Abstract
BACKGROUND:
Deregulated microRNAs (miRNAs) in breast and gynecological cancer might contribute to improve early detection of female malignancies.
OBJECTIVE:
Specification of miRNA types in serum and urine as minimally-invasive biomarkers for breast (BC), endometrial (EC) and ovarian cancer (OC).
METHODS:
In a discovery phase, serum and urine samples from 17 BC, five EC and five OC patients vs. ten healthy controls (CTRL) were analyzed with Agilent human miRNA microarray chip. Selected miRNA types were further investigated by RT-qPCR in serum (31 BC, 13 EC, 15 OC patients, 32 CTRL) and urine (25 BC, 10 EC, 10 OC patients, 30 CTRL) applying two-sample t-tests.
RESULTS:
Several miRNA biomarker candidates exhibited diagnostic features due to distinctive expression levels (serum: 26; urine: 22). Among these, miR-518b, -4719 and -6757-3p were found specifically deregulated in BC serum. Four, non-entity-specific, novel biomarker candidates with unknown functional roles were identified in urine (miR-3973; -4426; -5089-5p and -6841). RT-qPCR identified miR-484/-23a (all
CONCLUSIONS:
Promising miRNAs as liquid biopsy-based tools in the detection of BC, EC and OC qualified for external validation in larger cohorts.
Introduction
Among all cancer types in women, breast and gynecological cancers classify as some of the most frequent malignancies. According to an assessment of the World Health Organization (WHO) in 2018, the incidence of breast cancer (BC) in women is 2,088,849 with 626,679 BC related deaths annually worldwide [110]. Endometrial cancer (EC) is the sixth most common cancer in women with 382,069 women being diagnosed every year [110]. However, most EC subtypes show relatively good prognosis rates. Mortality accounts for 89,929 women each year allocating EC the 14
These still high numbers of cancer-related deaths caused by BC, EC and OC illustrate the unmet demand for sufficient cancer management. Facing the limitations in anti-cancer therapy, early detection of circumscribed malignant changes could represent one of the most compelling efforts to improve clinically implemented diagnosis and treatment strategies to significantly reduce BC, EC and OC-related mortality as well as morbidity. However, to date solely BC guidelines provide elaborate screening recommendations. A vast lack of evidence persists for sufficient and non- or minimally-invasive standard methods for early and subtype-specific EC and OC detection [7, 25, 81, 101, 109].
Mammography forms the centerpiece of BC screening. There is a consensus not to screen women under the age of 40, whereas the details of international screening programs vary [83]. A meta-analysis conducted by Nelson et al. attested an overall benefit of BC screening for average and high-risk women, in particular at the age of 60–69 with 21.3 prevented deaths in 10,000 patients (CI, 5.9 to 19.5) [77]. However, Miller et al. did not find a significant benefit by mammographic BC screening in women between 40–59 [71]. In summary, the value of mammographic screening remains questionable due to variations in diagnostic features caused by practitioner-based interpretations, individual breast tissue densities and false positive findings leading to over-diagnosis and subsequent unnecessary biopsies [83]. Hence, further, ideally non-invasive, screening means should be introduced aiming to narrow the gaping benefit-harm ratio of BC screening.
In contrast to the known positive impact of BC screening on successful therapy, EC routine screening studies did not reveal beneficial effects in asymptomatic women [10, 23, 73, 99]. The diagnostic features of endometrial malignancies comprise initial transvaginal ultra-sonography and subsequent endometrial biopsy or hysteroscopic sampling of the endometrium [89]. These biopsy methods represent an invasive procedure that needs general anesthesia and carries both, surgical and anesthetic risks. Laboratory biomarkers facilitating the detection of EC are not implemented in routine clinical EC management to date [10].
The screening for OC does not only comprise imaging-based procedures but also serum biomarkers, namely cancer antigen 125 (CA 125). However, German guidelines [24] as well as the U.S. Preventive Services Task Force recommendations [74] negate a significant benefit on mortality and morbidity by the performance of both screening methods neither in average nor in high risk groups [70, 119]. The significance of CA 125 as biomarker in OC is not completely understood with serum CA 125 being significantly elevated in only 50% of early stage OC [4, 119]. Moreover, varying individual expression levels of CA 125 complicate the definition of an appropriate threshold for OC detection. In spite of its limitations as a biomarker, CA 125 still outplaces upcoming biomarkers like human epididymis protein 4 (HE4) and mesothelin in terms of discriminatory potential [4, 41]. The discussed guidelines and recommendations demonstrate the lack of sufficient screening methods that provide satisfying sensitivity, easy accessibility and acceptable complication rates.
Characteristics of BC patients stratified by tumor molecular subtype and healthy controls
Characteristics of BC patients stratified by tumor molecular subtype and healthy controls
BC: breast cancer; Lum A: luminal A molecular subtype; Lum B: luminal B molecular subtype; Her2+: Her2 enriched molecular subtype; TNBC: triple negative molecular subtype; * defined by St Gallen Consensus 2012; n.a.: no information available; ER: estrogen receptor expression; PR: progesterone receptor expression; CTRL U40: healthy women at the age of
Being involved in overall physiological and pathophysiological pathways, microRNAs (miRNAs) are crucial regulators of carcinogenic processes [1, 14, 88]. In that context they can either act as tumor suppressors or exert oncogenic functions [120]. Growing evidence underlines the involvement of miRNAs in BC [21, 59, 98], EC [43, 104]and OC [9, 22, 63, 90]. Furthermore, both circulating miRNAs in the blood stream as well as secreted miRNAs in urine might serve as early biomarkers in screening, diagnostics and therapy response evaluation of various tumor entities [30]. The promising potential of miRNAs as novel, minimally-invasive biomarkers for the detection of BC has been shown by an increasing number of studies [6, 112, 124]. Before unraveling the auspicious potential of using urine as a diagnostic matrix for detecting BC specific miRNA patterns, the feasibility of reliably detecting miRNAs in urine first needed to be determined. A pilot study of Erbes et al. identified four urinary miRNAs that could distinguish between BC patients and healthy controls [29]. Hence, miRNA signatures might serve as innovative biomarkers for BC diagnosis. miRNA expression pattern might even serve as a prospective tool for classifying BC into molecular subtypes and as useful parameters in tumor grading [49]. In gynecological tumors such as EC and OC, miRNA patterns were found to be altered in different body fluids, respectively [32, 72]. However, evidence of miRNA dysregulation in cancer is conflicting and depends to a large degree on isolation methods, applied analytic approach and particularly on the identification of stable endogenous controls in definite settings.
Characteristics of EC and OC patients
EC: endometrial cancer; OC: ovarian cancer; n.a.: no information available; FIGO: Fédération Internationale de Gynécologie et d’Obstétrique. All patients showed no metastasis at the time of diagnosis (M0). * one patient diagnosed with a benigne ovarian lesion in retrospect.
The present study focuses on the discovery of blood- and urine-based miRNA biomarkers for screening and early detection of BC, EC and OC. Such easily accessible, minimally-invasive biomarkers could be beneficial for both patients and clinicians and could help to reduce health care costs. In the conducted discovery phase, we first investigate altered miRNA expression signatures in BC, EC and OC as reflected in body fluids using a state-of-the-art microarray chip analysis. Thereby, we aim to narrow the number of miRNA types as defined by miRBase V21 [35] to identify potential, robust biomarkers. An additional approach focuses on the identification of stably expressed miRNAs to be established as robust comparative values for quality control and data normalization in a qPCR setting. Subsequently, selected miRNAs identified in the discovery phase are validated via qPCR.
Patient cohort and sampling procedures
The initial cohort (discovery phase) comprised of 27 patients, the validation cohort comprised of 82 patients undergoing therapy for BC, EC or OC at the University Hospital Freiburg. Ten (discovery phase) and 62 (validation) proven healthy women attending routine screening procedures such as ultrasound, palpation or mammography were recruited as controls. Patients and controls were enrolled from December 2015 to November 2018 after the study was approved by the local Ethics Committee (#386/16; #607/16) and written informed consent was obtained from each participant. BC subtypes were defined according to St. Gallen Consensus 2012 [33]. For patient characteristics see Tables 1 and 2.
Collection of blood and urine samples was performed by physicians prior to treatment (surgery or neoadjuvant chemotherapy) following standard protocols. Whole blood was incubated for 20 min at room temperature allowing coagulation and then centrifuged (10 min, 3500 rpm, 4
RNA isolation
Total RNA was isolated from 200
miRNA microarray chip analysis
For miRNA microarray chip analysis, RNA from both, serum and urine samples, was pooled by tumor entity. The EC and OC pool consisted of five patients each. The entirety of the BC group contained 17 patients diagnosed with definite BC molecular subtypes. While the BC group was formed by five luminal A (Lum A), luminal B (Lum B) and triple-negative BC (TNBC) patients, respectively, it contained only two patients with Her2 enriched (Her2+) molecular subtype. BC samples were applied in four pools grouped by molecular subtype. Comprising five healthy women each, two control pools (CTRL) were set up separated by age (
The following steps were conferred and processed by OakLabs (OakLabs GmbH, Hennigsdorf, Germany) [80]. From each pool, 100 ng of total RNA was compiled. Human miRNA microarray chip analysis (Agilent-070156 Human miRNA V21.0 Microarray 046064, 8
Deregulated miRNAs in serum of breast and gynecological cancer patients discovered by microarray chip analysis. Log
(FC) and
-values
Deregulated miRNAs in serum of breast and gynecological cancer patients discovered by microarray chip analysis. Log
FC: foldchange; T: tumor (BC
Applying RT-qPCR, miR-484 (serum) and -205-5p (urine) were further analyzed for their diagnostic potential for EC and OC and miR-10b-5p (urine) as a non-specific biomarker for BC, EC and OC. Additionally, miR-23a-3p was investigated for its housekeeping potential. Poly(A) tailing-based RT was performed in a reaction mix containing 4
Workflow of data processing: Agilent-070156 Human miRNA V21.0 Microarray (046064, 8 x 60K) was performed assessing 2549 miRNAs. Dark grey blocks represent serum data set, light grey blocks represent urine data set. Data processing includes the removal of miRNAs at the level of detection (LOD 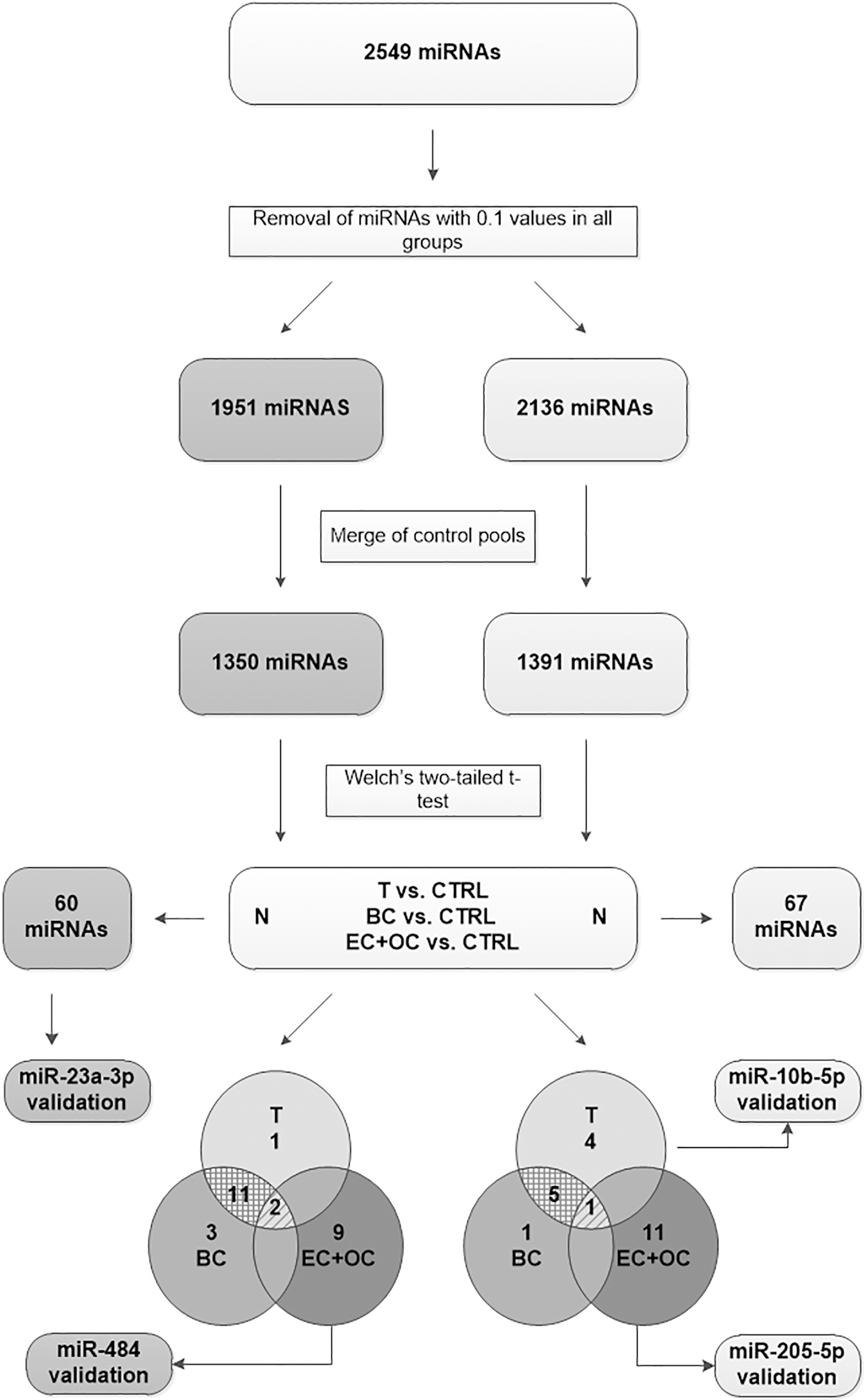
1
Expression levels of 2549 miRNAs in total (miRBase Release 21 [35]) were assessed by microarray chip analysis. In a first essential step, raw data obtained by OakLabs was processed by excluding miRNAs that exclusively exhibit expression values of 0.1 (assay limit of detection, LOD) in all investigated pools. An expression value of 0.1 can represent either a miRNA that is not present in the investigated specimen or one that shows a rather weak expression and is therefore not suitable to be assessed reliably by this analysis. This analytical approach reduced the serum data set to 1951 miRNAs and the urine data set to 2136 miRNAs, respectively. Raw data of the remaining targets was subsequently quantile normalized and log
miR-484 expression levels in serum of breast (BC), endometrial (EC) and ovarian cancer (OC) patients vs. healthy controls (CTRL). (A) Expression levels of miR-484 were significantly elevated in combined EC and OC vs. CTRL (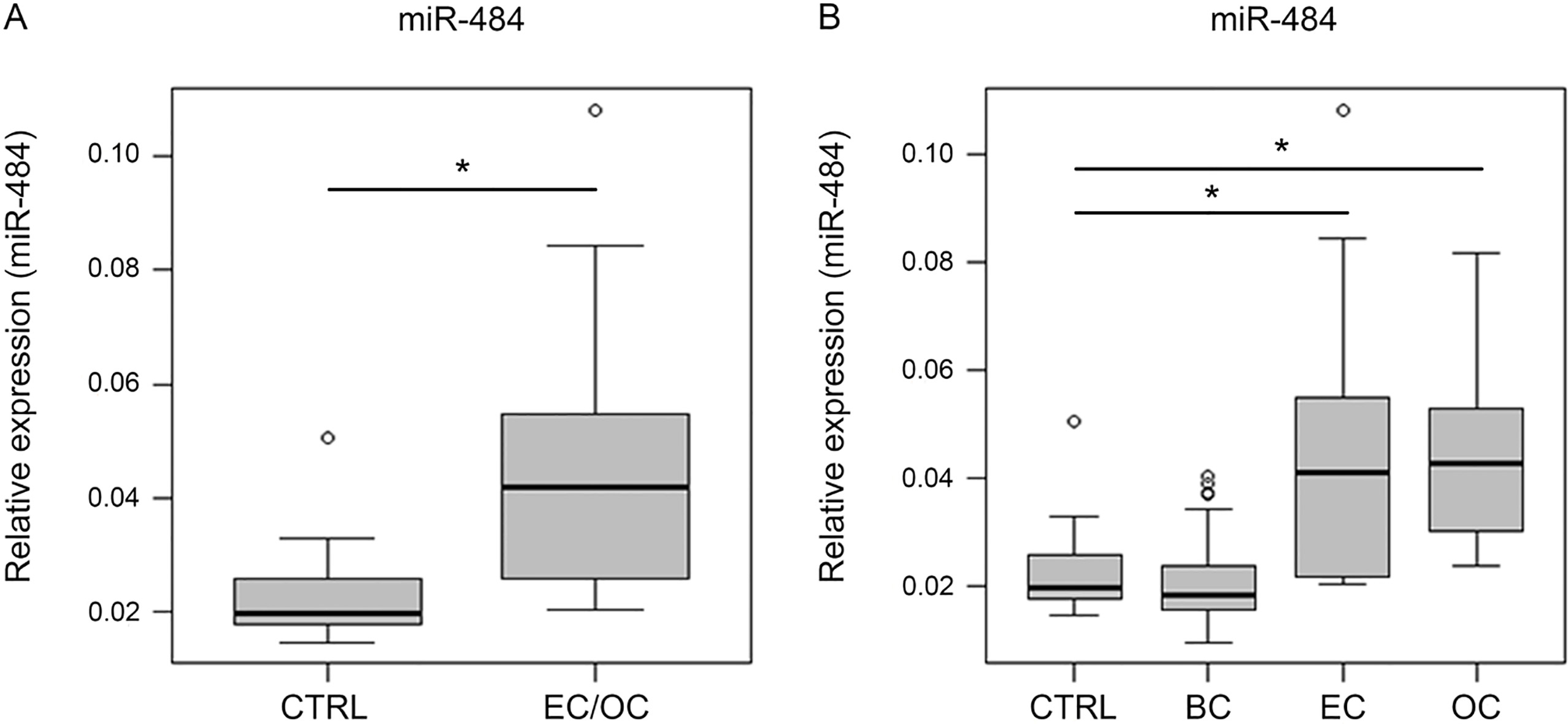
Discovery phase: miRNA biomarker candidates in serum
In serum, 14 miRNAs were discovered by microarray analysis that discriminated between cancer patients and healthy women. Furthermore, the expression levels of 16 miRNAs were altered in the BC group vs. CTRL, eleven in the EC
Deregulated miRNAs in urine of breast and gynecological cancer patients discovered by microarray chip analysis. Log
(FC) and
-values
Deregulated miRNAs in urine of breast and gynecological cancer patients discovered by microarray chip analysis. Log
FC: foldchange; T: tumor (BC
miR-10b-5p and -205-5p expression levels in urine of 25 breast (BC), 10 endometrial (EC) and 10 ovarian cancer (OC) patients vs. 30 healthy controls (CTRL). (A) No expression level alterations of miR-10b-5p between CTRL and tumor (T). T 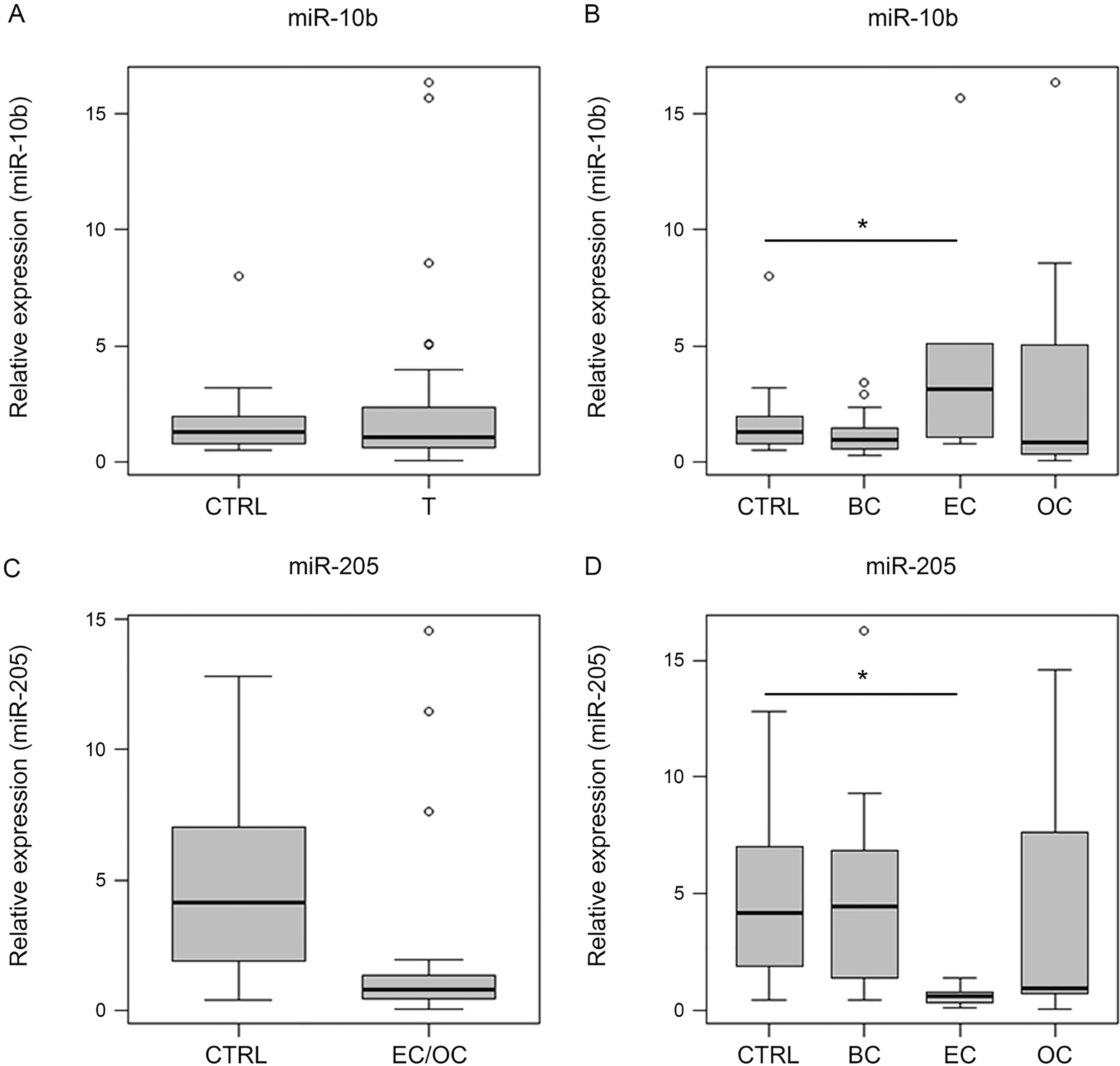
Statistical testing of 1391 miRNAs expressed in urine revealed ten miRNAs with tumor-specific attributes. While seven miRNAs discriminated BC from CTRL, only miR-526b-3p did not show overlapping traits with the overall tumor group. In the gynecological tumor group (EC
Discovery phase: Promising miRNAs for normalization
Stably expressed miRNAs are of great benefit for sample quality assessment and for data normalization when a methodological approach is restricted to a relative expression analysis (e.g. in RT-qPCR). In the present study, a miRNA type was considered a normalization candidate if foldchanges (FC) between all applied pools did not exceed a value of 2. In addition, a cut-off for quantile normalized expression values was set at a threshold of 0.2, which equals twice the LOD of the applied measurement technique.
Data analysis revealed 60 miRNA normalization candidates in the serum set and 67 in the urine set (Supplementary Table S3). Of these, 14 miRNAs showed consistent expression levels for both compartments, respectively (miR-3656; -3960; -6087; -6807-5p; 6752-5p; -188-5p; -6089; -4763-3p; -3194-5p; -3940-5p; -1249-3p; 6757-5p; -6088; -6756-5p). Since it has been proposed as a suitable miRNA for data normalization in other contexts, miR-23a-3p was chosen for validation in serum [8, 34, 94].
Validation phase
MiR-484 showed significant differences via RT-qPCR in serum of 13 EC, 15 OC and 32 BC patients vs. 32 CTRLs (Fig. 2A and B). In alignment with the discovery a two-sample
In urine, miR-10b-5p and -205-5p were analyzed in 25 BC, 10 EC, 10 OC patients and 30 CTRL samples (Fig. 3). However, miR-10b-5p could not be confirmed as a non-tumor entity-specific biomarker in this context (
Furthermore, miR-23a-3p was analyzed in the serum validation cohort to determine its potential as endogenous control. While miR-23a-3p showed a relatively stable expression in the discovery cohort (maximum FC
miR-23a-3p expression levels in serum of 32 breast (BC), 13 endometrial (EC) and 15 ovarian cancer (OC) patients vs. 32 healthy controls (CTRL). Expression levels of miR-23a-3p were significantly decreased in EC (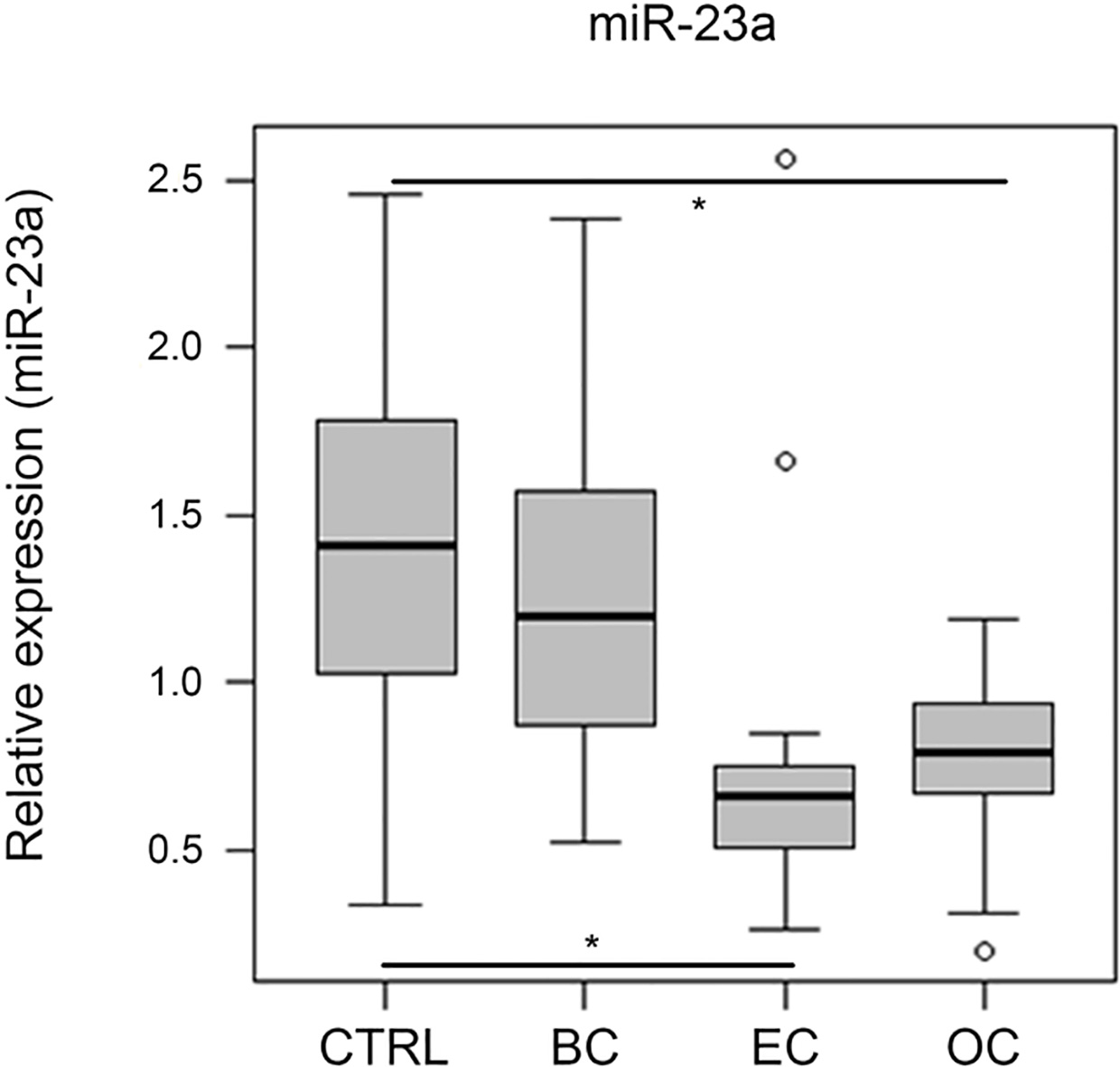
The present study contributes to the knowledge about miRNA expression alterations in serum and urine of BC, EC and OC patients compared to healthy women. Thereby, it underlines the immanent potential of miRNAs to serve as non- and minimally-invasive biomarkers in the field of breast and gynecological cancer detection. However, our findings also accentuate the complexity of biomarker research and unravel methodological limitations in miRNA biomarker screening.
Discovery of potential serum miRNA biomarkers for BC, EC and OC
In serum, 14 miRNAs were discovered by microarray analysis that discriminated between breast and gynecological cancer patients and healthy women (Table 1). Furthermore, 16 BC-specific liquid biopsy biomarker candidates were discovered. While 13 of these miRNAs overlapped with the overall tumor group, three miRNAs (miR-518b, -4719, -6757-3p) were found to be specifically deregulated in BC serum. These three miRNAs have not yet been set into context with BC and could be interesting new players in BC blood screening. Testing for combined EC
Contrariwise, miR-484 was reported to exert oncogenic functions by targeting MAP2 (microtubule-associated protein 2) and consecutively activating ERK1/2 (extracellular-signal regulated kinases) leading to a poor prognosis in glioma patients [114]. It was furthermore found to be upregulated in non-small-cell lung cancer tissue and promoted proliferation and migration in this carcinoma by downregulating APAF-1 (apoptotic protease activating factor 1) [53]. A correlation of high miR-484 expression and unfavorable clinicopathological characteristics was reported for various tumor entities [46, 85]. Interestingly, miR-484 was predicted to play a role in BC as well, with increased expression levels being associated with a worse prognosis [48, 95]. While we did not observe different expression levels between BC patients and CTRL in serum or urine specimen, Zearo et al. found elevated levels of miR-484 in serum of early BC patients compared to healthy women [117].
While a clear involvement of miR-484 in carcinogenic processes is certain – its role being highly context-dependent – this miRNA seems to interact in further pathological conditions (e.g. tuberculosis, carotid plaque development and axial spondyloarthritis) as well [3, 15, 31]. On the other hand, miR-484 was proposed as a housekeeping miRNA by some authors in liquid and tissue samples [40, 82, 87, 91, 127].
A number of miRNAs (miR-451b; -548ao-3p; -3921; -6757-3p; -6859-5p; -7844) identified during the discovery phase in serum samples have not been described in the literature, yet. As their physiological and pathophysiological impact remains unknown, their aberrational expression in serum of gynecological cancer patients suggests a discrete involvement in carcinogenic processes. Various miRNAs identified by this study have been investigated in the context of several malignancies and other diseases before and will raise the question of specificity. MiR-143-3p represents a concrete example. This miRNA seems to play a role in numerous cancer entities [27, 37, 125] and other medical conditions such as acute ischemic stroke [102], amyotrophic lateral sclerosis [107] and autoimmune diseases like multiple sclerosis [86]. A growing body of literature attributes a tumor-suppressive role to miR-143-3p which is downregulated in malignant tissues [13, 27, 37, 125]. Furthermore, it was drastically downregulated in serum of OC and estrogen receptor positive BC patients [20, 47]. These findings are congruent to our own conclusions as miR-143-3p was not detectable in serum of all gynecological cancer patients but an expression was observed in both control pools (Supplementary Table S1). Hence, an association between reduced expression levels and the existence of a malignancy can be expected. However, since miR-143-3p seems to be deregulated in various diseases, it lacks specificity in regard to the distinction of tumor entities. Consequently, miRNAs such as miR-143-3p should be combined with entity-specific biomarkers within the framework of liquid biopsy.
Discovery of potential urinary miRNA biomarkers for BC, EC and OC
Ten miRNAs with general tumor specific characteristics were identified (Table 2). Of these, miR-10b-5p was further analyzed via RT-qPCR due to a reliable detection in urine samples tested, but could not be confirmed as a urine-based non-specific diagnostic marker for BC, EC or OC. There seems to be an overall consensus on miR-10b-5p displaying oncogenic functions in BC by targeting, among others, HOXD10 (homeobox D10), KLF4 (kruppel-like factor 4) and PTEN (phosphatase and tensin homolog) [5, 96]. These functions, however, seem not to be represented in urine. Despite being described as an oncomiR, miR-10b-5p was also found to be downregulated in BC tissue in some studies [16, 55]. Furthermore, clinicopathological parameters could be correlated with miR-10b-5p expression in BC [19, 57, 123]. Focusing on EC, the role of miR-10b-5p is insufficiently understood. On the one hand, it was found downregulated in an invasive form of EC cells and in endometrial EC vs. normal endometrial tissue while no differential expression was observed in plasma [26, 103]. On the other hand, Chen et al. described elevated expression levels in EC tissue compared to CTRL [17]. Our RT-qPCR analysis indicated elevated miR-10b-5p levels in urine samples from EC patients compared to healthy women. In OC, the knowledge about miR-10b-5p function is even more scarce. An oncogenic role is proposed by the targeting of HOXD10 and a subsequent increase of MMP14 (matrix metalloproteinase-14) and RHOC (ras homolog family member C) [75]. While no studies focusing on miR-10b-5p expression in urine of BC, EC or OC patients could be identified for comparison with our own results, literature review revealed few serum-based diagnostic studies. Therein, miR-10b-5p showed potential to discriminate BC patients from CTRLs [16, 66]. To date, miR-10b-5p expression analysis in urine was described to help detect bladder cancer and acute rejection of renal transplants [58, 76]. Based on our own results, we could not confirm miR-10b-5p as a urine-based non-specific diagnostic marker for BC, EC or OC.
While six of the deregulated miRNAs discovered in the BC group overlapped with the overall tumor group comparison, miR-526b-3p remained as an exclusive candidate. It has been reported as a tumor supressor in colon cancer [126] and was found to be elevated in BC cell lines by COX-2 [65]. Downregulation in urine from BC patients as observed in this study suggests tumor-suppressive effects of miRNA-526b-3p.
Furthermore, twelve miRNAs were identified as potential markers for combined detection of EC and OC (Table 2). In analogy to our findings in serum, a subset of these miRNAs has been associated with various diseases and malignancies while another subset was found related to EC and OC by other investigators before [80, 81, 82, 83, 84, 85, 86]. A widely studied example is miR-205-5p that was found to be upregulated in OC tissue [50, 78], to act as an OC invasion promotor [51], to be associated with OC tumor stage, grade and poor survival [50, 78], to trigger EMT in EC cells [42] and to be linked to a poor prognosis in EC [64]. Researchers seem to agree on an oncogenic function of miR-205-5p in EC and report predominantly elevated levels in EC tissue [38, 61, 103]. The same assumption accounts for OC, while few studies also suggest tumor-suppressive traits [18, 52]. Targets of miR-205-5p include tumor suppressors such as FOXO1 (forkhead box O1), PTEN and ZEB1 [50, 61, 78, 121]. In our discovery microarray analysis, miR-205-5p showed reduced expression levels in the EC and OC groups. Consequently, miR-205-5p was chosen for validation. Although this miRNA failed to reach statistical significance in a two-sample
In contrast, literature search did not account for any known functional role for miR-3973; -4426; -5089-5p and -6841 offering novel perspectives to a potential applicability as liquid biomarker candidates for urine-based BC, EC or OC diagnosis.
Identification of stably expressed miRNAs
Regarding the identification of novel miRNA candidates for normalization and quality control, the present study provides promising results. Considering RT-qPCR, stably expressed miRNAs are extremely important to obtain reliable comparative values when relative quantification is applied. Through normalization to so-called housekeeping miRNAs or other “stable genes” as endogenous controls, biological fluctuations are being eliminated. To date, only a limited panel of housekeeping miRNAs is recommended by scientists in the field [68, 87]. There is a shortage of implemented choices of stable normalizers for the especially demanding compartments serum and urine. Working with biological materials, many variables influence sample quality and subsequent results of downstream applications [45, 118]. It is widely accepted that sample quality control constitutes the foundation of any analysis in the field of liquid biopsy [45]. Therefore, expression levels of endogenous controls should be assessed irrespective of the chosen analytical method and there is a great demand for reliable and robust housekeeping candidates. However, both, high percentage of GC bases in many of the here identified miRNAs and repetitive sequences bear methodological challenges, e.g. primer design and effective amplification rates, at current state-of-the-art RT-qPCR. Due to variable miRNA expression patterns in biofluids concerning distinct malignant conditions, an individual set of endogenous controls according to the investigated issue might need assessment and a subsequent validation.
In the conducted discovery phase, miR-23a-3p was identified as a potential endogenous control due to a rather stable expression in patient’s and control’s serum samples. Supporting this finding, miR-23a-3p was proposed for RT-qPCR data normalization by other researchers before, focusing on plasma, tissue and extracellular vesicle analysis [8, 34, 94]. Additionally, a published method for the assessment of hemolysis in serum samples suggests evaluation of miR-451a/-23a-3p ratio by RT-qPCR. Herein, miR-23a-3p serves as normalizer while miR-451a expression levels change according to the degree of hemolysis due to its enrichment in red blood cells [92]. Validation of miR-23a-3p by RT-qPCR in an independent cohort, however, resulted in the rejection of this miRNA as an endogenous control in serum of breast and gynecological cancer patients. In specific, we detected a downregulation of miR-23a-3p expression levels in EC and OC patients compared to CTRLs. In this context, miR-23a-3p expression levels were found decreased in EC vs. normal endometrium and in an invasive form of EC cells, as well [26, 56]. Targeting SMAD3 (mothers against decapentaplegic homolog 3), miR-23a-3p represses EMT and might function as a tumor suppressor in EC [56]. To the best of our knowledge, miR-23a-3p deregulation was not investigated specifically in serum of EC patients, yet. In regard to OC, miR-23a-3p was described to promote cell migration and EMT leading to malignant progression [113, 129]. Beyond these oncogenic functions, its expression in OC tissue was associated with clinicopathological parameters and survival [84, 100]. While our study identified lower miR-23a-3p expression levels in serum of OC patients vs. CTRLs, Pan et al. did not find a difference in plasma expression levels between a group of epithelial OC patients and healthy women. However, they were able to show decreased levels in ovarian cystadenoma vs. epithelial OC patients [82]. Microarray miRNA chip analysis as well as RT-qPCR validation of serum specimen did not detect expression level alterations of miR-23a-3p between BC patients and CTRLs. In the narrower sense, it might be possible to apply miR-23a-3p as an endogenous control for BC serum analysis. Still, literature review revealed an upregulation of miR-23a-3p in BC tissue and Wu et al. even reported elevated serum levels obtained by next generation sequencing [28, 54, 111]. Focusing on its functional role in BC, miR-23a-3p was described to activate the oncogenes FOXM1 (forkhead box protein M1) and HRG (histidine-rich glycoprotein), promoting EMT by targeting E-cadherin and causing tumor migration via SPRY2 (sprouty homolog 2) repression [28, 54, 62]. Generally speaking, miR-23a-3p manifested to be deregulated in several carcinomas, therein either contributing to the malignancy or exerting tumor-suppressive functions [108]. Summarizing, by means of the results of this study, we propose miR-23a-3p as a diagnostic serum-based biomarker for EC and OC while it might serve as an endogenous control in BC serum studies.
Methodological limitations
Microarray chip analysis is a convenient and efficient way to screen for novel biomarkers as it enables simultaneous investigation of vast numbers of targets. In the field of miRNA research we are faced with serveral challenges when applying this method. As described above, some housekeeping candidate and discovery phase miRNAs contain challenging repetitive sequences. Due to a short length of approximately 18–22 nucleotides [36], such sequences can not be fully avoided when designing probes or primers. These demanding sequences could result in artifacts masking results from microarray analysis through cross reactions of probes with similar miRNA types. One could assume this as one of the reasons leading to incongruent results of different studies [12]. Diverse analytical methods such as NGS, RT-qPCR or microarrays can lead to aberrational findings [118]. Even within one performed analytical method a risk of differing results exists due to slight variations in assay protocols and sample handling [12, 79]. These variable parameters create challenges for data comparability and might explain why the results of this work differ partly from those of other studies as outlined above. Hence, standardized methods and normalization procedures are urgently needed.
We acknowledge some limitations of the present study. Results of the discovery phase could have been impacted due to the small patient cohort and the pooling of samples. The OC pool contained one patient diagnosed with a benigne ovarian lesion in retrospect. This fact could have influenced the results from the OC cohort.
Another restriction of this study could be a high percentage of LOD measurements in the microarray chip analysis. Approximately 57% of all meassurements exhibit expression values at the LOD. Noteworthy, miRNAs identified by the performed t-tests characterize as being either present in samples of control probands while being absent or weakly expressed in patients specimen (LOD meassurements) or vice versa. In case of the overall tumor and the BC group, miRNAs discovered by this study could for the most part not be detected both in serum and urine of patients but were identified in samples of the control group. This fact indicates a downregulation of these biomarker candidates in tumor patients and their absence could possibly detect the existence of a carcinoma. Contrariwise, testing for combined EC and OC identified miRNAs that were predominantly expressed in patients specimens but not in healthy women.
Regarding the age distribution of our control group as described in the ‘Materials and methods’ section, the CTRL O 40 pool (women
Conclusions
miRNA liquid biopsy research has proven its potential in clinically challenging aspects such as screening, diagnostics and therapy response prediction of various tumor entities. Supporting this aspect, the present study proposes several liquid biopsy-based biomarker candidates to identify female malignancies while highlighting at the same time the complexibility of this approach. These as well as other miRNAs discovered in this work should be validated in larger cohorts to create a clearer picture of their eligibility as minimally invasive diagnostic tools in breast and gynecological cancer screening.
Footnotes
Conflict of interest
The authors declare no potential conflicts of interest.
