Abstract
BACKGROUND:
Gefitinib-resistance in lung cancers has become an intractable clinical problem. However, the mechanisms underlying this resistance are not fully understood.
OBJECTIVE:
Present study aims to investigate the roles and underlying mechanism of miR-153 in modulating gefitinib resistance in lung cancers.
METHODS:
In the present study, genes expression of miR-153, MDR-1 and ABCE1 were detected by qRT-PCR and western blot. The cell viability was examined by MTT assays. The regulation of miR-153 on ABCE1 was examined by luciferase reporter gene assays. The interaction of miR-153 and ABCE1 was detected by gene over-expression and siRNA interference technology.
RESULTS:
The mRNA level of miR-153 was significantly down-regulated in gefitinib-resistance (GR) tissues and HCC827 cells, while the protein level of ABCE1 was up-regulated in GR tissues and HCC827 cells. Besides, miR-153 over-expression evidently increased miR-153 level and suppressed cell viability and multi drug resistance gene (MDR-1) expression in HCC827/Gef cells, while silence of miR-153 caused adverse alterations in HCC827 cells. Luciferase reporter assay results showed that miR-153 directly targeted ABCE1. Further studies showed that ABCE1 over-expression improved the expression of ABCE1 and MDR-1 and increased cell viability in HCC827/Gef cells, while ABCE1 silencing resulted in contrary trends in HCC827 cells. What’s more, miR-153 over-expression inhibited tumorigenesis and ABCE1 expression, while increased miR-153 level in tumor tissues.
CONCLUSIONS:
MiR-153 regulates gefitinib resistance by modulating expression of ABCE1 in lung cancers. Our findings may provide a worthwhile therapeutic target to reverse gefitinib resistance in lung cancers in the future.
Introduction
Evolving clinical data implies that lung cancer has been the major reason of cancer deaths in the worldwide, and approximately 16.7% cancer deaths were caused by lung cancer [7, 16]. Thus, many researchers devoted to elucidate the pathogenesis and did their best to explore the effective therapies of it. Because of those efforts, gefitinib was observed and widely used in treatment of lung cancer as a specific inhibitor of epidermal growth factor receptor tyrosine kinase [2]. However, acquired gefitinib-resistance in lung cancers has become an intractable clinical problem along with the extensive application of gefinitib [21].
ATP-binding cassette E1 (ABCE1) belongs to ATP-binding cassette transporters family and implicates in viral infection, cell proliferation, anti-apoptosis, tumor growth and other multiple biological processes [3, 12]. ABCE1 has been considered as a candidate tumor suppressor protein, and takes part in the development of various cancers, including breast cancer, lung cancer, and esophageal carcinoma [8, 9, 10]. There were lots of studies indicated that ABCE1 was related to drug resistance in lung cancers. For instances, the down-regulation of ABCE1 influenced the sensitivity against chemotherapeutic drugs in non-small cell lung cancer (NSCLC) A549 cells [11]; knockdown of ABCE1 could inhibit the cells proliferation and invasiveness in NSCLC cells [8, 23]. Previous studies also showed the over-expression of ABCE1 in lung carcinoma tissues, and highlighted that ABCE1 could promote the viability and migration of lung cancer cells [17, 18]. Taken together, those studies revealed the crucial roles of ABCE1 in proliferation and migration of lung cancer cells and drugs resistance. Nevertheless, it was still largely unknown how ABCE1 affects the sensitivity of gefitinib in lung cancers.
It is well known that miRNAs has been recognized as a novel regulator in gene expression. Generally, miRNAs is short noncoding sequence and inversely modulates gene expression by binding to specific sites at 3’ UTR of target genes [5]. According to previous document, miRNAs plays important roles in the development of many cancers [1]. Furthermore, it has been confirmed that considerable miRNAs were associated with gefitinib-resistance in lung cancers, such as miR-30b-c, miR-221/222, miR-374a, miR-153 and miR-548b [6, 19]. Particularly, recent studies implied that miR-153 was low expressed in NSCLC tissues and cells [4], and could alleviate radiation tolerance in glioma [22]. Meanwhile, miR-153 suppressed the proliferation and migration of lung cancer cells [15], and exhibited the activity against tumor in lung cancers by affecting on AKT [24]. Interestingly, bioinformatics software analysis predicted that there was a potential miR-153 binding site at the 3’UTR of ABCE1 gene, which suggested the intimate relationship between miR-153 and ABCE1. Thus, we speculated that miR-153 may influence gefitinib resistance by regulating ABCE1 expression in lung cancers.
In order to explore the functions and underlying mechanism of miR-153 in regulating the sensitivity of lung cancers to gefitinib, we detected genes expression of miR-153, multi drug resistance gene (MDR-1) and ABCE1 by qRT-PCR and western blot, examined the viability of lung cancer cells by MTT assays, evaluated the regulation of miR-153 on ABCE1 by luciferase reporter gene assays, and assessed manipulation of miR-153 and ABCE1 by siRNA interference technology.
Primers used in the real-time quantitative PCR
Primers used in the real-time quantitative PCR
Human lung cancer tissues
Human lung cancer tissues were obtained from 40 clinical NSCLC patients. Of those tissues, twenty of them were sensitive to gefitinib, and the others were resistant to gefitinib. Hence, lung cancer tissues were divided into gefitinib-sensitive (GS) and gefitinib-resistance (GR) groups. We have informed the the purpose of this study to all patients involved and have obtained their consents. Our study received approval (NO. 2018009) from the institutional review board of the Affiliated Hospital of Inner Mongolia Medical University.
Animals
Nude mice (5 weeks old, 20.0
Cell lines and cell culture
Human NSCLC cell line HCC827 was obtained from American Type Culture Collection and cultured in RPMI-1640 medium (Invitrogen, USA) at 37
RNA isolation and real-time quantitative PCR
Total RNA was extracted from lung cancer tissues or NSCLC cells with TRIzol Reagent (Invitrogen, USA). RNase-free DNase I (Promega, USA) was used to wipe off genomic DNA contamination. The concentration and quality of RNA samples were determined by a spectrophotometer and 1% agarose gels electrophoresis, respectively. RNA (500 ng) was used to synthesize cDNA. Primer of miR-153 used in quantitative real-time PCR (qRT-PCR) was purchased from Guangzhou RiboBio (Guangdong, China). The information of other primers used in qRT-PCR was shown in the Table 1. The relative expression of ABCE1 and miR-153 were calculated by 2
Western blot analysis
Lung tissues or NSCLC cells were used to extract total protein. Tissues or cells were lysed by RIPA lysis buffer (CWBIO, China) on ice. The supernatant was collected to obtain total protein after centrifugation. BCA kits (CWBIO, China) were used to detect the concentration of protein. Protein was separated by 10% SDS-PAGE, and then was transferred to polyvinylidene fluoride membrane (PVDE). Further, the proteins were incubated with primary antibodies: anti-ABCE1 (1/1000, Abcam), anti-MDR-1 (1:500, Boster), or anti-
Luciferase reporter assay
HCC827 cells were co-transfected ABCE1-UTR-pISo (wild-type ABCE1) and Mu-ABCE1-UTR-pIS0 (mutant ABCE1) with miR-153 mimic and miR-153 inhibitor using Lipofectamine 2000 (Invitrogen, USA). Subsequently, Dual-Luciferase Reporter Assay System (Promega, USA) was used to detect the luciferase activity.
Cell transfection
NSCLC cells were cultured in 96-well plates with a concentration of 2
Expression profiles of miR-153 and ABCE1 in lung cancer tissues (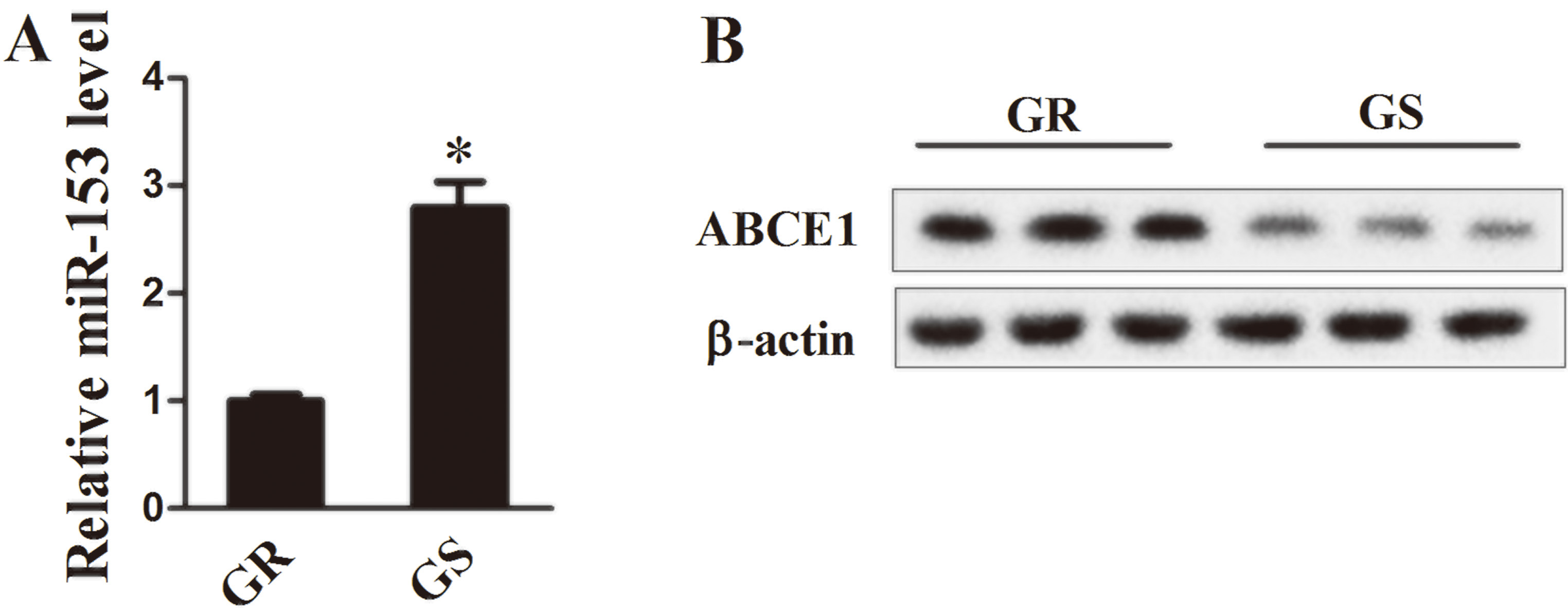
MiR-153 mimic, miR-153 mimic and their respective control were transfected into NSCLC cells. After then, cells were exposed to gefinitib at concentrations of 0, 0.1, 1, 10 and 20
Xenograft models
The nude mice were divided into 2 groups (
Statistical analysis
Data was analyzed using SPSS 20.0. Student’s
Results
Expression of miR-153 and ABCE1 in clinical lung cancer tissues
We firstly detected the expression of miR-153 and ABCE1 in patient’s lung cancer tissues. Compared with GR tissues, the mRNA level of miR-153 was remarkably down-regulated in GR tissues (Fig. 1A,
Expression of miR-153 and ABCE1 in NSCLC cells
As well, we evaluated the expression of miR-153 and ABCE1 in NSCLC cells. In the HCC827/Gef cells, transcript expression of miR-153 was significantly down-regulated (Fig. 2A,
Gene expression of miR-153 and ABCE1 in NSCLC cells. A: The mRNA expression of miR-153 measured by qRT-PCR (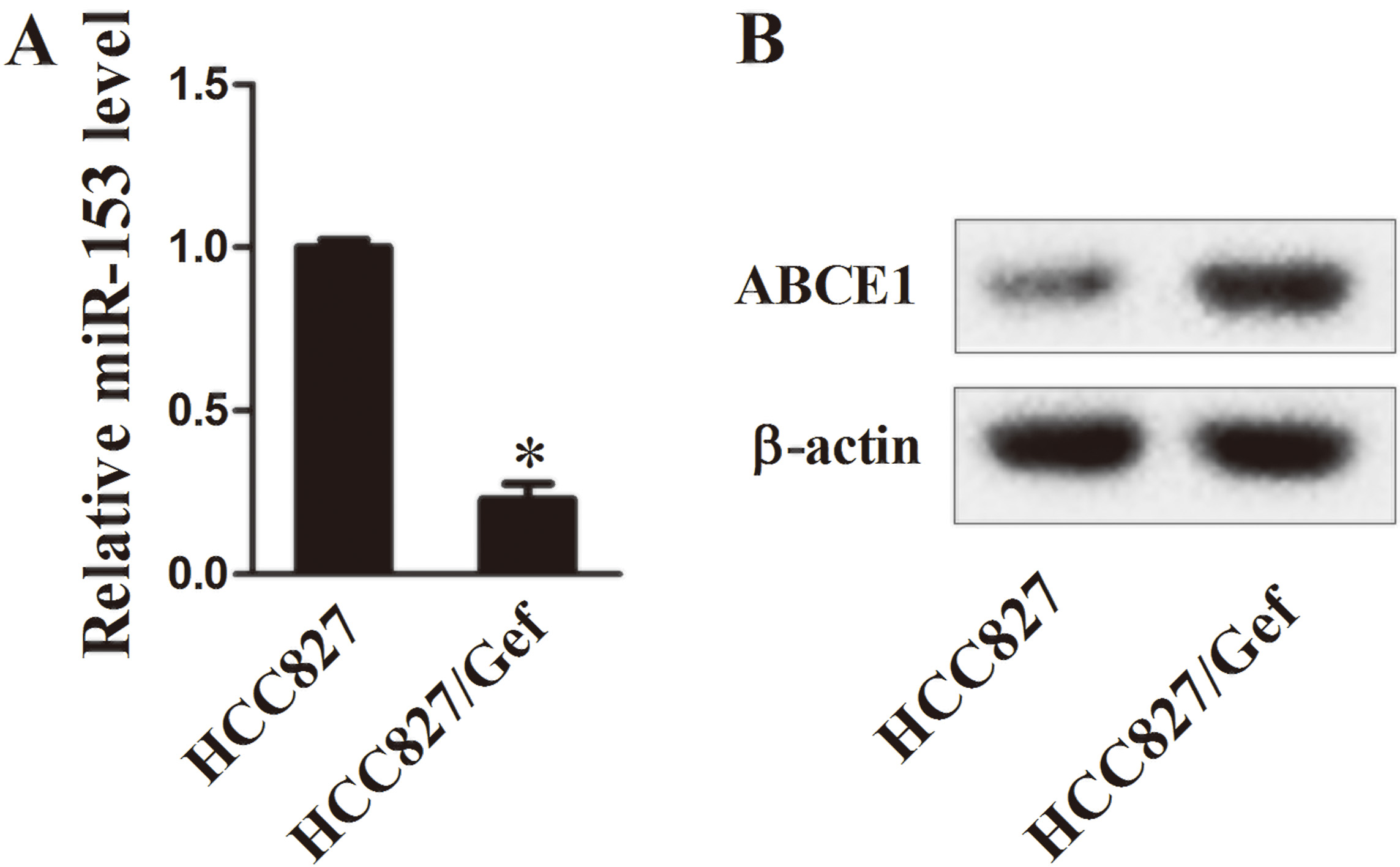
To verify the effects of miR-153 on gefitinib resistance in lung cancers, miR-153 mimic and miR-153 inhibitor were transfected to NSCLC cells. As is shown in Fig. 3, the transcript expression of miR-153 was significantly improved in HCC827/Gef cells by miR-153 mimic transfection (Fig. 3A,
Effects of miR-153 on gefitinib resistance in vitro. A: The mRNA expression of miR-153 in HCC827/Gef cells measured by qRT-PCR. B: Cell viability of HCC827/Gef cells following (0, 0.1, 1, 10, 20) 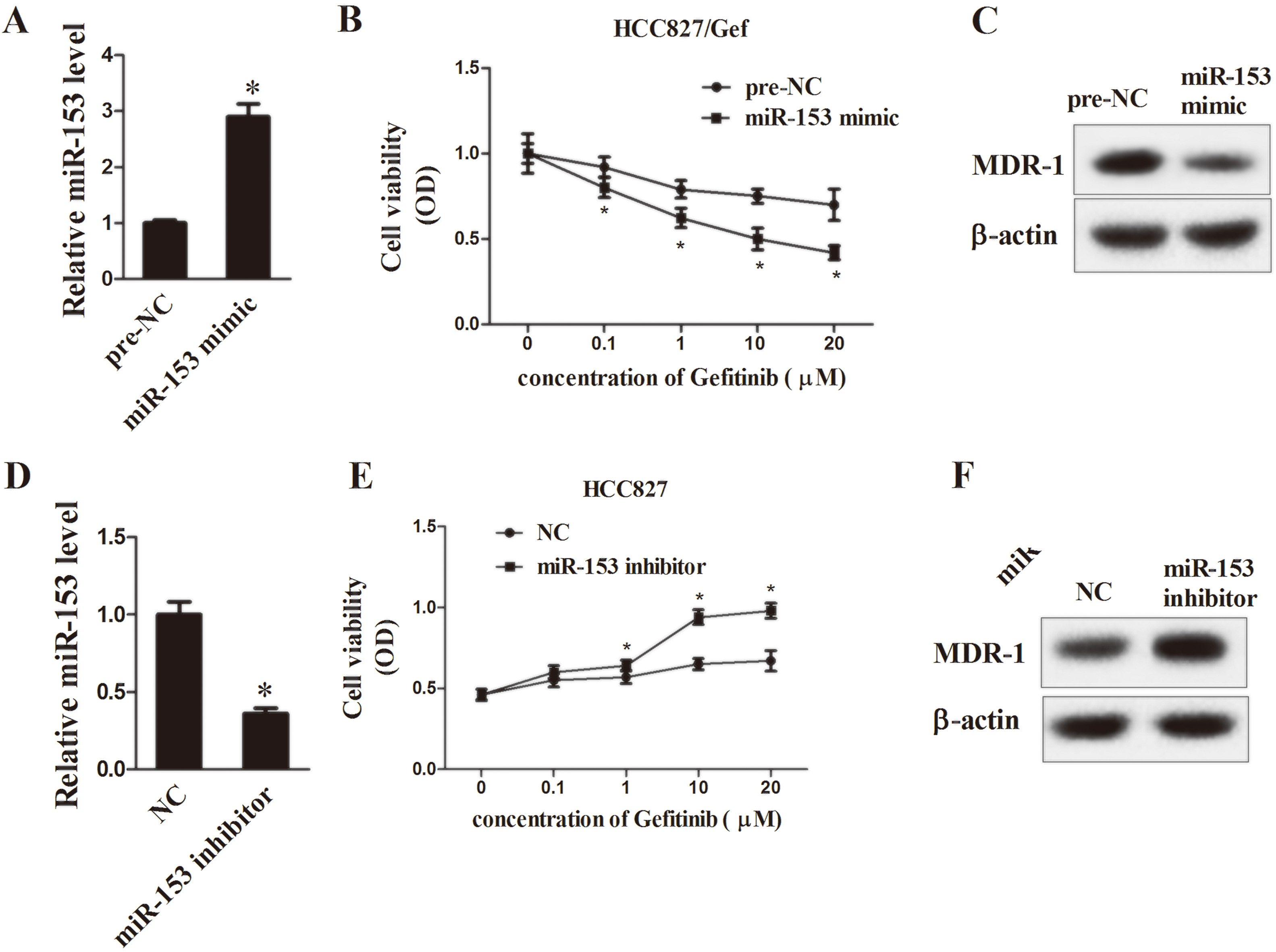
We searched for mRNA targets of miR-153 using bioinformatics softwares. As shown in Fig. 4A, we observed a potential miR-153 binding site at 3’ UTR of ABCE1. To investigate whether miR-153 directly interact with ABCE1, HCC827 cells were co-transfected with ABCE1-UTR-pISo and Mu-ABCE1-UTR-pIS0, miR-153 mimic, miR-153 inhibitor or controls. The luciferase activity of ABCE1-WT was evidently increased in miR-153 inhibitor group (Fig. 4B,
MiR-153 targets ABCE1. A: The predicted binding sites of mRNA targets with miR-153 and sequences of wild-type (WT) and mutant (MUT) ABCE1 were shown. B: Effects of co-transfection of miR-153 inhibitor with WT or MUT ABCE1 3’UTR plasmid vector constructs on luciferase reporter assays and expression of ABCE1. C: Effects of co-transfection of miR-153 mimic with WT or MUT ABCE1 3’UTR pGL3 constructs on luciferase reporter assays and expression of ABCE1. 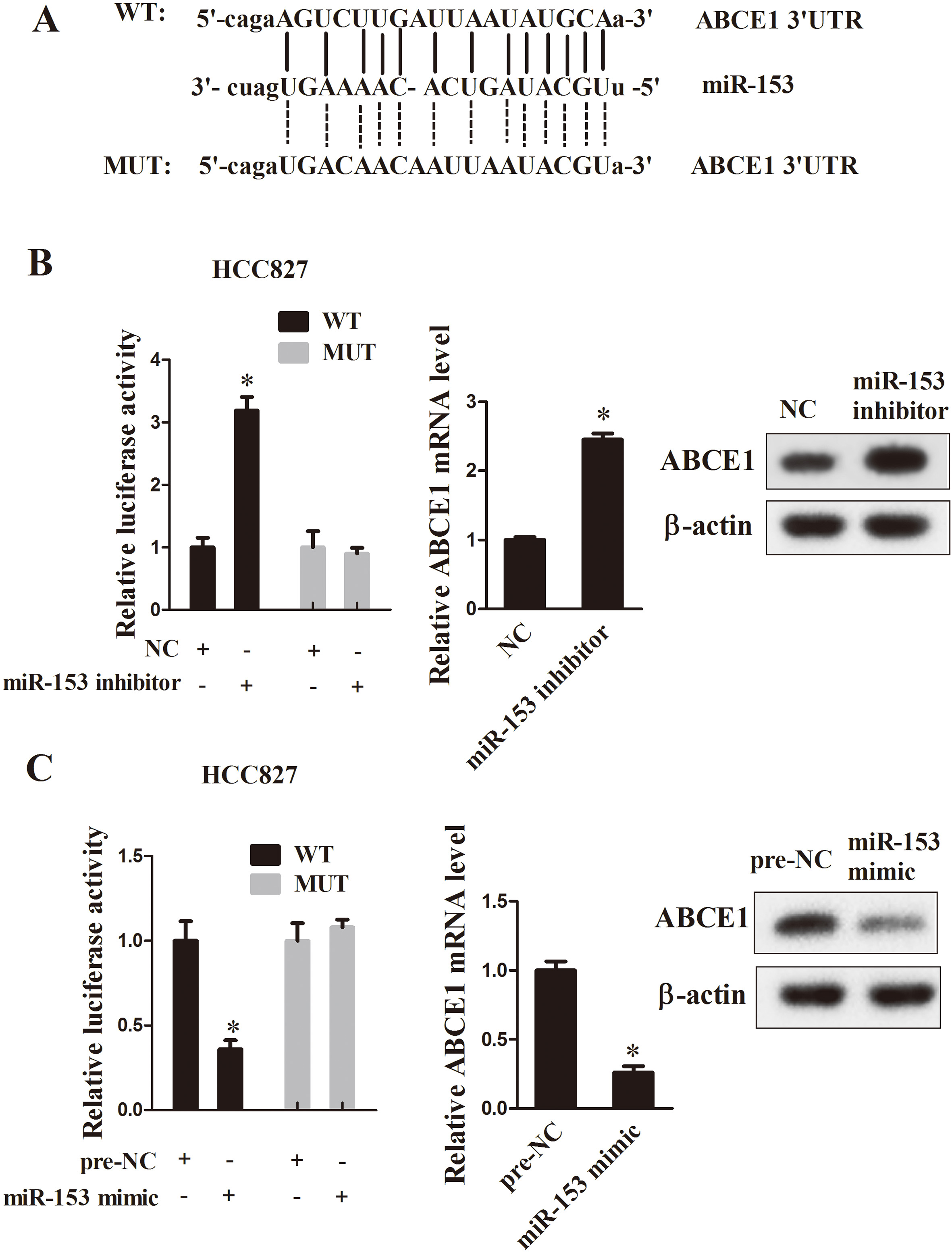
For HCC827/Gef cells, miR-153 over-expression induced increase of ABCE1 protein level, while pcDNA-ABCE1 transfection abolished the effect (Fig. 5A). The cell viability was remarkably inhibited by miR-153 over-expression following gefitinib stimulation, while the suppressed effect was abrogated by pcDNA-ABCE1 (Fig. 5B,
For HCC827 cells, the protein level of ABCE1 was up-regulated by miR-153 silencing, while ABCE1 interference removed the trend (Fig. 5D). As well, miR-153 silence notably improved the cell viability after gefitinib treated, and the impact was eliminated by interfering ABCE1 (Fig. 5E,
MiR-153 regulates of gefitinib-resistant in lung cancer cells by modulating expression of ABCE1. NSCLC Cells were transfected pre-NC, miR-153 mimic, miR-153 mimic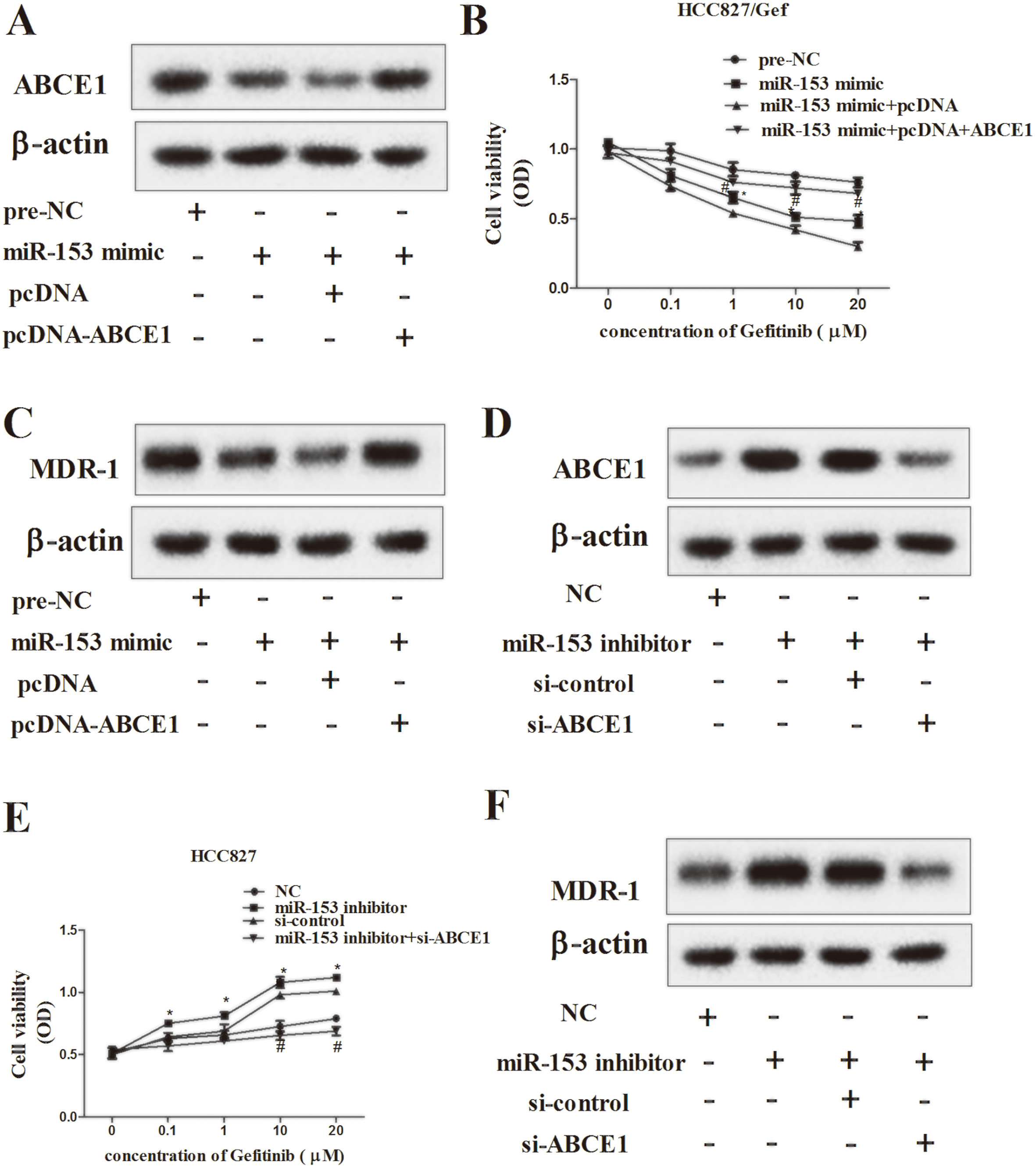
Finally, we assessed whether miR-153 can modulate gefitinib-resistant in lung cancers in vivo. We stably over-expressed miR-153 in gefitinib-resistant HCC827/Gef cells. Then HCC827/Gef-miR-153-mimic or HCC827/Gef-pre-NC cells were subcutaneously injected into nude mice. Compared to HCC827/Gef-pre-NC mice, the tumor volume was significantly decreased in HCC827/Gef-miR-153-mimic mice after 2 weeks injection (Fig. 6A,
MiR-153 regulates gefitinib resistance in lung cancer cells in vivo by modulating expression of ABCE1 (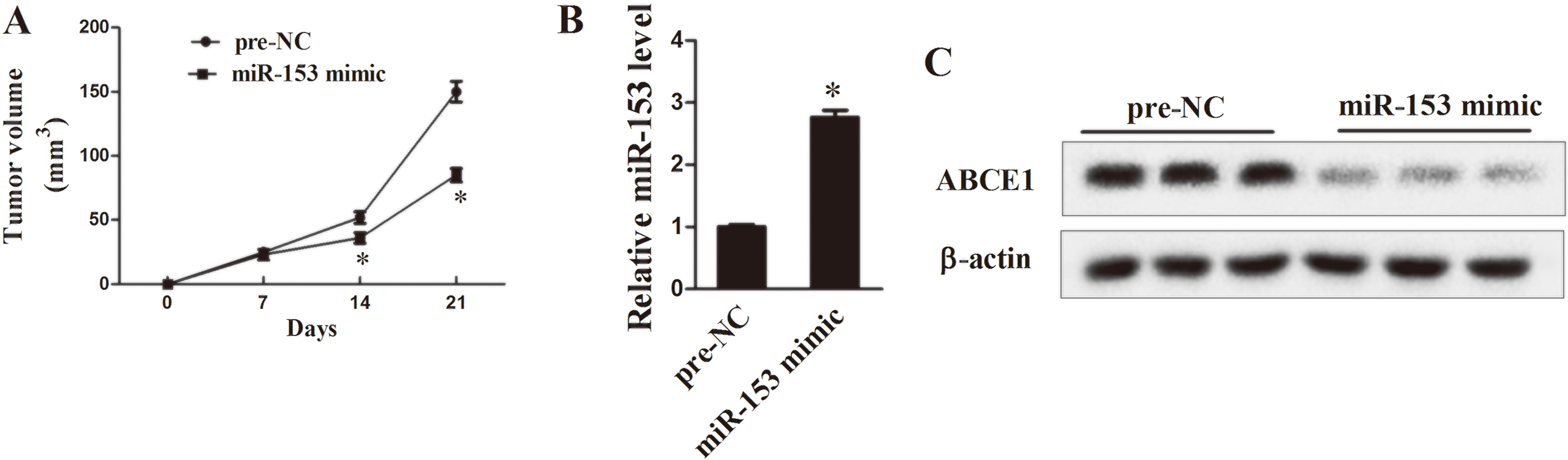
As known, gefitinib has been extensively used in therapy of lung cancer, while the acquired gefitinib-resistance has become a burning question. Recently, the roles of ABCE1 in regulating drug resistance have been reported in many studies. Previous study showed that ABCE1 expression was abundant in lung carcinoma tissues [25]. As well, study also indicated that down-regulation of ABCE1 improved sensitivity of chemotherapeutic agents in lung cancer cells [11]. Furthermore, the proliferation and migration of lung cancer cells were depressed on account of low expression of ABCE1 [8]. In current study, the expression of ABCE1 was higher in GR tissues than in GS tissues. In parallel, the expression of ABCE1 was more abundant in HCC827/Gef cells than in HCC827 cells. Our results were consistent with previous studies. Thus, these results suggest that ABCE1 is a momentous factor in regulating gefitinib resistance in lung cancers, and may be a therapeutic target to regulate gefitinib resistance.
Recent researches demonstrated that miRNAs plays crucial roles in the resistance to gefinitib in lung cancers. Thirteen miRNAs involved in acquired gefitinib-resistance of lung adenocarcinoma were found through microarray analysis [13]. Moreover, studies have confirmed that miR-30b-c, miR-221/222, miR-21, miR-374a and miR-548b took parting in modulating the resistance to gefinitib in lung cancer cells [6, 19, 20]. In this study, the expression of miR-153 was notably up-regulated in GS tissues. As well, the protein level of miR-153 was significantly increased in HCC827 cells. Those results were in common with previous study which showed low expression of miR-153 in NSCLC tissues and cells [6]. As a member of multi drug resistance genes, MDR-1 is important for the development of multi drug resistance which has been considered a major obstacle for cancer therapy [14]. However, over-expression of miR-153 resulted in evident decrease of cell viability and MDR-1 expression in HCC827/Gef cells, and miR-153 silencing remarkably improved the cell viability and the expression of MDR-1 in HCC827 cells in the present study. The results indicated the effects of miR-153 on modulating the sensitivity of gefinitib in lung cancers. Additionally, our results observed that the tumor volume was significantly cut down in nude mice model when miR-153 over-expressed. These results were in consistent with previous studies which revealed the suppressed effects of miR-153 on proliferation, migration and anti-tumor activity of lung cancer cells [15, 24]. These results further indicated that miR-153 inversely regulated the gefinitib-resistance in lung cancers.
What’s more, the expression patterns of miR-153 and ABCE1 were opposite both in vivo and in vitro. That might suggest the reverse regulatory effect between them. According to previous studies, miRNAs regulate genes expression by combining with 3’ UTR of target mRNAs. For examples, MiR-214 works by modulating expression of PTEN and p-AKT [20], miR-299-3p by targeting ABCE1 [24], and miR-548b by affecting on CCNB1 in lung cancer cells. Similarly, our results of luciferase reporter assay implied that miR-153 targets ABCE1 in lung cancer cells. Besides, current studies showed that pcDNA-ABCE1 notably reversed the inhibitor effects of miR-153 over-expression on ABCE1 and MDR-1 expression and cell viability. However, ABCE1 silencing abrogated the changes of ABCE1 and MDR-1 expression and cell viability caused by miR-153 inhibitor. Taken together, those results demonstrated that miR-153 re-sensitized HCC827/Gef cells to gefitinib by down-regulating ABCE1 expression in lung cancer cells.
In summary, the up-regulation of miR-153 inhibits the expression of ABCE1 and thus influences the resistance of lung cancers to gefinitib in vivo and in vitro. The present results may provide a worthwhile therapeutic target to alleviate gefitinib resistance in lung cancer cells in the future.
Footnotes
Acknowledgments
This study was supported by grant from the Natural Science Foundation of Inner Mongolia Science and Technology Department (NYFY YB 2014009).
