Abstract
Non-small cell lung cancer (NSCLC) , as the most prevalent type of lung carcinoma with high severity, is of urgent necessity to be investigated for novel therapeutic strategies. Long non-coding RNAs (lncRNAs) are notable for their participation in cancer regulation, and lncRNA long intergenic non-protein coding RNA 641 (LINC00641) has been found to have an inhibitory influence on bladder cancer, but its role in NSCLC has not yet been studied. In this research, we launched an investigation into the biological functions and the underlying molecular mechanisms of LINC00641 in NSCLC. At first, downregulation of LINC00641 was identified in NSCLC cells. Functionally, LINC00641 suppressed cell proliferation and induced cell apoptosis in NSCLC, indicating that LINC00641 exerted tumor-suppressive role in NSCLC. Through mechanism investigation, we determined that LINC00641 acted as a competing endogenous RNA (ceRNA) in NSCLC by sponging miR-424-5p to upregulate phospholipid scramblase (PLSCR4) expression. Further rescue assays indicated that miR-424-5p and PLSCR4 could reverse LINC00641-mediated cellular processes. Taken together, it is demonstrated in our study that LINC00641 can function as a tumor suppressor in NSCLC via a ceRNA network.
Keywords
Introduction
Lung cancer, is the most common cause of carcin-oma-related death among both male and female patients all over the world, which has remained to be one of the global public health issues for many years [1]. Non-small cell lung cancer (NSCLC), as the most prevalent occurrence of all lung cancer types, is responsible for roughly 75–80% of all cases of lung carcinoma [1]. Although progresses have been continuously made in recent years, the patients with NSCLC still suffer from high rates of recurrence and metastasis, and the five-year survival rate remains at a low level of less than 15% [2]. For these reasons, it is urgently needed to investigate the carcinoma-related molecular mechanism in NSCLC.
Long non-coding RNAs (lncRNAs) are known as a class of non-coding RNA molecules containing more than 200 nucleotides in length [3]. Numerous investigations have been conducted to identify the functions and regulatory roles of lncRNAs in NSCLC. For instance, LINC00968 was found to be overexpressed in NSCLC tissues and cells, and tumor progression was greatly inhibited by its downregulation [4]. LINC00961 was reported to be an inhibitory factor for cell invasion and metastasis in NSCLC [5]. Another lncRNA, PCAT6, was also studied for its oncogenic activity in NSCLC by binding to the repressor EZH2 to epigenetically suppress LATS2 [6]. Therefore, exploring the underlying biological functions and molecular mechanisms of lncRNAs is of considerable necessity for a better development of novel therapeutic methods for NSCLC. In a recent report, long intergenic non-protein coding RNA 641 (LINC00641) has been revealed to be an inhibitory regulator participating in bladder cancer progression [7]. Nevertheless, the potential roles of LINC00641 contributed to NSCLC have not yet been clarified. Many recent studies have demonstrated that lncRNAs can serve as competing endogenous RNAs (ceRNAs) via sponging specific microRNAs (miRNAs) and facilitating the expression of target messenger RNAs (mRNAs) [8, 9, 10, 11, 12]. Our research herein investigated whether LINC00641 can exert functions in NSCLC by cooperating with miRNAs and mRNAs. In this study, we analyzed the expression level of LINC00641 and its biological functions in NSCLC. Mechanistically, LINC00641-dominant ceRNA network was analyzed by bioinformatics analysis and mechanism investigation. In summary, this study reported the function and mechanism of a ceRNA network in NSCLC.
Materials and methods
Cell culture
Cell lines for the experiments, as NSCLC cell lines A549, H1299, H1650 and H1975, were purchased beforehand from American Type Culture Collection (ATCC, Manassas, VA, USA). Normal human bronchial epithelial cell line BEAS-2B as a group of non-tumor control, together with human embryonic kidney cells HEK-293T, were also purchased from ATCC. The cells for the following assays were cultured in Dulbecco’s modified Eagle medium (DMEM; Gibco, Waltham, MA, USA), which was supplemented with 10% fetal bovine serum (FBS; Gibco) and mixed with 100 U/ml penicillin (Gibco) and 100 U/ml streptomycin (Gibco), as instructed by the supplier. For cell preservation, the cultured cells were stored in a humidified environment with 5% of CO
Cell transfection
The miRNAs involved in the experiments, namely miR-NC and miR-424-5p mimic, were previously purchased from Genepharma (Shanghai, China). The small interfering RNAs (siRNAs) as si-NC, si-LINC00641 #1, #2 and #3, si-PLSCR4 #1 and #2 were likewise bought from Genepharma. The target sequence of LINC00641 for transfection was cloned into pcDNA3.1 vector (Invitrogen, Carlsbad, CA, USA), constructing the recombinant plasmid pcDNA3.1/LINC00641. Cell transfection was performed by treating the cultured cells with the transfection reagent Lipofectamine 2000 (Invitrogen) containing the specific miRNAs, siRNAs or plasmids, according to the protocol from the manufacturer. The cells were harvested after 48 h of transfection for subsequent experiments.
Quantitative real-time PCR (qRT-PCR) analysis
Total RNAs were extracted from the cultured cells by processing the cells with TRIzol regent (Invitrogen) as instructed by the manufacturer. RNA concentration was then measured with NanoDrop 2000 spectrophotometer (Thermo Scientific, Waltham, MA, USA). The extracted RNAs subsequently went through reverse transcription into cDNAs with Reverse Transcription Kit (Takara, Dalian, China) or Taqman Advanced miRNA cDNA Synthesis Kit (Applied Biosystems, Foster City, CA, USA). The levels of LINC00641 and PLSCR4 were measured through qRT-PCR analysis with SYBR
LINC00641 forward, 5
Cell proliferation assay
EdU assay was performed to detect cell proliferation by treating the cells incubated for 48 h with the fluorescent stain 5-Ethynyl-2’-deoxyuridine (EdU) as instructed by the supplier. The treatment lasted for 90 min through the use of the Click-iT
Cell apoptosis assay
JC-1 assay was conducted to detect mitochondrial transmembrane potential as an indication for cell apoptosis level. The assay involved the fluorescent probe JC-1 from the mitochondria staining kit (Sigma-Aldrich, St. Louis, MO, USA). The cells cultured for 48 h were stained with 0.5 mL JC-1 staining working solution for 20 min of incubation. The JC-1 aggregates generating red fluorescence, representing high membrane potential, and the monomers generating green fluorescence, representing low membrane potential were respectively detected through a fluorescence microscope (Olympus), and the relative ratios of red/green fluorescence were calculated accordingly. All the experiments involved were repeated three times.
Wound healing assay
The cells were cultured in 6-well plates until occupying 80% of the area of the well bases. An artificial wound was produced among the cells in each well by scratching the well base with the tip of a 200-
Western blot (WB) analysis
Radioimmunoprecipitation assay (RIPA) buffer (Gibco) supplemented with protease inhibitor was adopted to extract total proteins from the cultured cells, and protein concentrations were quantified with Bicinchoninic Acid Kit (BCA-kit; Takara). Proteins of different molecular weights were separated through the assay of sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE). Polyvinylidene difluoride (PVDF) membranes (Millipore, Burlington, MA, USA) were used for transferring separated proteins after electrophoresis. After coated with skim milk, the membranes were enclosed with primary antibodies assigned for the analysis overnight at 4
RNA immunoprecipitation (RIP) assay
RIP assay was performed according to the instructions from the manufacturer of the Magna RNA-binding protein immunoprecipitation kit (Millipore). The cultured cells were lysed previously to obtain whole-cell extracts in lysis buffer (Gibco) supplemented with protease inhibitor and RNase inhibitor, and the supernatant was gathered for immunoprecipitation. The lysates for the assay were incubated with protein A/G Sepharose beads conjugated with antibodies against Ago2 (8
RNA pulldown assay
RNA pulldown assay was prepared by biotin-label-ing miR-424-5p-WT, miR-424-5p-Mut and negative control respectively with Biotin RNA Labeling Mix (Roche, Basel, Switzerland). The labeled RNAs were previously co-incubated with streptavidin agarose beads (Life Technologies, Gaithersburg, MD, USA) overnight at 4
Luciferase reporter assay
Luciferase reporter assay was prepared by amplifying and cloning the sequences of LINC00641 or PLSCR4 3’ untranslated region (3’-UTR) containing miR-424-5p normal binding site or mutant binding site into pGL3 vectors (Promega, Madison, MI, USA). To perform the assay, specific exogenous miRNAs and/or siRNAs were respectively co-transfected into HEK-293T cells together with the recombinants plasmids. After 48 h of culturing, the relative luciferase activities were measured with Dual-Luciferase Reporter Assay System (Promega). All the experiments involved were repeated three times.
Statistical analysis
All the experimental data were demonstrated as the mean
Results
LINC00641 was downregulated and affected proliferation, apoptosis and migration in NSCLC cells
Searching from online database GEPIA (
LINC00641 was downregulated and affected proliferation, apoptosis and migration in NSCLC cells. (A) qRT-PCR analysis showed that LINC00641 expression level in NSCLC cell lines was significantly lower than in normal cells. (B) qRT-PCR was applied to confirm knockdown and overexpression of LINC00641 in H1299 and A549 cell lines, respectively. (C) EdU assay illustrated the inhibitory effect of LINC00641 on NSCLC cell proliferation. (D) JC-1 assay showed the promotional effect of LINC00641 on NSCLC cell apoptosis. (E) Wound healing assay illustrated that LINC00641 could suppress migration capacity of NSCLC cells. 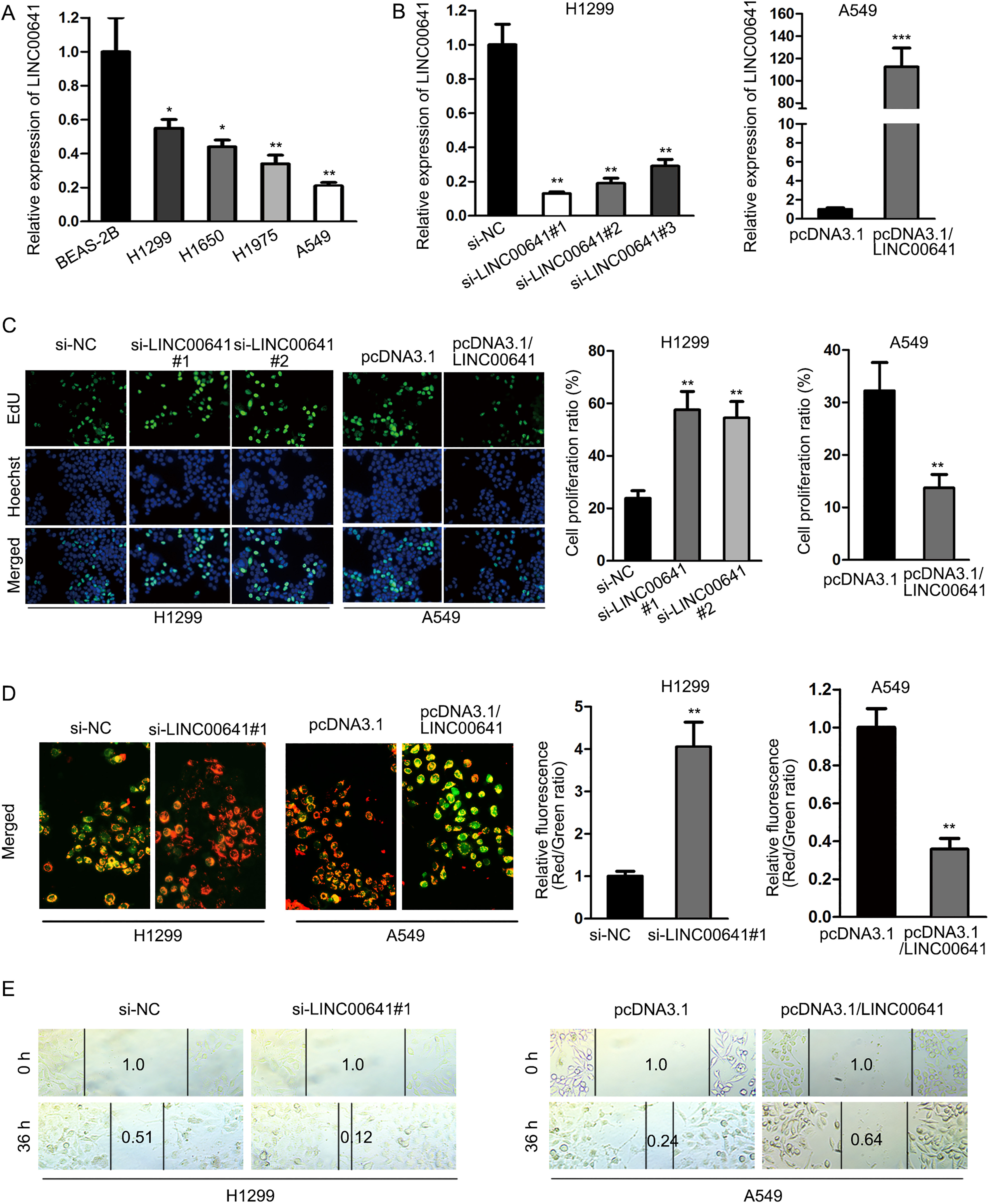
LINC00641 acted as a molecular sponge of miR-424-5p in NSCLC cells. (A) The binding site of LINC00641 and miR-424-5p was identified via starBase prediction. LINC00641 with the wild-type or mutant binding site (red) was correspondingly designed. (B) The expression level of miR-424-5p was upregulated in NSCLC cells compared with normal cells. (C) LINC00641 knockdown increased miR-424-5p expression, while LINC00641 overexpression decreased miR-424-5p expression. (D) Coexistence of LINC00641 and miR-424-5p in RISCs was identified by RIP assay. (E) The direct bond between LINC00641 and miR-424-5p with the wild-type binding site was confirmed by RNA pulldown assay. (F) Luciferase reporter assay confirmed the binding capacity between miR-424-5p and LINC00641 with the wild-type binding site. 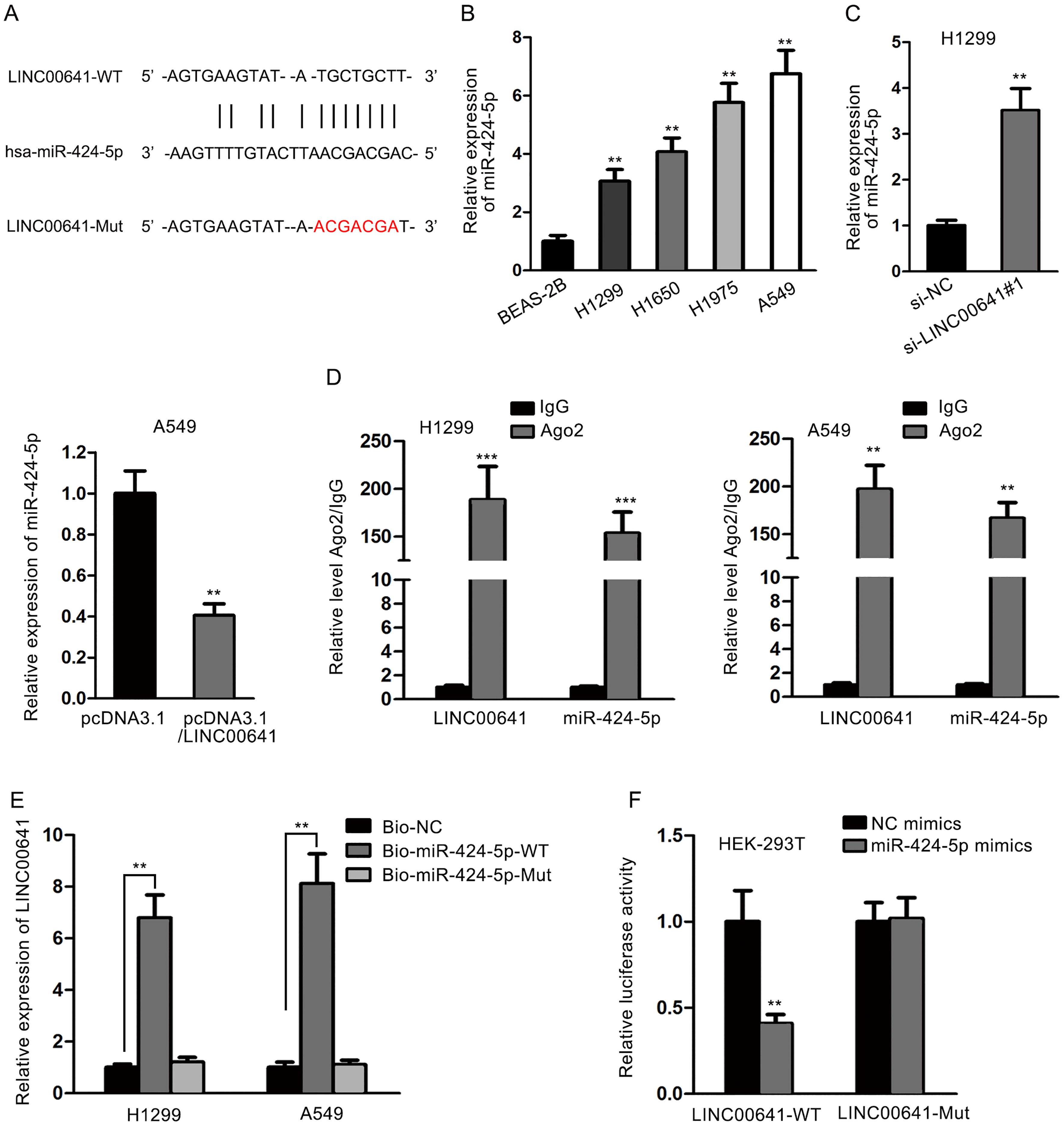
The underlying regulatory mechanism of LINC00 641 in NSCLC cells was further investigated by examining the potential target miRNA. Numerous discoveries have verified that lncRNAs can serve as molecular sponges to regulate expression of specific genes by sponging target miRNAs [13, 14, 15, 16, 17]. MiR-424-5p was selected as the predicted target of LINC00641 via searching potential binding miRNAs in starBase online database (
LINC00641-induced regulations in NSCLC cells were reversed by overexpression of miR-424-5p
To verify that the biological functions of LINC00641 affecting NSCLC cells were implemented via sponging miR-424-5p, we transfected miR-424-5p mimics or the negative control vector into A549 cells with overexpression of LINC00641. We found that the addition of miR-424-5p mimics attenuated the downregulation of miR-424-5p expression level caused by LINC00641 (Fig. 3A). Moreover, the inhibitory effect on cell proliferation, the promotional effect on cell apoptosis and the inhibitory effect on cell migration caused by LINC00641 were respectively rescued by overexpression of miR-424-5p, as revealed in EdU assay (Fig. 3B), JC-1 assay (Fig. 3C) and wound healing assay (Fig. 3D). To make further confirmation, we conducted rescue assays in LINC00641-downregulated H1299 cells. It was found that the expression of miR-424-5p increased by LINC00641 knockdown was rescued by the inhibition of miR-424-5p expression (Fig. 3E). Cell proliferation and migration promoted by si-LINC00641#1 was also recovered after co-transfection with miR-424-5p inhibitor (Fig. 3F–H). However, apoptosis reduced by silenced LINC00641 was recovered after downregulation of miR-424-5p (Fig. 3G). Taken together, miR-424-5p reversed the biological functions of LINC00641 in NSCLC cells.
LINC00641-induced regulations in NSCLC cells were reversed by overexpression of miR-424-5p. (A) qRT-PCR showed a decrease of miR-424-5p expression level in A549 cell line with overexpressed LINC00641, which was upregulated by transfection of miR-424-5p mimics. (B) EdU assay showed that miR-424-5p rescued the inhibitory effect of LINC00641 on cell proliferation. (C) JC-1 assay illustrated that miR-424-5p rescued the promotional effect of LINC00641 on cell apoptosis. (D) miR-424-5p rescued the inhibitory effect of LINC00641 on cell migration, as shown by wound healing assay. (E) Relative expression of miR-424-5p was examined in cells transfected with si-NC, si-LINC00641#1 or co-transfected with si-LINC00641#1+miR-424-5p inhibitors. (F-G) Cell proliferation, apoptosis and migration were measured in indicated H1299 cells. 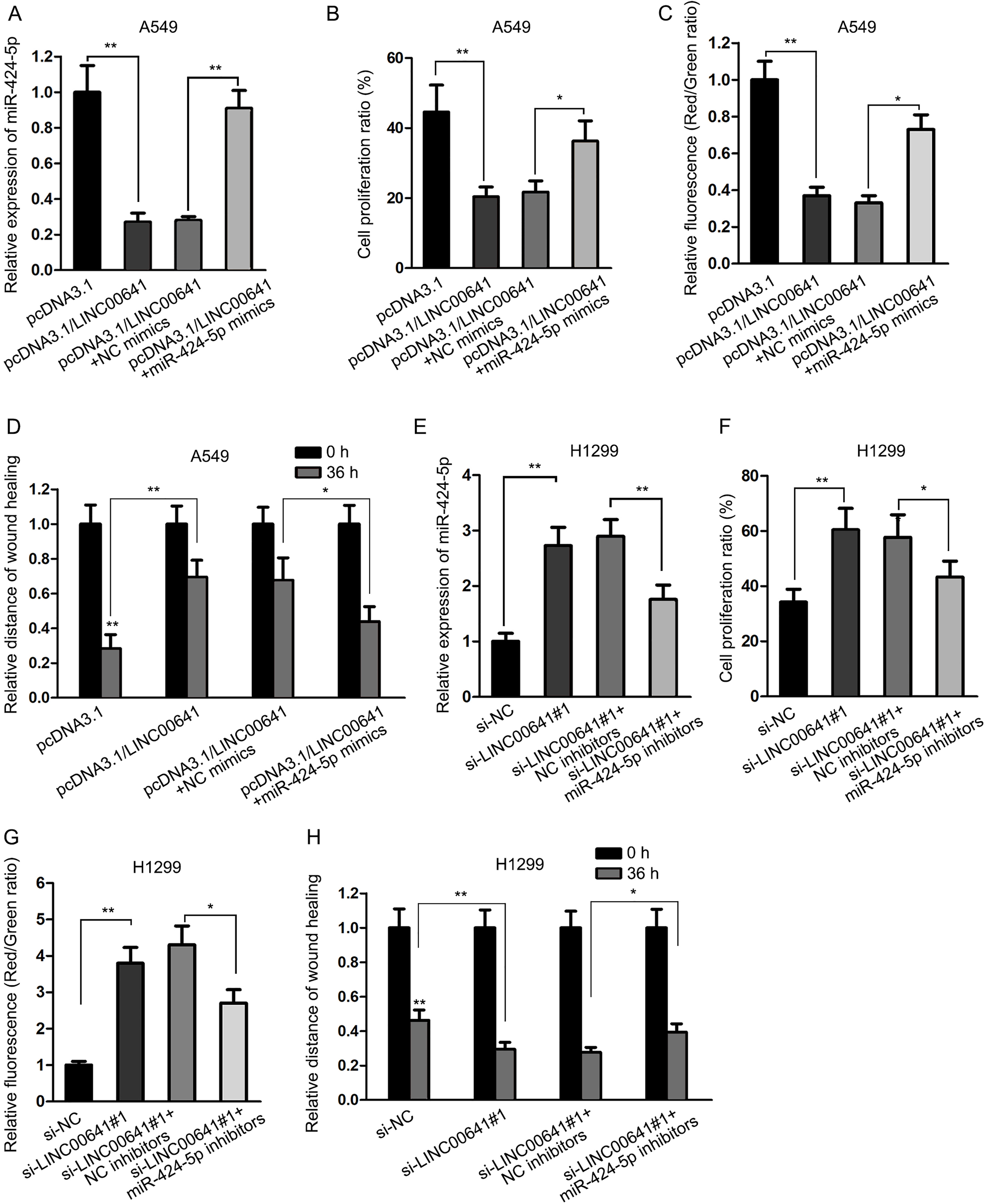
It has been validated by accumulated evidence that miRNAs can regulate the expression of specific genes via binding to the 3’-UTR of the target mRNAs [14, 15, 16, 17], and lncRNAs can protect these mRNAs via acting as ceRNAs to compete for miRNAs. Therefore, the mechanism by which LINC00641 and miR-424-5p functioned in NSCLC cells was further investigated via searching for potential target genes of miR-424-5p. Using the bioinformatics tool TargetScan (
LINC00641 functioned as a ceRNA to regulate PLSCR4 expression by competing for miR-424-5p. (A–B) The expression levels of PLSCR4 mRNA and protein were lowered in NSCLC cells in comparison with normal cells, as illustrated by qRT-PCR and western blot, respectively. (C-D) The expression levels of PLSCR4 mRNA and protein were downregulated and upregulated by LINC00641 knockdown and overexpression, respectively. (E) RIP assay illustrated that LINC00641, miR-424-5p and PLSCR4 coexisted in RISCs. (F) RNA pulldown assay illustrated the direct bond between PLSCR4 mRNA and miR-424-5p with the wild-type binding site. (G) miR-424-5p mimics suppressed the luciferase activity of PLSCR4 3’-UTR wild-type reporter, which was reversed by LINC00641 overexpression, while the luciferase activity of mutant PLSCR4 3’-UTR reporter did not vary significantly. 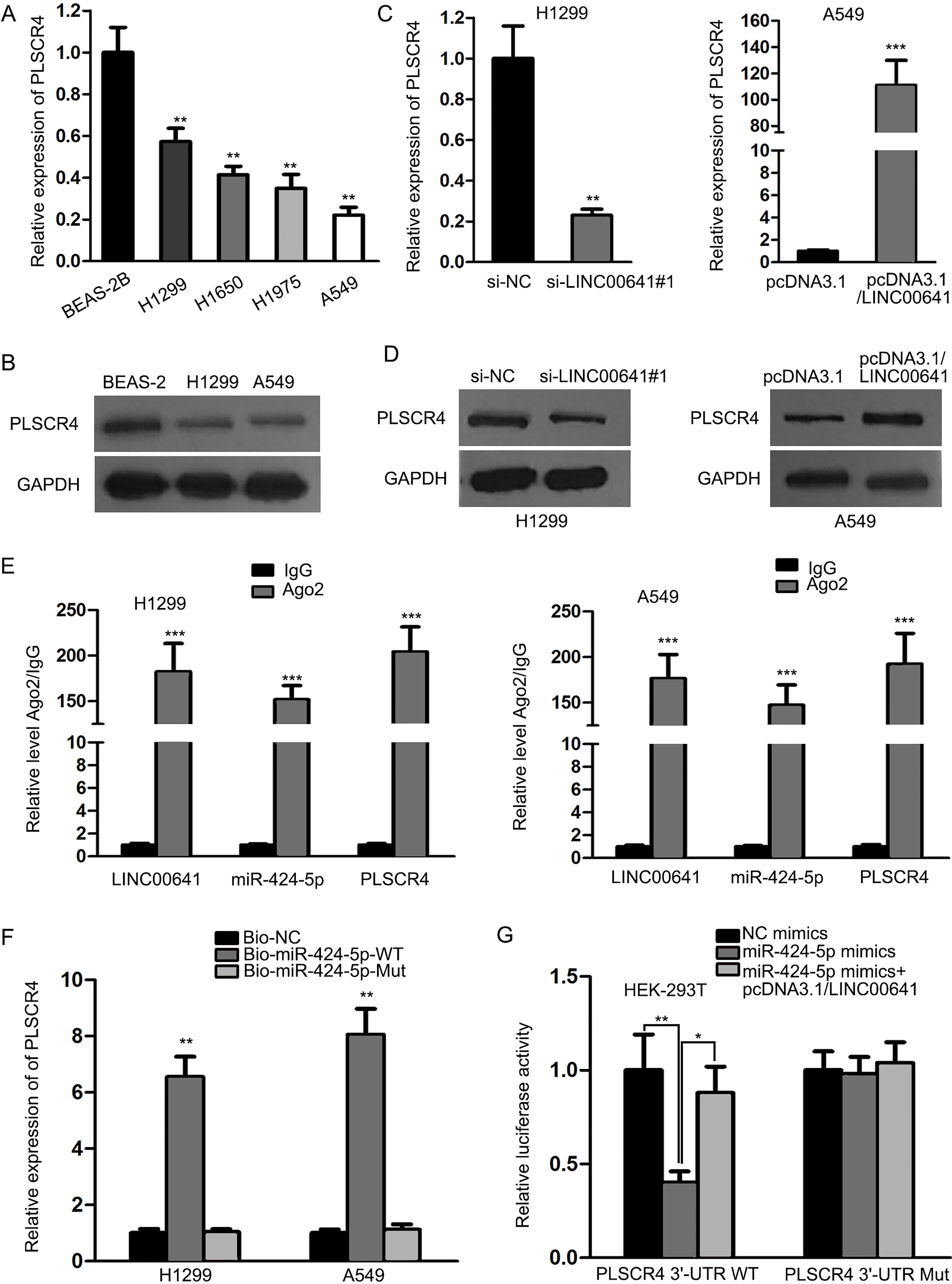
LINC00641-induced regulations in NSCLC cells were reversed by knockdown of PLSCR4. (A) PLSCR4 expression level enhanced by overexpressed LINC00641 was downregulated by transfection of PLSCR4-specific siRNAs. (B) Cell proliferation inhibited by LINC00641 was reversed by PLSCR4 knockdown. (C) Cell apoptosis promoted by LINC00641 was reversed by PLSCR4 knockdown. (D) Transfection of si-PLSCR4 rescued the inhibitory effect of LINC00641 on cell migration. 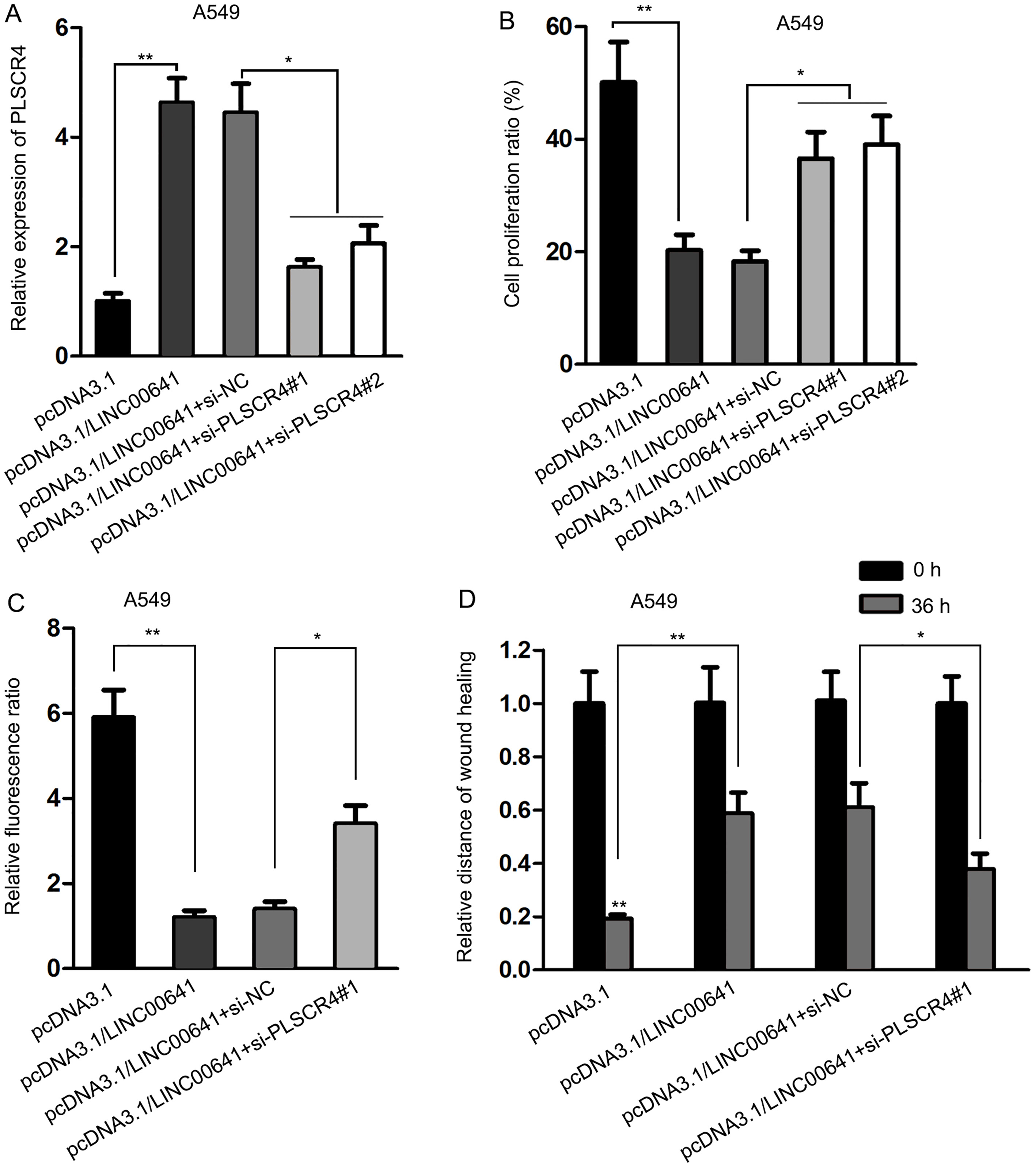
To confirm that the functions of LINC00641 were exerted through PLSCR4, siRNAs of PLSCR4 or the negative control were transfected into LINC00641-overexpressed A549 cells. The results demonstrated that the expression level of PLSCR4 promoted by LINC00641 was downregulated by silenced PLSCR4 (Fig. 5A). And as a result of PLSCR4 knockdown, the results of EdU assay and JC-1 assay illustrated that the level of cell proliferation reduced by LINC00641 was reversed (Fig. 5B), and cell apoptosis enhanced by LINC00641 was attenuated (Fig. 5C). Similarly, as shown in wound healing assay, the inhibitory effect of LINC00641 on cell migration was also rescued by si-PLSCR4 (Fig. 5D). To sum up, PLSCR4, as a target gene of miR-424-5p, could reversely regulate the biological functions of LINC00641 in NSCLC cells.
Discussion
It has been widely known that abnormal expression of cancer-related gene is of prevalent existence within tumors. LncRNAs are essential regulators in tumorigenesis and tumor progression [18, 19, 20]. In our present study, we determined that LINC00614 is downregulated in NSCLC cell lines. Previous studies showed that downregulated lncRNAs is correlated with cellular processes, including proliferation, apoptosis and migration [21, 22, 23, 24]. In this study, data of functional assays revealed that LINC00641 expression was negatively correlated with NSCLC cell growth, indicating the tumor suppressive role of LINC00641 in NSCLC.
Mechanistically, lncRNAs can regulate their downstream genes at transcriptional or post-transcriptional level [25, 26, 27, 28, 29]. It has been widely reported that lncRNAs can exert their functions via regulating miRNA/mRNA axis [30, 31]. In this regard, we applied bioinformatics analysis to search out miRNAs that can interact with LINC00641. Due to highest binding score, miR-424-5p was chosen to do further study. The expression of miR-424-5p was negatively correlated with LINC00641 in NSCLC cells. Mechanistically, we determined that miR-424-5p could interact with LINC00641 in NSCLC cells. Through rescue assays, we validated that miR-424-5p reversed LINC00641-medicted cell proliferation and apoptosis. Thus, we confirmed the regulatory relationship between LINC00641 and miR-424-5p in NSCLC cellular processes.
Furthermore, the target mRNA of miR-424-5p was also searched and analyzed. PLSCR4, an oncogenic protein, which was upregulated lung cancer samples in GEPIA database. In our current study, PLSCR4 was found to be interact with miR-424-5p. Moreover, PLSCR4 was negatively regulated by miR-424-5p but was positively regulated by LINC00641. Hence, we demonstrated that LINC00641 acted as a ceRNA in NSCLC by sponging miR-424-5p to upregulate PLSCR4. Similarly, PLSCR4 was involved in LINC00 641-mediated NSCLC cell proliferation and apoptosis. In conclusion, our research indicated that LINC00641 acted as a ceRNA to suppress NSCLC cell proliferation and migration by cooperating with miR-424-5p and PLSCR4. The tumor-suppressing function of LINC00641 may be beneficial for improving the treatment of NSCLC. Lacking of clinical analysis and animal study is the deficient of our present study. Thus, we may analyze the clinical significance of LINC00641 and perform in vivo experiment in our future study.
Footnotes
Acknowledgments
Thank you for all involved in the study.
Conflict of interest
The authors declare that there is no competing interest in this article.
Supplementary data
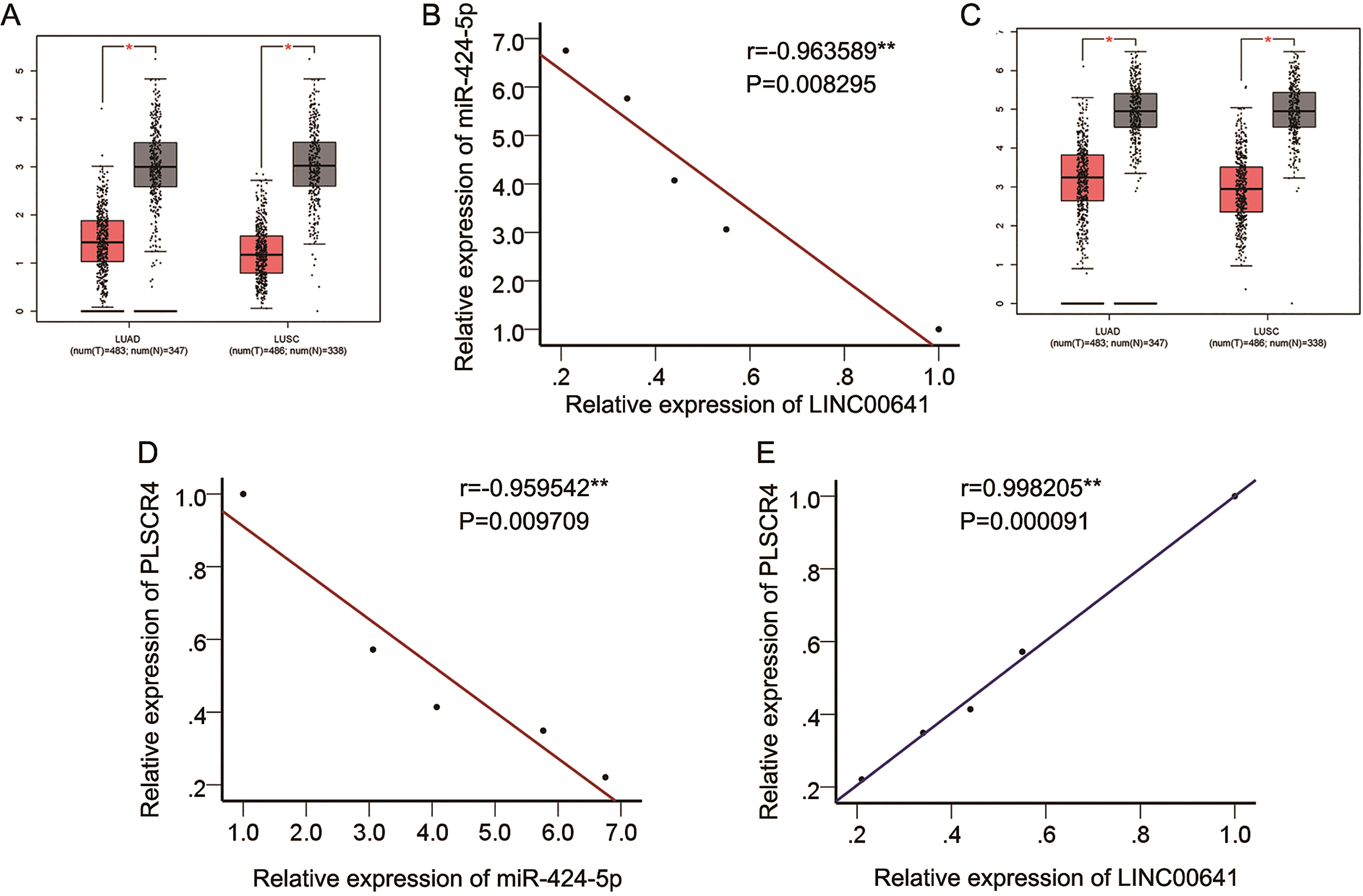
(A) LINC00641 expression in lung cancer and adjacent normal samples in GEPIA database. (B) Expression correlation between LINC00641 and miR-424-5p in NSCLC cells. (C) PLSCR4 expression in lung cancer and adjacent normal samples was obtained from GEPIA database. (D-E) Expression association between LINC00641 and miR-424-5p as well as between LINC00641 and PLSCR4 in NSCLC cell lines.