Abstract
BACKGROUND AND OBJECTIVE:
Chemotherapy remains the basis of the treatment of lung cancer, and screening biomarkers with predictive value for chemotherapy is of great interest. The present study focused on status of genes methylation in NSCLC patients receiving pemetrexed- or gemcitabine-based chemotherapy.
PATIENTS AND METHODS:
Promoter methylation of Ras association domain family (RASSF1A) and short stature homeobox 2 (SHOX2) was examined in bronchoalveolar lavage (BAL) from 117 NSCLC patients treated with chemotherapy. Multivariate analysis was used to identify the predictive value of gene methylation. Progression-free survival (PFS) rather than overall survival (OS) was used as the clinical outcome to minimize the impact of chemotherapy on gene methylation.
RESULTS:
The methylation of RASSF1A and SHOX2 was significantly associated with shorter PFS (RASSF1A: HR
CONCLUSION:
Unmethylated RASSF1A is a favorable prognostic indicator for patients receiving pemetrexed doublets. Because of the promoting effect of most chemotherapeutic drugs on gene methylation, unmethylated RASSF1A is not suitable as a predictor for gemcitabine doublets.
Introduction
Despite recent advances in molecular-targeted therapies and immunotherapy, chemotherapy remains the standard treatment for patients with advanced lung cancer [1, 2]. Docetaxel or pemetrexed are viable options for non-small cell lung cancer (NSCLC) patients who have been unsuccessfully treated with, or are not eligible for, targeted therapy or immunotherapy [3]. To further improve the efficacy of chemotherapy, it is necessary to know the molecular mechanisms influencing lung cancer chemosensitivity and to identify factors associated with improved patient survival after chemotherapy [4, 5, 6, 7, 8, 9, 10].
Methylation of cytosine-guanine (CpG) islands in promoter regions generally blocks gene transcription in a wide variety of tumors [11]. Alterations in gene expression induced by aberrant genomic methylation comprise a major epigenetic feature of multiple tumors, including lung cancer [5, 7]. Considering that these methylation abnormalities are associated with various biological characteristics of lung cancer, the detection of such molecular alterations from specific specimens is helpful for lung cancer diagnosis, treatment efficacy prediction and survival prognosis. Evidences showed that promoter methylation of Ras association domain family 1A (RASSF1A) and short stature homeobox 2 (SHOX2) have been identified as diagnostic and prognostic biomarkers for lung cancer [12, 13, 14, 15]. Among those studies, SHOX2 methylation is mostly used as diagnosis marker for lung cancer. Studies about RASSF1A methylation as prognostic factor focused on early stage lung cancer patients receiving surgical resection [16]. Because chemotherapy was administered in the neoadjuvant setting, the predictive role for chemotherapy could not be assessed accurately. Thus, sufficient confirmatory studies are still lacking to assess the reproducibility and stability of these molecular predictors, especially for advanced stage NSCLC.
In our study, modified sodium bisulfite and real-time methylation-specific PCR with fluorescence hybridization probes were used to determine promoter methylation of RASSF1A and SHOX2 in bronchoalveolar lavage (BAL) from 117 NSCLC patients. Real-time methylation-specific PCR has made it possible to detect gene methylation from ultratrace tumor DNA and has the advantage of increasing specificity via an internally binding probe for each gene [17]. Collected through fiberoptic bronchoscopy operation, BAL represents an alternative source of biomarker detection by minimally invasive methods. Due to its vicinity to tumor cells, BAL is superior for DNA detection compared with traditional analytes, such as plasma or sputum [18]. DNA methylation in non-neoplastic cells from plasma or sputum may make it difficult to interpret the final results.
Although evidences have revealed that methylation of the RASSF1A and SHOX2 were correlated with lung cancer diagnosis or prognosis, the predictive role of these genes’ methylation for prognosis of advanced lung cancer patients receiving chemotherapy has hardly been reported. In the present study we focused on the predictive potential of RASSF1A and SHOX2 methylation in NSCLC patients treated with pemetrexed or gemcitabine doublet chemotherapy.
Materials and methods
Ethics statement
This study was approved by the Medical Ethics Committee of Shanghai Pulmonary Hospital. All patients in our study provided written informed consent. This study was conducted in accordance with the Declaration of Helsinki. The patients’ records were anonymized and de-identified prior to analysis.
Study subjects and samples
Between August 2016 and December 2018, a total of 129 NSCLC patients from Shanghai Pulmonary Hospital were enrolled in our cohort analysis. The following criteria were used to distinguish patients with lung cancer from those without lung cancer: (a) diagnosis of lung cancer based on histological analysis of a biopsy specimen or (b) cytologic diagnosis of lung cancer was considered to be a positive reference standard if this diagnosis was supported by other clinical findings (i.e., imaging or bronchoscope). Of the 129 patients, 3 patients received radiotherapy, 2 received targeted therapy, 5 received surgical resection and 2 did not receive any treatment. Finally, only 117 patients who received chemotherapy alone were included in the statistical analysis.
Clinicopathology characteristics were obtained from the Oncology Department or Pathology Department. Radiographic tumor responses were evaluated according to the Response Evaluation Criteria in Solid Tumors: complete response (CR), disappearance of all target lesions; partial response (PR),
The BAL samples of all patients were used not only for cytological diagnosis, but also for detection of gene methylation. All tumor tissues including primary tumors, lymph nodes and distant metastases were used for pathological diagnosis. In order to verify the reliability of gene methylation detection in BAL samples, gene methylation in primary tumor tissues was also detected, and the results were used as a reference. Methylation status of genes was determined at the time of diagnosis.
Cytological and pathological analysis
Bronchial biopsy samples were processed and stained with hematoxylin and eosin. BAL samples were collected during bronchoscopy by aspiration with a flexible bronchoscope from the region of the lesion after injecting 40 mL of saline solution. Precipitation of 10 mL BAL was achieved following centrifugation at 20000 rpm for 10 min. The supernatant was removed after centrifugation, and the cellular precipitate was resuspended with 10–15 mL of Scott’s tap water substitute. Next, the specimen was centrifuged again at 2000 rpm for 10 min, and the cellular precipitate was used to make slides using alcohol fixation and hematoxylin-eosin staining.
DNA Extractions and bisulfite modification of genomic DNA
DNA methylation levels in BAL samples were determined using the Lung-Me
Precipitation of 10 mL BAL centrifuged at 10000 rpm or tumor tissues stored in liquid nitrogen were used for DNA extraction. Genomic DNA was isolated using standard phenol and chloroform extraction. Genomic DNA was treated with sodium bisulfite to modify unmethylated cytosine into uracil. The concentration of each sample was quantified by Qubit 3.0 fluorometer_Invitrogen. A total of 200ng DNA was modified with sodium bisulfite and 50ng of modified DNA was used for PCR analysis. DNA samples were analyzed immediately or rapidly frozen in liquid nitrogen and stored at
Methylation specific real-time fluorescent PCR analysis
PCR amplification targeting sequences modified with sodium bisulfite was detected using TaqMan probes. Sequences of primers and probes are as follows: SHOX2 forward primer (5’-TTGTTTTTGGGTT CGGGTT-3’) and reverse primer (5’-CATAACGTAAA CGCCTATACTCG-3’); RASSF1A forward primer (5’- CGGGGTTCGTTTTGTGGTTTC-3’) and reverse pri- mer (5’-CCGATTAAATCCGTACTTCGC-3’). Probe for RASSF1A (FAM-TCGCGTTTGTTAGCGTTTAA AGT-BHQ) and probe for SHOX2 (VIC-ATCGAACA AACGAAACGAAAATTACC-BHQ). The reactions were performed in a 40
The unmethylated cytosine bases were modified to uracil base by sulfite, and then the uracil bases were converted to thymine bases in the process of PCR amplification, but the methylated cytosine bases remain, so the methylated and unmethylated cytosine bases can be distinguished. Specific primers targeting sequences after bisulfite modification were designed for PCR. Products of PCR amplification were detected by TaqMan probes (85nM). Plasmid containing corresponding sequence of methylated genes was as positive control and purified water as negative control. The
Consistency of methylation detection between BAL and primary tumor
Consistency of methylation detection between BAL and primary tumor
The methodological consistency was analyzed using McNemar’s test and Kappa test. The characteristics of patients with genetic methylation were compared using Chi-square or Fisher exact tests as appropriate. Correlation was analyzed using Spearman’s correlation coefficient. The Cox model was applied to adjust for potential confounders. Survival curves were generated using the Kaplan-Meier method, and the comparison of curves was performed by the log-rank method. Internal validation of the models was achieved by bootstrap resampling (1,000 samples), with all the steps of the model development process being replicated on each bootstrapped sample.
Statistical significance was set at
Results
Evaluation of genes methylation detected in BAL
Of the 117 patients, 51 were diagnosed by cytopathology and the remaining by histopathology. Among those with histopathological biopsies, 22 were from primary tumors, 25 were from lymph nodes and 19 were from distant metastases. All patients were routinely examined by tracheoscopy and alveolar lavage fluid samples were collected.
We tested the promoter methylation pattern of RASSF1A and SHOX2 in 22 primary lung tumors and corresponding matched BAL DNA. The methylation status in tumor tissue and BAL was compared for consistency evaluation. Compared with the methylation detection in tumor specimens, the overall concordance of RASSF1A and SHOX2 promoter methylation were both 86.36% (19/22). The consistency results in BAL for RASSF1A (Kappa
Clinicopathological characteristics of patients and genes methylation
After applying a series of exclusion criteria, including cytology or histological examination and treatment regimens, a total of 117 NSCLC patients were included in statistical analysis. The median age at diagnosis was 62 years (range: 33–82 years). The median follow-up time was 10 months, and the median PFS was 5.2 months (95% CI: 4.819–8.381). Overall, RASSF1A methylation was detected in 60.68% of the patients, and SHOX2 methylation was detected in 59.83% of the patients. Demographic factors were similarly distributed in patients with and without methylation, except for histology and smoking status, as squamous cell carcinoma was more frequently observed in patients with RASSF1A methylation (78.57% vs 21.43%,
Characteristic of 117 NSCLC patients with genes methylation analysis
Characteristic of 117 NSCLC patients with genes methylation analysis
Since age has been reported to influence patterns of DNA methylation [19], we analyzed the correlation of methylation with age using Spearman’s correlation coefficient. However, no significant correlations were found (RASSF1A methylation and age: Rho
Before potential confounders were analyzed via Cox regression analysis, we first tested the assumption of hazard proportion. Test results showed that chemotherapy doublet regimen (HR
Predictors on PFS in cox regression analysis for 117 NSCLC patients
Predictors on PFS in cox regression analysis for 117 NSCLC patients
Predication on PFS by interaction between genes methylation and chemotherapy for 117 NSCLC patients
The Kaplan-Meier curve of PFS against methylation status showed that patients in unmethylated groups had a significantly longer median PFS than patients with methylated RASSF1A (7.1 vs. 4.0 months, HR
PFSs according to genes promoter methylation status. A. RASSF1A promoter methylation status; B. SHOX2 promoter methylation status. RASSF1A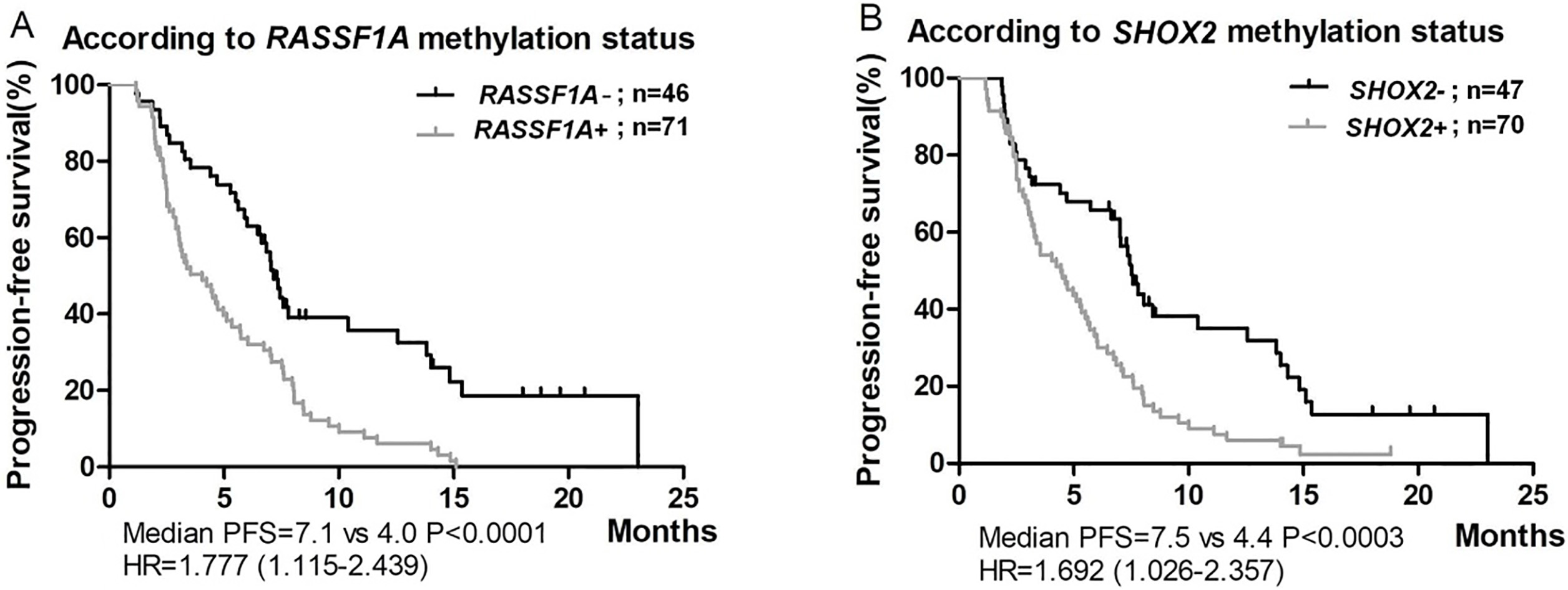
The prognostic value of RASSF1A was further validated with a resampling bootstrap procedure (1,000 replications), in which the statistical analysis (i.e., univariable selection of genes and selection of and adjustment for clinical confounders) were replicated on each bootstrapped sample. RASSF1A methylation was associated with significantly shorter PFS in 70% of the samples, with 0.76 (95% CI: 0.58–0.68) corrected c-indexes, confirming the predictive accuracy of RASSF1A methylation for worse prognosis.
Among the 117 NSCLC patients, 58 patients (49.57%) were treated with chemotherapy of pemetrexed plus carboplatin (or nedaplatin), and 59 patients (50.43%) were treated with gemcitabine plus carboplatin (or nedaplatin). Overall, chemotherapy partial response (PR) was observed in 15 patients (12.82%), whereas stable disease (SD) was found in 49 (41.88%) patients and progressive disease (PD) was found in 53 patients (45.30%). Response to chemotherapy was similarly distributed in patients with methylated and unmethylated status (
PFSs according to chemotherapy regimen and genes methylation status. A. RASSF1A promoter methylation status; B. SHOX2 promoter methylation status. RASSF1A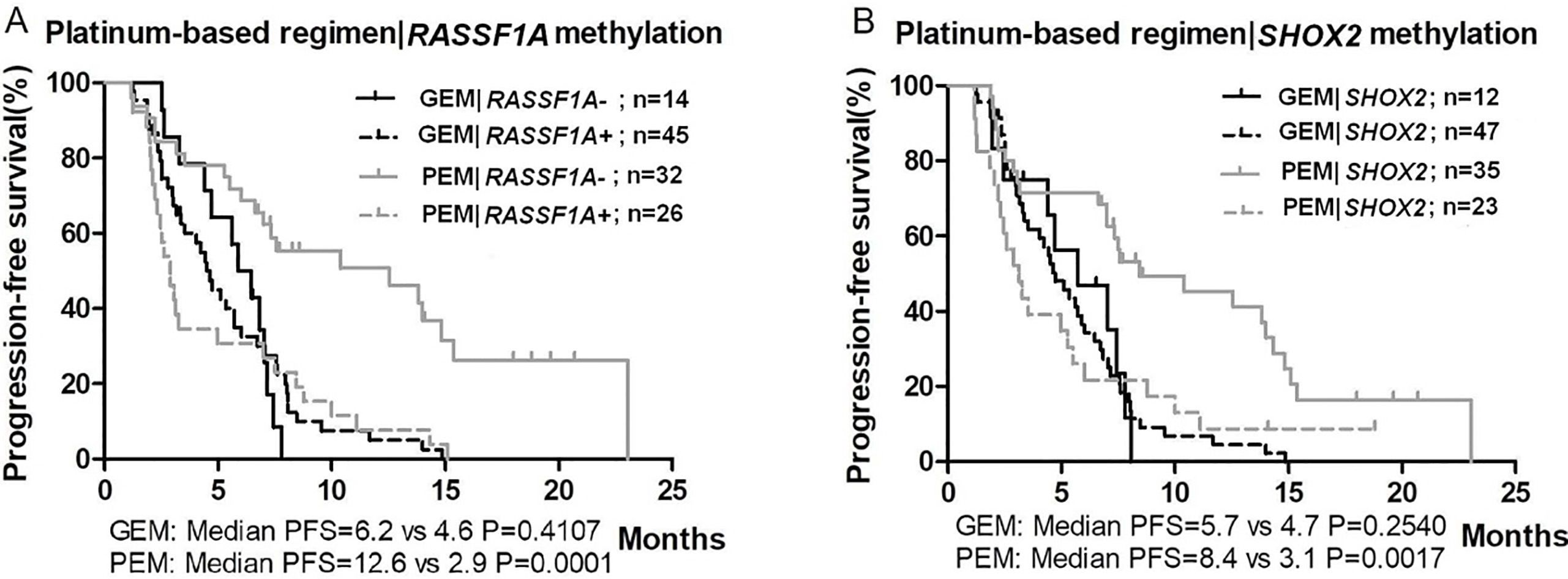
In univariate analysis, the interaction between methylation status and chemotherapy regimens (pemetrexed doublets vs. gemcitabine doublets) on PFS was statistically significant (HR
After adjustment for confounding factors, the statistical interaction between RASSF1A methylation and chemotherapy regimens on PFS remained significant (
Platinum-based chemotherapy has long been the standard treatment for advanced-stage NSCLC. The choice of a suitable regimen for NSCLC is mainly dependent on the treatment benefit based on pathological types, which is far from meeting the requirements of individualized precision treatment. At the same time, the nonnegligible cytotoxic effect and the reduction of quality of life limits the application of chemotherapy. Therefore, analysis of molecular factors for predicting the chemosensitivity and the prognosis of NSCLC patients is considered important for selecting better therapeutic strategies [20, 21, 22]. We detected promoter methylation of RASSF1A and SHOX2 in BAL using real-time methylation-specific PCR and analyzed the correlation with PFS in NSCLC patients treated with chemotherapy. Results in our study showed that RASSF1A methylation was an independent predictor of disease progression. In the pemetrexed doublet group, the PFS of patients with unmethylated RASSF1A was significantly longer than those with methylated RASSF1A, whereas no significant difference was found in the gemcitabine doublet group.
In this study, a commercial technique based on real-time fluorescence quantitative PCR was used to detect abnormal methylation of RASSF1A and SHOX2 in BAL. However, the advantages of quantitative analysis are obvious in specimens with uniform distribution of various cells, for the methylated degree of tumor can be truly reflected by quantitative data detected in such specimens as tumor tissue or serum [23, 24]. The distribution of various cells in BAL is greatly affected by external factors, so the quantitative results of methylation can only reflect whether there is abnormal methylation in tumor, but not the degree of methylation. Therefore, the continuous variables obtained by quantitative PCR in our study were converted into binary variables for qualitative analysis. Although the advantages of quantitative detection can not be realized due to the specimens, the technology based on real-time fluorescent PCR still has such advantages as high sensitivity and specificity, totally closed reaction system, integration of detection and analysis, automation and high throughout [17, 25]. In addition, the technique platform in our study was based on multiplexing real-time fluorescent PCR. RASSF1A, SHOX2 and
Promoters of genes, including RASSF1A and SHOX2, are frequently methylated in lung cancer [11, 12, 13, 14, 15, 16]. In our study, the methylation of these genes in primary tumors was first compared with corresponding BAL samples. The results showed that methylation detection of RASSF1A and SHOX2 between BAL and tumors have good consistency, which indicated that gene methylation detected in BAL better reflect the methylation status of tumors under our laboratory conditions. We then explored the predictive value of RASSF1A and SHOX2 methylation for chemotherapy response and disease progression. Although no significant association was found between gene methylation and overall response to chemotherapy, RASSF1A methylation status was an independent predictor of PFS for patients treated with chemotherapy. Univariate analysis showed that SHOX2 methylation was associated with PFS in lung cancer patients, but SHOX2 methylation was not an independent predictor of PFS after adjustment for confounding factors. In general, PFS will be further assessed only when the responses to treatment were observed in early clinical trials. Thus, an ideal response to chemotherapy was often demolished by poor PFS [26, 27]. However, unlike clinical trials for assessment of new therapies, our study explored the predictive value of gene methylation for standard chemotherapy. As a tumor suppressor gene [28], methylation-induced silence of RASSF1A may promote the tumor progression, but may not necessarily be related to the pharmacological effect of chemotherapeutic drugs.
RASSF1A, a negative effector of RAS, generally functions as a tumor suppressor in cancer through several distinct pathways, including apoptosis, genomic and microtubule stability, and cell cycle regulation [11]. Promoter methylation of RASSF1A is frequently found in a variety of human cancers, including lung cancer [13]. There have been many studies on RASSF1A methylation as a prognosis factor for lung cancer patients [29]. Previous studies have reported that RASSF1A methylation is a poor prognostic factor for lung cancer, and our findings were generally consistent with these reports [30, 31]. A study by Fischer et al. believed that RASSF1A methylation is a favorable prognostic factor [32]. There may be several reasons that contribute to this contradictory conclusion, e.g., the different subgroups of patients enrolled, varieties of the specimens for methylation detecting and that the different endpoint setting may be more likely to be related to the impact of treatment regimens on the final outcome. As an important epigenetic mechanism, DNA methylation is very sensitive to environmental factors, such as radiation, nutrition and temperature [33, 34]. In particularly, the impact of chemotherapeutic drugs on gene methylation cannot be ignored.
One study examined the variations of gene methylation in cancer cells from patients administered with chemotherapy drugs. Among the 14 genes and 800 CpG islands analyzed, nearly half of the genes tested did not show any alterations in DNA methylation level [35]. However, we cannot exclude the possibility that more aberrant methylation occurred when cancer cells were exposed to chemotherapeutic drugs for a long time. It has been reported that promoter methylation of tumor suppressor genes plays an important role in tumor resistance to chemotherapeutic drugs, targeted drugs and radiotherapy [36, 37, 38, 39, 40], which indicates that various therapies, including chemotherapy, do promote gene methylation. Since gene methylation status can be changed by chemotherapy, it is usually difficult to evaluate the prognostic value of gene methylation for chemotherapy. We assumed that a shorter follow-up time may help to minimize the effect of chemotherapeutic drugs on gene methylation [35, 41], though it is not clear whether this effect is time-dependent or dose-dependent. Therefore, PFS rather than overall survival (OS) was used in our study as the primary clinical outcome measure.
The stratified survival curves of PFS in our study showed that the median PFS of patients with methylated genes in the pemetrexed doublet group was similar to that in the gemcitabine doublet group, whereas the median PFS of patients with unmethylated genes in the pemetrexed group was significantly better than that in other patients (Fig. 2). This may be related to the effect of chemotherapeutic drugs on gene methylation. Insufficient intake of folic acid affects the production of S-adenosylmethionine, which is the methyl donor of cytosine methylation [42]. Pemetrexed is a multitargeted antifolate that inhibits several enzymes involved in the folate pathway [43]. In the pemetrexed doublet group, gene methylation induced by chemotherapeutic drugs may be inhibited, so that unmethylated RASSF1A in lung cancer remains unmethylated. However, unmethylated RASSF1A tends to be methylated in the gemcitabine doublet group due to the lack of inhibition on methylation-promoting by chemotherapy. The inference suggests that RASSF1A methylation status is not appropriate as prognostic marker for most chemotherapy [36, 38, 40], except for pemetrexed-based regimen. From the perspective of guiding clinical treatment, methylation status of RASSF1A can be used as an indicator of pemetrexed-based regimen, because those NSCLC patients with unmethylated RASSF1A who received pemetrexed chemotherapy had longer PFS. For NSCLC patients with methylated RASSF1A, whether pemetrexed- or gemcitabine-based chemotherapy has no significant effect on PFS.
It is worth noting that the validity of the hazard proportional assumption is required in the Cox regression model. We tested confounding factors for hazard proportional assumption and found that the number of chemotherapy cycles, the chemotherapy regimen and overall response were all time-dependent covariates. We used the extended Cox regression model to deal with the time-dependent confounding factors, which has received little attention in previous studies on methylation [16, 44]. Due to the small sample size in our study, analysis of data without significant differences should be interpreted with caution. Future studies using larger samples are needed to verify whether SHOX2 methylation is not an independent factor for PFS in patients receiving pemetrexed-based chemotherapy. After all, pemetrexed, as an inhibitor of folic acid metabolism, should not merely target the methylation of specific genes. In further studies, patients treated with more chemotherapy regimens will also be analyzed to expand the applicability of the conclusions.
Our study focused on the predictive value of gene methylation in NSCLC patients receiving pemetrexed or gemcitabine doublet chemotherapy and observed that RASSF1A methylation is an independent predictor of PFS. The median PFS of patients with unmethylated RASSF1A in the pemetrexed doublet group was significantly longer than that of the patients with methylated RASSF1A. No predictive effect of unmethylated RASSF1A was found in gemcitabine doublets group. In summary, unmethylated RASSF1A may predict disease progression for NSCLC receiving pemetrexed doublets though no predictive value was observed for overall response to chemotherapy. BAL is only a non-invasive alternative to lung cancer biopsy in clinical practice, so results in the study still need to be validated with lung cancer tissue as detection specimen. Future study will collect more lung cancer tissues as test samples to assess the predictive value of RASSF1A, SHOX2 and many other genes that were reported frequently methylated in lung cancer.
Footnotes
Acknowledgments
We gratefully acknowledge the contributions of Huangfu for his help in language modification. We also greatly appreciate technicians from Tellgen Corporation for their support of research.
Conflict of interest
The authors declare that they have no competing interest.
