Abstract
BACKGROUND:
Hepatocellular carcinoma (HCC) is the second most common cause of cancer death worldwide and the search for clinically useful biomarkers is ongoing. Neighbor of Punc E11 (NOPE) is an established biomarker of murine HCC that remains undetectable in normal liver and at preneoplastic stages.
OBJECTIVE:
The aim of our study was to evaluate the presence of NOPE in human HCC.
METHODS:
Histologically confirmed HCC and corresponding non-tumor liver samples from 20 patients were analyzed for expression of NOPE using qRT-PCR and mRNA-in-situ technology in a conserved tissue context.
RESULTS:
In our cohort, 30% of HCC samples were expressing NOPE which proved particularly useful in non-cirrhotic HCC samples with up to 155-fold higher expression than in adult liver. Using mRNA-in-situ technology, NOPE was clearly identified within epithelial tumor cells of NOPE positive human HCCs. In our analyzed cohort, the combination of AFP with NOPE did not reach more than 40% sensitivity while GPC-3 and NOPE were complementary to each other reaching a combined sensitivity of 85.7%.
CONCLUSIONS:
This is the first characterization of NOPE as a potential biomarker for human HCC. Our results underline the value of NOPE as a complementing biomarker for human HCC.
Introduction
The World Health Organization (WHO) lists hepatocellular carcinoma (HCC) as the second leading cause of cancer death worldwide [1]. In general, HCC development is based on the interaction between genetic and environmental factors and is diagnosed primarily in patients with chronic hepatitis B virus (HBV) infection, nonalcoholic steatohepatitis (NASH) or in patients with a chronic liver disease that has already progressed to liver cirrhosis [2].
According to the classification of the Barcelona Clinic Liver Cancer (BCLC), the prognosis strongly depends on the stage of HCC at the time of diagnosis. Patients with HCC in very early and early stages can be curatively treated with a median overall survival of 60 months and a 5-year-survival rate of 40–70% [3]. In patients diagnosed with terminal HCC, treatment is limited to best supportive care and overall survival is reduced to less than 3 months [4, 5].
Biomarkers have become a very valuable tool in diagnosis and prognosis as well as in prediction and monitoring of treatment outcome in clinical oncology. In HCC, the most prominent biomarker is alpha-fetoprotein (AFP), which is strongly expressed during fetal liver development and diminishes after birth and in adulthood. The AFP serum level may increase again with evolving HCC progresses [6], but 80% of small HCCs are not identified by AFP in serum [7]. In addition, the marker shows a low distinctiveness since patients with HBV and cirrhosis also show high levels of AFP without HCC [8] and the distinction between HCC and intrahepatic cholangiocarcinoma is not reliable [9]. Due to its low sensitivity and specificity in serum, both the European and American guidelines for HCC do not recommend AFP for screening or diagnosis. The membrane protein Glypican 3 (GPC-3) is an additional marker that is widely used in histological analysis [10]. It has been studied as a tissue marker mainly for the diagnosis of hepatitis B virus (HBV)/HCV-related HCC [11]. A diagnostic marker panel combining GPC-3 with heat shock protein 70 and glutamine synthetase reaches a sensitivity of 58.7% and a specificity of 100% respectively for the diagnosis of virus-based HCC population [12].
MicroRNAs such as mi-R122 and various proteins such as Golgi protein 73, AFP-L3 and DCP have also been described to identify HCC at different stages [13, 14, 15]. So far, they are all in the process of further research into combinational diagnostics, but none of them are used in clinical routine. Therefore, the search for clinically useful biomarkers, in particular for screening and early diagnosis of HCC, is not yet complete.
The oncofetal stem/progenitor cell marker Neighbor of Punc E11 (NOPE) was first identified in the liver by microarray analysis of murine fetal liver [16]. Like AFP and GPC-3, NOPE is strongly expressed during fetal liver development, but not in healthy adult hepatocytes, but reappears when the liver is exposed to severe damage. Recently, we have shown that NOPE is expressed in adult depolarized hepatocytes of mice after cholestatic liver injury [17]. Our results suggest a possible role for NOPE as a histochemical biomarker for stage-specific stratification of cholestatic liver diseases with or without parenchymal hepatocyte involvement. In a mouse model of HCC development, we have shown that NOPE is specifically expressed in HCC but remains undetectable in normal liver and at preneoplastic stages [18]. In contrast to AFP and GPC-3, NOPE was detectable in all HCC specimens. These results were more recently confirmed by Kim et al. in a cell culture HCC mouse model with diethylnitrosamine (DEN) for the induction of HCC [19]. In our study, we could also show that murine hepatoma cell lines like Hepa1-6 and Hepa 126 and human hepatoma cell lines such as HepG2, Hep3b, Huh7, Huh6, Huh1, PLC/PRF/5, WRL68 and KMCH also express NOPE up to 229 times higher than adult liver [18]. First data from Oncomine Microarray predicted the applicability to the human system. Therefore, we suggested that NOPE could also be a promising marker for clinical application in humans [18].
The present study is the first evaluation of NOPE in human HCC samples. We analyzed NOPE mRNA levels in comparison to AFP and GPC-3 in human HCC and the corresponding non-tumor liver samples of a cohort of 20 patients. Since specific immunohistochemically-based antibodies against human NOPE are not commercially available, we evaluated the sensitivity and specificity of NOPE expression through a histological approach using qRT-PCR and RNA-Scope technology to specifically detect NOPE mRNA on-slide in a conserved tissue context.
Patient cohort
Patient cohort
Abbreviations: LCI: liver cirrhosis; HCC: hepatocellular carcinoma; TNM: tumor, nodus, metastasis; T: tumor; Y/N: Yes/No; f/m: female/male.
Patients and specimens
Cryopreserved tumor samples and the corresponding non-tumor liver specimens were taken from 20 patients who had resected histologically confirmed HCC in a single center (University Hospital Cologne) between 1997 and 2006. None of the selected patients underwent preoperative cancer treatment. The surgical samples were cut into small pieces and immediately stored under aseptic conditions in liquid nitrogen. The procedures were performed in accordance with the ethical standards formulated in the Helsinki Declaration of 1975 (and revised in 1983). Patients gave their written consent to the use of their tumor samples, an approval was obtained by the Ethics Committee of the University of Cologne (reference number: 13-091).
RNA extraction
The total RNA from all samples was extracted with the RNeasy Mini Kits (RNeasy Minikit (50)-Set Qiagen Shredder and RNAse-Free Dnase, QIAGEN Cat No. 74104) according to the manufacturer’s instructions. The amount of RNA was measured with a Nanodrop ND-1000 spectrophotometer (NanoDrop Technologies, Wilmington, USA).
Reverse transcription of isolated RNA to cDNA
The reverse transcription of the isolated RNA into cDNA was performed by Super ScriptTm III First Strand Synthesis for RT-PCR kit (Invitrogen Cat. No.18080-051) according to the manufacturer’s instructions. For each sample, 5
Quantitative real-time reverse transcriptase polymerase chain reaction
The differential expression of NOPE, AFP and GPC-3 was analyzed using quantitative real-time reverse transcriptase polymerase chain reaction (qRT-PCR). We performed a two-step qRT-PCR with SYBR Green Master Mix and the iQ5 System (BioRad Laboratories, Munich, Germany). Hypoxanthine Phosphoribosyltransferase (HPRT) was used as reference gene. Oligonucleotide primers are listed in Supplementary Table 1 (Table S1). The amplification protocol was as follows: 95
The relative expression level of each gene was calculated using formula 2
Expression of NOPE by qRT-PCR. A) The expression levels of NOPE were analyzed in HCC (T, black bars) and NON-T(umorous, white bars) tissue samples by qRT-PCR. A horizontal dashed line indicates the NOPE expression level in the healthy adult liver (AL), defined as 1. With regard to the cut-off level of 4 times higher expression than AL, indicated by the continuous horizontal line, 30% of tumor samples were NOPE positive, while all NON-T samples except one were NOPE negative. NOPE expression was not measurable in 5 NON-T samples. B) The median expression levels of NOPE in Non-T samples were calculated in cirrhotic (LCI) and non-cirrhotic (NON LCI) samples showing a higher expression level of NOPE in cirrhotic liver samples. The median is below the cut off of 4 in cirrhotic and non-cirrhotic NON-T tissue, confirming its utility for differentiation of NON-T from tumor samples. 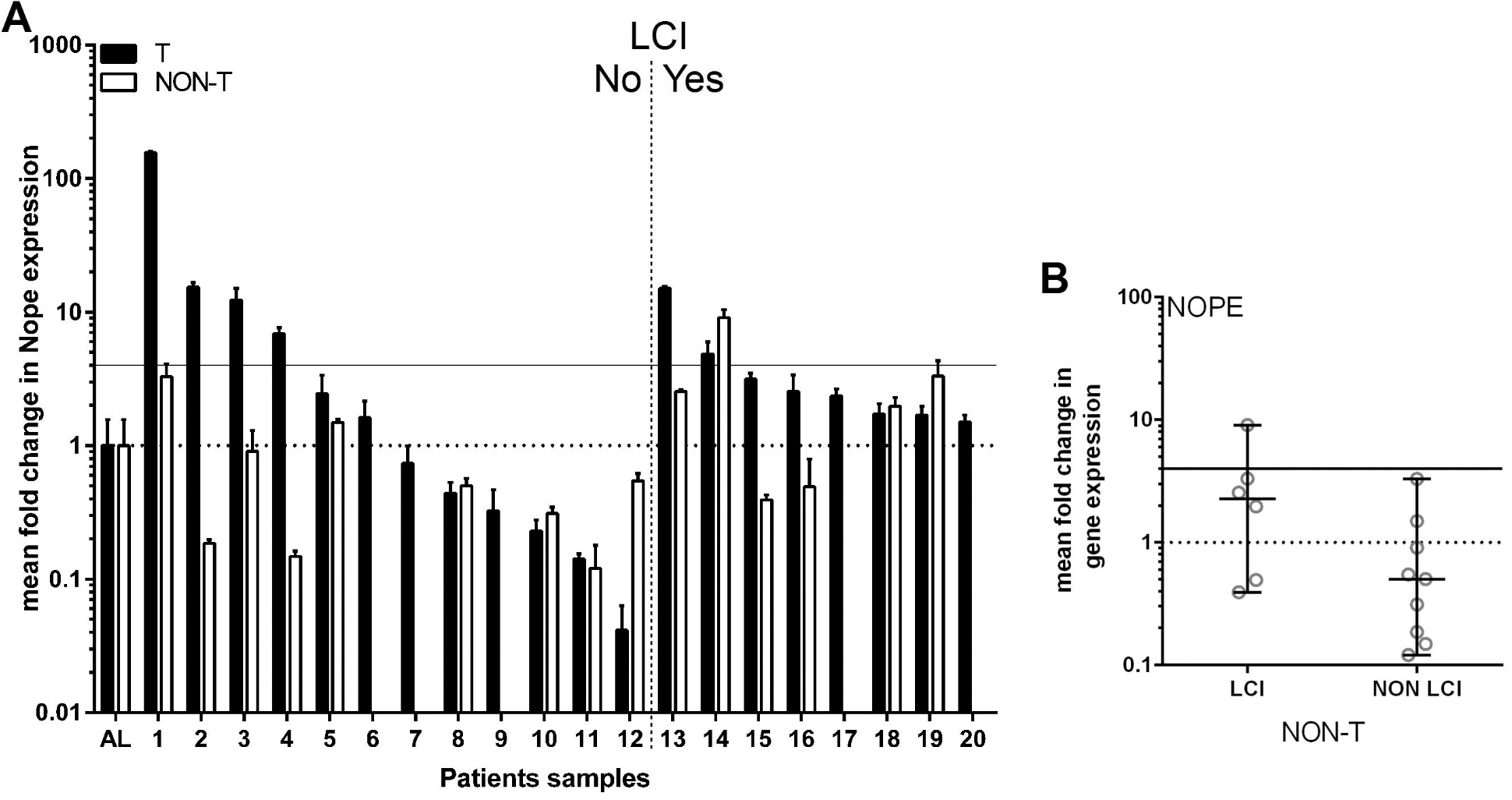
The RNA-Scope assay was performed according to manufacturer’s instructions [21, 22]. In brief, paraffin-embedded TMA blocks were cut into 5
Statistical analysis
The differential gene expression was analyzed by unpaired t-test analysis using GraphPad Prism (version 6.07 for Windows, La Jolla California USA, www.graphpad.com). The
Results
Patient characteristics
The patient cohort consisted of 70% male and 30% female patients aged 38 to 77 (median 68.5) years (Table 1). According to histological studies, 40% of the patients developed HCC in the background of a cirrhotic liver, while 60% showed varying degrees of liver fibrosis but no established cirrhosis. In non-cirrhotic patients, 25% of HCC developed due to chronic hepatitis B (Table 1). Tumor differentiation grade II was identified in 60% of specimens, followed by differentiation grade III in 20% of the samples and differentiation grade II-III in 15% of the samples.
Expression level of NOPE in tumor and non-tumor specimens by qRT-PCR
The expression level of NOPE was measured in all tumor and corresponding non-tumor specimens by qRT-PCR and adult liver served as reference (Fig. 1A).
Across all 20 tumor samples, the maximum expression level of NOPE was 155 times higher than that of AL (Fig. 1A). NOPE was not measurable in 5 non-tumor samples excluded from further analysis. The remaining non-tumorous liver samples from cirrhotic and non-cirrhotic livers were examined for their expression level of NOPE. The non-cirrhotic non-tumorous liver samples showed no significant difference to AL (
Identification of NOPE by mRNA-in-situ technology. A) With mRNA-in-situ technology, NOPE mRNA (red dots) can be specifically identified in carcinoma cells in a conserved tissue context (400-fold magnification, arrows point to NOPE positive cells). However, carcinoma cells reveal only low NOPE mRNA expression levels with an mRNA-Score of 1, i.e. 1-3 dots per tumor cell (see inlet). B) Carcinoma cells which were evaluated to be negative for NOPE were surrounded by NOPE mRNA-positive tumor-related inflammatory cells. 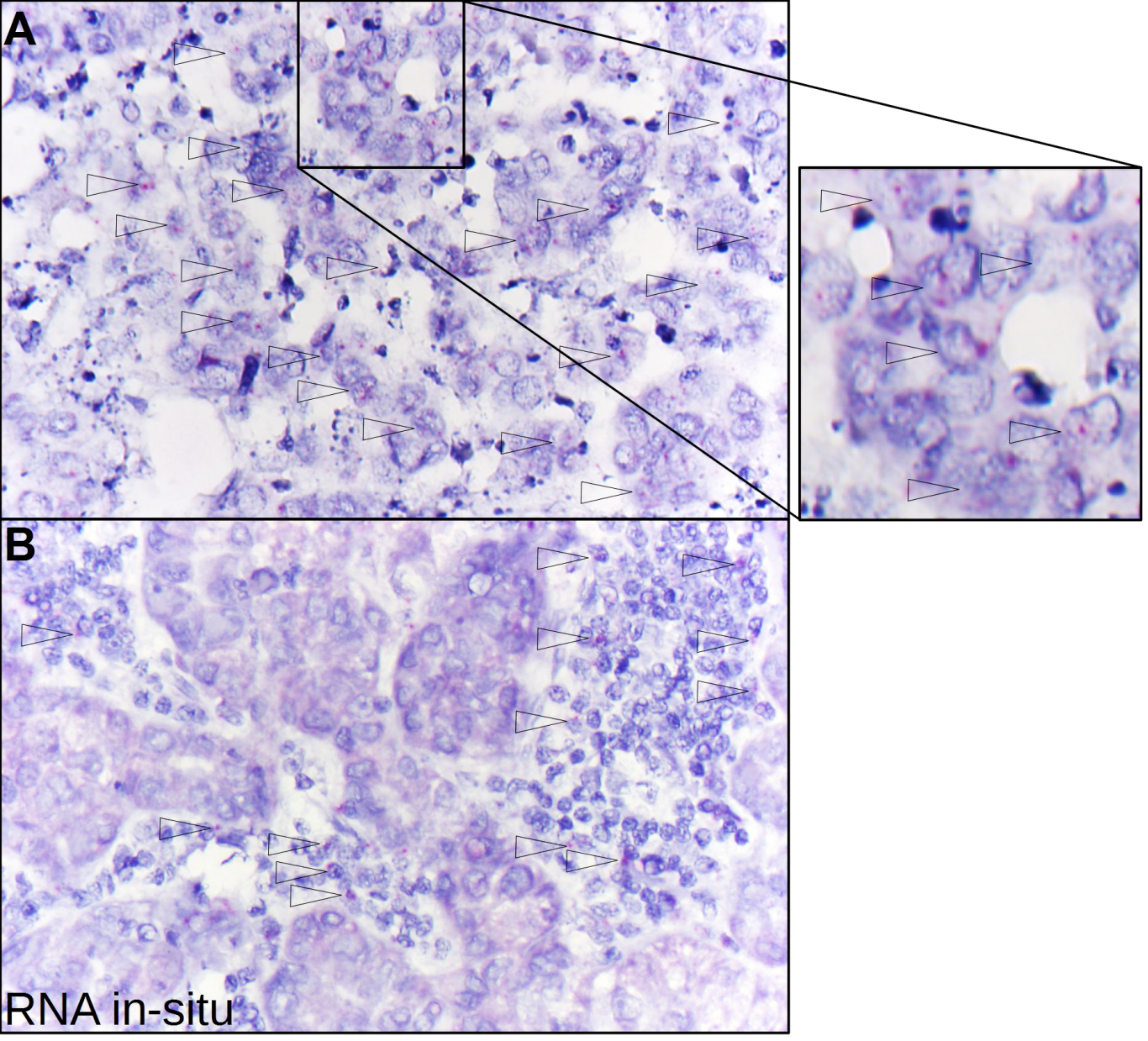
In addition, we analyzed five different NOPE positive human HCCs using mRNA-in-situ technology to measure NOPE mRNA in a conserved tissue context. Three out of five HCCs were evaluated positive for NOPE, i.e. they confirmed signals of NOPE mRNA within the epithelial carcinoma cells (Fig. 2A). However, all carcinoma cells showed only low NOPE mRNA expression levels with an mRNA-Score of 1, i.e. 1–3 dots per tumor cell (Supplement Fig. 2). In addition, we found NOPE mRNA in tumor-related inflammatory cells surrounding carcinoma cells that were evaluated to be negative for NOPE by mRNA-in-situ technology (Fig. 2B).
Comparison of NOPE with AFP
At a 4-fold cut-off compared to AL, 5 of the tumor specimens were AFP positive (sensitivity 25%) (Fig. 3A), but 2 non-cirrhotic non-tumor samples were also AFP-positive (specificity 86.7%). 60% of AFP positive tumor samples came from HCC in cirrhotic livers, resulting in 37.5% sensitivity and 100% specificity. In non-cirrhotic livers, 22.2% of the HCC samples were AFP positive (
Correlation of NOPE, AFP and GPC-3 by qRT-PCR. A horizontal dashed line indicates the expression level of NOPE and AFP in healthy adult liver (AL) defined as 1. Samples originating in a cirrhotic liver are highlighted in black. With regard to the cut-off threshold of 4 times higher expression than AL indicated by the continuous horizontal line, 6 HCC samples were identified by NOPE, 2 of which originating in a cirrhotic liver. AFP was expressed in 3 of the NOPE positive samples, 1 of these originating in a cirrhotic liver. However, AFP identifies only 2 NOPE negative HCC samples, both originating in a cirrhotic liver. 5 HCC samples were identified by NOPE, 2 of which originating in a cirrhotic liver. GPC-3 was expressed in 3 of the NOPE positive samples, 1 of these originating in a cirrhotic liver. However, GPC-3 identifies additional 6 NOPE negative HCC samples, originating from cirrhotic as well as non-cirrhotic liver. 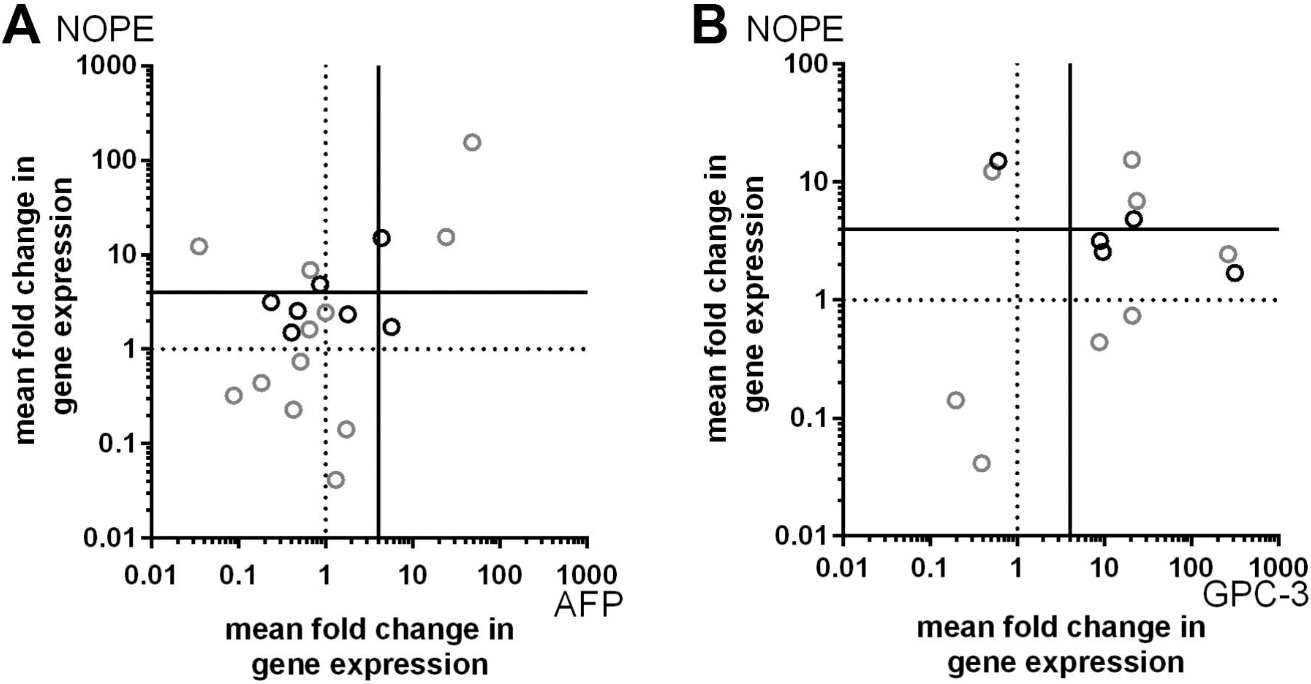
50% of NOPE-positive samples were AFP-positive and 50% were AFP-negative. AFP identified two additional HCC samples (both 6-fold), which were NOPE negative (1.7-fold) (Fig. 3A). In contrast, NOPE identified 3 additional HCC samples (5-, 7-and 12-fold) which were AFP negative (0.9-, 0.7- and 0.03-fold). On average, AFP expression in positive samples was 17.6 times higher than in AL, while NOPE expression in positive tumor samples was 35 times higher than in AL (Supplementary Fig. 1).
Nevertheless, the correlation coefficient between both markers had a value of 0.93 which suggests a very high correlation.
At a 4-fold cut-off compared to AL, 9 of the tumor specimens were GPC-3 positive (sensitivity 60%) (Fig. 3B), but 2 cirrhotic non-tumor samples were also GPC-3-positive (specificity 91.6%). 44.44% of GPC-3 positive tumor samples came from HCC in cirrhotic livers, resulting in 66.66% sensitivity and 83.3% specificity. In non-cirrhotic livers, 55.55% of the HCC samples were GPC-3 positive (
60% of NOPE-positive samples were GPC-3-positive and 40% were GPC-3-negative. GPC-3 identified six additional HCC samples (average 104-fold), which were NOPE negative (average 1.8-fold) (Fig. 3B). In contrast, NOPE identified 2 additional HCC samples (12-and 15-fold) which were GPC-3 negative (0.5-and 0.6-fold). On average, GPC-3 expression in positive samples was 77 times higher than in AL, while NOPE expression in positive tumor samples was 35 times higher than in AL (Supplementary Fig. 1).
The correlation coefficient between both markers had a value of
Discussion
In this study, we confirm NOPE as a potential biomarker for human HCC by qRT-PCR and RNA-Scope analysis. With the RNA-Scope analysis, we were able to describe for the first time the expression of NOPE in epithelial tumor cells of NOPE-positive human HCC. Furthermore, we were able to demonstrate a limited expression of NOPE mRNA in inflammatory cells (Kupffer cells) surrounding NOPE negative carcinoma cells.
In the analyzed cohort of HCC samples, NOPE expression in HCC has been shown to be up to 155 times higher than in AL, which served as a control. With a stringent cut-off of 4-fold higher expression than AL, we identified 30% of the HCC samples as NOPE positive. We had previously calculated a similar detection rate of NOPE in human HCC using preliminary data from Oncomine microarray studies [18].
The ideal biomarker for HCC in cirrhotic and non-cirrhotic liver has not been identified. Besides the high sensitivity and the reliable and specific detection of early stages in carcinogenesis, the limited expression in normal tissue and the independency from the underlying disease are further attributes of ideal tumor markers. Interestingly, NOPE is expressed during fetal liver development and not in the healthy adult liver, which is also true for the established oncofetal marker protein AFP and GPC-3. However, AFP identified only 50% of the NOPE positive samples. Our current data show that HCC derived from cirrhotic or non-cirrhotic liver tissue has differences in the sensitivity of biomarkers: NOPE had a higher sensitivity in non-cirrhotic tissue and AFP a higher detection rate in cirrhotic tissue. 62.5% of these cirrhotic samples had a chronic hepatitis-C-virus (HCV) infection as the underlying disease, which correlates with high serum levels of AFP according to literature [23, 24]. Indeed, geographical differences in underlying liver diseases causing HCC, e.g. predominantly HBV in Asia versus NASH in western countries, also reflect differences in the reliability of biomarkers used in diagnostics [25]. AFP not only has limited sensitivity, but also limited specificity, since high levels of expression also occur in damaged and existing pathological diseases [8]. In contrast, NOPE was not detectable in preneoplastic stages of murine HCC, allowing a precise identification of tumor tissue [18].
GPC-3 is described to be a more reliable marker than AFP in HCC diagnosis [12]. Indeed, our data show 60% sensitivity for HCC samples. The most striking result was the combination of NOPE and GPC3 resulting in an 85.7% sensitivity. Thereby, the combination shows a higher sensitivity than combinations with hock shock protein 70 and glutamine synthetase (58.7–72%) [26, 27, 28] or the triple combination with arginase-1, hepatocyte paraffin-1 and GPC-3 (79.3%) [29].
Although NOPE is not assigned to any enzyme class, the cytosolic domain shows possible binding sites for kinases of the serine/threonine kinase class. One of the predicted possible interacting kinases is Protein kinase C (PKC). NOPE is located at the plasma membrane and conventional PKC isoenzymes are translocated to the plasma membrane upon activation, making an interaction probable. The isoenzyme PKC alpha has been described as highly expressed in murine mesenchymal, chemoresistant HCC cells that show sensitivity to PKC inhibitors [30].
Interestingly, while we have shown that NOPE is expressed in fetal and adult liver stem/progenitor cells [31], the expression of PKC has also been described in stem/progenitor or stem-cell enriched (de-differentiated) cell populations that retain their stem-cell properties [32].
Although NOPE appears to be promising, we have not yet developed a valid method to detect NOPE in human sera. The identification of NOPE is therefore based on tissue samples that are not routinely available, as most HCC can be diagnosed by imaging alone. A further limitation is the fact that we had to use RNA-Scope as a histological method with specially developed probes responding to NOPE-mRNA to confirm our findings. Therefore, investigations on a specific antibody for serum detection and immunohistochemically analysis would complement and strengthen our results.
NOPE appears to be more useful in non-cirrhotic HCC tissue, admittedly, the majority of our analyzed cohort was non-cirrhotic HCC. The analyzed cohort presented a broad range of possible underlying diseases and reflected the gender distribution of HCC patients, but does not reflect the distribution pattern in the real world. To confirm our results, further assessment with larger cohorts is required.
The poor prognosis for patients underlines the urgent need for practicable biomarkers in clinical diagnostics. Since all potential biomarkers for HCC are deficient in meeting the requirements for an optimal biomarker, combined screening settings with other early-stage biomarkers appear to be a reasonable solution at this time.
In conclusion, this is the first characterization of NOPE as a potential biomarker for human HCC samples with high specificity, in particular for non-cirrhotic HCC samples. Our results underline the additional value of integrating NOPE as a complementing biomarker. Further investigations with extended cohorts need to be done.
Footnotes
Acknowledgments
We are grateful to Gisela Holz, Elisabeth Konze and Wiebke Jeske for excellent technical assistance.
Supplementary data
Human oligonucleotide primer sequences used for quantitative RT-PCR
Gene name
ID
Primer sequence
Amplicon length (bp)
Previous publication
NOPE
57722
5’-TGGGCCAAATTCGAAATAAC-3’
109
Marquardt 2007
5’-GATGCTTTCAGGGTTTGAGG-3’
AFP
174
5’-AAGGCCAGGAACAGGAAGTC-3’
127
Marquardt 2007
5’-CACACCGAATGAAAGACTCG-3’
GPC-3
2719
5’-CTACAGCACCCTGTGGTCTTC-3‘
107
Marquardt 2007
5’-TCCTCTACTTCATGGCTGGAG-3’
HPRT
3251
5’-GACCAGTCAACAGGGGACAT-3’
95
5’-GTGTCAATTATATCTTCCACAATCAAG-3’
Expression of NOPE, AFP and GPC-3 in HCC positive samples. A horizontal dashed line shows the level of expression of NOPE and AFP in healthy adult liver (AL) defined as 1. In terms of the cut-off level of 4 times higher expression than AL, indicated by the continuous horizontal line, 6 HCC samples were identified by NOPE. AFP was expressed in 3 of the NOPE positive samples. However, AFP identified only 2 NOPE negative HCC samples, both originating in a cirrhotic liver (samples 13 – 20). GPC-3 was also expressed in 3 of the NOPE positive samples. Interestingly, GPC-3 identified 6 NOPE negative HCC samples, originating from cirrhotic as well as non-cirrhotic liver.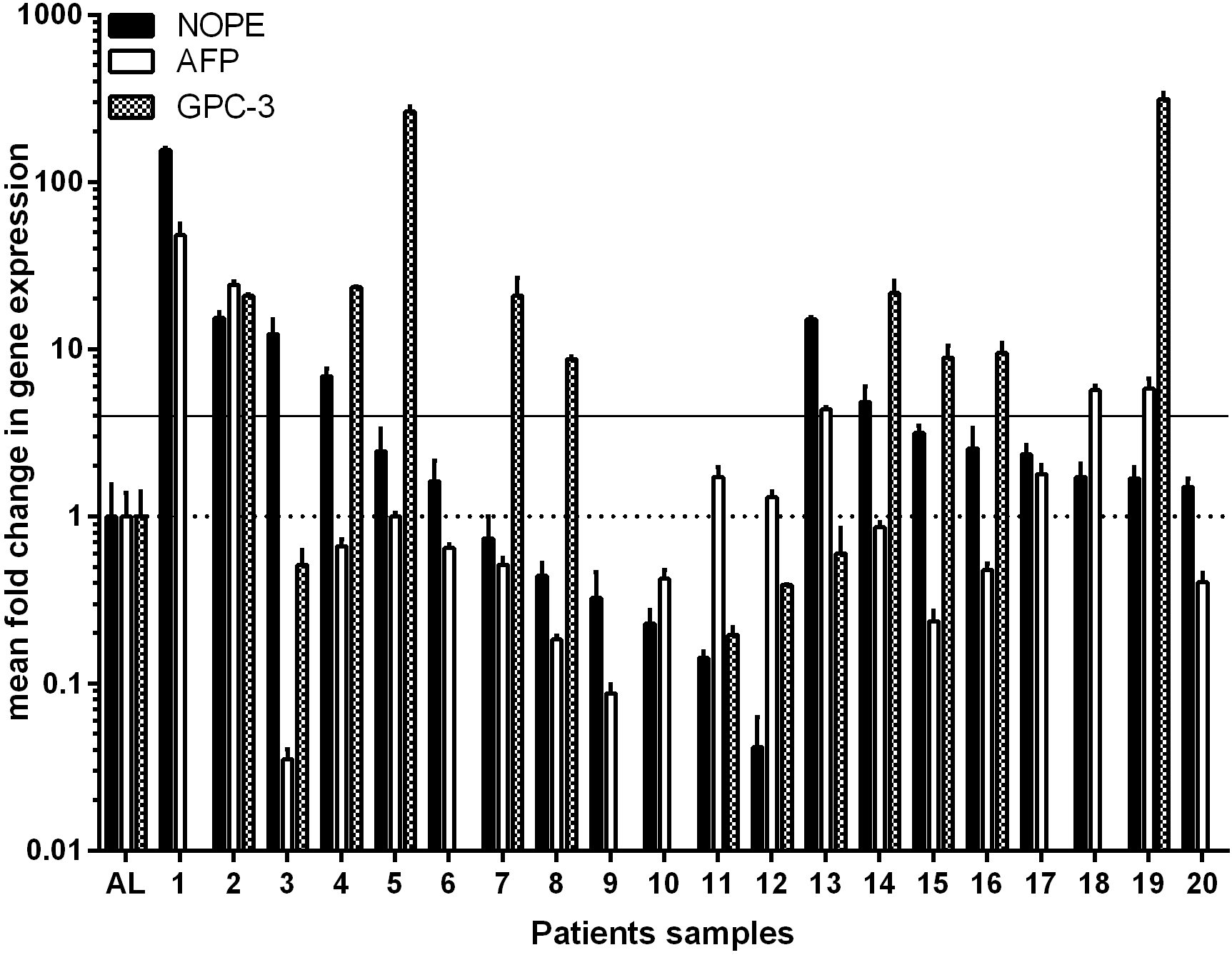
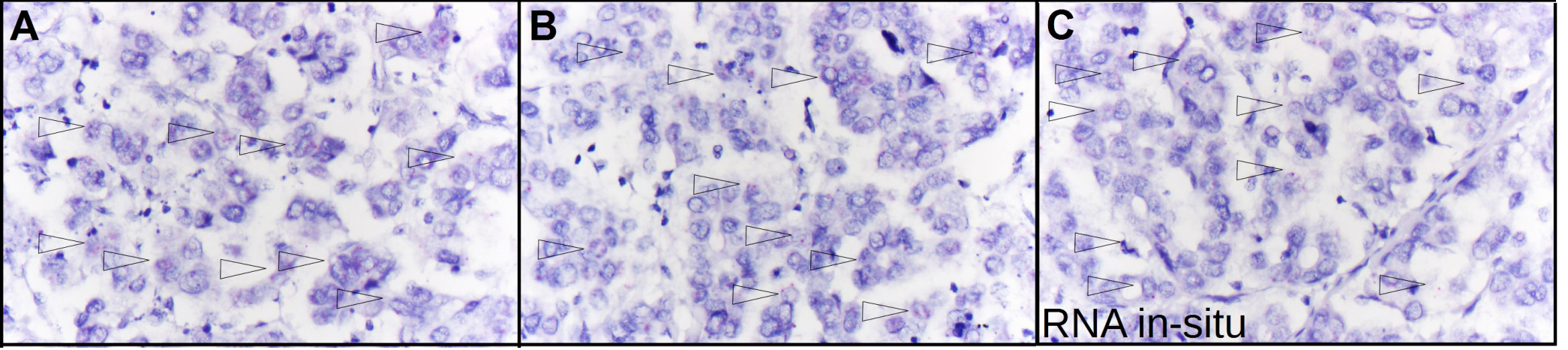
Identification of NOPE by mRNA-in-situ technology. A–C) mRNA-in-situ analysis of NOPE-positive HCC tissues corroborate low NOPE mRNA expression levels with an mRNA-Score of 1, i.e. 1-3 dots per tumor cell. The arrows point to NOPE positive cells.