Abstract
Long non-coding RNAs (lncRNAs) are implicated in the development of carcinomas, containing renal carcinoma. The competing endogenous RNA (ceRNA) network is well-known in modulating the pathological and physiological processes of tumors. Still and all, the function role of oncogenic lncRNA PCGEM1 prostate-specific transcript (PCGEM1) in renal carcinoma was undefined till now. This paper aimed to figure out the role and mechanism of PCGEM1 in renal carcinoma. In this study, PCGEM1 was observed to be lifted in renal carcinoma cells. Loss-of-function experiments displayed that silencing of PCGEM1 repressed cell proliferation and migration, and activated apoptosis in renal carcinoma. FISH assay and subcellular fractionation assay indicated that PCGEM1 was largely located in the cytoplasm. As demonstrated, PCGEM1 interacted with microRNA433-3p (miR-433-3p). Subsequently, luciferase reporter and RIP experiments together with qRT-PCR certified that PCGEM1 and fibroblast growth factor 2 (FGF2) functioned as ceRNA for miR-433-3p, leading to the upregulation of FGF2 expression. Finally, rescue assays exhibited that FGF2 overexpression rescued the inhibited cell progression caused by PCGEM1 downregulation. MiR-433-3p inhibitor could reverse the cell growth and migration caused by PCGEM1 downregulation. The present research investigated the molecular mechanism underlying PCGEM1 in renal carcinoma, exposing a PCGEM1-mediated therapy for the treatment of patients with renal carcinoma.
Introduction
Renal carcinoma is an usual malignant disease in the urinary system and the second primary cause of urinary cancer-interrelated death [1]. Owing to the lack of emblematic clinical symptoms or useful diagnostic markers in the early stage of renal cancer, the initiation of renal cancer is hard to inspect [2]. The majority of patients taking targeted drugs finally develop drug resistance within 6 to 15 months, counteracting the improvement of the survival rate [3, 4].
The sequencing from the human genome indicated that more than 80% of genes are incapable to encode proteins, among which long noncoding RNAs (lncRNAs) above 200 nucleotides in length are important transcripts [5, 6]. Recent studies have demonstrated that lncRNAs play momentous parts in tumor progression via the regulation of gene expression, such as transcription and translation of genes, post-transcriptional modification and so on [7, 8]. For example, lncRNA EGFR antisense RNA 1 (EGFR-AS1) activates cell growth and metastasis in renal cancer by influencing HuR-stabilized mRNA stability of epidermal growth factor receptor (EGFR) [9]. silencing tumor susceptibility 101 (TSG101) represses proliferation and cell cycle in renal cell carcinoma, with the involvement of cyclin E1 (CCNE1), transcriptional regulator Myc-like (c-myc) and cyclin dependent kinase 2 (CDK2) [10]. A DExH-box helicase 9 (DHX9)/lncRNA CCDST/MDM2 proto-oncogene (MDM2) interaction modulates cell invasion and angiogenesis in cervical cancer [11]. LncRNA PCGEM1 prostate-specific transcript (PCGEM1) is a prostate-specific gene in prostate cancer, which has also been researched in several studies and unveiled to be oncogenic [12, 13, 14]. However, the involvement of PCGEM1 in renal carcinoma was unspecified.
The current research inquired the expression profile of PCGEM1 and presented that PCGEM1 was heightened in renal carcinoma cells. Knockdown of PCGEM1 inhibited cell proliferation and migration, as well as induced apoptosis in renal carcinoma. Besides, PCGEM1 functioned as a competing endogenous RNA (ceRNA) to promote fibroblast growth factor 2 (FGF2) expression via sponging miR-433-3p in the cytoplasm. In rescue assays, FGF2 overexpression rescued the repressed cell progression by PCGEM1 downregulation. Suppression of miR-433-3p could reverse the cell growth and migration caused by PCGEM1 downregulation. This study aimed to explore the possible mechanisms underlying PCGEM1 in renal carcinoma.
Materials and methods
Tissue specimens
Human tissue samples used in current study were obtained from 47 renal carcinoma patients who underwent surgical resection in Taizhou Hospital of Zhejiang Province, Wenzhou Medical University. Patients received nothing therapy before surgery. After surgical resection, tissues were rapidly frozen in liquid nitrogen and then stored at
Cell lines and cell culture
All cell lines containing human renal tubular epithelial cells (HK-2) and renal cancer cells (OSRC-2, ACHN, A498 and 786O) were all acquired from the American Type Culture Collection (ATCC, Manassas, VA, USA). RPM11640 medium (Gibco) containing 10% fetal bovine serum (HyClone) was prepared for the culture of 786O and OSRC-2 cells and MEM medium (Gibco) containing 10% fetal bovine serum for the incubation of A498 and ACHN cells, in an incubator at 37
Cell transfection
For transfection, short hairpain RNAs (shRNAs) specifically targeting PCGEM1 (shPCGEM1#1/2/3), miR-433-3p mimics, miR-433-3p inhibitor and pcDNA3.1/FGF2 with relative negative controls (shNC, NC mimics, NC inhibitor and pcDNA3.1) were purchased from Shanghai GenePharma Co., Ltd. (Shanghai, China). 786-O and A498 cells (10
Quantitative real-time polymerase chain reaction (qRT-PCR)
The extraction of total RNA from 786O and A498 cells was performed by use of TRIzol reagent (Invitrogen, USA) under the distributor’s instructions, followed by reverse transcription into cDNA with a First Strand cDNA Synthesis Kit (TOYOBO Life Science, Shanghai, China). A reaction mix of SYBR Green (Takara, China) was used to detect gene expression. The relative expression of PCGEM1 was normalized to that of GAPDH, and miR-433-3p to U6. The used primer sequences were as follows: 5’-TGCCTCAGC CTCCCAAGTAAC-3’ (sense) and 5’-GGCCAAAAT AAAACCAAACAT-3’ (antisense) for PCGEM1; 5’-C GATCATGATGGGCTCCTCG-3’ (sense) and 5’-GTG CAGGGTCCGAGGT-3’ (antisense) for miR-433-3p; 5’-ACTGGCTTCTAAATGTGTTACG-3’ (sense) and 5’-TTGGATCCAAGTTTATACTGCC-3’ (antisense) for FGF2; 5’-CTCGCTTCGGCAGCACA-3’ (sense) and 5’-AACGCTTCACGAATTTGCGT-3’ (antisense) for U6 and 5’-CACCCACTCCTCCACCTTTGA-3’ (sense) and 5’-ACCACCCTGTTGCTGTAGCCA-3’ (antisense) for GAPDH. The results were analyzed using the -
Microarray analysis
The microarray profiling was carried out using HK-2 cell and four kinds of renal cancer cells. RNA extraction and sequential microarray hybridization were performed by Biotechnology Company (Shanghai, China). Human lncRNA array V6.0 (4
Cell Counting Kit-8 (CCK-8) assay
For the CCK-8 assay, we firstly inoculated cells (0.2
5’-Ethynyl-2’-deoxyuridine (EdU) incorporation assay
An EdU assay kit (Guangzhou Ribobio Co., Ltd., Guanzhou, China) was manipulated following the supplier’s protocol. In brief, cells were cultivated in culture media complemented with 10% FBS with 5
Flow cytometry
Cell apoptosis was determined using the Annexin V-fluorescein isothiocyanate (FITC)/propidium iodide (PI) apoptosis detection kit (R&D Systems Europe, Ltd., Abingdon, UK) and analyzed by FACSCalibur flow cytometer, following the quantification with CellQuest 6.0 software (BD Biosciences, Franklin Lakes, NJ, USA).
Caspase-3 activity assay
Caspase-3 assay kit (Beyotime, Haimen, China) was applied to assess the caspase-3 activity following the protocols by the supplier. Stably transfected cells (5
Transwell assay
The migratory capacity was evaluated through the migration number crossing Transwell inserts. Routinely, cell (3
Fluorescence in situ hybridization (FISH)
Fluorescence-conjugated PCGEM1 probes were utilized for RNA FISH experiment. DNA probe sets (Ribobio, Guangzhou, China) were performed for the hybridization, and an NA1.4 inverted Leica DMI6000 microscope (Leica, Heidelberg, Germany) for the observation according to the distributor’s instructions. The images were visualized under a Hamamatsu ORCA-R2 camera (Hamamatsu Photonics, Hamamatsu, Japan) and analyzed by LAS AF software (Leica).
Subcellular fractionation assay
As the supplier’s recommendation directed, subcellular fractionation assay was conducted using PARIS
Luciferase reporter assay
PCGEM1 full length which included the wild-type or mutant binding site for miR-433-3p were inserted into the pmirGLO vector (Promega, Madison, WI, USA), forming the reporter plasmids PCGEM1-WT and PCGEM1-Mut. FGF2 3’UTR containing miR-433-3p-binding sites or mutated miR-433-3p-binding sites were also inserted into pmirGLO vector (Promega, Madison, WI, USA) to construct FGF2-WT or FGF2-Mut reporter. A total of 2
RNA immunoprecipitation (RIP)
RNA immunoprecipitation was carried out by employing RIP kit (Millipore, USA, 17-700) under the franchiser’s instructions. Briefly, the cell lysate by RIP lysis buffer was co-cultured with magnetic beads conjugated with human antibody against Argonaute2 (Ago2) or a negative control mouse Immunoglobulin G (IgG). Antibodies Ago2 and IgG were acquired from Abcam. qRT-PCR was adopted to detect the levels of PCGEM1, miR-433-3p and FGF2 in the RISC complex and also the levels of predicted miRNAs.
Tumor xenograft analysis
Male nude mice were obtained from Shanghai Institute of Material Medical (Chinese Academy of Science). With the approval of Taizhou Hospital of Zhejiang Province, Wenzhou Medical University, the animal experiments were conducted as previously documented [15]. Immunohistochemical staining was conducted based on previous report for Ki-67 expression [16].
Statistical analysis
Data are demonstrated as the mean
Results
High level of PCGEM1 was a potential prognostic factor in patients with renal carcinoma
According to the results from the microarray, we focused on the top five overexpressed lncRNAs (LINC00958, LINC00899, SBF2-AS1, NR2F2-AS1, PCGEM1) in renal cancer cells (Fig. 1A). Besides, we probed their expression in human renal cancer tissues and normal control via qRT-PCR. No significant alterations were discovered in LINC00958, LINC00899, SBF2-AS1, and NR2F2-AS1 expressions (Fig. 1B–F). Of note, PCGEM1 was obviously overexpressed in renal carcinoma tissues. The cellular expression of PCGEM1 was also probed and results suggested its abundance in renal cancer cells, particularly in A498 and 786O cells (Fig. 1G). Meanwhile, highly expressed PCGEM1 predicted poorer survival in renal carcinoma patients (Fig. 1H). More importantly, we disclosed the close correlation of PCGEM1 intensity with TNM stage, tumor size and distant metastasis (Table 1).
Correlation between PCGEM1 expression and clinical features (
47)
Correlation between PCGEM1 expression and clinical features (
Low/high by the sample median. Pearson
High level of PCGEM1 was a potential prognostic factor in patients with renal carcinoma. (A) Expression of lncRNAs in renal carcinoma cells and normal cells were screened by microarray analysis and the results were converted into the heatmap. (B)–(F) The expression of top five lncRNAs in human renal cancer tissues and control samples in qRT-PCR analysis. (G) qRT-PCR detection of PCGEM1 expression in renal carcinoma cell lines (OSRC-2, ACHN, A498 and 786O) and human renal tubular epithelial cell line (HK-2). (H) Kaplan-Meier survival curves demonstrated the outcomes of renal cancer patients with high or low PCGEM1 expression. Group evaluation was analyzed by paired Student’s 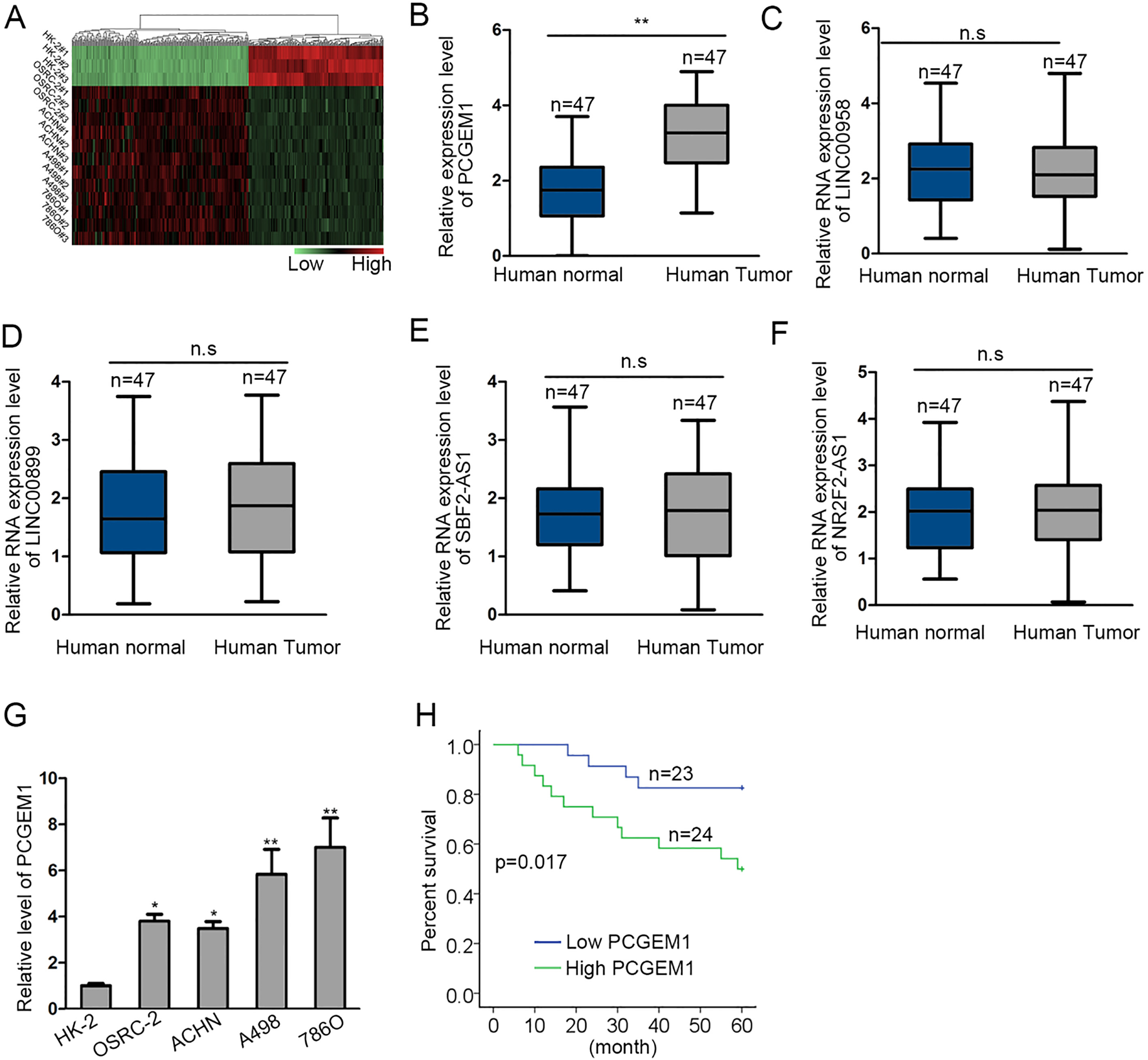
Knockdown of PCGEM1 repressed cell proliferation and migration, and stimulated apoptosis in renal carcinoma. (A) The inference efficiency of shPCGEM1#1/2/3 was detected through qRT-PCR. (B) In shPCGEM1#1/2-transfected 786O and A498 cells, cell viability was examined by using CCK-8 assay. (C) Cell proliferation influenced by shPCGEM1#2 was confirmed by EdU assay. (D) and (E) Detection of caspase-3 activity and flow cytometry analysis were performed to analyze cell apoptosis under transfection of shPCGEM1#2. (F) Transwell assay was employed to dissect cell migration under the treatment of shPCGEM1#2. (G) and (H) Tumor size and tumor volumes in shNC group and shPCGEM1#2 group were detected. (I) Tumor weights in shNC group and shPCGEM1#2 group were examined. (J) Expression of PCGEM1 in tumors developed from shNC cells or shPCGEM1#2 cells. (K) Immumohistochemical staining of Ki-67 level. Statistically significant differences between two groups were analyzed by Student’s 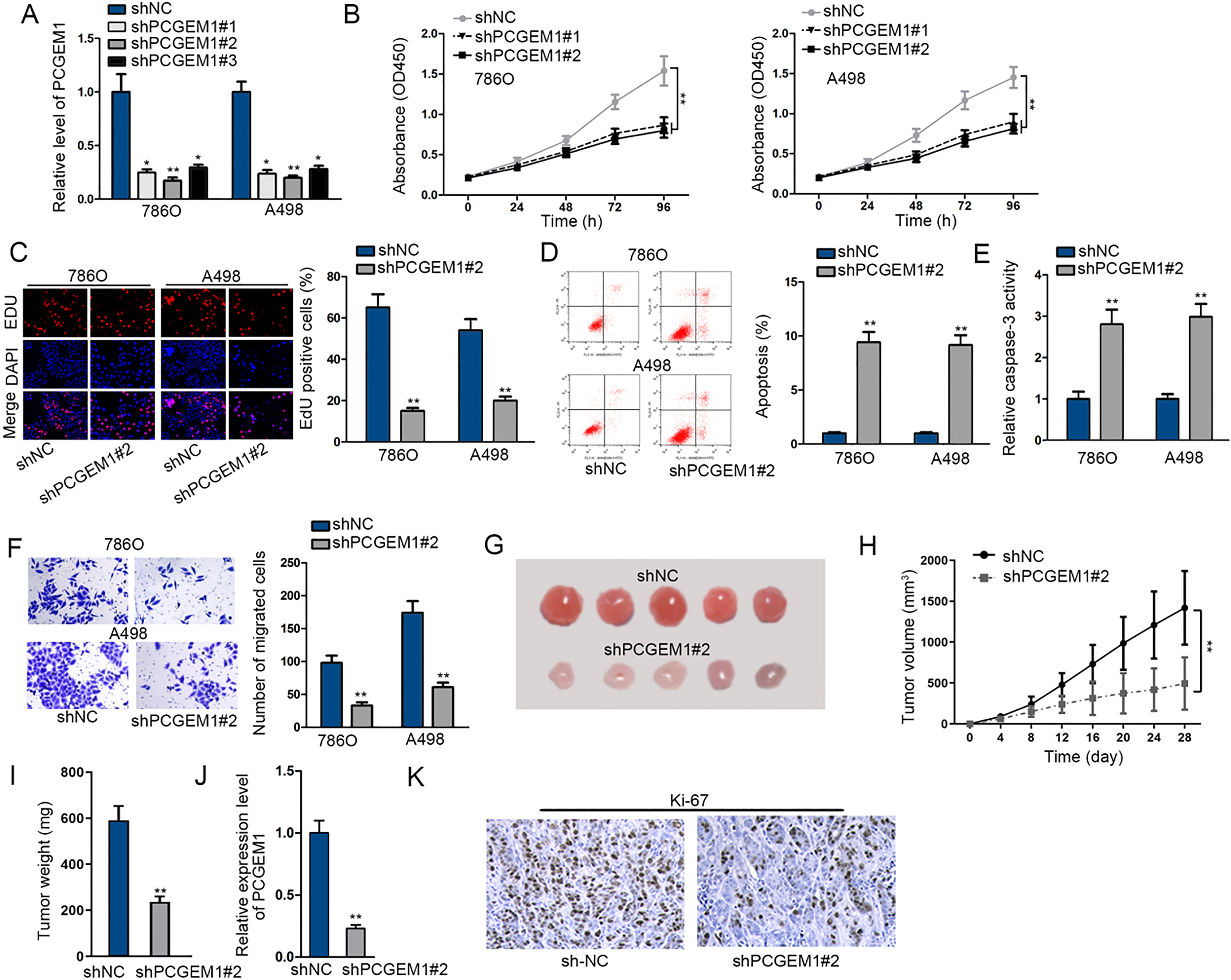
PCGEM1, located in the cytoplasm, interacted with miR-433-3p. (A) and (B) FISH assay and subcellular fractionation assay were applied for the location of PCGEM1. (C) RIP assay was employed for searching the miRNA mostly enriched in Ago2 group. (D) The potential binding sites between PCGEM1 and miR-433-3p and the mutated binding site was illustrated. (E) The expression of miR-433-3p by miR-433-3p mimics was evaluated by qRT-PCR. (F) and (G) Luciferase reporter and RIP results of the interplay of PCGEM1 with miR-433-3p. (H) qRT-PCR analysis of miR-433-3p expression in renal carcinoma cells and human renal tubular epithelial cells. (I) The influence of shPCGEM1#2 on miR-433-3p expression was estimated by qRT-PCR. Unpaired Student’s 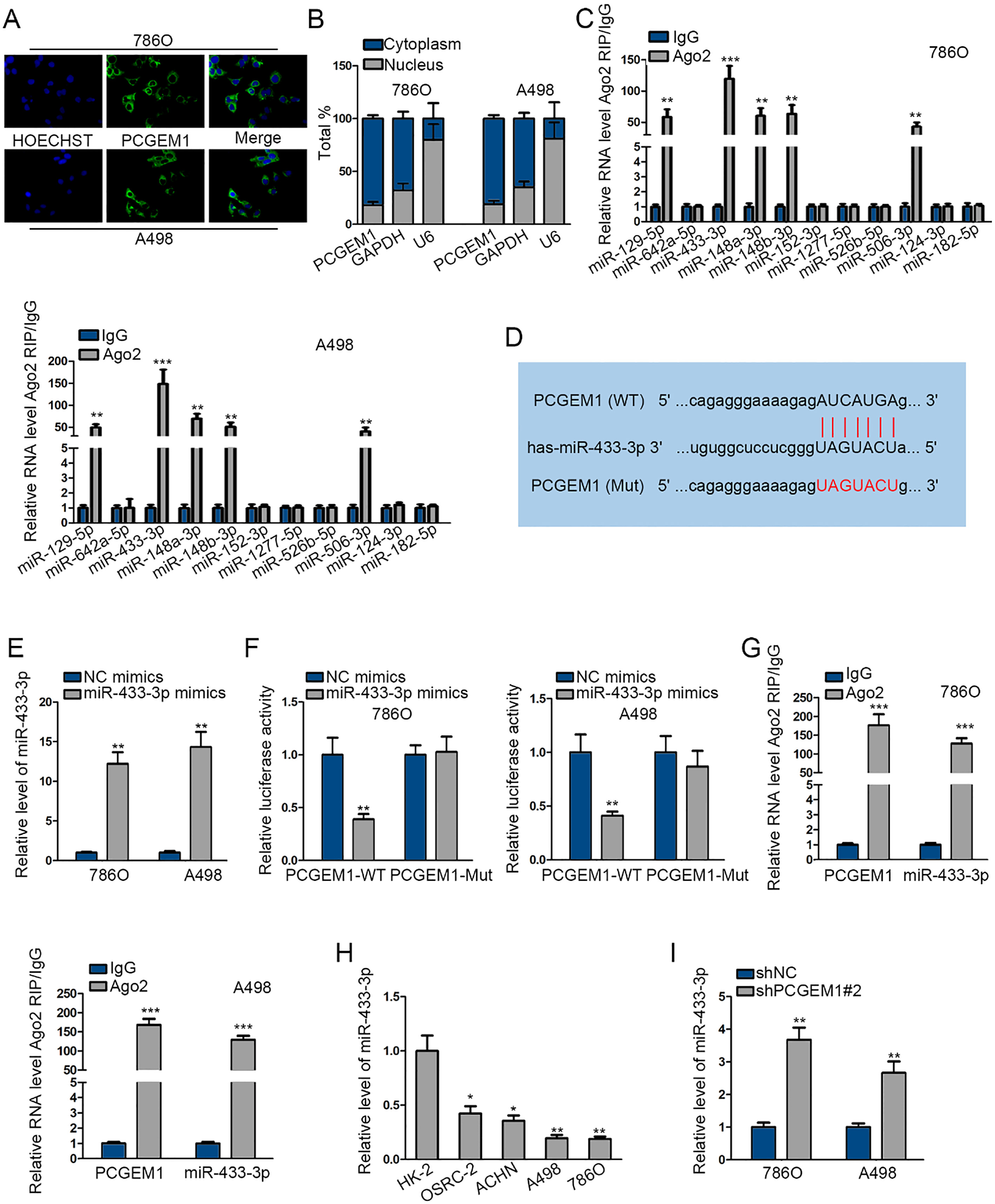
PCGEM1 functioned as a ceRNA to upregulate FGF2 via sponging miR-433-3p. (A) The Venn diagram showed that there are 498 mRNAs both predicted to be targeted by miR-433-3p via miRmap and PITA websites. (B) The expression levels of 498 mRNAs by miR-433-3p mimics were analyzed, as shown in the heatmap. (C) The possible binding sites between FGF2 and miR-433-3p and variant binding site were shown. (D) and (E) The interaction between miR-433-3p and FGF2 was affirmed through luciferase reporter and RIP assays. (F) qRT-PCR and western blotting results of FGF2 expression in renal carcinoma cells and normal HK-2 cells. (G) FGF2 mRNA and protein levels after transfection of miR-433-3p mimics were assessed via qRT-PCR and western blotting. (H) and (I) The co-regulation of PCGEM1 and miR-433-3p on FGF2 was investigated through transfection of shNC, shPCGEM1#2, shPCGEM1#2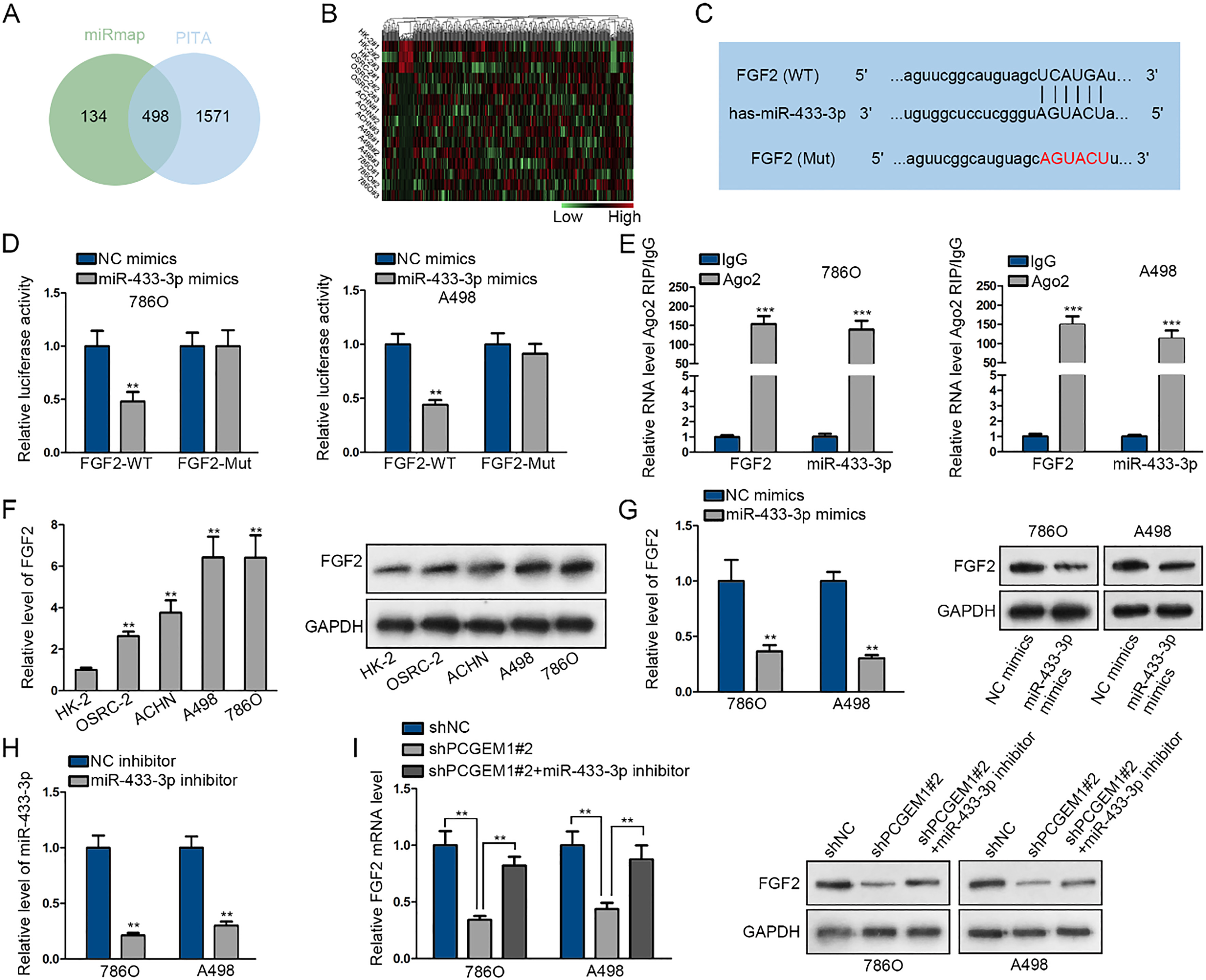
PCGEM1/miR-433-3p/FGF2 axis regulated cell proliferation, apoptosis and migration in renal carcinoma. (A) 786O cells were transfected with shNC, shPCGEM1#2, or shPCGEM1#2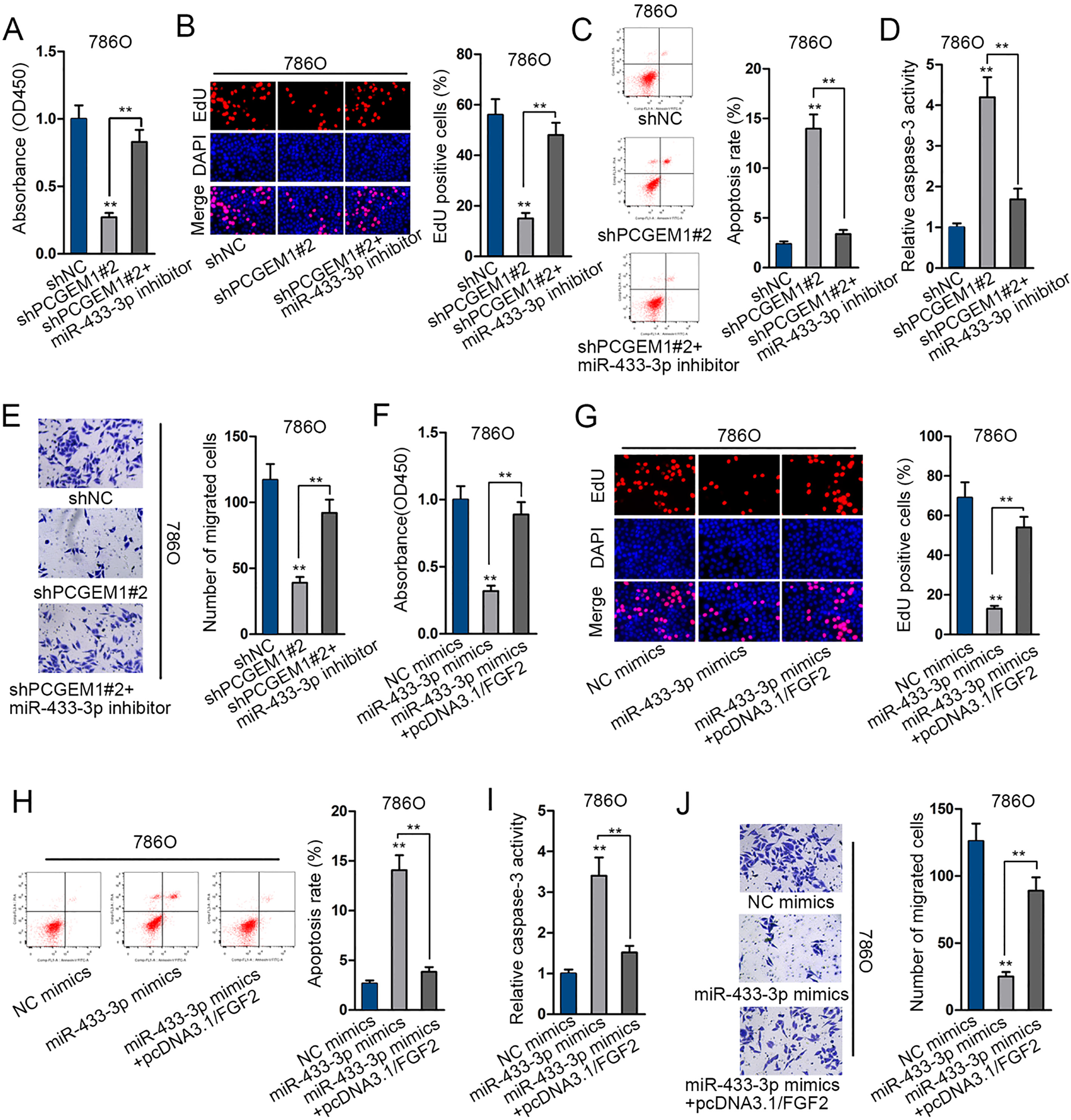
FGF2 overexpression rescued the inhibited cell progression by PCGEM1 downregulation. 786O cells were respectively infected with shNC, shPCGEM1#2 and shPCGEM1#2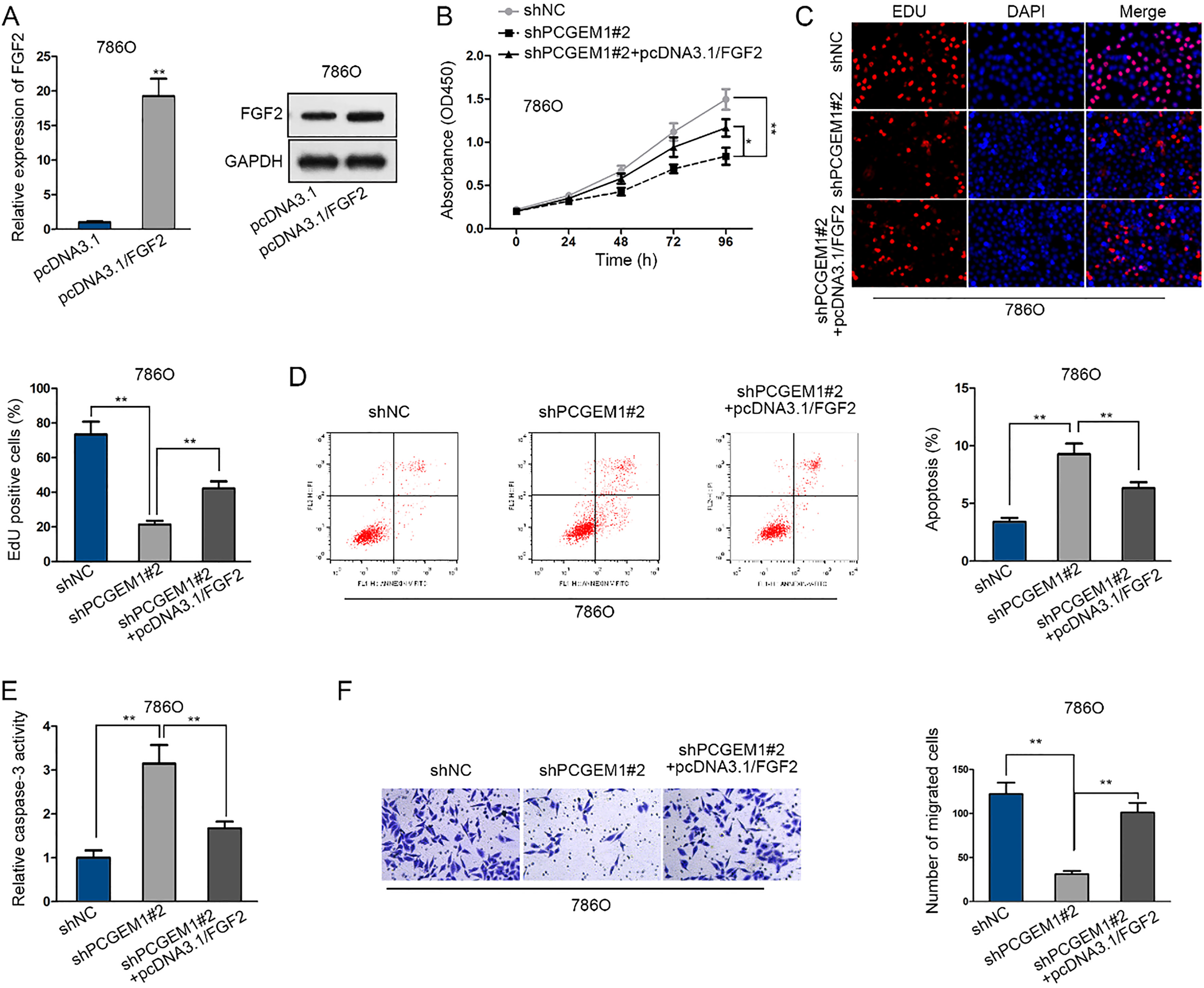
To inquire whether PCGEM1 took part in the initiation and development of renal carcinoma, we attempted to knock down PCGEM1 by transfecting shPCGEM1#1/2/3 into 786O and A498 cells for loss-of-function assays. PCGEM1 expression was decreased after transfection, among which shPCGEM1#1/2 exhibited higher transfection efficacy than shPCGEM1#3 (Fig. 2A). It was revealed by CCK-8 and EdU assays that PCGEM1 silencing restrained cell proliferation (Fig. 2B and C). Flow cytometry analysis exhi- bited that more apoptotic cells were produced when PCGEM1 was silenced (Fig. 2D). Caspase-3 activity assay uncovered that silencing PCGEM1 promoted cell apoptosis (Fig. 2E). And tranwell assay demonstrated that cell migration was inhibited when PCGEM1 was silenced (Fig. 2F). Besides, in vivo animal experiments were conducted. Tumors developed from shPCGEM1#2 cells were smaller and lighter than those from shNC cells (Fig. 2G–I). Besides, qRT-PCR assay also demonstrated the suppressed level of PCGEM1 in shPCGEM1#2-induced tumors in comparison to shNC-induced tumors (Fig. 2J). Immumohistochemical staining showed an inhibited Ki-67 level in shPCGEM1#2-induced tumors (Fig. 2K). To sum up, silencing of PCGEM1 repressed the growth and migration of renal carcinoma cells.
PCGEM1, located in the cytoplasm, interacted with miR-433-3p
LncRNAs located in the cytoplasm have been widely recognized to serve as ceRNAs. It was viewed that PCGEM1 was mainly situated in the cytoplasm, as suggested in FISH assay and subcellular fractionation assay (Fig. 3A and B). We searched on starBase and obtained the predicted miRNAs as target of PCGEM1. Ago2-RIP assay was performed to examine the enrichment of all these miRNAs and indicated the highest enrichment of miR-433-3p in RNA induced silencing complex (RISC) (Fig. 3C). The wild-type and mutant binding sites of PCGEM1 and miR-433-3p sequence were presented in Fig. 3D. Then miR-433-3p was overexpressed by miR-433-3p mimics for follow-up luciferase reporter experiment (Fig. 3E). In luciferase reporter experiment, only the luciferase activity of PCGEM1-WT was reduced by miR-433-3p mimics while PCGEM1-MUT had no change (Fig. 3F). In RIP experiment, the enrichment of PCGEM1 or miR-433-3p was found in Ago2 group (Fig. 3G). In addition, miR-433-3p expression was significantly low in renal carcinoma cells, and could be increased by shPCGEM1#2 (Fig. 3H and I). Collectively, PCGEM1 was located in the cytoplasm and targeted miR-433-3p.
PCGEM1 functioned as a ceRNA to upregulate FGF2 via sponging miR-433-3p
To further probe the regulation mechanism underlying PCGEM1, we continued to browse starBase v3.0 and gained 498 shared mRNAs from miRmap and PITA datasets (Fig. 4A). Heatmap illustrated that FGF2, a noted oncoprotein, exhibited the highest down-regulation in response to miR-433-3p promotion (Fig. 4B). The feasible wild-type and mutated binding sites between miR-433-3p and FGF2 were presented in Fig. 4C. As expected, the luciferase activity of FGF2-WT was dramatically suppressed by miR-433-3p overexpression, and both miR-433-3p and FGF2 were abundant in the mixture precipitated by anti-Ago2 (Fig. 4D and E). Besides, FGF2 expression in mRNA and protein levels were overtly elevated in renal carcinoma cells and obviously lessened by miR-433-3p upregulation (Fig. 4F and G). In order to explore the co-regulation of miR-433-3p and PCGEM1 on FGF2, miR-433-3p expression was abated by transfecting miR-433-3p inhibitor (Fig. 4H). FGF2 mRNA and protein expression was decreased by PCGEM1 knockdown but recovered by miR-433-3p inhibition (Fig. 4I). Taken together, PCGEM1 functioning as a ceRNA sponged miR-433-3p to regulate FGF2.
PCGEM1 sponged miR-433-3p to promote renal cancer cell growth and migration via upregulating FGF2
Finally, to certify whether miR-433-3p or FGF2 could function in PCGEM1-regulated renal cancer growth and migration, we performed a series of cell biological experiments. Cell proliferation, apoptosis and migration regulated by silenced PCGEM1 were rescued by the inhibition of miR-433-3p (Fig. 5A–E). Similarly, miR-433-3p induced inhibition on cell proliferation and migration as well as the promotion on cell apoptosis were rescued after co-transfection with FGF2 expression vector (Fig. 5F–J). Further, we employed rescue assays to verify the mechanism of PCGEM1 in renal carcinoma. FGF2 mRNA and protein expression were promoted by pcDNA3.1/FGF2 in 786O cells (Fig. 6A). As examined by CCK-8 and EdU experiments, cell proliferation was repressed by PCGEM1 downregulation, whereas regained by FGF2 overexpression, as shown in the promotion of cell viability and increase of EdU positive cells in three groups (Fig. 6B and C). As for cell apoptosis, flow cytometry revealed that apoptotic cells were firstly increased with PCGEM1 silencing and then restrained with FGF2 enhancement (Fig. 6D). Further, in caspase-3 activity assay, the PCGEM1 silence-activated cell apoptosis was abolished by FGF2 promotion (Fig. 6E). It was unveiled by transwell assay that cell migration retarded by shPCGEM1#2 was relieved by co-transfection of pcDNA3.1/FGF2 (Fig. 6F). In conclusion, PCGEM1 modulated cell progression in renal carcinoma by FGF2.
Discussion
The ceRNA network explains a hypothesis where lncRNAs and messenger RNAs (mRNAs) can interact with microRNAs. This “competing endogenous RNA” (ceRNA) activity develops into a large-scale modulatory network, greatly revealing the functional information in the human genome and playing crucial roles in pathophysiological conditions, such as carcinomas [17, 18].
LncRNAs are recognized to impact cellular activities of tumors via ceRNA model. For instance, E2F1-promoted lncRNA UFC1 produces oncogenic activities via serving as a ceRNA of FOXP3 [19]. NORAD activates TGF-
From starBase, we found the target miRNAs of PCGEM1 and screened miR-433-3p for further study in renal carcinoma. In recent research, miR-433-3p exerts inhibitive activities in tumors. MiR-433-3p hampers proliferation and invasion of esophageal squamous cell carcinoma via targeting GRB2 [25]. MiR-433-3p obstructs cell growth and improves chemosensitivity through targeting CREB in glioma [26]. It was viewed in this study that PCGEM1, located in the cytoplasm, interacted with miR-433-3p. Hence, we assumed that PCGEM1 might be involved in the ceRNA network with miR-433-3p. Also, through the Venn diagram and the heatmap, we obtained fibroblast growth factor 2 (FGF2), a member of the fibroblast growth factor (FGF) family which is implicated in diverse biological activities, such as tumor growth and wound healing. As numerous reports indicated, FGF2 boosts cellular processes in multiple cancers. For instance, miR-497-5p obstructs proliferation and invasion in non-small cell lung carcinoma via modulating FGF2 [27]. MiR-23a-3p impairs cell proliferation in oral squamous cell cancer through targeting FGF2 and associates with a better prognosis [28]. Stromal fibroblast growth factor 2 lowers the efficiency of bromodomain inhibitors in uveal melanoma [29]. Our study confirmed that PCGEM1 sponges miR-433-3p to modulate FGF2 in renal carcinoma.
Rescue experiments validated that PCGEM1 modulate cell progression of renal carcinoma though miR-433-3p/FGF2 pathway. The current work probed the modulation mechanism of PCGEM1 in renal carcinoma for supporting efforts to recognize reliable and effective biomarkers and therapeutic targets for the survival rate of renal carcinoma.
Footnotes
Acknowledgments
We sincerely appreciate the supports from all parti- cipators. The Science and Technology Plans of Taizhou City (Fund Number: 111KY0609).
Conflict of interest
The authors declare no conflicts of interest.
