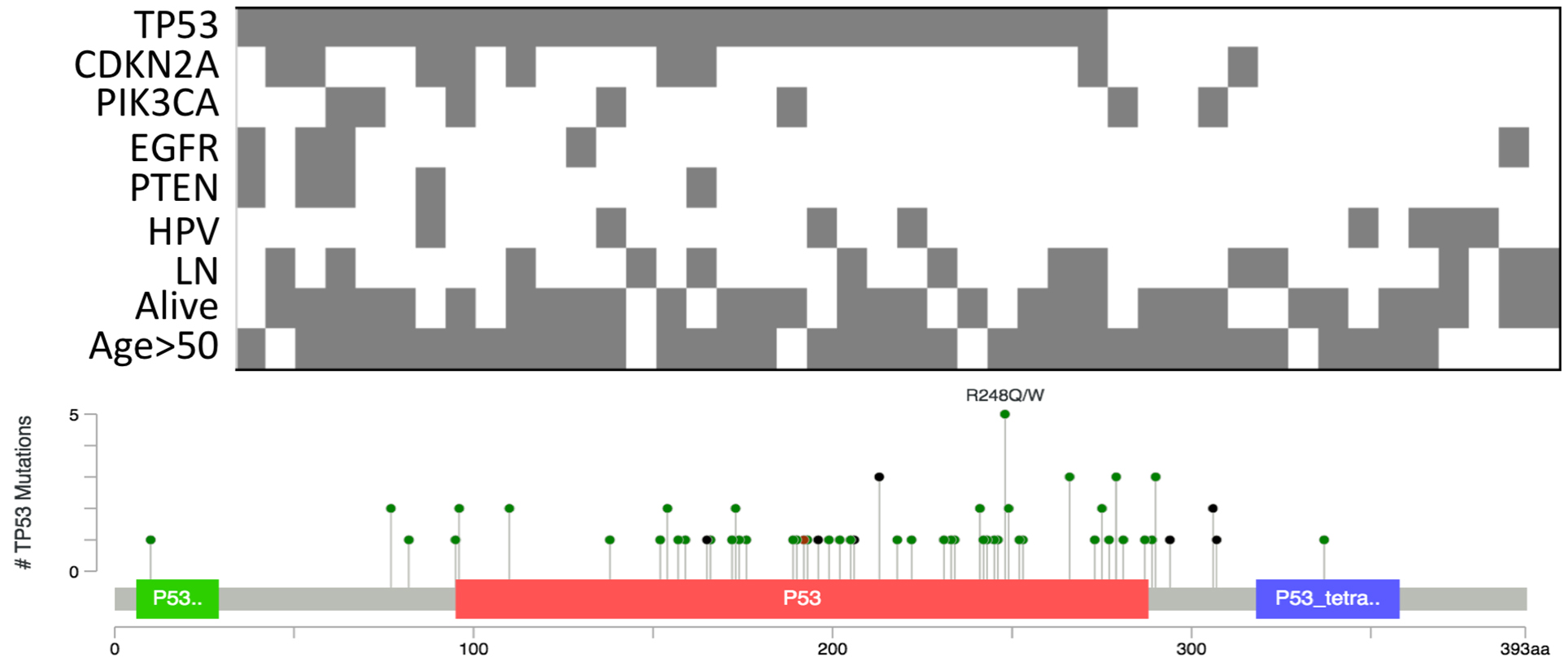
Formalin Fixed Paraffin Embedded (FFPE) Blocks were cut at 4
Statistical analysis
All data analysis was carried out using IBM SPSS Statistics version 24. Fischer’s exact test (two-sided) was applied to evaluate the statistical significance of the association between, genetic and the clinical variables (gender, age, site, LN involvement, tumor size, EPV). Kaplan-Meier method was used to estimate the impact of events on overall survival. Log-rank (Mantel-Cox) comparison method was tested between the strata. Overall survival was defined as the time (in month) from the date of diagnosis to the last recorded date of being alive or death from any cause. Two-sided
Results
We screened 44 OTSCC patients elected on the basis of the availability of tissue block and patient data (Table 1). The average age at diagnosis was 56 years (range, 21–93 years); 35 (80%) patients were
To identify somatic mutations in the oral tongue tumor tissue samples, Ion ampliseq cancer hotspot panel V2 and the ion torrent PGM platform were utilized. This Panel contains 207 primers and is designed to target 2,800 COSMIC (Catalogue Somatic Mutation in Cancer) mutations in the following 50 important oncogenes and tumor suppressor cancer genes: ABL1, AKT1, ALK, APC, ATM, BRAF, CDH1, CDKN2A, CSF1R, CTNNB1, EGFR, ERBB2, ERBB4, FBXW7, FGFR1, FGFR2, FGFR3, FLT3, GNAS, HNF1A, HRAS, IDH1, JAK3, KDR, KIT, KRAS, MET, MLH1, MPL, NOTCH1, NPM1, NRAS, PDGFRA, PIK3CA, PTEN, PTPN11, RB1, RET, SMAD4, SMARCB1, SMO, SRC, STK11, TP53, and VHL. Variant calling and quality-based filtering of the results were performed, and sequencing reports were generated identifying and visualizing the genomic variants present in the samples. All the variants annotated as known mutations due to their presence in the COSMIC, 1000 genomes and dbSNP. The variants not registered in any genomic database were annotated as novel variants and excluded from further analysis (Fig. 1).
Clinicopathological characteristics of the OTSCC cohort used in this study
Clinicopathological characteristics of the OTSCC cohort used in this study
Expression of EBNA1 in OTSCC tissues as detected by immunohistochemistry. (A-B) Non-tumourous tissue, (C-D) Tumour tissue, magnification 20x.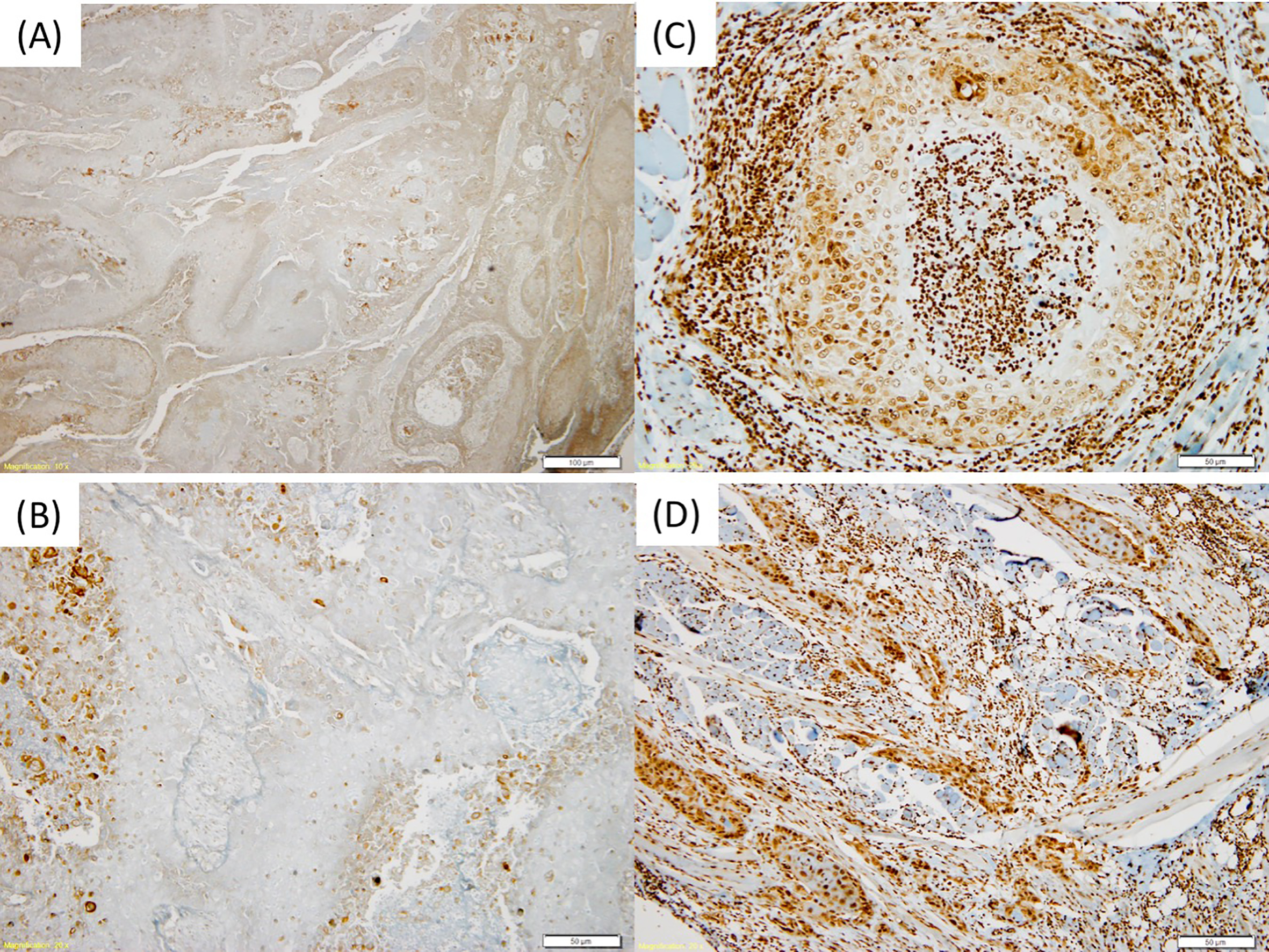
Frequent mutations for the top genes were identified in TP53 (66%), CDKN2A (25%), PIK3CA (27%), EGFR (18%), PTEN (14%), FGFR3 (9%). Less frequent mutations were additionally identified in 40 other genes. In total, 396 somatic mutations were registered across OTSCC tissue samples. Altogether, different missense, nonsense, frameshift mutation, as well as splice site were detected in our cohort. Table 1 illustrated all the protein changes with respect to mutated genes. The mutations within each gene were varied between most of the samples. Majority of the TP53 mutations were missense mutations (82%), while truncating mutations (including nonsense, splice site mutations and frameshift mutations) constituted 18% of the total number of mutations. The most frequent mutation in CDKN2A was the p.Trp110Ter nonsense variant (20%). Overall, truncating mutations constituted 30% of CDKN2A mutations identified in our cohort of OTSCC. The pathogenic change p.Glu545Lys was the most recurrent variant identified in the PIK3CA gene in our cohort (30%) followed by the lesser known p.Ala1066Thr variant which was identified in 3 cases in our cohort. In term of correlating the mutation event with relevant clinicopathological parameters (gender, age, grade, lymph node status, and tumor size) of the OTSCC patients, we observed that somatic mutations in CDKN2A gene were largely associated with lymph node metastasis (
Oncoviruses such as HPV and EBV are major risk factors for OTSCC. Using PCR-based method for the detection of the L1 ORF of the HPV virus we have identified total of 10/44 of the OTSCC samples (23%) as positive for HPV-DNA and 34/44 (77%) as negative. We also observed that viral genomic DNA was significantly dominated in male gender with tongue cancer (
The strongest predictor for survival was positive HPV infection (Kaplan-Meier log-rank test;
Overall survival Kaplan-Meier analysis of the impact of HPV positivity in OTSCC.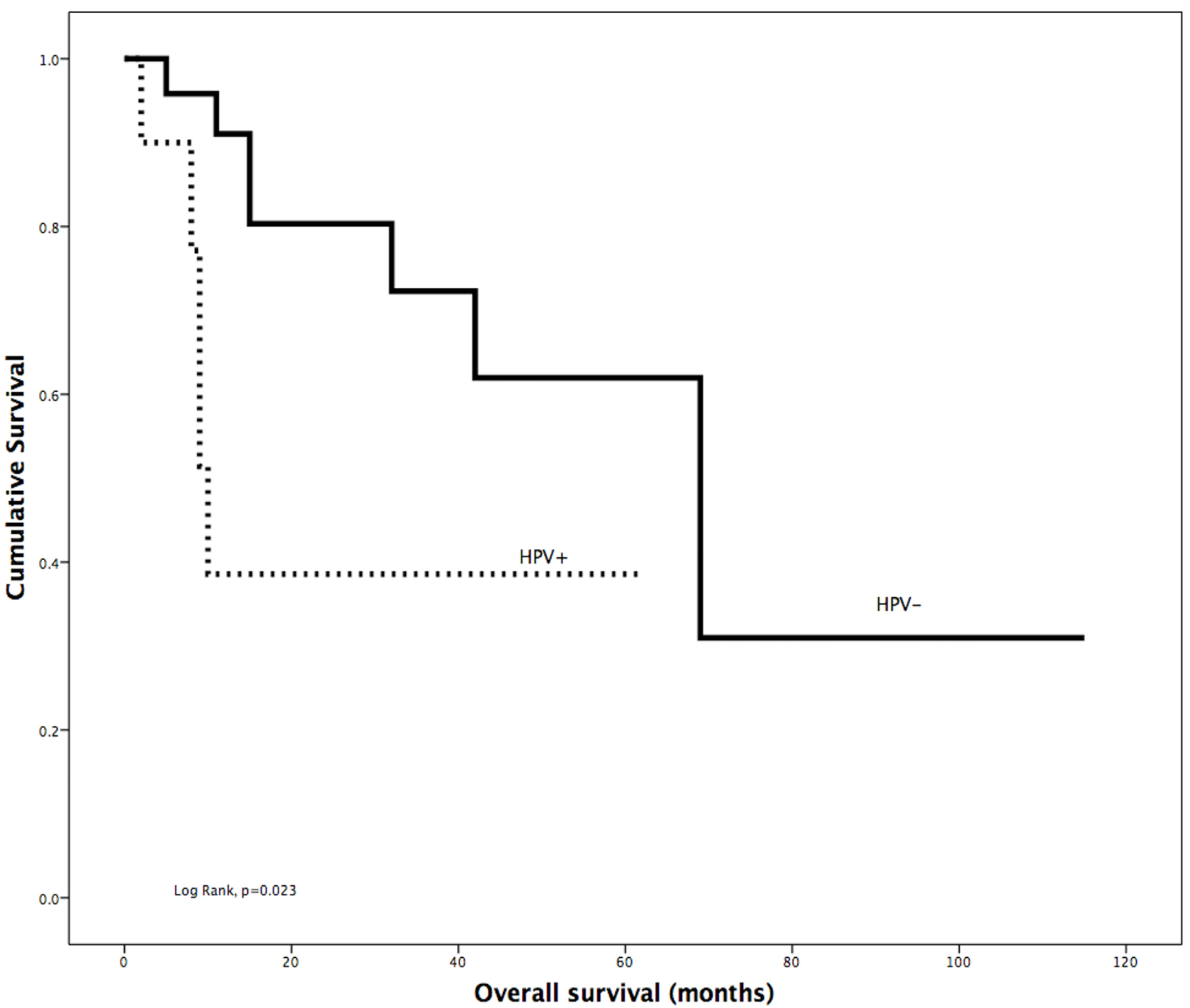
Oral tongue squamous cell carcinoma is the most aggressive cancer in the oral cavity characterized by higher rate recurrence and poor survival. In the current study, we performed a high-resolution genomic sequencing in 44 OTSCC in patients from Saudi Arabia using the Ion Ampliseq Cancer hotspot Panel v2 to sequence 737 loci in 50 cancer-related genes, mainly oncogenes and tumor suppressor genes. We identified TP53 mutations in 66% of tongue tumors as the most commonly mutated gene. This percentage was higher than reported in oral cavity squamous cell carcinomas by COSMIC database [11]. Hotspot mutation TP53 R248Q within the DNA-binding domain of the TP53 protein recurred in our samples. This mutation results in oncogenic gain-of-function phenotype, decrease transactivation of TP53 targets and interference with wild-type TP53 transactivation in cell culture, as well as increased Akt activation and enhanced tumor onset and growth in mouse models [12]. Targeting mutant TP53 is showing promise as a form of universal precision therapeutic approach to cancers. Using small molecule therapeutics such as PRIMA-1/APR-246 small molecule have been shown a promising result in phase I/II clinical trials [13]
PIK3CA mutations in exons 9 and 20 were identified in 27% of 44 tumors. This mutation rate was higher than that detected in COSMIC (7%) and other previous studies in HNSCC tumors (12% [14]). PIK3CA is a critical gene in the phosphatidylinositol 3-kinase (PI3K) signaling pathway. Five patients were found to had the same c.3196G
Cyclin-dependent kinase inhibitor 2A (CDKN2A/ p16INK4A) is another gene that is mutated at a relatively high frequency in our cohort (25%). Other studies reported genomic alterations of CDKN2A in a range between 4 to 74% [18]. Interestingly, tumors harboring CDKN2A mutations were also the most prone to lymph-node metastasis which could indicate a separate subclass of OTSCC. EGFR mutational rate detected in this study was 18%. Deregulation of the EGFR pathway has been described in 80–90% of HNSCC [19].
In terms of incidence and prognosis, the role of HPV in oral tongue carcinogenesis still not well understood unlike in oropharyngeal cases. In the present study, the prevalence of HPV DNA was screened in embedded tissue samples that had been taken from 44 patients with OTSCC. We could detect HPV DNA in 10 tumor samples (22%). Although there is no consensus on rate of HPV incidence among oral tongue tumor patients, it is usually thought that it is low compared to oropharyngeal tumors [20]. We found overall survival for patients with HPV-positive was significantly worse than that of patients. The rate of EBV positivity in our cohort was remarkable. Strong EBV EBNA1 protein expression in the epithelial compartment was detected in 20 out of 21 OTSCC tissues. The high frequency of EBV positivity in OTSCC is expected as it is estimated that around 90% of adults are infected with the virus in different manifestations [21]. The impact of EBV positivity in our cohort is not clear. However, the observation that tumor tissues apparently harbor higher levels of EBNA1 expression may indicate an active role in OTSCC carcinogenesis that warrants further analysis.
Footnotes
Acknowledgments
We acknowledge the valuable support of the technical staff at the Center of Excellence in Genomic Medicine Research.
Conflict of interest
The authors report no conflict of interest.
